Abstract
Baboon orthoreovirus (BRV) is associated with meningoencephalomyelitis (MEM) among captive baboons. Sporadic cases of suspected BRV-induced MEM have been observed at Southwest National Primate Research Center (SNPRC) for the past 20 years but could not be confirmed due to lack of diagnostic assays. An immunohistochemistry (IHC)–based assay using an antibody against BRV fusion-associated small transmembrane protein p15 and a conventional polymerase chain reaction (PCR)–based assay using primers specific for BRV were developed to detect BRV in archived tissues. Sixty-eight cases of suspected BRV-induced MEM from 1989 through 2010 were tested for BRV, alphavirus, and flavivirus by IHC. Fifty-nine of 68 cases (87%) were positive for BRV by immunohistochemistry; 1 tested positive for flavivirus (but was negative for West Nile virus and St Louis encephalitis virus by real-time PCR), and 1 virus isolation (VI) positive control tested negative for BRV. Sixteen cases (9 BRV-negative and 7 BRV-positive cases, by IHC), along with VI-positive and VI-negative controls, were tested by PCR for BRV. Three (of 9) IHC-negative cases tested positive, and 3 (of 7) IHC-positive cases tested negative by PCR for BRV. Both IHC and PCR assays tested 1 VI-positive control as negative (sensitivity: 75%). This study shows that most cases of viral MEM among baboons at SNPRC are associated with BRV infection, and the BRV should be considered a differential diagnosis for nonsuppurative MEM in baboons.
Spontaneous viral encephalitis is a rare pathology in baboons 51 and is seldom diagnosed clinically. So far, the only reported outbreak of viral meningoencephalomyelitis (MEM) among baboons occurred during 1993–1994 at the Southwest National Primate Research Center in San Antonio, TX (SNPRC). 51 During this outbreak, 8 juvenile baboons developed neurological symptoms and showed histopathological lesions of acute nonsuppurative MEM. 51 A novel double-stranded RNA (dsRNA) virus with 10 genomic fragments and syncytium-inducing capability was isolated from 5 affected baboons. 28,51 The isolated virus was inoculated into 2 healthy baboons to reproduce similar lesions of MEM. 51 Based on the morphological and genomic characteristics, the viral isolate was classified as a novel species (Baboon orthoreovirus; BRV, isolate 10895) of the genus Orthoreovirus, family Reoviridae. 26,28,51
The family Reoviridae is the largest and the most diverse family of the nonenveloped dsRNA virus families. 24 It has 2 recognized subfamilies, based on the presence (subfamily Spinareovirinae) or absence (subfamily Sedoreovirinae) of spikes or turret proteins at each of the icosahedron vertices of the viral capsid. 68,69 The subfamily Spinareovirinae includes 9 genera (Orthoreovirus, Aquareovirus, Oryzavirus, Fijivirus, Mycoreovirus, Cypovirus, Idnoreovirus, Dinovernavirus, and Coltivirus), and subfamily Sedoreovirinae includes 6 genera (Orbivirus, Rotavirus, Seadornavirus, Phytoreovirus, Cardoreovirus, and Mimoreovirus). 68,69
The genus Orthoreovirus includes viruses affecting a wide variety of vertebrate hosts, including fish, birds, reptiles, and mammals. Distinguishing features include the following: (1) a segmented genome consisting of 3 large (L1, L2, and L3), 3 medium (M1, M2, and M3), and 4 small (S1, S2, S3, and S4) linear dsRNA segments, which code for 3 λ, 3 μ, and 4 σ translation products; 27,69 (2) transmission by respiratory or fecal-oral route; 69 (3) isolated only from vertebrate hosts; and (4) all members of all species groups, except the mammalian orthoreovirus (MRV), forming syncytia. 24,26,27,69 Furthermore, depending on syncytium formation and the phylogenetic analysis of the viral capsid proteins, 2 subgroups have been proposed under the genus Orthoreovirus, the fusogenic and the nonfusogenic subgroups. The nonfusogenic clade/subgroup consists of the prototypical MRV species and its member isolates, while the fusogenic subgroup consists of the baboon orthoreovirus (BRV), Nelson Bay virus (NBV), avian orthoreovirus (ARV), and reptilian orthoreovirus (RRV). 19,26
MRV is the only member of the nonfusogenic subgroup of the Orthoreovirus genus, and it affects a wide variety of hosts (including humans); however, it causes significant pathology in only a few species (most notably in mice). MRV usually causes mild self-limiting respiratory and gastrointestinal symptoms in children and infants. 59,60 MRV isolates have been isolated from human cases of nonbacterial fatal pneumonia, 64 hepatitis-encephalitis, 44 necrotizing encephalopathy, 57 meningitis, 37,42,43 and myocarditis. 62 Overall, it appears that MRV may at times cause serious/fatal infections in humans; however, in most cases, it causes mild self-limiting disease. In contrast to humans, MRV isolates cause severe lesions in infant laboratory mice, characterized by encephalitis, hepatitis, biliary atresia, 4,9,29 pancreatitis, myocarditis, and bronchiolitis obliterans organizing pneumonia. 10,52,53
The fusogenic subgroup of Orthoreovirus consists of BRV, ARV, NBV, and RRV. ARV mostly causes subclinical infections among birds and is associated with an important economic disease of young chickens, known as viral arthritis syndrome (tenosynovitis). 11 NBV was first isolated in 1968 from the blood of a gray-headed flying fox (Pteropus poliocephalus, a fruit bat) from the Nelson Bay in New South Wales, Australia. 24,35 So far, only 1 isolate of NBV is known. Recently, a number of Orthoreovirus isolates have been identified in human patients with a history of recent exposure to bats, in Malaysia, Hong Kong, and China. 16 –19 These isolates are genetically closely related and are strongly suspected to be of bat origin. It has been proposed that, based on the uniformity of the carrier species (Pteropine spp, bats), NBV and the other aforementioned novel isolates be put together in 1 species by renaming NBV as pteropine orthoreovirus (PRV). 19 It is very likely that these viruses are capable of species jumping from bats to humans. 19 RRV was first isolated from a python (Python regius) with hemorrhagic kidney lesions, 2,24 and subsequently, a number of RRVs have been isolated from various types of snakes, 1,25,50,67 lizards, 25,54 tortoise, 25,54 chameleon, 25 and iguana. 13,25,36 It has been suggested that in wild conditions, subclinical infections with RRVs are common. 24 Experimental infection of snakes with RRV resulted in fatal proliferative interstitial pneumonia. 47,50 At least 3 species are known among identified RRVs that are not species restricted, making it a distinct possibility that future isolates will be discovered. 13,27
As stated earlier, BRV was first isolated from an outbreak of meningitis, encephalitis, and myelitis (MEM) among baboons housed at SNPRC during 1993–1994. There is limited antigenic similarity and sequence conservation between BRV and other members of the genus Orthoreovirus. Phylogenetic analysis indicates that BRV evolved in parallel to ARV and NBV. 28 BRV-specific antibodies have been detected in serum samples of >95% tested baboons (n > 100) from 3 captive bred populations from the United States and Egypt, as well as in baboon sera preserved at the SNPRC from the early 1970s, suggesting that BRV may have coevolved with baboons in nature and is not geographically restricted. 3,51 During the previous outbreak at SNPRC, BRV antibodies were not detected in the serum samples of human caretakers who worked with infected baboons or among other nonhuman primate species, including macaques, langurs, and chimpanzees housed within the same facility, 3 indicating the species specificity of the virus.
Over the past 20 years at SNPRC, there have been sporadic cases of meningitis, encephalitis, and myelitis (MEM) among baboons. Some of the cases had confirmed bacterial etiology, but most cases were categorized as suspected viral/BRV etiology based on clinical history, gross, and histopathological lesions. Although BRV was always high on the list of suspected etiologies, it could not be confirmed due to lack of a proper diagnostic assay. Since the isolation and molecular characterization of BRV, several recent reports have studied the BRV fusion-associated small transmembrane (FAST) family of proteins, specifically p15, 20 –23,61,65,66 which is a nonstructural protein required for syncytium formation. Using polyclonal antibody specific for the C terminus of BRV-p15 protein (residues 90–140) 23 and primers specific for the S4 genome segment (which codes for BRV-p15 protein), we developed immunohistochemistry (IHC) and conventional polymerase chain reaction (PCR)–based assays to detect the presence of BRV in archived formalin-fixed tissue samples of baboons. This article presents the findings from a retrospective study of 68 cases of suspected viral/BRV-induced MEM at SNPRC from 1989–2010 that were tested by IHC and PCR for BRV, Alphavirus, and Flavivirus genera (which are known to be present in the geographical vicinity of SNPRC).
Materials and Methods
Case Selection
An internal pathology database (apath) at the SNPRC was searched for the terms encephalitis, meningoencephalitis, encephalomyelitis, myelitis, and meningitis as the morphologic diagnoses at the time of necropsy from 1989 to 2010. Cases with a determined bacterial or traumatic etiology were excluded. Only spontaneous neurological cases (n = 68) for which no specific etiology was determined were selected. The majority of these cases had histologic lesions of lymphocytic or nonsuppurative meningitis, encephalitis, and myelitis, sometimes with hemorrhage and necrosis. Four cases from the previous outbreak, from which BRV was detected by virus isolation (VI), were used as positive controls. Confirmed cases of Eastern equine encephalitis (EEE) and West Nile virus (WNV) were used as positive controls for alphavirus and flavivirus, respectively (gift from Dr Matti Kiupel, Michigan State University, Diagnostic Center for Population and Animal Health, Lansing). Four healthy baboons that were sacrificed for routine colony management that had no clinical symptoms and no gross or histological lesions of any neurological disease were selected as virus-negative control animals. The original medical records, gross necropsy reports, and histopathology reports were retrieved and reviewed for confirmation or clarification, as needed.
Primary Antibodies
Baboon Orthoreovirus
Rabbit anti-BRV primary polyclonal antibody was a generous gift from Dr Roy Duncan (Department of Microbiology and Immunology, Faculty of Medicine, Dalhousie University, Halifax, Nova Scotia, Canada). 22 The antibody is specific for the C terminus of the p15 protein (residues 90–140), which is a member of FAST protein family and helps in the dissemination of virus to the adjacent cells by syncytium formation. 22 There is no significant sequence identity between BRV FAST p15 protein and the FAST proteins identified in other members of the genus Orthoreovirus. 22
Alphavirus
Commercially available mouse anti–alphavirus monoclonal antibody clone 3581 (cat. SC-58088; Santa Cruz Biotechnology, Santa Cruz, CA) raised against alphavirus was used.
Flavivirus
Commercially available mouse anti–flavivirus monoclonal antibody clone 3571 (cat. SC-58128; Santa Cruz Biotechnology) raised against purified St Louis encephalitis virus (SLE) was used.
Immunohistochemistry
Formalin-fixed, paraffin-embedded (FFPE) nervous tissue blocks of the selected cases were obtained from the tissue archives, and 3.5-μm sections were cut and placed on propylsilane-coated glass slides. Tissue sections were paired with a primary antibody control (diluent only), virus-positive controls (BRV, EEE, and WNV), and nervous tissue sections from 4 normal baboons as negative controls. Formalin fixed, paraffin-embedded tissue sections were deparaffinized (Varistain Gemini ES; Thermo Scientific, Waltham, MA), and antigen unmasking was done by immersing in antigen retrieval solution (cat. S1669; DAKO, Glostrup, Denmark) in a decloaking chamber (cat. DC2002; Biocare, Concord, CA) at 94°C for 35 minutes followed by 90°C for 10 minutes. For BRV, tissue sections were quenched in 0.3% hydrogen peroxide (in methanol) for 10 minutes, washed twice with distilled water, and blocked in nonfat dried milk (1%) for 30 minutes followed by another round of washing with washing buffer (cat. S3006l; DAKO). Rabbit anti-reovirus anti-p15 antibody was used at 1:20 14;000 dilutions (dilution buffer, cat. AAK1; Prohisto, Columbia, SC). Tissue slides were placed in a slide box (cat. AAK1; Prohisto), and 3 ml of primary antibody solution was poured over each slide. The slides were gently rocked overnight at 4°C. For alphavirus and flavivirus, following antigen retrieval, tissue sections were incubated with mouse anti–alphavirus monoclonal antibody (1:50 dilution) or mouse anti–flavivirus monoclonal antibody (1:100 dilution) for 1 hour at room temperature in a humidified chamber. Subsequent steps were common for all 3 viruses. Following primary antibody incubation, slides were washed twice in washing buffer (cat. S3006; DAKO), incubated with a horseradish peroxidase link (20 minutes; DAKO), and then washed twice in washing buffer. Next, slides were incubated with horseradish peroxidase enzyme (20 min; DAKO) and washed twice, and then freshly prepared 3,3′-diaminobenzidine tetrahydrochloride (DAB) chromogen substrate solution (DAKO) was added to develop the color. Finally, slides were washed, counterstained in Mayer’s hematoxylin using a standard protocol, cover-slipped, and examined microscopically by 2 board-certified pathologists (MAO and EJD) in a blind manner. Sections were identified as positive or negative based on the presence of discrete staining.
PCR
For BRV
A PCR-based assay was designed as a test to detect BRV genome in formalin-fixed tissues. Twenty-four cases were selected for this assay, which included 4 VI-positive controls, 4 negative controls, and 16 (of a total 68) unknown/suspect cases. Among the suspect cases, all cases that tested negative for BRV by IHC (n = 9) and 7 cases that tested positive for BRV by IHC were selected. Total RNA was extracted from FFPE brain tissue sections using the RNeasy FFPE kit (QIAGEN, Valencia, CA). Reverse transcription (RT) was done using the RETROscript kit (Life Technologies, Carlsbad, CA). For amplification of viral DNA, PCR was carried out using primer sequences based on the BRV genome segment S4 (complete sequence; GenBank accession AF406787). The primer sequences are as follows: Forward primer: 5′-ATG GGT CAA AGA CAT TCA ATA G-3′ Reverse primer: 5′-TCA AAC GTT GAT ACT TCC ATC TGG-3′
Touchdown PCR conditions were used for the PCR. Briefly, following an initial denaturation step at 94°C for 2 minutes, the annealing temperature was set to 59°C for 2 cycles and thereafter decreased by 1 degree for every 2 cycles until the temperature reached 52°C. At those conditions, the samples were run for an additional 20 cycles. The PCR product was loaded on 1% agarose gels and electrophoresis was performed. They were then stained with ethidium bromide, visualized under UV illumination, and photo-documented. A 430–base pair (bp) band on a gel was determined as positive (Fig. 1). All reactions were run with positive and negative controls. PCR products from 1 sample were cloned into the pCR2.1-TOPO vector (Life Technologies, Carlsbad, CA) and sequenced to confirm if the PCR products were indeed from BRV (Table 1). In separate reactions, a 244-bp sequence of the 16S ribosomal RNA (rRNA) gene was amplified from the baboon samples as a DNA-positive control that would make it possible to distinguish between PCR failure and truly negative results using the primer set L2513 and H2714 (data not shown). 45
Sequence of the Amplified Polymerase Chain Reaction Product of the S4 Genome Segment of Baboon Orthoreovirus.

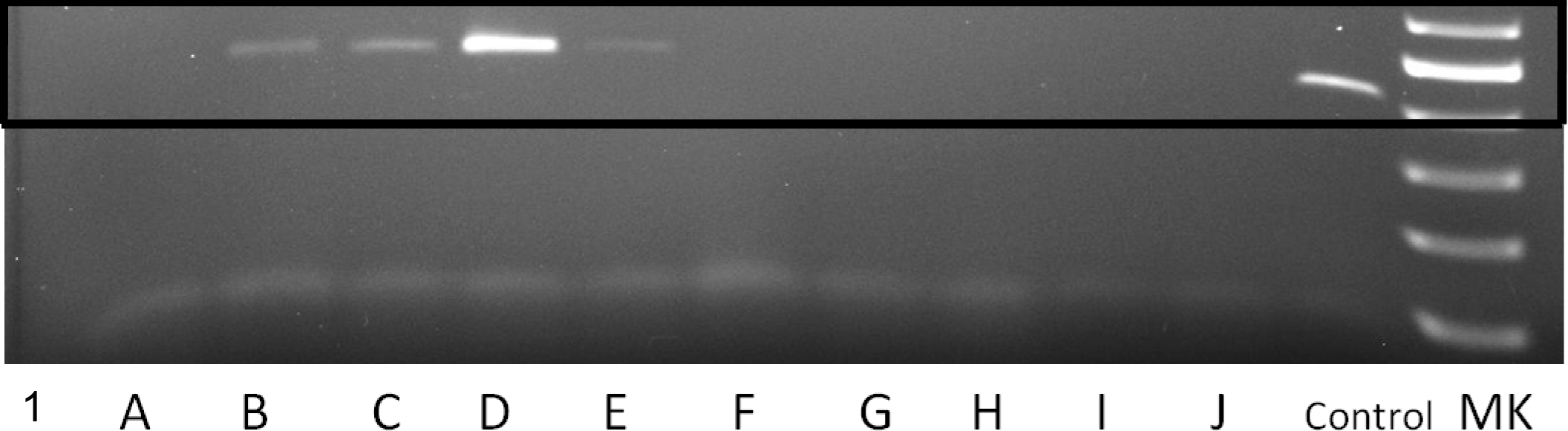
Agar gel electrophoresis image of the amplified sequence of the S4 genome segment of baboon orthoreovirus. A 430–base pair band on the gel was considered positive. Columns with case numbers: A, 23; B, 63; C, 44; D, virus isolation positive control; E, 32; F, 58; G, 20; H, 1; I, 27; J, 61; Control, 49 (sequenced); MK, size marker.
For alphavirus and flavivirus
The confirmatory tests for alphavirus (Western equine encephalitis [WEE]) and flavivirus (WNV and SLE) were done by a quantitative RT-PCR (qRT-PCR) assay at a diagnostic laboratory as per established protocols. 46,48,49
Results
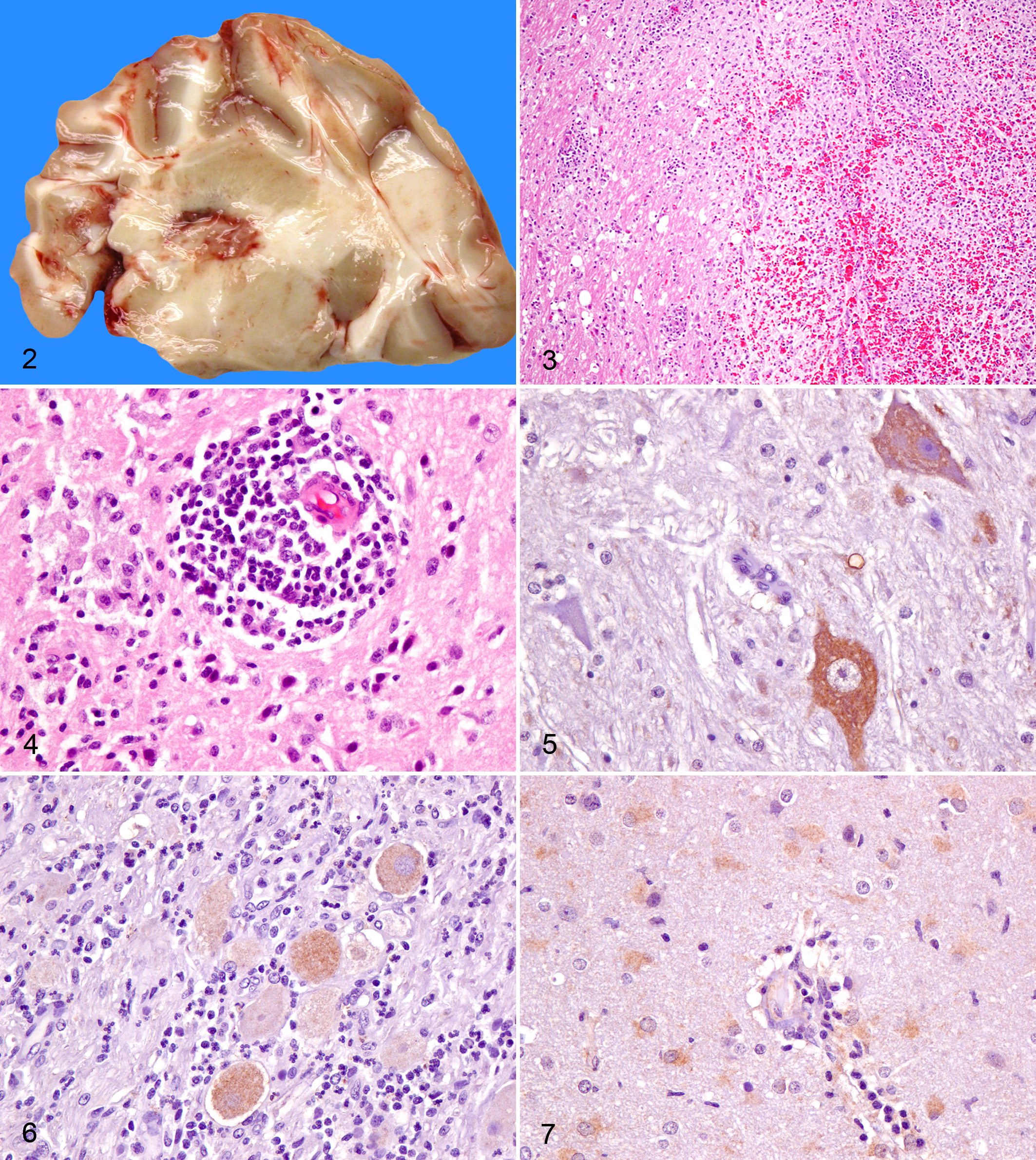
Table 2 lists all of the cases, along with the signalment, year of necropsy, reported relevant clinical history, distribution of lesions in the central nervous system (CNS), histopathologic changes, BRV immunohistochemistry (IHC), and PCR results. Of the 68 suspect baboons, the average age at the time of necropsy was 9.27 years (range, 0.01–25.33 years), and 62% (42 baboons) were younger than 10 years. The male to female ratio was 0.94. Few cases (21/68) had gross lesions, but when present, there were typically discrete foci of hemorrhage and necrosis, randomly scattered throughout the brain (Fig. 2) and spinal cord. The typical microscopic lesions of perivascular lymphocytic nonsuppurative encephalitis and multifocal necrosis and hemorrhage were similar to those as previously reported 51 (Figs. 3, 4); a few cases had primarily suppurative inflammation. Table 3 summarizes the IHC, PCR, and qRT-PCR testing results.
Clinical, Pathology, IHC, and PCR Results of All Cases Tested for Baboon Orthoreovirus.
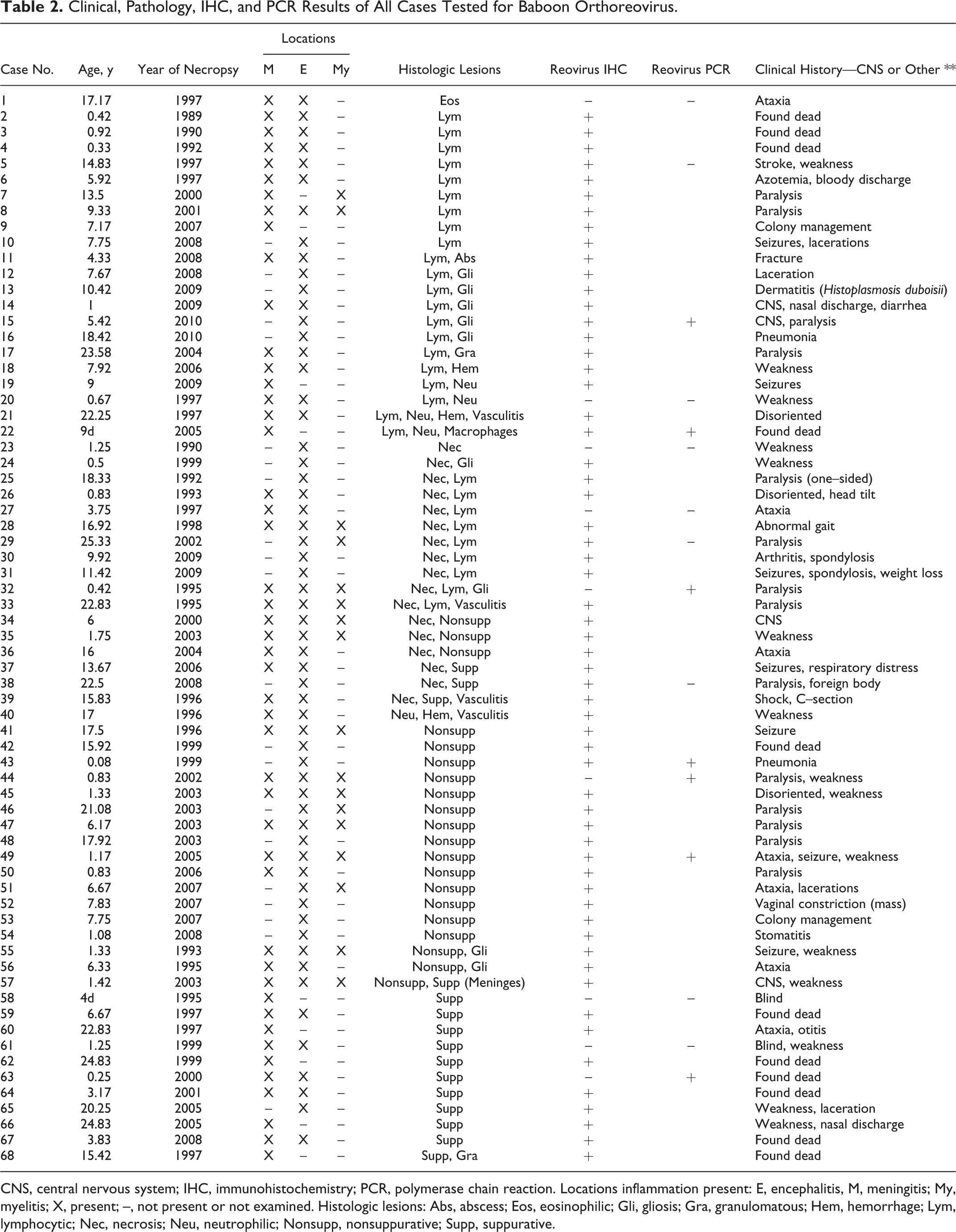
CNS, central nervous system; IHC, immunohistochemistry; PCR, polymerase chain reaction. Locations inflammation present: E, encephalitis, M, meningitis; My, myelitis; X, present; –, not present or not examined. Histologic lesions: Abs, abscess; Eos, eosinophilic; Gli, gliosis; Gra, granulomatous; Hem, hemorrhage; Lym, lymphocytic; Nec, necrosis; Neu, neutrophilic; Nonsupp, nonsuppurative; Supp, suppurative.
Summary of Immunohistochemistry and PCR-Based Results.
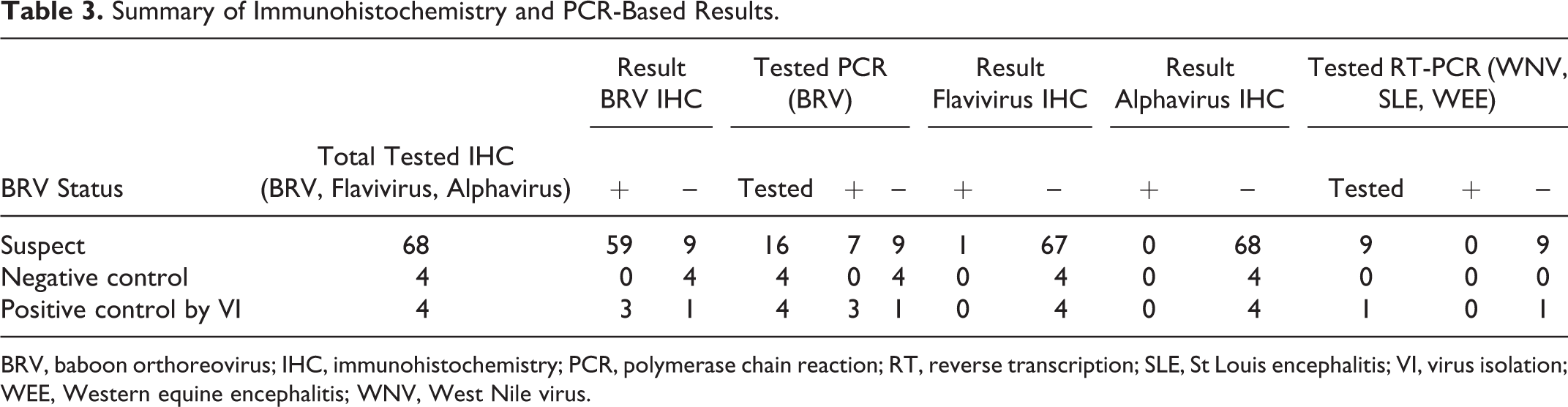
BRV, baboon orthoreovirus; IHC, immunohistochemistry; PCR, polymerase chain reaction; RT, reverse transcription; SLE, St Louis encephalitis; VI, virus isolation; WEE, Western equine encephalitis; WNV, West Nile virus.
Immunohistochemistry for BRV (Figs. 5–7) demonstrated diffuse to granular cytoplasmic labeling of neurons multifocally within the cerebral cortex, brainstem (Fig. 5), ganglia (Fig. 6), and midbrain; labeling of small neurons in the cerebellar Purkinje cell layer; and few to many glial cells in the cerebrum (Fig. 7), with fewer numbers of labeled glial cells in the cerebellum, brainstem, and spinal cord. The type and number of positive cells varied between cases. IHC indicated that of 68 unknown cases, 59 were positive for BRV; 1 was negative for BRV but positive for flavivirus. The other 8 tested negative by IHC for BRV, alphavirus, and flavivirus. Three of 4 BRV VI-positive controls were positive by IHC; 1 BRV VI-positive control tested negative by IHC for BRV and also for alphavirus and flavivirus. The 10 cases (9 unknown and 1 VI-positive control), which tested negative by IHC for BRV, all tested negative for WNV, WEE, and SLE viruses by qRT-PCR. No mixed infections were detected.
Twenty-four cases were tested for BRV using PCR, which included 4 VI-positive controls, 4 negative controls, and 16 unknown cases (9 BRV-negative cases and 7 BRV-positive cases, by IHC). All 4 negative controls tested negative by PCR, and 3 of 4 VI-positive controls were positive by PCR. One VI-positive control that was positive by IHC tested negative, whereas 1 VI-positive control that was negative for BRV by IHC was positive by PCR. Of 16 unknown cases, 3 (of 9) cases that were negative for BRV by IHC tested positive by PCR, including 1 case that was actually positive for flavivirus by IHC but negative for WNV, SLE, and EEE by qRT-PCR. Three (of 7) cases that were positive for BRV by IHC tested negative by PCR. The PCR products (Table 1) amplified using BRV-specific primers matched with 99% sequence identity to a previously published BRV S4 genome segment sequence (Genbank accession: AF406787).
Discussion
Using the recently developed FAST p15 polyclonal antibody, an IHC methodology was developed to detect BRV in archived formalin-fixed baboon tissues, revealing that 59 of 68 cases (∼87%) of suspected viral MEM at the SNPRC were positive for BRV by IHC. A second assay (PCR) was also developed to detect BRV in formalin-fixed tissues. Of 16 unknown cases, 3 cases that were negative for BRV by IHC were positive by PCR, whereas 3 cases that were positive by IHC tested negative by PCR. One of the VI-positive controls that tested negative by IHC was positive by PCR, while one other VI-positive control that was positive by IHC was negative by PCR. All the negative controls tested negative by both IHC and PCR. If virus isolation is considered the gold standard, then on the basis of limited numbers of samples, both IHC and PCR have a sensitivity of 75% and a specificity of 100%. The main objective of this study was to demonstrate that most cases of nonsuppurative MEM in baboons are associated with BRV infection. The results from this study suggest that BRV is associated with nonsuppurative viral MEM in baboons at the SNPRC and that the presumptive diagnosis of BRV infection can be made by the characteristic lesions, pending further confirmation. The overall prevalence of CNS diseases in baboons is extremely low. In a recent publication that documented 10,883 macroscopic or microscopic morphologic diagnoses in 4297 baboons at SNPRC over a 20-year period, nervous system lesions accounted for only 4.21%, which included encephalitis (n = 57), meningitis (n = 27), meningoencephalitis (n = 17), scoliosis (n = 7), kyphosis (n = 6), myelitis (n = 5), encephalomyelitis (n = 3), porencephaly (n = 2), meningioma (n = 1), and glioblastoma (n = 1). 14 This further emphasizes the importance of BRV among baboons.
BRV is capable of being highly pathogenic, yet the presence of BRV-specific antibodies in asymptomatic animals suggests that the virus is often subclinical in baboons. 28 Over the past 20 years at SNPRC, sporadic cases of MEM have been suspected to be BRV induced but could not be confirmed. These cases were more or less evenly distributed over the period (Table 2). Many of these suspected cases had clinical histories suggestive of CNS lesions, such as paralysis, ataxia, and seizures (Table 2). Gross lesions indicative of CNS disease were not present in most cases but, when present, were typically hemorrhage and necrosis; this is in contrast to a previous report that reported no gross lesions in infected baboons. 51 Most cases had nonsuppurative or lymphocytic meningoencephalitis, with occasional myelitis. It is possible that myelitis is more common than reported in Table 2, because the spinal cord was not always taken for histology. In contrast to these more typical changes, a few cases had primarily suppurative inflammation, which is important because these would be more likely to be presumed bacterial in origin and overlooked as potential BRV cases. Vasculitis was reported as a significant histologic lesion in a few cases and may account for the hemorrhage and necrosis seen histologically and on gross examination. Myocarditis was the most common non-CNS inflammatory lesion, occurring in 9 animals, followed by pneumonia in 6 animals; these are common lesions in this baboon colony, and the low number of cases suggests they are not major components of BRV infection and may be unrelated lesions. 14
Besides BRV, other potential etiological agents of viral MEM in baboons exist, including members of the genera Flavivirus (WNV, SLE), Alphavirus (Eastern equine encephalitis [EEE], Venezuelan equine encephalitis [VEE], and WEE), Herpesvirus (Herpesvirus papio 2 [HVP2], simian agent 8 [SA8]), and Cardiovirus (encephalomyocarditis virus, EMC70 strain). WNV-specific antibodies were isolated from 36% of baboons tested, housed at Tulane National Primate Research Center (TNPRC), following an outbreak of WNV infection in the neighboring human population, 58 but clinical signs or histopathological lesions of encephalitis were not reported among the baboon population. 58 Filatenkov et al 34 reported that virulent strains of VEE can induce clinical symptoms of MEM in baboons. Brack et al 15 reported experimentally induced encephalitis in Kenya baboons due to herpesvirus SA8. Dzhikidze et al 30 reported the occurrence of encephalomyocarditis in hamadryas baboons due to the EMC70 strain of virus; however, during an outbreak of EMCV in the SNPRC baboon colony, the findings were all related to the cardiovascular system, and no lesions were found in the brain. 40 Isoun et al 41 observed hemorrhagic encephalitis in a wild baboon; however, the etiology was not confirmed. Benveniste et al 12 did not observe any clinical signs of encephalitis in baboons due to experimental inoculation of simian immunodeficiency virus.
Out of these viruses, only a few have epidemiological data suggestive of their prevalence in the state of Texas. WNV is relatively more frequent in both humans and animals in the geographical vicinity of SNPRC. 8,63 WNV affected Texas beginning in 2002, and although 34 baboons in this study were necropsied between 2002 and 2010, only 1 was positive on IHC for flavivirus (but was negative for WNV and SLE qRT-PCR) and, in turn, was confirmed positive for BRV by PCR. SLE is very prevalent in Texas; from 1964 to 2010, among all 48 contiguous states, the highest number of confirmed cases has been reported from Texas (total 1021; high incidence in 1964–1968 followed by decreased annual incidences). 7,39 Although there are only sporadic reports of EEE among horses in Texas, 38 it is much more prevalent among humans in the neighboring state of Louisiana (157 confirmed cases in humans from 2003–2011) compared with Texas (43 cases from 2003–2011). 6 VEE is unlikely given that no recent confirmed cases have been reported in humans since the outbreak among humans and equines in 1971, 70 although, due to the close proximity with Mexico and frequent cross-border transport of equids and reports of occurrence of VEE in Mexico, it is always a concern. 33,56 There are not much epidemiological data to suggest the prevalence of WEE in Texas (no confirmed cases in humans from 2000–2011). 5 HVP2 has been shown to be common in both captive and wild-caught baboon populations. 31,32 HVP2, originally reported as SA8, has been shown to be present in our colony; however, lesions were mostly restricted to mucocutaneous herpetic lesions and not associated with MEM. 55
Our testing of other potential causes of MEM was limited to the use of a pan-flavivirus antibody and a pan-alphavirus antibody on all cases, as well as qRT-PCR for WNV, SLE, and WEE on the cases that tested negative for BRV by IHC (all of them tested negative by qRT-PCR). One case tested positive for flavivirus by IHC, but we were unable to determine if this was a true positive or a false positive, as the most likely flaviviruses (WNV and SLE) were excluded by the qRT-PCR testing. Overall, in this report, 68 suspected cases of BRV-induced MEM over a period of 20 years in the captive baboon population housed at SNPRC were tested using a novel IHC-based assay to identify BRV in FFPE tissues. The results indicate that 87% of the suspected cases were positive for BRV by IHC. A conventional PCR-based assay was designed as a confirmatory test to detect BRV, but both IHC and PCR assays failed to pick 1 of 4 VI-positive controls (1 false negative; sensitivity, 75%). Thus, neither the IHC- nor the PCR-based assay seems to be perfect, and both have some limitations. However, a negative result does not completely rule out infection. Also, in retrospective studies such as the present study, variations in fixation conditions can result in false negatives. This study has 2 major implications; first, it presents 2 diagnostic assays (IHC and PCR) that (with some reservations) can be used for the detection of BRV infection in nervous tissues. Second, the results from this study suggest that BRV is associated with the majority of MEM in baboons, and BRV should be considered an important differential for cases of nonsuppurative MEM in baboons. To prove whether BRV is the causative agent or an innocent bystander in these lesions would be a subject of future investigations.
Footnotes
Acknowledgements
We acknowledge Michaelle Hohmann for her assistance with the immunohistochemistry. We also acknowledge Dr Jim Cooley (Mississippi State University) for critically reading the manuscript. We sincerely thank Dr Roy Duncan (Dalhousie University) for the generous gift of p15 antibody, as well as Dr Matti Kiupel at Michigan State University for providing positive controls of alphavirus (EEE) and flavivirus (WNV). We also thank Marie Silva, Jesse Martinez, and Jacob Martinez for pathology support and the veterinary clinical staff at the SNPRC.
Declaration of Conflicting Interests
The authors declare no conflict of interests with respect to research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship and/or publication of this article: This study was supported in part by NCRR grant P51 RR013986 to the Southwest National Primate Research Center.
