Abstract
Congenital skeletal abnormalities may be genetic, teratogenic, or nutritional in origin; distinguishing among these different causes is essential in the management of the disease but may be challenging. In some cases, teratogenic or nutritional causes of skeletal abnormalities may appear very similar to genetic causes. For example, chondrodysplasia associated with intrauterine zinc or manganese deficiency and mild forms of hereditary chondrodysplasia have very similar clinical features and histologic lesions. Therefore, historical data are essential in any attempt to distinguish genetic and acquired causes of skeletal lesions; as many animals as possible should be examined; and samples should be collected for future analysis, such as genetic testing. Acquired causes of defects often show substantial variation in presentation and may improve with time, while genetic causes frequently have a consistent presentation. If a disease is determined to be of genetic origin, a number of approaches may be used to detect mutations, each with advantages and disadvantages. These approaches include sequencing candidate genes, single-nucleotide polymorphism array with genomewide association studies, and exome or whole genome sequencing. Despite advances in technology and increased cost-effectiveness of these techniques, a good clinical history and description of the pathology and a reliable diagnosis are still key components of any investigation.
Keywords
Skeletal abnormalities are commonly encountered in farm animals; in one study of congenital abnormalities in lambs in Australia, 52% of abnormalities involved the skeletal system. 29 The abnormalities may be sporadic or occur as outbreaks, and they may be genetic or teratogenic in origin or secondary to intrauterine nutritional deficiency. Pathologists must be careful not to suggest a genetic basis without good reason: this can have important consequences for the farmer and stud breeder, and it has the potential to result in litigation.
There are a large number of potential genetic skeletal abnormalities in humans; 456 syndromes, organized into 40 groups, have been described on the basis of clinical features, radiographic abnormalities, and molecular defects. 109 As of 2011, mutations had been found in 226 genes to explain 316 diseases in humans. 109 Genetic skeletal diseases of animals are not described and classified in such detail; however, with the increasingly widespread availability of screening techniques, such as single-nucleotide microarrays and whole genome sequencing, the recognition of disease-causing mutations in animals has been revolutionized. The OMIA (Online Mendelian Inheritance of Animals) currently lists disease-causing mutations in 99 of 199 genetic disorders of cattle, 44 of 97 disorders of sheep, and 26 of 54 disorders of pigs. 74 Genetic skeletal diseases with known mutations and the proposed mechanisms are listed in Table 1, with descriptions and illustrations of a number of these recently reviewed. 55,101 A number of genetic mutations, such as developmental duplication in Angus cattle have not been published, as they have become the basis of patented commercial tests. 13 Unfortunately, this leads to a delay in reporting of the gene associated with the defect and limits our ability to establish mechanisms behind these diseases, complete bone development pathways, and advance scientific knowledge.
Inherited Skeletal Diseases of Livestock With Known Mutations, OMIA Numbers, 74 and Proposed Mechanisms.

This review outlines an approach to investigating skeletal diseases in farm animals, with emphasis placed on investigating genetic causes.
Clinical History
A thorough clinical history is often key to distinguish between abnormalities due to genetic and nongenetic causes. A good example lies in investigating cases of chondrodysplastic dwarfism in cattle. A number of hereditary forms of chondrodysplasia with varying levels of severity are recognized, including relatively mild forms—such as “snorter” dwarfism in Herefords, 56 long-nosed dolichocephalic dwarfism in the Angus breed, 60 Ellis van Creveld 2 syndrome in Japanese Brown, 71 and Grey Alpine breeds 73 —and the severe form, Dexter bulldog dwarfism. 45 Mild forms of hereditary chondrodysplasia may be clinically and histologically indistinguishable from the disproportionate dwarfism associated with nutritional deficiencies of manganese or zinc (Figs. 1, 2). The mild genetic forms and the nutritional forms are both associated with a mild-moderate shortening of long bones and a slight decrease in thickness of the hypertrophic zone of the physis. It is seldom possible on the basis of clinical features alone to be confident of the etiology, but information such as pedigree details, knowledge of the number of animals affected, and maternal diet during pregnancy may provide useful clues.

Chondrodysplasia.
In general, a high incidence of skeletal disease in a cohort of newborn animals suggests a non-genetic etiology, unless the disease is caused by a new, dominantly inherited mutation in germ cell lines of a newly introduced sire, as occurs in severe forms of osteogenesis imperfecta. 10 Recessively inherited diseases will only affect 1 in 4 offspring of a heterozygous dam mated to a heterozygous sire, and unless the defective gene is widespread throughout a herd or flock, the incidence is typically low. Even if 1 or more heterozygous sires are used excessively and mated to their offspring, an incidence close to 25% is highly unlikely. If the defect is due to recessive inheritance, then the affected animals are likely to be the offspring of the same sire or a closely related animal mated to related dams. In such cases, examination of pedigrees will be of considerable value.
If the skeletal abnormality has simple autosomal dominant inheritance, then evidence of disease might be expected in approximately 50% of the progeny of similarly affected sires or dams. However, some dominantly inherited skeletal diseases of animals are characterized by incomplete penetrance or codominance, and the abnormality is only partly expressed in heterozygous individuals. Spider lamb syndrome in Suffolk and Hampshire sheep 95 and bulldog chondrodysplasia in Dexter cattle 21 are well-recognized examples of codominance in farm animals. In spider lamb syndrome, heterozygous animals are slightly taller than those not carrying the defective gene, while homozygous mutant animals express the characteristic lesions of the disease. In bulldog chondrodysplasia of Dexter cattle (Fig. 3), heterozygotes have short stature and are considered desirable by many lifestyle farmers, but homozygous animals with 2 copies of the defective gene are nonviable.
Skeletal deformities have been reported in cattle subjected to drought conditions in early pregnancy and fed an inappropriate diet, such as apple pulp, 107 or forced to eat potentially teratogenic plants that would otherwise be avoided or constitute an insignificant percentage of the ration. 111 In such cases, the percentage of newborn animals affected may be very high but the lesions highly variable due to exposure to the nutritional deficiency or teratogenic agent at different times during pregnancy. Insults during embryogenesis (first trimester) are likely to induce more significant defects. If the disorder is due to a teratogenic agent or intrauterine deficiency of a nutrient, such as manganese, mildly affected animals may recover following exposure to a normal ration after birth. This would not be expected if the cause were of genetic origin.
Infection of naïve dams with teratogenic viruses during pregnancy may result in outbreaks of congenital abnormalities on several properties in a region, depending on the distribution of their insect vectors, as seen regularly in parts of Eastern Australia with Akabane virus 59 and more recently with Schmallenberg virus in Europe. 1 However, when the virus is endemic in a region, then only occasional affected animals may be produced. The lesions associated with intrauterine exposure to Schmallenberg virus and other potentially teratogenic viruses, such as Akabane virus, bluetongue virus, and bovine viral diarrhea virus, typically involve multiple organ systems but can resemble skeletal diseases of genetic origin, such as complex vertebral malformation. In cases where a skeletal abnormality is confined to an individual animal, the cause is usually impossible to determine unless the lesions are consistent with a recognized genetic defect in a particular breed and the defective gene can be identified.
Examination of Affected Animals
It is important to examine as many affected animals as possible to accurately characterize the lesions and assess their variability. During autopsy, bones of endochondral and intramembranous origin should be examined with attention to rapidly growing physes, such as costochondral junctions, distal radius, proximal humerus, distal femur, and proximal tibia. Bone fragility should also be assessed, as increased fragility is a feature of certain inherited skeletal diseases, such as osteogenesis imperfecta and osteopetrosis.
Radiography and other forms of imaging may also provide valuable information and can be used either antemortem or postmortem to assess such features as bone shape, cortical thickness, and mineral density or to locate pathologic fractures.
Not all skeletal abnormalities will be associated with an abnormality in bone shape; therefore, sagittal sectioning of representative bones is recommended. In a recent investigation of gingival hamartomas in Belgian blue cattle, it was not until a mutation was found in a gene associated with osteopetrosis that the bones were examined and found to be osteopetrotic. 93
Although skeletal abnormalities of nutritional or toxic origin tend to be variable in presentation, inherited skeletal defects are typically more consistent. An exception to this paradigm is chondrodysplasia of Texel sheep (Fig. 4), where severely affected animals succumb to dyspnea and sudden death as a result of tracheal collapse, while mildly affected animals survive to adulthood and may go on to breed. 103
The trace element status of affected and control animals may provide useful information, but it is important to be aware that test results at the time of birth may not reflect those present several weeks or months previously when the skeleton was at an earlier stage of development. Liver samples from affected animals could be collected for manganese or zinc analysis if the animals have a short stature or for copper analysis if the bones are fragile.
If a genetic etiology is suspected, then whole blood, liver, ear notch, or tail hair samples for DNA extraction should be collected from all affected animals and from the dam, sire, and unaffected siblings (if there are any) to maximize the opportunity to detect a new mutation. We are often contacted about a suspected genetic abnormality near the end of a lambing/calving season, by which time there may be only 1 or 2 affected animals available and no samples have been collected from any others. In such cases, the chances of gaining enough information to distinguish between an acquired and a genetic abnormality or identifying a new genetic defect are remote.
Table 2 lists a number of differential diagnoses for increased bone fragility; Table 3 lists differential diagnoses for short stature; and Table 4 contains other possible presenting signs. Further details are included on the features of each disease that may help distinguish them.
Differential Diagnoses of Increased Bone Fragility in Livestock.

Differential Diagnoses of Short Stature in Livestock.

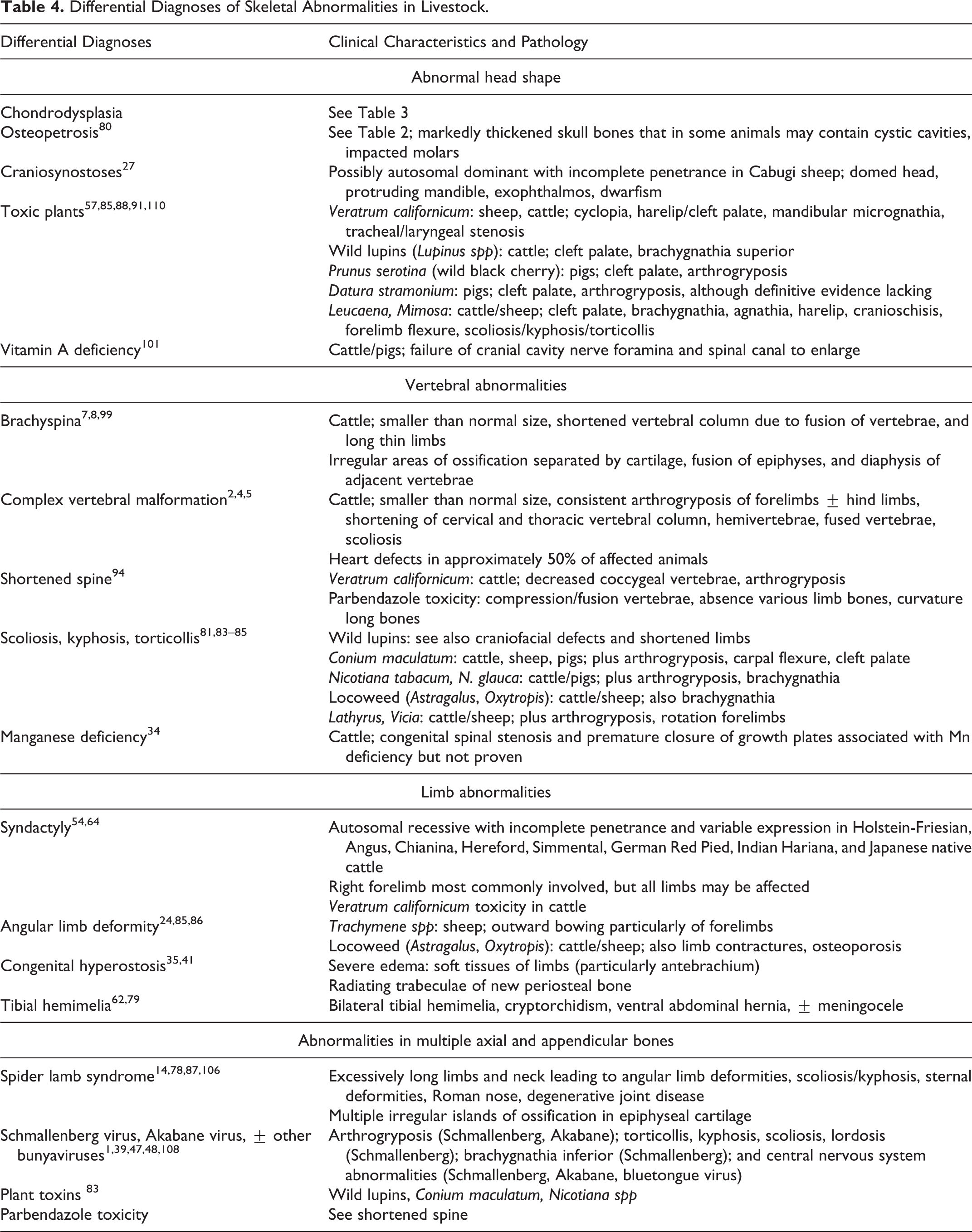
Differential Diagnoses of Skeletal Abnormalities in Livestock.
Methods of Mutation Detection
In cases where nutritional, toxic, and infectious causes of the skeletal abnormalities are considered unlikely or can be excluded, investigation of a genetic etiology should be considered. Historically, this would involve a breeding trial aimed at producing affected animals. This is most efficiently achieved by mating the sire of affected animals to dams that also sired affected animals. However, more long-term trials may be required whereby the sire is mated to unrelated dams and then mated to the F1 females the following year. If the disease has recessive inheritance, it would be expected that 12.5% of the progeny in the second year would be affected.
Mutation detection, while easier than it was even 5 years ago, is still challenging, but potential exists for detection of a mutation without the expense, inherent variability, and potential welfare concerns involved in breeding defective animals. The type of gene mutation analysis used may vary depending on the inheritance pattern, determined from either pedigree analysis or breeding trials. With the decreasing costs of DNA technology, the temptation exists to neglect a thorough clinical and pathologic description of the disease. However, the recognition and characterization of new genetic diseases are often based on the reporting of specific abnormalities in a small number of animals on widely separated properties, sometimes over a period of several years, and it is important that veterinary clinicians and pathologists continue to accurately identify and record their findings.
Despite advances in technology, even in human medicine for nearly half of the described monogenic disorders, the molecular basis of the disease is still unknown. 96 There are a number of different methods of mutation detection, the 3 most commonly used methods are each discussed in turn.
Candidate Gene Approach
This technique is used when the clinical signs and histologic changes are consistent with a previously described disease of the same species with a known mutation or when the disease is closely analogous to a human disease or mouse model with a known affected gene. 28 The candidate gene can be sequenced either in one hit—using long-range polymerase chain reaction and next-generation sequencing—or by sequencing each exon of the gene individually. Although exonic sequencing may miss disease-causing intronic mutations, such as splice site mutations, this risk can be mitigated by including in the sequencing 100 to 200 base pairs of intron on either side of the exon. DNA of a poorer quality, such as that obtained from formalin-fixed paraffin-embedded sections, is suitable for this approach.
We used this approach to investigate a chondrodysplasia in Scottish Highland cattle. The mating of 2 purebred Scottish Highland cattle resulted in the birth of a calf with severely shortened limbs, a ventral abdominal hernia, domed head, shortened muzzle, and protruding tongue, consistent with bulldog chondrodysplasia of Dexter cattle. On histologic examination, intramembranous bone formation was normal, but endochondral ossification was severely restricted (Fig. 5). Testing showed that the calf was homozygous for the Dexter BD1 mutation, 21 while both parents were heterozygous carriers. 30
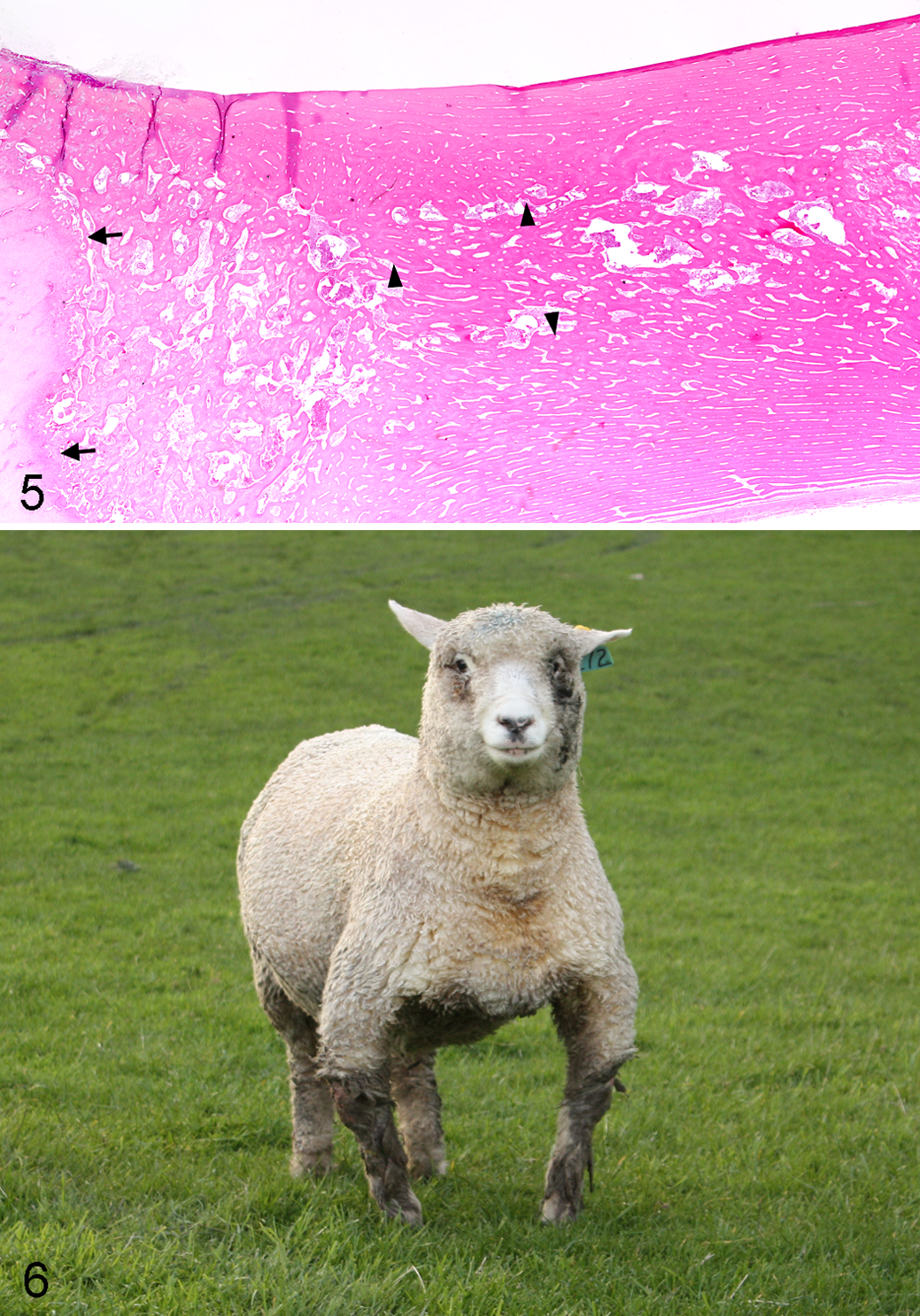

Bulldog chondrodysplasia, radius, bovine. Severely restricted endochondral ossification (arrows) with a disproportionate amount of bone formed by intramembranous ossification beneath the periosteum (arrowheads). HE.
Single-Nucleotide Polymorphism Array and Genomewide Association Studies
Single-nucleotide polymorphisms (SNPs) are single base pair changes in DNA that occur commonly in the genome. 19 The SNPs may either be silent (ie, cause no change in the amino acid sequence) or lead to functional changes if, for example, there is a change in an amino acid or if they affect a mRNA splice site. Genome-wide association studies make use of the concept of linkage disequilibrium, which, at its most basic, is where sections of DNA containing 2 or more SNPs are transferred together during a recombination event in meiosis and the sequence of DNA is inherited intact. 19,61 These areas of linkage disequilibrium become smaller with increasing numbers of generations. Using homozygosity mapping (in the case of recessive inheritance), we can determine regions that are inherited by descent (IBD) and associate them with the affected phenotype, thus focusing the area of DNA that must be searched for a mutation.
This technique requires a relatively large amount of high-quality DNA. While it is possible to do SNP arrays on formalin-fixed paraffin-embedded tissue without optimization, base calling may be decreased, thereby reducing the effectiveness of the tool. Chips are available for a number of different species. For cattle, Illumina arrays are available for a number of different levels of coverage across the bovine genome. Illumina beadchips are also available for the dog, sheep, and pig. Affymetrix, the other major SNP array producer, manufactures a SNP array for the cow, chicken, and dog.
We used this technique to determine the mutation causing inherited rickets in Corriedale sheep (Fig. 6). With use of the Illumina ovine 50-K chip and a combination of known heterozygotes, homozygous affected animals, and sire and dam DNA followed by homozygosity mapping, a large 6-Mb IBD region was found on OAR 6. By aligning this area with the annotated bovine genome, 35 potential candidate genes were found. Of the 35 genes in the area, several had an association with bone development or skeletal dysplasias; however, mutations in one of these—dentin matrix protein 1 (DMP1)—had recently been described as causing autosomal hypophosphatemic rickets in humans and mice. 68 Sequencing of this gene in our sheep demonstrated a nonsense mutation in exon 6 of the DMP1 gene, which led to a premature stop codon and truncation of the protein. 114
The chances of successfully detecting a mutation using this technique is increased if the inheritance of the disease is known and if DNA from family members, including unaffected siblings, dams, and sires is available. Accurate recognition of phenotype is also important, as misdiagnosis of affected animals will obviously affect the analysis. Cost may be a limiting factor, as is the availability of tissue samples from related animals in diseases that take time to develop if sires, siblings, or dams are no longer available. The use of SNP chips for investigating diseases with dominant inheritance is more challenging. Not only are more affected animals required in the analysis, but in diseases with incomplete penetrance, accurate determination of affected animals can be difficult.
For cattle, a custom SNP chip that contains the genetic diseases of cattle with known mutations has been developed by Irish researchers. Known as the International Dairy and Beef chip, this provides the ability to scan bovine DNA samples for all known genetic diseases at one time. 72
Whole Exome Sequencing or Whole Genome Sequencing
Whole exome sequencing involves selectively sequencing only the coding sequences or exons in the genome by the process of exome capture (see Bamshad et al for a description of the process 12 ), which allows enrichment of the sample. This may be a more cost-effective method for mutation detection than whole genome sequencing, particularly for Mendelian disorders. However, whole exome sequencing is not an option for all species. Exome capture is available for cattle, and an exome-wide DNA capture based on Bos taurus has been used for Bos indicus and Bison species, 26 indicating that there may be enough homology between some ruminant species and the bovine exome to allow its use in other ruminants.
A large quantity of good-quality DNA is required for exome sequencing, which may limit its usefulness in archived samples. In addition, approximately 5% to 10% of the exome (in humans) is poorly covered by the exome capture process. 12
Whole genome sequencing, as the name suggests, involves sequencing the whole genome, including exons and introns, and the technology is now relatively affordable. Although it might seem that whole genome sequencing is the ultimate answer for determining a disease-causing mutation, there are limitations. In reality, sequencing of the “whole” genome is not necessarily possible, and the results may be skewed by poor coverage of GC-rich areas. 11 A large volume of information is obtained from whole genome sequencing, and appropriate computing and bioinformatics resources are required to process and interpret the data.
In a recent investigation of primordial dwarfism in Angus cattle, we have used SNP array with genome-wide association studies, followed by whole genome sequencing. Due to the distant familial relationships of affected animals, large numbers of small IBD regions were obtained on SNP array, and since many older samples were only formalin-fixed paraffin-embedded sections, base calling on these was unreliable. Basic analysis of whole genome sequencing from 2 affected animals produced a large number of variants between affected animals and the reference bovine genome, and data analysis is continuing.
One investigation in which this technique of combined SNP array and whole genome sequencing has been used successfully is an investigation into a chondrodysplasia in Grey Alpine cattle. SNP array and genome-wide association studies showed a 1.6-Mb IBD region on bovine chromosome 6, and whole genome sequencing of an affected calf revealed a 2–base pair deletion in the Ellis van Creveld 2 (EVC2) gene. 73
Conclusion
Distinguishing between genetic and nongenetic causes of congenital skeletal abnormalities in farmed livestock remains a challenge but has been greatly enhanced in recent years by advances in molecular genetic technology. Many techniques previously used only for research purposes because of their complexity and expense are now becoming available for commercial use. This trend will inevitably continue, but progress in molecular methods must be underpinned by accurate diagnosis and description of disease syndromes based on traditional methods, including epidemiology, clinical characteristics, pathology, and imaging. Pathologists will continue to play an important role in the recognition of these disorders and are ideally placed to recommend what techniques should be applied in an effort to make a definitive diagnosis.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
