Abstract
We report the identification of a novel papillomavirus, Fulmarus glacialis papillomavirus 1 (FgPV1), present within an interdigital foot mass of a Northern Fulmar (Fulmarus glacialis). The mass of interest was composed of normal stratified and keratinized epithelium and dense mesenchymal cells with central cartilaginous islands. Within the nuclei of many chondrocytes were loose aggregates or paracrystalline arrays of virions approximately 50 nm in size. Degenerate polymerase chain reaction was used to identify the virus as a putative papillomavirus, and the entire viral genome of 8132 base pairs was subsequently amplified and sequenced. Analysis revealed canonical papillomavirus architecture, including the early open reading frames E6, E7, E1, and E2 and the 2 late proteins L1 and L2. FgPV1 is most closely related to a cluster of avian and reptilian papillomaviruses as visualized by phylogenetic trees. This observation suggests that papillomavirus virion production can occur in mesenchymal cells.
Keywords
The Papillomaviridae family continues to expand with the ongoing discovery of previously unidentified species occupying over 30 different genera. 46 Avian papillomaviruses have been described in more than 12 avian species and 6 orders 27 since the 1970s. 28,33 In that time, only 3 have been fully sequenced: Francolinus leucoscepus papillomavirus 1 (FlPV1), Psittacus erithacus papillomavirus (PePV), and Fringilla coelebs papillomavirus (FcPV). 31,41,44 Among species described so far, avian papillomaviruses and mammalian papillomaviruses share epitheliotropism and similar tissue distribution. In psittacines, virus has been detected in intestinal mucosal cells, 13,16,19,23,27,30,40 nasolacrimal epithelium, 13,16,22 and oral mucosa at the commissure of beaks. 22 There is also a report of papillomavirus-associated growths on the legs and feet of two related finch species, Fringilla. coelebs and Fringilla montifringilla. 31 To date, there have been no reports of papillomaviruses or any neoplastic lesions in free-ranging seabirds despite the identification of papillomaviruses in numerous marine mammals, 5,35 including dolphins, 36,37 porpoises, 43 and sea turtles. 18
Regardless of host species, productive papillomavirus infection (formation of infectious particles) has been invariably tied to the terminal differentiation of epithelial cells. A small number of papillomaviruses are capable of infecting and transforming fibroblasts, but these infections are considered nonproductive. The members of the Deltapapillomavirus genus such as bovine (Bos taurus) papillomaviruses 1 and 2 (BPV1, BPV2), for example, can infect fibroblasts in cattle, horses, and cats and is associated with transformation in these cells, causing the formation of fibropapillomas, also known as sarcoids in nonbovid species. 20 In these cases, virion production is considered limited to infected cells within the overlying epithelium. 4,10 –12,47
Papillomaviruses are most notable for causing cutaneous papillomas or warts with hyperplasia of the statum spinosum and/or neoplasia of the cutaneous stratified epithelium. 26,32 The viruses are also capable of causing lesions in the alimentary tract and within oral mucosa in the case of bovine (Bos taurus) papillomavirus 4 4 and canine (Canis familiaris) oral papillomavirus, 9 respectively. There are also a few papillomaviruses capable of causing a fibropapilloma, which is characterized by both a cutaneous papilloma and an underlying fibroma of the connective tissue. 9,26,32
Northern Fulmars, Fulmarus glacialis, are seabirds of the family Procellariidae and are surface feeders that spend the majority of their life at sea covering vast migratory ranges. They have been used as biomonitors of plastic, 2 halogenated organic compounds, 17,24 and heavy metal pollution, 17,24 but otherwise research on this species of seabird is limited. In this study, we set out to characterize masses identified on the left foot of a Northern Fulmar that was rescued due to severe emaciation.
Materials and Methods
Histology
Formalin-fixed tissues were routinely processed and embedded in paraffin within 1 day of necropsy, sectioned at 5 μm, and routinely stained with hematoxylin-eosin (HE) for histologic evaluation. Select 5-μm sections were stained with Alcian blue solution (pH 2.5). Immunohistochemistry was performed on 4-micron-thick, formalin-fixed, paraffin-embedded tissue sections processed routinely for deparaffinization (for full methods, see Supplemental Material). Antigen retrieval (AR) and primary antibody sources are as follows. List of immunohistochemistry (IHC) specifics (primary antibody type, clone [source], dilution, antigen retrieval, diluent, and block): Mouse anti-cytokeratin cocktail, clone AE1 and AE3 (MU071; BioGenex, San Ramon, CA), 1:300, HIER steamer (S1699) + citrate–30′, phosphate-buffered saline (PBS)–Tween, normal horse serum (NHS) Mouse anti–pan-cytokeratin, clone LU-5 (CM043; BioCare Concord, CA, USA), 1:100, HIER Steamer (S1699)–30′, PBS-Tween, NHS Mouse anti-vimentin, clone 3B4 (M7020; Dako, Carpinteria, CA), 1:300, HIER steamer (S1699)–30′, PBS-Tween, NHS Rabbit anti–bovine papillomavirus type 1 (Dako B0580, Carpinteria, CA), 1:800, no antigen retrieval, Dako Antibody Diluent (S0809), Dako Serum Free Protein Block (X0909)
Sections were counterstained (Mayer’s hematoxylin). Nonspecific background was evaluated in duplicate sections receiving normal serum or an isotype-matched control in place of the primary antibody. Histologically normal tissue within the sections served as our internal control for all IHC with the exception of the rabbit anti–bovine papillomavirus type 1 antibody (rb-antiBPV1), for which a section of bovine esophageal papilloma served as the positive control.
Electron Microscopy
Formalin-fixed tissue from the lesion was fixed and prepared for transmission electron microscopy (see Supplemental Material). For direct electron microscopy, fresh tissue from the lesion was scraped directly into a small amount of 2% phosphotungstic acid stain and deposited onto a formvar/carbon-coated grid and examined as above.
Nucleic Acid Isolation and Sequencing
Nucleic acids were extracted from a 0.25mg lesion using the DNeasy Blood and Tissue Kit with overnight Proteinase K digestion in Buffer ATL (Qiagen, Valencia, CA) and stored at −20°C. Then, 2.5 ng of extracted nucleic acid and 1 ng of pUC19 (positive control) or water (negative control) were used as a template for rolling circle amplification (RCA; illustra TempliPhi 100 Amplification Kit; GE Healthcare Life Sciences, Piscataway, NJ). Digestion of the RCA product with 4-cutter, Sau3AI, resulted in multiple DNA fragments that were gel purified (QIAquick Gel Extraction Kit; Qiagen) and ligated into dephosphorylated BamHI-digested pUC19. Plasmid inserts were Sanger sequenced with a BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, Foster City, CA). Alternatively, 2.5 ng of extracted nucleic acid or a 1:10 dilution of RCA product served as the template for degenerate polymerase chain reaction (PCR) by using published papillomavirus primers. 1,15 Two primer sets were then designed to span the remaining gaps between E1 and L1 (Supplemental Material) and were amplified using AccuPrime Taq (Invitrogen, Carlsbad, CA). Samples were cloned and sequenced as described above. As we obtained more sequence, new primers were designed to primer walk around the inserts and complete the genome.
Sequence Analysis
Sequenced fragments were analyzed for high-quality sequence, vector sequence was trimmed, and the genome was assembled and confirmed by CAP3 21 (see Supplemental Material for full description). The full-length genome was deposited in GenBank (KJ452243). Open reading frames were predicted and analyzed using Basic Local Alignment Search Tool (BLAST) algorithms (BLASTn and BLASTp) to assess similarity to known papillomavirus sequences. The L1 nucleotide sequence was examined using the Pavic L1 Typing Tool. 29,34 All E1 and L1 protein sequences were downloaded from the reference clone collection of 274 full-length animal and human papillomaviruses available at PaVE. 34,45 Multiple sequence alignments (MSAs) were performed with FgPV1 putative proteins using ClustalW in MEGA5.2.2. 42 MSAs were used to construct neighbor joining (NJ), 39 maximum likelihood (ML), minimum evolution (ME), and maximum parsimony (MP) phylogenetic trees in MEGA5.2.2. The trees were drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the p-distance method 48 and are in the units of the number of amino acid differences per site. The analysis involved 275 amino acid sequences. All ambiguous positions were removed for each sequence pair. There were a total of 405 (E1) and 616 positions (L1) in the final data set. To estimate confidence, bootstrap resampling was performed with 1000 replicates for NJ and ME trees; only values with greater than 75% are displayed as significant.
Results
Clinical and Gross Findings
A Northern Fulmar was admitted for rehabilitation after stranding on Dillon Beach, California, with severe emaciation. The bird was euthanized due to significant plumage damage associated with ulcerative dermatitis that would have necessitated an untenably prolonged stay in captivity. Unrelated to the feather damage were 2 discrete masses present on the left foot. One 2.5 × 2 × 1.5cm mass protruded from the lateral surface of the fourth digit, near the junction of the second and third phalanx, and the second (∼3 cm diameter) bulged from the plantar and dorsal surfaces of the intermediate web apposed to the third digit (Fig. 1). All but a stalk of tissue extending to the latter mass dropped off the bird shortly after arrival in care, leaving a ∼1cm hole in the webbing. This investigation focuses on the former mass, although both were similar. The overlying epidermis was elevated and centrally ulcerated but otherwise blended indiscriminately with the surrounding skin. Radiographs (data not shown) revealed that the bones of the foot were smooth and intact except for mild periosteal proliferation in segmental regions near the masses. The masses were within soft tissues only, with no detectable connection to or involvement of the joints or phalanges. On cut surface, the mass consisted of wavy, slightly translucent, grayish lobules of tissue with dull, linear regions of friable material at the center and a peripheral rim of firm, tan fleshy tissue.
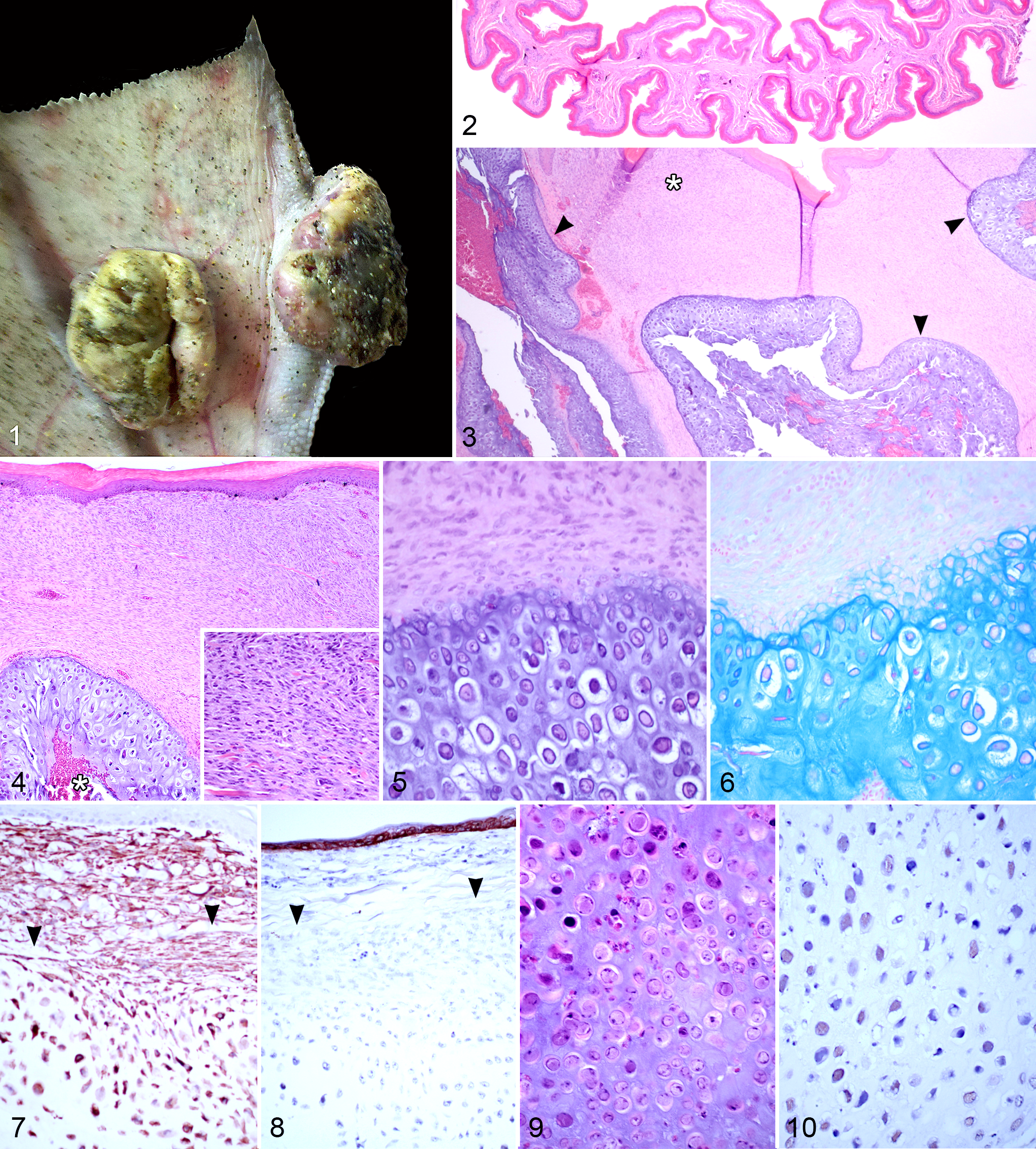
Papillomatous masses, left foot; Northern Fulmar. Two pedunculated and ulcerated masses bulge from the plantar surface of the interdigital webbing and the lateral surface of the third phalanx of digit 4.
Histologic Findings
The normal dermis of interdigital foot webbing in the Northern Fulmar is thin (<0.1 cm) and contains scant numbers of fibroblasts, a modest amount of loosely arranged collagenous tissue, and small caliber vessels (Fig. 2). The papillomatous masses expanded the interdigital space, apposing a mostly intact keratinizing surface epithelium (Fig. 3, top), and were composed of a spindle cell component (asterisk) and central cartilage (arrowheads). In a few areas, there was regional extension to an ulcerated surface. The composition of the mass at its periphery was a dense spindle to stellate cell population, with cells arranged haphazardly or in streams parallel to the dermal-epidermal junction (Fig. 4 and inset). The range of mitotic metaphase figures in this region was from 1 to 5 per 10 high-power fields, and there were rare, scattered, bizarre stellate cells. Embedded in, and comprising over half of the mass, were lobular to serpentine regions of cartilage (Figs. 3, 4), with central, irregular, blood-filled clefts (Fig. 4, asterisk). There was a smooth yet indiscrete progression from densely packed spindle to oval cells at the cartilage periphery to isolated, round, and multiangular cells surrounded by abundant matrix in the central regions (Figs. 4, 5). The cartilaginous matrix stained characteristically with both Alcian blue (Fig. 6) and toluidine blue (data not shown). Both the spindle cell portion of the mass and the chondrocytes were positive for vimentin by immunohistochemistry (Fig. 7). Cytokeratin-positive cells were limited to the overlying epidermis (Fig. 8). Most central chondrocytes contained large, amphophilic to lightly basophilic intranuclear inclusions (Fig. 9). Inclusions were reactive with a polyclonal antibody generated against bovine papillomavirus 1 (Fig. 10). There was marked inflammation of the connective tissue subjacent to regions of ulceration. As suggested by the antemortem radiographs, neither the joints nor the phalangeal bones were contiguous with the tumor, but there was a regional hyperplastic subperiosteal reaction that eccentrically radiated from the diaphysis toward the tumors (data not shown).
Electron Microscopy
By transmission electron microscopic examination of fixed tissue, within chondrocytes (and targeting a region of the histologically evident inclusions), virions were present within both the cytoplasm and nuclei of some infected cells (Fig. 11). Many of the affected cells also contained paracrystalline arrays of virions with diameters ranging from 46 to 48 nm (Figs. 12, 13). The tissue was consistent with hyaline cartilage by the following ultrastructural characteristics: there were no cytoplasmic connections between cells, and chondrocytes contained lipid droplets and very few mitochondria or other organelles (Figs. 11, 12). Also, the pericellular region was empty space except for a feltwork halo of loosely arranged, scattered, delicate, short fibrils (Fig. 11). 38 In the immediate pericellular region of some cells were irregular, perpendicular fibrillar extensions into the surrounding matrix (Fig. 11). Viruses identified by direct electron microscopic examination of negatively stained preparations of tissue from the mass were composed of capsomeres arranged in icosahedral symmetry in the size range of 56 to 58 nm (Fig. 14).
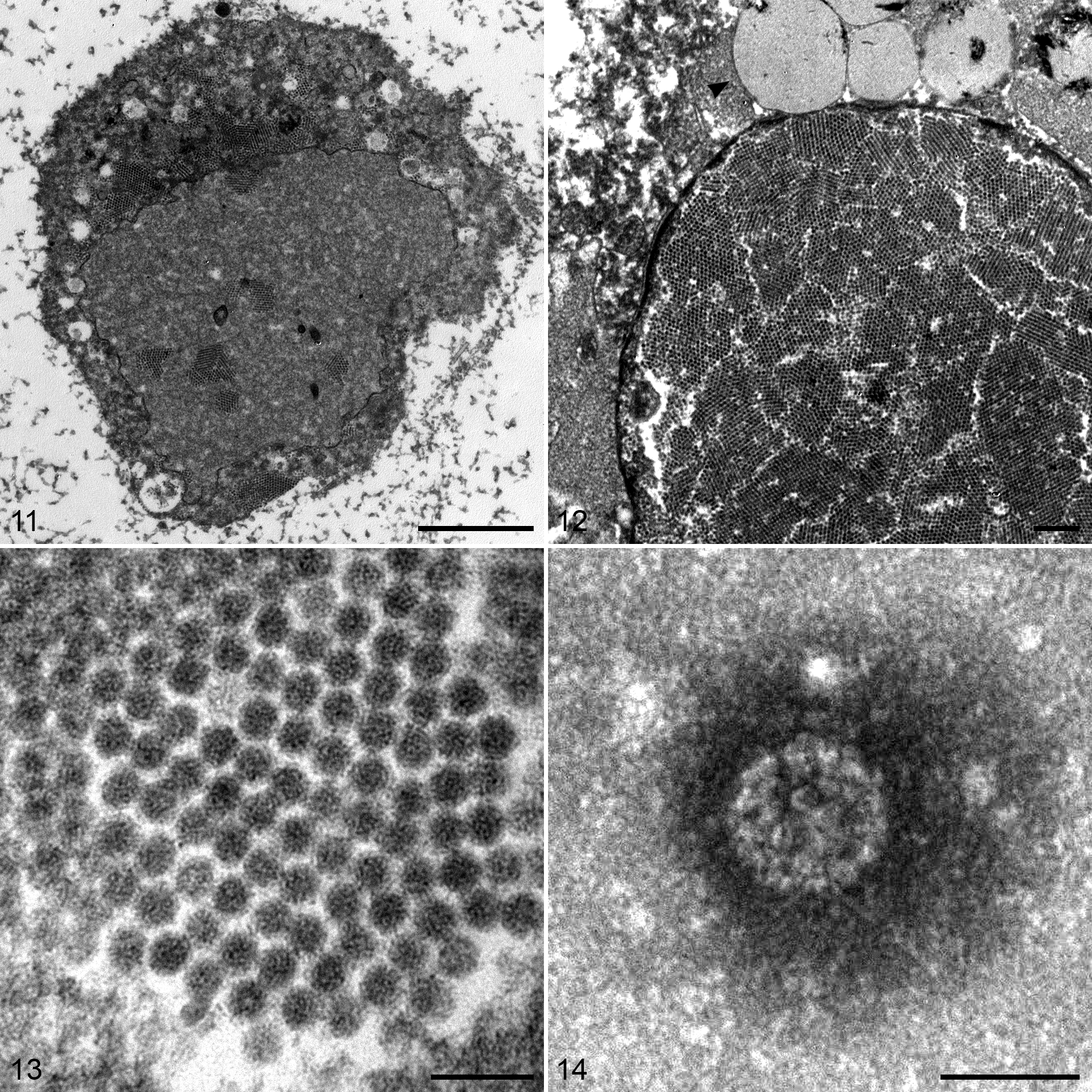
Papillomatous masses, chondrocytes, left foot; Northern Fulmar. Transmission electron microscopy, uranyl acetate.
Virus Identification
Total nucleic acid was extracted from a portion of the mass, and degenerate PCR for papillomaviruses was performed. Sequencing of the resultant amplicon and analysis by BLASTx revealed that Mus musculus papillomavirus 1 (MmuPv1) was the most significant hit and shared only 52% amino acid identity. The remainder of the genome was amplified and sequenced as described in the Materials and Methods. The level of taxonomic uniqueness was determined by analyzing the L1 coding region using the L1 Typing Tool. 29 This analysis demonstrated that the virus we identified shared only 26.45% identity with Francolinus leucoscopus papillomavirus (FlPV1) and can be considered a new papillomavirus species in a new papillomavirus genus. Using the proposed classification and taxonomy previously established, 3 we have named the virus based on the host species from which it was identified, Fulmarus glacialis papillomavirus 1, FgPV1 (KJ452243).
The complete genome of FgPV1 is 8132 base pairs (bp), and analysis of the DNA sequence revealed genomic features characteristic of animal papillomaviruses. 20,47 First, the size was within the range previously reported for papillomaviruses, 6.9 to 8.6 kb. In addition, the GC content of 48.1% was within the range described for avian and reptilian papillomaviruses (47.2%–52.9%). 18,30,41,44 Using open reading frame (ORF) prediction software, we identified 12 predicted ORFs open on the single coding strand. Four of the predicted ORFs corresponded to the early (E) proteins, E6, E7, E1, and E2, and 2 corresponded to the late (L) proteins, L2 and L1 protein (Fig. 15). A noncoding region of 1501 bp (18.5% of genome) was identified between the stop codon for L1 and the initiation codon for E6. There was no intergenic region between the early and late proteins, since the E2 and L2 genes had an overlapping stop and start.
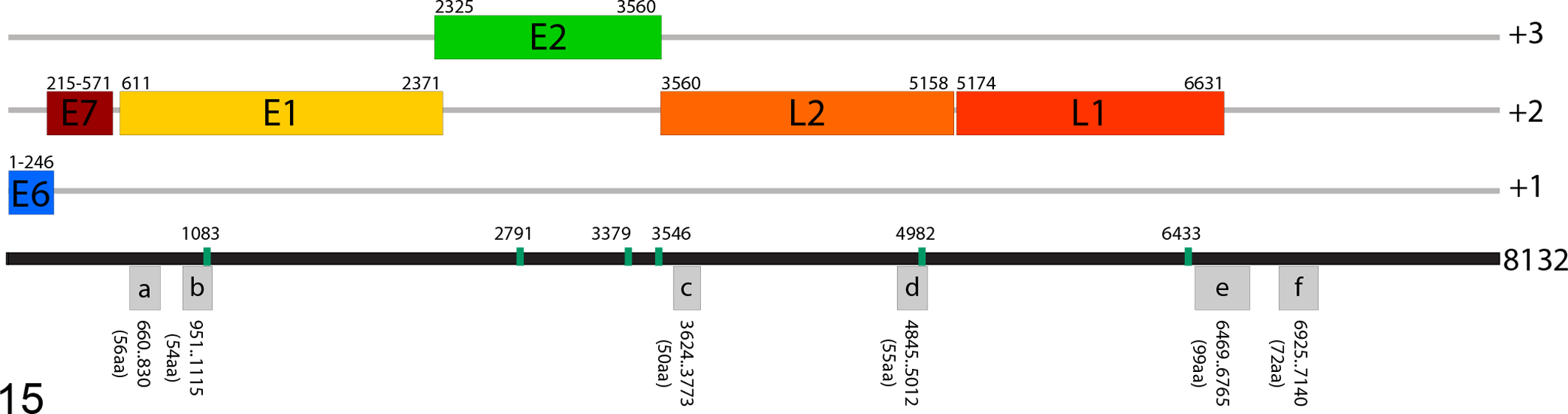
Schematic of the Fulmarus glacialis papillomavirus 1 (FgPV1) genome. Each predicted open reading frame (ORF) is represented as a rectangle in the appropriate reading frame with the denoted nucleotide positions. The gray boxes represent predicted ORFs, denoted with a lowercase letter, with no homology to viral proteins. The vertical green bars represent the E2 binding sites.
A putative E6 (82aa) and E7 (119aa) were identified based on genomic location and predicted sizes, and no significant identity to papillomavirus (PV) genomes or gene products was identified using BLASTp. The putative E6 initiation codon has also been used to identify the first nucleotide of the genome. The predicted E7 protein contained characteristic features, including a putative Rb binding domain (LxCxE) of LRCYE at positions 33–37 and 2 putative zinc-binding domains (CxxC) in the C-terminal region. The remaining 6 putative ORFs did not share identity with papillomavirus coding sequence in the nr (NCBI) database, and no determination has been made regarding functionality.
The predicted E1 protein was 587 amino acids (aa) long and contained the conserved N-terminal and C-terminal domains annotated in the Pfam database. The putative E2 protein was 412 aa and similar to several PV genera, and no E4 ORF was predicted within the E2 coding region. A total of 6 E2 binding sites (ACCN6GGT) were found starting at positions 1083, 2791, 3379, 3546, 4982, and 6433 (Fig. 15). The papillomavirus capsid proteins L2 (533 aa) and L1 (486 aa) were present and contained the conserved Pfam domains.
To examine the phylogenetic relationship of FgPV1 in relation to the currently sequenced human and animal viruses, multiple sequence alignments were performed on L1 and E1 protein sequences and used as the input for phylogenetic tree construction (Fig. 16, Suppl. Fig. S1). From these analyses, FgPV1 is most closely related to other avian and reptilian papillomaviruses. Using the neighbor-joining method, FgPV1 E1 and L1 primarily clustered with the 3 avian species (FlPV1, FcPV1, and PePV1) and were in a larger cluster including the 2 reptilian species (Caretta caretta papillomavirus [CcPV1], Chelonia mydas papillomavirus, [CmPV1]) (Fig. 16, Suppl. Fig. S1). The exact topology varied slightly with FgPV1 E1 depending on the method used; with ME, the topology matched that of the NJ tree, but when ML or MP methods were used, FgPV1 E1 appeared as an out-group to the avian-reptilian cluster but was still within that clade and did not cluster with any other clades of animal papillomaviruses (data not shown). FgPV L1 maintained the same relationships in all methods (data not shown).
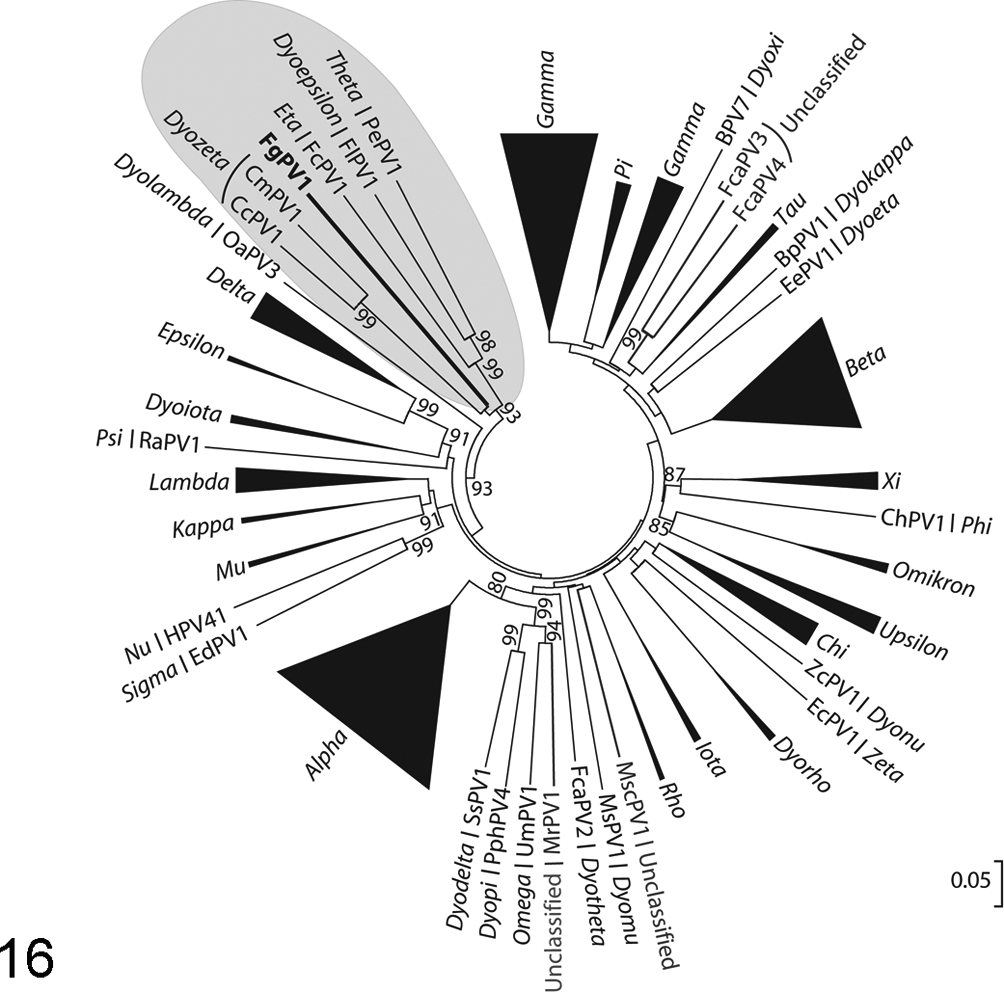
Neighbor-joining phylogenetic tree of the predicted L1 protein. All genera have been denoted with italic text, and significant bootstrap values are present. Genera with more than one member were collapsed, with the triangle representing the diversity within that genera. The avian/reptilian cluster is highlighted in gray, with the Fulmarus glacialis papillomavirus 1 (FgPV1) denoted with bold font. The scale bar represents the number of amino acid differences per site.
Discussion
Using a combination of histopathology, electron microscopy, and degenerate PCR, we characterized interdigital papillomatous masses and identified a novel papillomavirus in a Northern Fulmar. FgPV1 has a circular genome of 8.1 kb, well within the range of published animal papillomaviruses (6.9–8.6 kb), and although highly divergent at the nucleotide level, it maintains a genomic organization expected for the family. The virus shares less than 65% nucleotide identity in the L1 coding region (26.4% nt identity to FlPV1) and is therefore a new papillomavirus species, and it represents a new taxonomic genera, tentatively Dyosigma.
The lesion we have characterized is unique with regard to papillomavirus biology. The papillomatous masses were composed of a proliferative mesenchymal cell population with regional cartilaginous differentiation that contained paracrystalline arrays of FgPV1 within mature chondrocytes. In classic papillomas, infected epithelial cells mature to their terminally differentiated phenotype in the highly keratinized stratum corneum. 14 Despite virus-induced hyperplastic responses within the layers of the epithelium, papillomavirus virions have been demonstrated to be fully formed and visualized only in paracrystalline arrays within the nuclei of degenerated cells in the keratinized layer. 26 This process begins with basal cell infection, with the rest of the life cycle modulated by differential viral protein synthesis tightly linked to epithelial cell differentiation. 20 The virus is shed with the differentiated epithelial squame that is released from the surface. In this case, the surface epithelium of the Northern Fulmar lesion lacked any histologic evidence of epithelial involvement. The proliferative changes were limited to an expanded dermal layer where dense spindle cells had a smooth but indistinct transition to chondrocytes with a progressive accumulation of surrounding chondroid matrix. Only the central-most chondrocytes (vimentin positive, cytokeratin negative, and anti-BPV1 positive) contained virions by transmission electron microscopy (TEM). We propose 2 potential explanations for this outcome. First, FgPV1 could be infecting an undifferentiated mesenchymal cell and driving their differentiation to chondroblasts and to chondrocytes, analogous to that in maturing epithelium infected by other papillomaviruses. Second, FgPV1 could be a secondary infection of a primary neoplastic lesion and infect these cells as they are actively dividing. Unfortunately, with only a single case and lack of sufficient tissue, we are unable to distinguish these possibilities.
In considering this unique lesion, the historical precedent of BPV-induced fibropapillomas deserves mention. Bovine fibropapillomas involve virus-induced hyperplastic responses within the epithelium along with transformation of the dermal fibroblastic population. Moreover, in BPV-infected equine sarcoids, virus production is limited to epithelium overlying the mass, despite the detection of episomal BPV and potentially viral replication within the proliferative spindle cells that form the mass. 4,6 –8,25,47 We observed minimal hyperplasia of the overlying epithelium, with no evidence of viral infection within the epithelial cells, but rather dermal expansion by dense spindle cells and virions within chondrocytes of central cartilaginous islands.
Analysis of the genome of FgPV1 revealed conservation of the general organization of papillomavirus genomes. The 4 most conserved proteins—E1, E2, L2, and L1—were all easily identified as ORFs and with sufficient sequence similarity for protein identification. In addition, ORFs corresponding to the genomic location and size of E6 and E7 were predicted and contained conserved motifs but did not have significant sequence similarity to other papillomaviruses based on BLASTp analysis. However, FgPV1 did not appear to have an E4 protein encoded within the E2 ORF (like the other avian species) or an E9 ORF (which 2 of the 3 avian papillomaviruses encode). 44 There were 6 additional predicted ORFs between 50 and 99 aa in length, but analysis by BLASTp revealed no significant matches to the Papillomaviridae family or any other protein sequence in the nr (nonredundant) database on NCBI. These proteins do not seem to have orthologs in any known papillomavirus based on location or size, and without functional or transcription analyses, it is difficult to assess whether they are produced during infection. Lastly, one major difference between FgPV1 and the other avian and reptilian species is the length between the stop codon of L1 and the start codon of E6, which in those 5 species ranges from ∼450 to 750 nt and comprises 6% to 10% of the genome. In FgPV1, the noncoding region (NCR) or upstream regulatory region (URR) comprises 1501 bp or 18.5% of the genome. However, if ORFe and ORFf are functional proteins, the NCR or URR would be smaller, but at 12% of the genome, it would still be the largest in this group. On average, it is from 7% to 11% in papillomaviruses and is known to contain a large number of cis-acting elements to control gene expression and replication.
In conclusion, we have identified a novel papillomavirus in a lesion of proliferating and differentiating mesenchymal tissue, perhaps indicating a novel mechanism of productive viral infection for papillomaviruses. To our knowledge, this is the first description of a productive papillomavirus infection in terminally differentiated mesenchymal tissue, the first identification of a papillomavirus in this species of bird, and the first member of a novel genus within the Papillomaviridae family.
Footnotes
Acknowledgements
We thank the CAHFS–Davis and UC Davis VMTH Histopathology Laboratories for their technical expertise; Drs John Munday, Jennifer Luff, Keith Thompson, and Steven Kubiski for their helpful discussions and input; and Nat Smith for the gross image.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research is partially funded by grants received for pathogen discovery efforts including The Bernice Barbour Foundation and Boerhinger-Ingleheim Vetmedica.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
