Abstract
The binding of influenza A viruses to epithelial cells in the respiratory tract of mammals is a key step in the infection process. Therefore, direct assessment of virus-host cell interaction using virus histochemistry (VH) will enhance our understanding of the pathogenesis of these new viruses. For this study, the authors selected viruses that represented the 4 main genetic clusters of North American swine H1 (SwH1) viruses, along with A/California/04/2009 H1N1 and a vaccine strain for the positive controls, and the virus label, fluorescein isothiocyanate (FITC), for the negative control. A group of 5 viruses containing a 2–amino acid insertion adjacent to the binding site of the hemagglutinin protein and their presumed ancestral viruses were also examined for changes in binding patterns. Viruses were bound to formalin-fixed paraffin-embedded, 6-week-old (6w) and adult pig tissues. Qualitative VH scores per respiratory zone ranged from + to +++, with bronchioles having the highest and most consistent scores, regardless of animal age. For the 6w bronchioles, a quantitative VH score was calculated using digital images of 5 bronchioles per tissue section using image analysis software. Significant differences in attachment were found among the SwH1 viruses (P < .0001) and among the ancestral and insertion viruses (P < .0001). These results provide new insights on virus binding to porcine respiratory epithelial cells and the usefulness of morphometric scores. The results also highlight limitations of in vitro techniques, including VH for predicting virulence and host range.
Influenza is an acute respiratory disease that occurs year-round in swine and is caused by influenza A virus (IAV) subtypes H1N1, H1N2, and H3N2. Although most IAV infections in swine are mild with clinical recovery usually occurring within 7 to 10 days of infection, the respiratory lesions can vary from mild tracheobronchitis to severe bronchoalveolar pneumonia. 29 To further characterize the process of infection with different IAV strains, we need to examine the interaction of various viruses with cells that they infect.
It has been generally accepted that avian and equine IAV have a host cell receptor preference for α2,3-linked sialic acids (SAα2,3), and human and swine have a host cell receptor preference for α2,6-linked sialic acids (SAα2,6). The distribution of these receptors in the human and swine respiratory tract has been examined using lectin histochemistry (LH) as an indirect measurement of IAV virus attachment. 15,19,23,37 LH uses the SAα2,3 binding preference of Maackia amurensis agglutinin (MAA) and the SAα2,6 binding preference of Sambucus nigra agglutinin (SNA) to identify the IAV receptors in formalin-fixed or frozen tissues. In LH, a virus-free system, the amount of MAA binding is a proxy measurement of SAα2–3 receptors and SNA binding indicates the presence of SAα2–6 receptors.
Some of the limitations of the LH technique are the binding of MAA-1 to nonsialic residues in contrast to poor recognition of alternative glycans by MAA-2 and cross-reactive binding of SNA to some SAα2–3 receptors. 19,20 These limitations have led to several discrepancies in LH studies, including the distribution of influenza receptors, the specific cell types involved, and the actual cells that were infected in culture compared with those predicted by LH in the human airway 15,19,20,23,37 and in the swine airway. 9,25,28
Another method that is used to characterize IAV virus interaction with host cells is virus histochemistry (VH). In contrast to the indirect methods of LH, VH allows for direct examination of the pattern of virus attachment (PVA) to tissues. VH has been used to visualize the PVA for avian-origin IAV H5N1, H5N9, and H6N1 and human-origin IAV H1N1 and H3N2 in paraffin-embedded respiratory tissues of human, ferret, mouse, macaque, cat, dog, and pig origin. 30,31 This technique has yet to be applied to swine influenza viruses and respiratory tissues of pig origin.
To provide new insights regarding the pathogenesis of IAV in swine, the purpose of this study was to examine the binding patterns of swine-origin IAV to swine respiratory tissues using viruses that have been definitively characterized by prior genetic and antigenic analyses and with known pathogenicity in vivo.
Materials and Methods
Viruses
Well-characterized viruses demonstrated to be virulent in previous studies were selected from the representative 4 genetic clusters (alpha, beta, gamma, delta) of North American swine H1 viruses. 14,15,32 –34 These included SwH1α A/swine/Illinois/02450/2008 (IL08), SwH1β A/swine/Iowa/00239/2004 (IA04), SwH1γ A/swine/Ohio/511445/2007 (OH07), SwH1γ A/swine/Kansas/77778/2007(KS07), SwH1δ1 A/swine/Texas/01976/2008 (TX08), and SwH1δ1 A/swine/Illinois/07003243/2007 (IL07) (Table 1). In addition, a group of H1N1 viruses isolated from pigs within a single farm system over a 2-year period that contained a 2–amino acid (AA) insertion in the Sb antigenic site of the hemagglutinin gene 2 (SG443, SG444, SG1094, SG1096, SG1150) and their presumed ancestral viruses without the insertion (SG445, SG1097a) were selected. 6 A/California/04/2009 H1N1 (pH1N1) and A/swine/Minnesota/001482/2007 H1N2 (SG239) 5 were used as positive controls, and the label, fluorescein isothiocyanate (FITC), was used as a negative control. OH07 and pH1N1 were kindly provided by the US Department of Agriculture, Agricultural Research Service, and KS07 was generously provided by Kansas State Veterinary Diagnostic Laboratory. All remaining viruses were retrieved from the influenza virus archive at the University of Minnesota Veterinary Diagnostic Laboratory (UMVDL).
Table of Viruses Selected for Virus Histochemistry
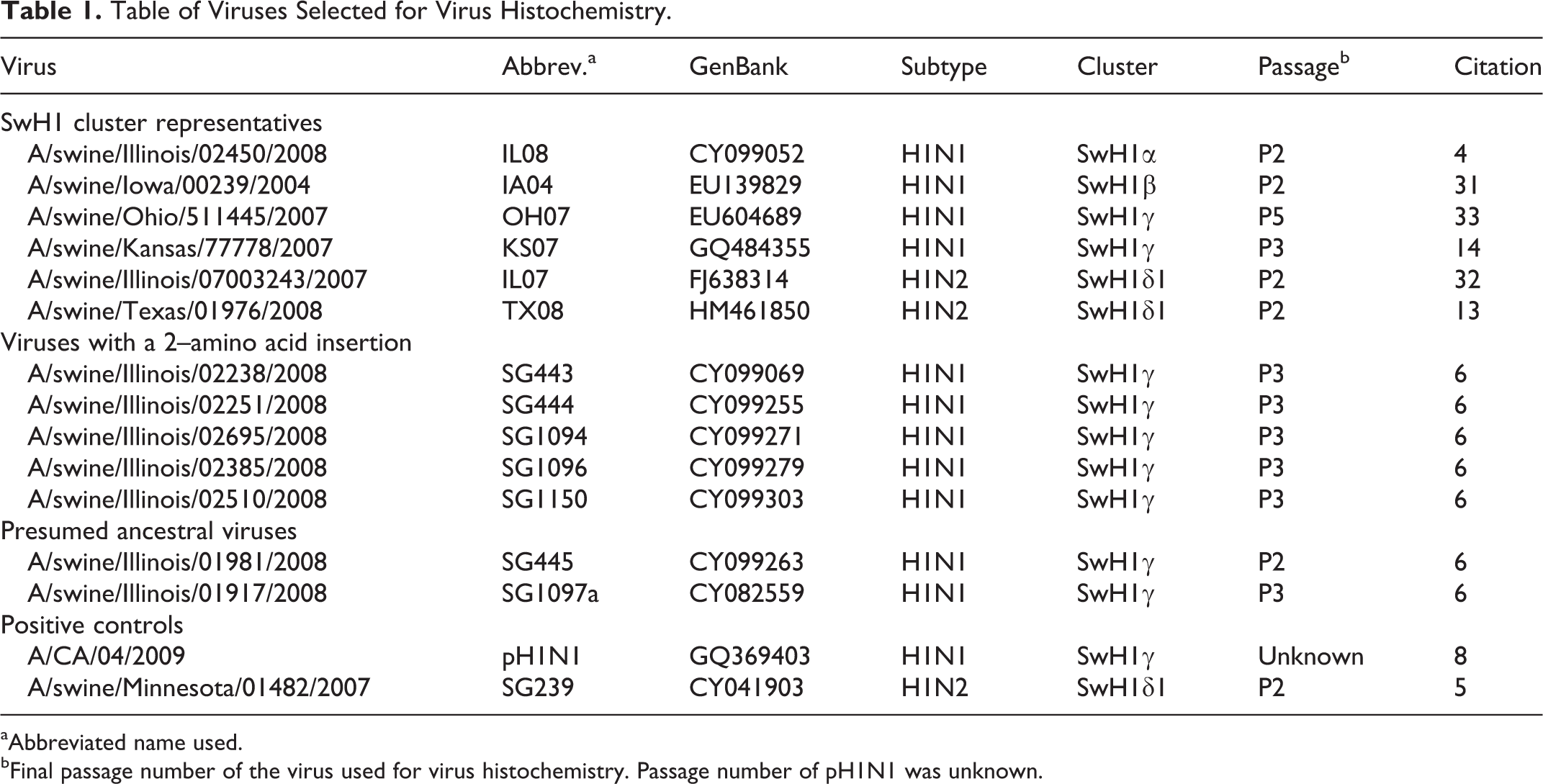
aAbbreviated name used.
bFinal passage number of the virus used for virus histochemistry. Passage number of pH1N1 was unknown.
Virus Sequence Analysis
Hemagglutination (HA) gene sequences for the viruses were available in GenBank (Table 1). Molecular evolutionary analyses were conducted using MEGA 5 with sequence alignments using Clustal W. 26,27 The evolutionary history was inferred using the neighbor-joining method, and evolutionary distances were computed using the maximum composite likelihood method. 7,22,26,27 The 3-dimensional structure of hemagglutinin protein was artistically rendered using the educational version of the PyMOL Molecular Graphics System, Version 1.5.0.4 (Schrödinger, LLC, Cambridge, MA) and A/South Carolina/1/18 (PDB code: 1RUZ) from the online protein database (www.rcsb.org/pdb).
Virus Propagation and Label Attachment
Viruses were propagated on monolayers of Madin-Darby canine kidney (MDCK) cells grown in flasks with Eagle’s minimum essential media containing 4% bovine serum albumin, trypsin, and antibiotics. 16 All cultures were incubated at 37°C under a 5% CO2 atmosphere and examined daily for up to 3 days under an inverted light microscope to observe cytopathic effects (CPEs). After 3 freeze-thaw cycles, the supernatant was harvested and cleared by centrifugation for 10 minutes at 2272 × g. The cleared supernatant was then centrifuged at 4°C for 2 hours at 85 000 × g, and the virus pellet was resuspended in 2 ml of phosphate-buffered saline (PBS). The resuspended virus was inactivated by dialysis against 1% formalin overnight at 4°C followed by 4 cycles of dialysis against PBS for 2 hours at 25°C. The virus was then inoculated in MDCK cells to confirm inactivation. The label was prepared by mixing 0.1 mg/ml FITC (Sigma, St. Louis, MO) in 0.5 mol/L bicarbonate buffer at pH 9.6 (Sigma). Equal volumes (1:1) of the inactivated virus and label solution were constantly stirred for 1 hour at 25°C followed by 4 cycles of dialysis against PBS for 2 hours at 25°C. The HA titers of all viruses were determined using turkey red blood cells 10 on propagated virus, concentrated virus before and after inactivation, and final FITC bound product. The final products were diluted to contain 50 to 100 hemagglutinating units/50 μl.
Respiratory Tract Tissues of Pigs
Formalin-fixed and paraffin-embedded (FFPE) respiratory tissues (nasal turbinate, trachea, and lung) were selected from the UMVDL tissue block archive. There was no macroscopic or microscopic evidence of respiratory disease, and all animals tested negative by routine diagnostic polymerase chain reaction (PCR) tests on the lung at UMVDL for influenza A virus, porcine respiratory and reproductive syndrome virus, porcine circovirus type 2, and Mycoplasma hyopneumoniae at the time of necropsy. Blood collected at necropsy from the 6-week-old (6w) pigs tested negative for influenza antibodies using hemagglutination inhibition titers against IA04, A/swine/North Carolina/2001H1N1, and A/swine/Missouri/069/2005 H3N2. Blood was not collected from the adult pig. One 6w pig and one 13-month-old sow (adult) were selected to test all of the viruses, and 3 additional 6w pigs were selected to compare variations in binding among individual pigs using positive (SG239) and negative controls. In addition, since the slide incubation container could only hold 20 slides at a time, the variability between staining runs was compared for the positive control (SG239) results.
Virus Histochemistry
The FFPE respiratory tissues were microsectioned (4 μm) and attached to adhesive glass slides. The tissues were deparaffinized with CitriSolv (Fisher Scientific, Waltham, MA) and hydrated using graded alcohols. After blocking with TNB buffer (PerkinElmer, Waltham, MA), the FITC-labeled influenza viruses were incubated on the tissues overnight at 4°C in a humidified staining chamber (StainTray; Simport Scientific, Beloeil, Québec, Canada). The virus label, FITC, was then detected using an anti-FITC antibody conjugated with horseradish peroxidase (Dako, Glostrup, Denmark), and this signal was then amplified using the tyramide signal amplification (TSA) biotin system (PerkinElmer). The amplified signal was then revealed using 3-amino-9-ethyl-carbazole (AEC; Dako), and the tissues were counterstained with Mayer’s hematoxylin (Dako) prior to cover slipping using glycergel mounting medium (Dako). The PVA to the apical surface of the epithelial cells was scored as follows: (–) no attachment, (+) rare or few cells, (++) moderate number of cells, and (+++) abundant or many cells.
Image and Statistical Analyses
To quantify the level of virus attachment in the 6w bronchiole, 5 bronchioles were selected per tissue section (1 section of lung per each of 4 pigs). The images were captured at 40× magnification with an Olympus BX45 and an Olympus DP70 digital camera (Olympus America, Inc., Center Valley, PA). Digital images were analyzed with Image-Pro Plus 6.2 image analysis software (Media Cybernetics, Silver Springs, MD) as previously described. 1 Thresholds (n = 3) were determined by using appropriate hues to identify positive staining for the red granular staining generated by the AEC chromogen. The total area (μm2) and circumference of the area analyzed per bronchiole were recorded. Using the total area of staining per bronchiole, the average log-transformed percentage of area containing granular red staining was calculated for the 3 threshold levels and evaluated by analysis of variance (ANOVA) using SAS (SAS Institute, Cary, NC) with post hoc Tukey test. A P value of <.05 was considered to indicate a statistically significant difference between groups.
Results
Virus Sequence Analysis
The inferred evolutionary history of the selected viruses is shown in the phylogenetic tree of the HA sequences, and the 4 SwH1genetic clusters are labeled (Fig. 1). The ancestral and insertion viruses fall into the SwH1γ cluster with pH1N1 but are divergent from pH1N1. Basic Local Alignment Search Tool (BLAST) search of the Influenza Sequence Database (www.fludb.org) revealed that the HA sequences of the ancestral and insertion viruses were most similar to OH07 and KS07. Using pairwise analysis using the maximum composite likelihood model for the first 1688 nucleotides, OH07 and KS07 were 99.9% similar to each other with 100% amino acid similarity. The ancestral viruses had 98.5% to 98.6% and the insertion viruses had 97.5% to 98.3% nucleotide similarity compared with OH07 and KS07. Pairwise analysis of the first 292 AAs of the ancestral and insertion viruses compared with OH07/KS07 revealed 16 to 18 and 18 to 28 AA differences, respectively.
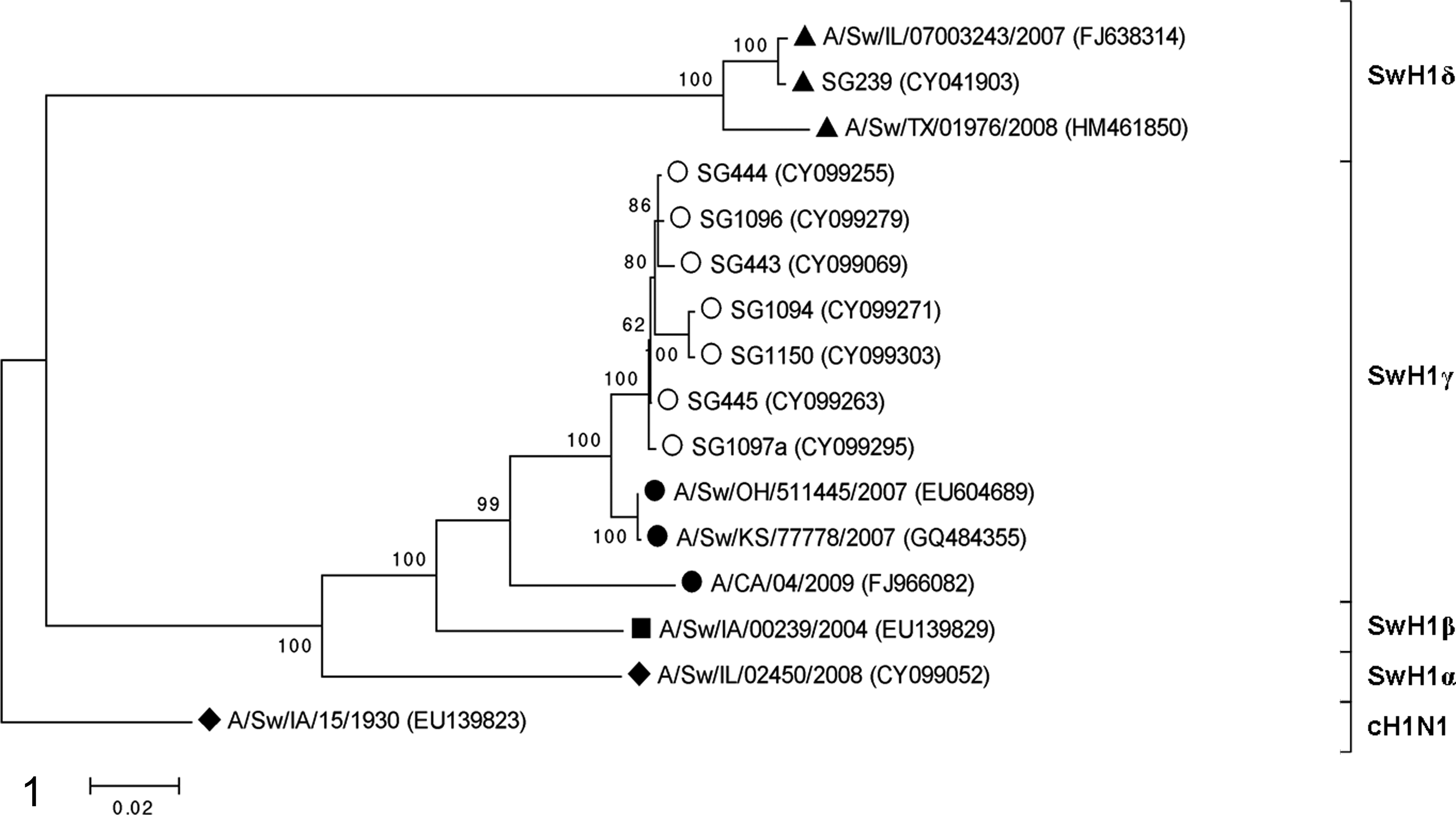
Virus Histochemistry
The HA titers of the propagated viruses 1:25 to 1:1600 and concentrating the virus resulted in titers of >1:3200 for all except pH1N1 and IA04, which had titers of 1:1600. After inactivation, the titers of the concentrated viruses dropped variably to between 1:50 and 1:3200 (data not shown). There was little to no drop in HA titer between the inactivated virus and the final FITC bound product (data not shown). The final products were confirmed to have titers between 1:50 and 1:100 after dilution.
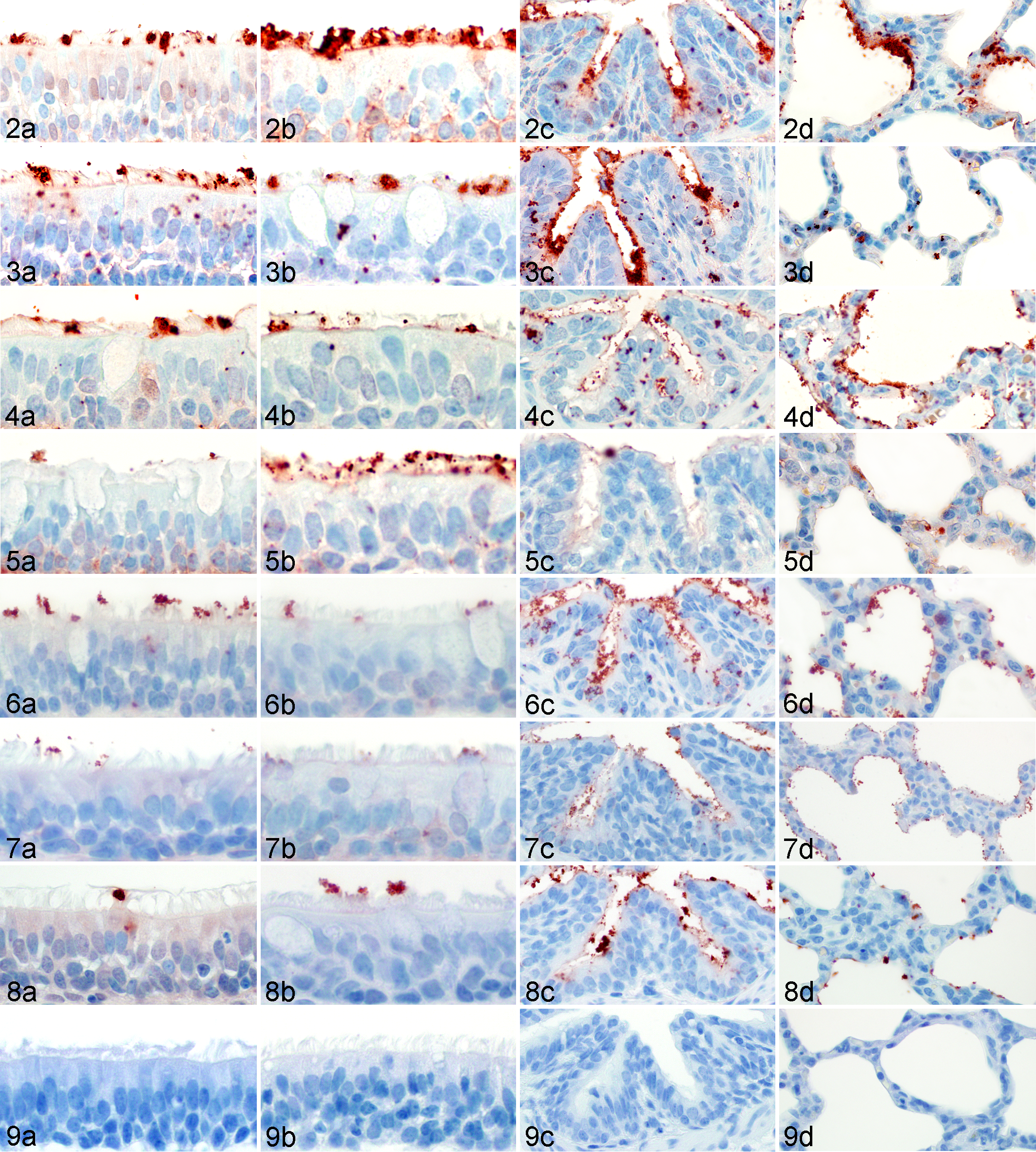
The VH scores per respiratory zone (nasal turbinate, trachea, bronchus, bronchiole, and alveoli) ranged from + to +++ for all viruses. With some exceptions, virus attachment appeared to be most abundant in the bronchioles, regardless of virus or animal age. Both positive control viruses had mild to moderate binding (+ to ++) throughout the upper and lower respiratory tracts, and there was no binding of the FITC alone for the negative controls (Table 2; Figs. 2 –9). The percentage of area containing granular red staining (AEC chromogen) was calculated for each of the 5 bronchioles using the cumulative area of the 3 thresholds divided by the total circular area analyzed. The results of the morphometric analysis are reported as the mean percent area of the 5 bronchioles ± standard deviation.
Distribution and Intensity of Virus Attachment to Respiratory Tract Detected by the Red Chromogen Signal
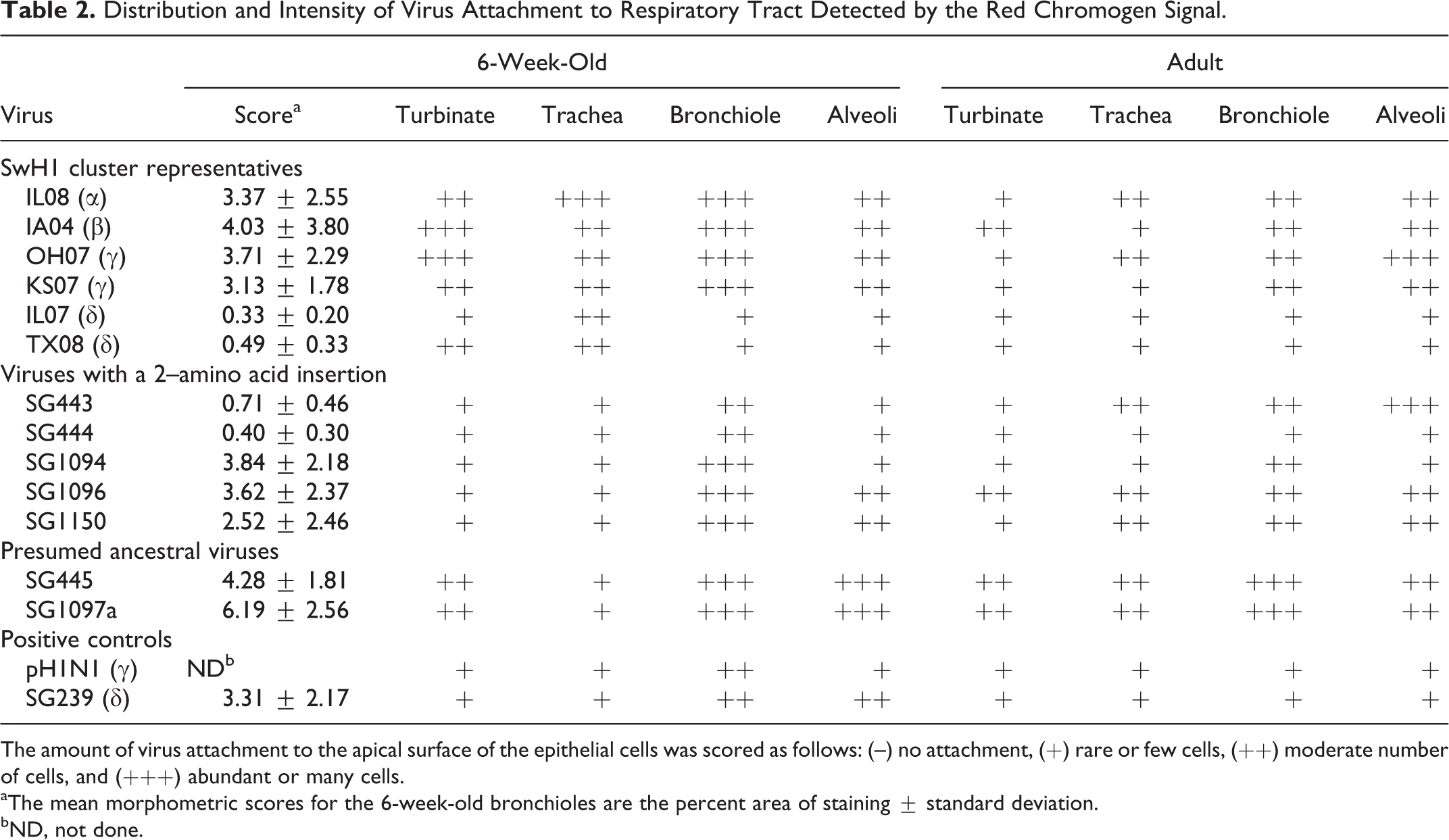
The amount of virus attachment to the apical surface of the epithelial cells was scored as follows: (–) no attachment, (+) rare or few cells, (++) moderate number of cells, and (+++) abundant or many cells.
aThe mean morphometric scores for the 6-week-old bronchioles are the percent area of staining ± standard deviation.
bND, not done.
Variation Between Staining Runs
Due to the space constraints of the slide incubation chamber, which holds a maximum of 20 slides, the VH technique was performed in several different staining runs. The variation in the PVA for the positive control, SG239, appeared to be minimal by the naked eye. However, the morphometric scores for SG239 used on the lung tissue of the same 6w pig in 4 different slide runs had mean percent staining in the bronchioles of 2.61% ± 1.01%, 3.82% ± 1.28%, 5.43% ± 2.76%, and 1.38% ± 0.93%. A statistically significant difference was detected between the last 2 means (P = .0038), indicating a limitation in comparing VH results from the different staining runs based on the analysis of 5 bronchioles per lung section. For the remaining comparisons, only results from the same staining run were compared.
Variation Among Individual Animals
Initial assessment of the variation in the amount of viral binding using the control virus SG239 was measured in 4 different 6w pigs with similar genetic background. The PVA in the upper and lower respiratory tract appeared to be the same for all 4 pigs (data not shown), and the morphometric scores were 2.61% ± 1.01%, 3.72% ± 1.62%, 3.70% ± 2.07%, and 2.27% ± 1.41%. No difference was detected between the means (P = .2789), indicating that VH results from these four 6w pigs in the same staining run were comparable.
Swine H1 Cluster Representatives
The SwH1α, β, and γ viruses had similar patterns of virus attachment with abundant binding in both the upper and lower respiratory tracts. The SwH1δ viruses had minimal binding throughout the respiratory tract, regardless of animal age. One of the SwH1γ viruses, KS07, had less binding compared with the genetically similar virus, OH07. One notable difference in the patterns of virus attachment for the 2 age groups was found in the nasal turbinates where there was more abundant binding to the 6w turbinate compared with the adult turbinate (Table 2; Figs. 2 –4). The computerized morphometric scores of the 6w bronchioles revealed more binding for the SwH1α, β, and γ viruses (>3%) compared with the SwH1δ viruses (<1%) that was demonstrated to be different in the post hoc Tukey test (P < .0001).
Viruses With and Without Insertion
The patterns of attachment for viruses that had the 2-AA insertion were compared with those of their presumed ancestors (viruses without the insertion). The most abundant virus attachment to the bronchiolar epithelium of the 6w and adult pigs occurred with the ancestral viruses. Within the pulmonary parenchyma, there was abundant bronchiolar attachment of the ancestral virus for the 6w and adult pigs, and there was moderate to abundant attachment to the alveoli. Comparatively, there was less abundant binding of the insertion viruses to the bronchioles and alveoli for the 6w and adult pigs. For both the ancestral and insertion viruses, there was less virus attachment in the upper respiratory tract than in the lower respiratory tract. Although there was only minimal (+) attachment to the 6w nasal turbinate and tracheal epithelium for both the insertion and ancestral viruses, there was moderate (++) attachment to the adult nasal turbinate and tracheal epithelium for the ancestral viruses and minimal to moderate attachment for the insertion viruses (Table 2; Figs. 5–6). The morphometric scores of the ancestral viruses, SG445 and SG1097a, revealed more attachment to the bronchiolar epithelium (4.28% ± 1.81% and 6.19% ± 2.56%, respectively) than insertion viruses SG443, SG444, SG1094, SG1096, and SG1150 (0.71% ± 0.46%, 0.40% ± 0.30%, 3.84% ± 2.18%, 3.62% ± 2.37%, and 2.52% ± 2.46%, respectively). According to the post hoc Tukey test, the detectable difference (P < .0001) was between the 2 viruses with the lowest binding (SG443 and SG444) and the other 5 viruses studied (SG445, SG1097a, SG1094, SG1096, and SG1150) with overlap between SG444 and SG1150.
Discussion
IAV had remained relatively stable in the North American swine population until the emergence of the triple reassortant H3N2 viruses in 1998. 36 The rapid evolution of North American swine lineage IAV over the past decade has resulted in remarkable genetic and antigenic variability and presented a major challenge to the swine industry. This has been particularly true for the HA gene of viruses in the SwH1 subtype, which have 4 genetically distinct clusters and 3 subclusters. 18 Although a number of studies have examined the genetic and antigenic characteristics of these emerging virus variants in swine, 13,14,32 –34 there have been limited studies to determine if genetic and antigenic variation has led to changes in attachment of virus to the epithelial cells during infection. The purpose of this study was to examine the PVA to porcine respiratory tissues for IAV isolated from North American swine.
Although limited replication of influenza viruses can occur in cell cultures without SA receptors present, 24 much of what is known about IAV attachment to epithelial cells in the respiratory tract focuses on α2–3 and α2–6 SA and has been determined by LH assays despite the discrepancies between studies and the limited specificity of SNA and MAA lectins. 15,19,23,37 Recent studies have indicated that α2–3 SA and α2–6 SA are not the sole receptors for avian and mammalian IAV viruses. 12,19,24 Glycan microarrays allow for avidity comparisons due to modifications in the SA (eg, total SA length and fucosylation, sulfation, and sialylation at position 2 or 3), and microarray analysis has found that influenza viruses bind to a wide spectrum of glycans, including non-SA residues. 3 This highlights the need to specifically identify glycans of importance in the respiratory tract that are overlooked by LH assays.
Virus Histochemistry Compared With In Vivo Infection
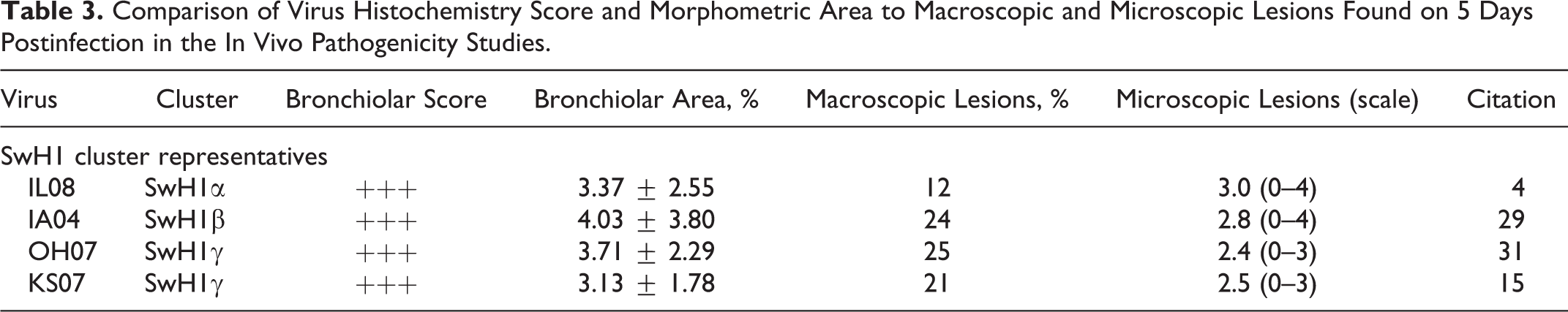
One of the viruses in a glycan array study by Chen et al, 3 A/Ohio/1/2007 (FJ986620), demonstrated high affinity for α2–6 SA and limited affinity to α2–3 SA. Since A/Ohio/1/2007 was collected from the same zoonotic event as OH07 and the HA nucleotide sequences are identical, it is likely that they would have similar binding affinity. Experimental infection of 4-week-old pigs with OH07 (n = 15) also demonstrated lung lesions typical of virulent strains of IAV in swine. Macroscopic lung lesions averaged 25% and microscopic lung lesions (0–3) were 2.4 on 5 days postinfection (DPI). 34 All 8 gene segments of KS07 have at least 99.7% nucleotide similarity to OH07, and experimental infection with KS07 (n = 20) resulted in similar lesions with 21% macroscopic and 2.5 (0–3) microscopic scores 5 DPI. 16 OH07 and KS07 had similar patterns of virus attachment to the respiratory epithelium and morphometric scores (3.71% ± 2.29% and 3.13% ± 1.78%, respectively).
The SwH1β virus, IA04, had similar lesions in vivo (n = 15) compared with the SwH1γ viruses with 24% macroscopic and 2.8 (0–4) microscopic lung scores 5 DPI. 33 Similar findings were demonstrated in vitro using VH with IA04 having a similar PVA to the respiratory epithelium and morphometric score (4.03 ± 3.80) as the SwH1γ viruses. The SwH1α virus, IL08, also had a similar attachment pattern and morphometric score (3.37 ± 2.55), and experimental in vivo infection (n = 9) with IL08 resulted in a similar average microscopic lung score of 3(0–4) 5 DPI, but macroscopic lung lesions averaged only 12%. 4 This may have been related to the D127E substitution that has been associated with increased α2–6 SA affinity described by Chen et al 3 or virulence factors found in the other gene segments that do not play a role in attachment to the cell surface (eg, polymerase basic 1 or 2 genes).
There was a stark contrast in virus attachment patterns between the strong binding of the SwH1α, β, and γ viruses compared with the minimal binding of the SwH1δ1 viruses throughout the respiratory tract of both 6w and adult pigs. This difference was detected by the ANOVA for the morphometric scores of the SwH1 cluster viruses (P < .001) and the post hoc Tukey test. The SwH1δ1viruses examined in the present study have the human-lineage T133 S residue and 131G deletion in HA. TX08 had the D187 N substitution, which resulted in higher α2–3 and α2–6 SA affinity in the human isolates. 3 Also, TX08 and SG239 have the NS1 residues (108R, 125E, and 189G), which have been associated with a loss of binding to the epithelial receptors. 13 The SwH1δ1 viruses had minimal attachment to the respiratory epithelium, but the morphometric score for control virus (averaged for the 4 staining runs) was 3.31% ± 2.17% compared with 0.33% ± 0.20% for IL07 and 0.49% ± 0.33% for TX08. Experimental in vivo infection with TX08 (n = 5) resulted in 16.8% gross lung lesions and microscopic lung lesions averaged 2.1 (0–3) at 5 DPI (J. R. Ciacci-Zanella, unpublished data).
The patterns of virus attachment and the morphometric scores in the respiratory tissues of the 6w pig appear to be highly correlated to the macroscopic and microscopic lung lesions at 5 DPI for the SwH1α, β, and γ viruses. However, the correlation between VH results and in vivo lung lesion did not hold true for the SwH1δ1 viruses, which had low levels of virus attachment throughout the respiratory tract and low morphometric scores in the bronchioles. This variation is likely due to other factors of virulence (eg, viral polymerase activity) for these SwH1δ1 viruses. Although in vivo pathogenesis studies have not been performed on SG239, it is important to note that this virus is also a SwH1δ1 virus, and this virus had more binding in the 6w bronchioles and alveoli than IL07 and TX08 along with a higher morphometric score (3.31% ± 2.17%).
Ancestral and Insertion Viruses
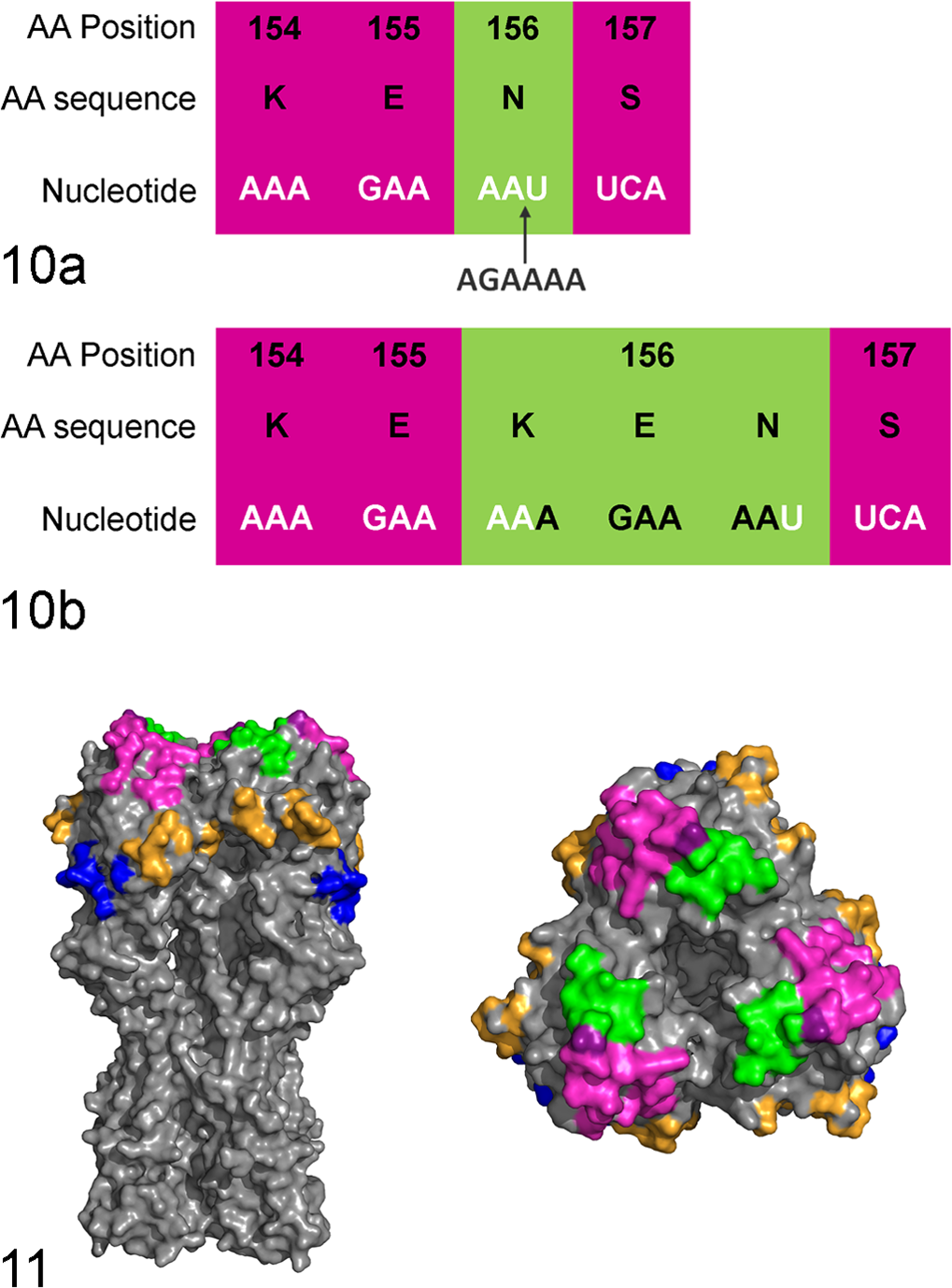
The ancestral and insertion viruses originated from a multisite production system with weaned pigs sent to multiple off-site nurseries. The ancestral viruses predated the insertion viruses and had 98.8% to 99.8% nucleotide similarity to the insertion viruses. All of these viruses were isolated from different nursery sites within the farm system over a 3-year period, but SG443 and SG444 were collected at the same time from 2 different barns. The infected animals were 3 to 7 weeks old (average age 4.6 ± 1.4 weeks) with 15% to 85% morbidity and up to a 4% increase in mortality compared with the other groups of pigs placed in other nursery sites at the same time. The origin of the insertion is not certain, but 6 inserted nucleotides within the codon for site 156 in the HA1 numbering system 2 are an exact repeat of the previous 6 nucleotides (Fig. 10). Site 156 is part of the Sb antigenic site, 2 which is adjacent to the SA binding pocket at the top of the HA protein trimer (Fig. 11). Predicted protein models of the ancestral and insertion viruses would be useful to visualize how an additional 2 AAs in all 3 HA in the trimer may change the shape of the binding pocket.
The PVA for the insertion and ancestral viruses showed strongest binding in the lower respiratory tract of both the 6w and adult pigs. Binding to the lower respiratory tract, particularly to the alveoli, and the subsequent diffuse alveolar damage, was the main pathologic abnormality in fatal cases of infection with pH1N1 in humans during the pandemic. 17 Further studies will be needed to examine which specific mutations within HA or other genes are associated with the apparent lower respiratory tract binding predilection of the gamma cluster viruses examined in this study (Tables 2 and 3). Further studies will also be needed to determine if these changes translate directly to in vivo binding and replication models or if these changes can be compensated for by synergistic effects of the virulence factors of the internal gene proteins.
Comparison of Virus Histochemistry Score and Morphometric Area to Macroscopic and Microscopic Lesions Found on 5 Days Postinfection in the In Vivo Pathogenicity Studies
Limitations
Some of the limitations of this technique are apparent in the results. Although minimal variation was found in the viral binding to the bronchiole of the four 6w pigs, these 4 pigs were from 3 different litters on the same farm and therefore had a similar genetic background. It would be useful to examine lung samples from 4 to 6 animals using a multicassette to get more genetic variation on a single slide while also increasing the number of bronchioles for the analysis. In addition, the dynamics of virus binding throughout the developmental stages should be examined using the respiratory tracts of pigs at different stages of development from birth to slaughter (6 months). One of the analyses that showed variation was the morphometric results between staining runs. This is a serious limitation that should be examined further. Until repeatability of the technique is improved, comparisons between staining runs or even laboratories and other publications for this technique would be limited. The virus was quantified by the HA titer for this study and in previous studies by other authors. 30,31
The HA titers were measured by an experienced technician who routinely runs HA and HI titers for swine-origin influenza viruses in the UMVDL serology lab using turkey erythrocytes. Multiple samples were taken during the virus preparation process to demonstrate that there was little to no change in the HA titer from before and after the FITC was attached. However, the drop in the HA titer after the formalin inactivation could indicate that the formalin inactivation caused structural changes in HA proteins that could have an effect on PVA. The viruses were inactivated for biosafety reasons, but if the technique was done in a location with enhanced biosafety measures, the viruses would not need to be inactivated. Another possibility would be to use a different method of inactivation, such as binary ethyleneimine (BEI) used for inactivated influenza vaccines, 11 to better preserve the protein structure.
Some of the other limitations of VH are similar to those of immunohistochemistry (IHC). Some of the sources of nonspecific background staining for IHC can result from endogenous biotin in the tissues and can be avoided by using a polymer-based IHC method, such as Dako’s EnVision+ system. 21,35 Background produced by endogenous antibodies present in plasma cells can be limited by adding a 15-minute, 10% normal serum step after the hydrogen peroxide step and before the antibody incubation. In addition, endogenous antibodies can be blocked by adding 2% to 4% normal swine serum to the rabbit EnVision+ solution. These modifications in the VH technique were tested using SG239 in the adult respiratory tissues, and although there was minimal background staining, there was strong nonspecific immunoreactivity in the plasma cells. Further modifications need to be tested to reduce the plasma cell staining.
Conclusions
In this study, computerized morphometry was used for the first time in combination with VH to provide a quantitative measure to compare the PVAs of multiple viruses within the bronchiole. The measurements were taken from only 5 bronchioles, resulting in a minimal detectable significant difference of 2.85%. To detect a difference of 2.02%, a sample size of 10 bronchioles would be needed, and 40 bronchioles would be needed to detect a 1.01% difference.
The present study also provided new insights on PVAs for IAV from swine to porcine respiratory epithelial cells. One of the new insights was a significantly lower morphometric score among the human-origin HA SwH1δ1 viruses compared with the other cluster representatives that have swine-origin HAs. This might indicate a difference in preference for swine or human SA receptors. Another important finding was that although not significantly different, morphometric scores were lower among the insertion viruses compared with the ancestral viruses. Since this insertion is still being found in swine 5 years later in the Midwestern United States, this finding should be further examined to demonstrate if and how this change could correspond with in vivo virulence or adaptation of the virus.
In addition, it is important to note that using the VH results alone cannot predict either virulence or host range. There was strong binding to the bronchioles and alveoli for the PVA of some of the cluster representative viruses previously described in the in vivo studies as causing severe lung lesions. However, for other viruses, the VH results did not always correlate with the reported in vivo virulence. This finding highlights the need to interpret VH results in combination with ex vivo or in vivo binding and replication models. A balanced interpretation of viral binding of swine viruses will lead to better understanding of the role of virus tropism and replication in IAV pathogenesis in pigs and may also provide new targets for more effective prevention and treatment measures.
Footnotes
Acknowledgements
The authors thank Debby van Riel, Chris Stuart, Josh Parker, Giordana Costa, Wendy Wiese, Jan Shivers, Laurie Landgren, and Michelle Nelson for their technical assistance and expertise, as well as Drs Hesse and Vincent for providing KS07 and OH07 viruses, respectively. The authors also thank Drs Ciacci-Zanella and Vincent for sharing their expertise and experimental results.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship and/or publication of this article: This work was funded by USDA Discretionary GAR Funds AES0060014 as part of the Signature Program Funding at the College of Veterinary Medicine, University of Minnesota. SD and MG were funded in part by federal funds from the NIAID, NIH, HHS, under Contract No. HHSN266200700007C.
