Abstract
Pathbase, the database of mouse histopathology images, was developed as a resource to provide free access to representative images of lesions in background and mutant strains of laboratory mice. When utilized with diagnostic workups or phenotyping of mutant mice, it can provide a “virtual second opinion” for those working without access to groups of experienced pathologists. This is a community resource, and it facilitates the sharing of expertise and data among members of the pathology community worldwide. MPATH—the mouse pathology ontology—was developed alongside Pathbase for the annotation of images and now represents an important resource for the coding of diagnoses, permitting sophisticated data retrieval and computational analysis of mouse phenotypes. In this article, the structure and use of MPATH is discussed, along with current and future challenges for the coding of mutant mouse phenotypes.
The major challenge of the postgenomic era is the attribution of function to genes and pathways. The use of model organisms such as the mouse to provide phenotype–genotype relations is now established as a key approach to establishing normal gene function, and traditional hypothesis-driven research has now been augmented by the coordinated large-scale mutagenesis and phenotyping projects established internationally over the last decade. 16 At the time of writing, Mouse Genome Informatics (http://www.informatics.jax.org/) lists 23,600 mutant alleles and 34,038 genotypes with phenotypic information. The International Knockout Mouse Consortium (http://www.knockoutmouse.org/) 8 aims to systematically generate in the order of 20,000 new null or conditional alleles within the next 2 years and, through the EUMODIC program, initially phenotype 650 new mutant mouse lines. The scale of the resources now available and promised is huge and the potential impact of using these new tools enormous. However, achieving this potential requires systematic annotation of phenotypes, integration of disparate and complex data types, and resources for sharing the data.
The pathological analysis of mutant mice is a key mode of investigation for both small- and large-scale approaches to phenotyping, generating what are often the key data needed to interpret and classify the phenotypic consequences of gene mutation or dysregulation. Pathbase and MPATH (mouse pathology ontology) represent developments made in response to community demand for a systematic, computable description framework for mouse pathology and access to primary data and reference images for mutant mouse phenotyping. These terms and codes can be integrated into diagnostic databases that, when linked to Pathbase online, can provide access to images and definitions, thereby providing a “virtual second opinion.” 22,26
Addressing the Expertise Gap in Mutant Mouse Pathology
The increasingly urgent need for trained pathologists to phenotype genetically engineered mutant mice highlights a now well-recognized deficiency.
2,6,11,19
Encouraging young pathologists to enter into this field as a full-time endeavor, beyond toxicological pathology and drug safety in industry, has been difficult. One reason is that few universities really specialize in genetics-based mouse research to the level of creating large groups to provide an environment to encourage career development. Alternatively, small pathology phenotyping services develop, often lacking a critical mass of experts, much less with a large and diverse-enough caseload to provide experience over time or accessibility to consultation.
24
A number of specialty courses have become available to help address some of the training issues,
23
as well as
A key problem is the dissemination of experts around the world and the lack of up-to-date resources for training and reference. Much of this can now be addressed through use of the World Wide Web. A number of highly specialized websites are available, some of which provide reference images of mouse histopathology. 5,27 There are also, gratifyingly, numerous books on the subject, with more being published annually. 27
Initially funded by the European Commission, Pathbase (http://www.pathbase.net/) was developed in 1999 by a group of European and North American pathologists as a community response to the above problems. It contains photomicrographs of representative images 17,18 annotated to a set of defined controlled vocabularies and ontologies that provide a public resource for the sharing of images of normal and abnormal tissues from mutant and background strains of laboratory mice (Fig. 1 ). 17 It currently holds more than 2,000 TIF and JPG images covering a range of normal, neoplastic, and nonneoplastic lesions from more than a hundred mutant and inbred background strains. Pathbase has recently included zoomable whole-slide images (virtual slides) in its collection, with plans to expand the proportion of these valuable images. The images are annotated with details of strain, genotype, anatomical location, and diagnosis, with key annotations derived from controlled vocabularies or OBO Foundry–compliant bio-ontologies: 20 MPATH, MA (mouse anatomy ontology), EMAP (Edinburgh Mouse Atlas Project; developmental anatomy ontology), CL (cell ontology), and GO (gene ontology). Nomenclature for mouse strains and mutant gene symbols are included, when provided, and follow the International Mouse Genetic Nomenclature Committee formats, 25 which allows for comparison among studies addressing modifier genes or strain-specific diseases that might confuse interpretation.
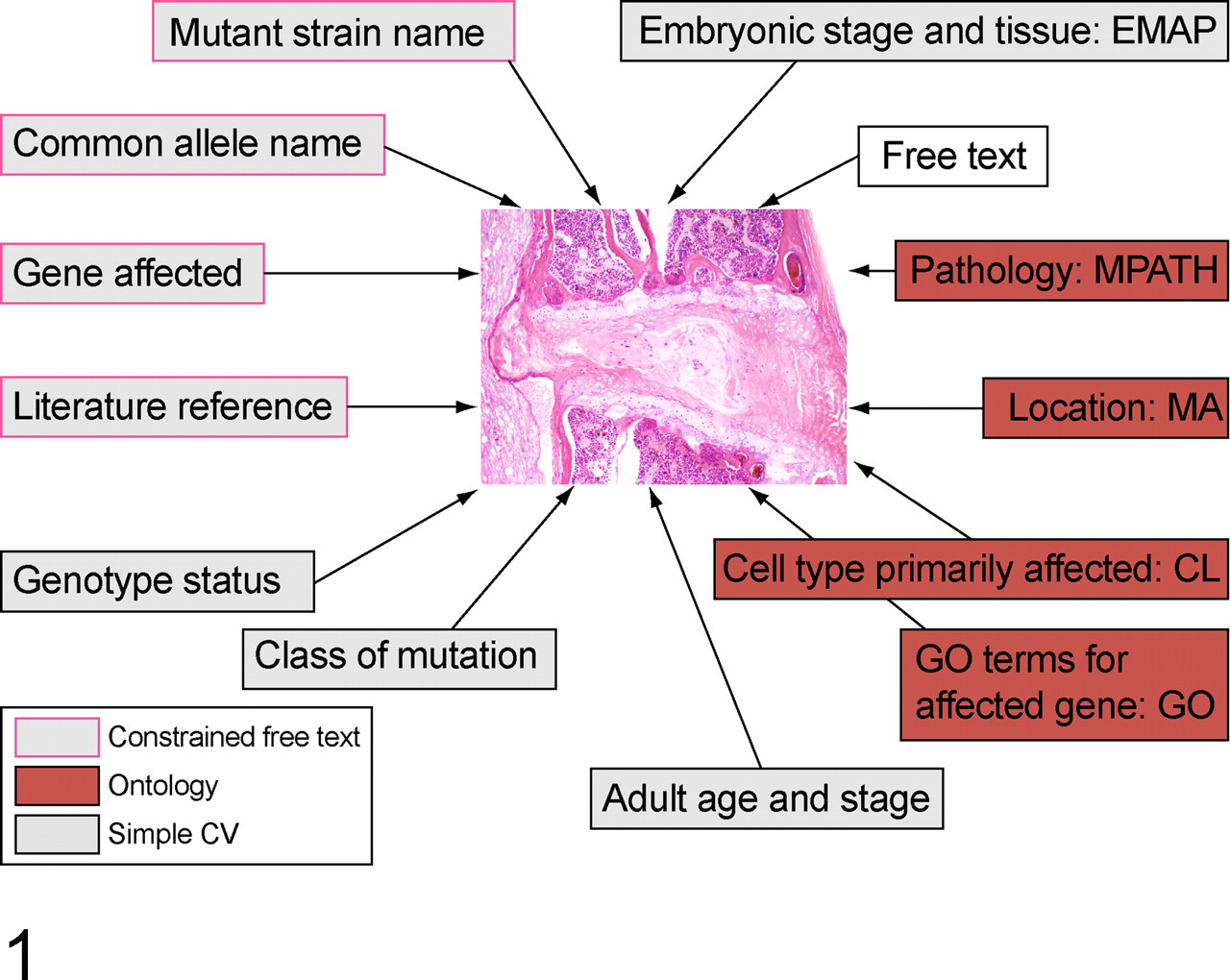
Pathbase image annotations. Each image is annotated to provide information on the animal in which the lesion originated and the lesion itself. The gene, allele name, and strain name are constrained free text (ie, using standard nomenclature). Genotype status and type of mutation are drawn from short controlled vocabularies (CV); the remainder use specified ontologies. There is an additional provision for a free text annotation to provide other nonstructured information. EMAP, Edinburgh Mouse Atlas Project, developmental anatomy ontology; MPATH, mouse pathology ontology; MA, mouse anatomy ontology; CL, cell ontology; and GO, gene ontology.
Using Pathbase
Pathbase is completely open access and is used for reference, training, and data sharing. Users range from individual investigators carrying out hypothesis-driven academic research based on the mouse as a model organism to large-scale mouse mutagenesis and phenotyping programs. The images are contributed by major research centers, individual investigators, and legacy sources (wherein rare and valuable material that is at risk of being lost is digitized and made available to the community). Pathbase is open to anyone to submit images directly through the webpage. These are checked by curators and reviewed by members of the pathology panel before being made available. Thereafter, a feedback facility (available though each record page) enables users to comment on the images and share their expertise.
The use of Pathbase as a data-sharing resource is becoming increasingly important. Histopathology images on which the conclusions of publications are based are frequently placed on the “supplementary information” sites of journals or are unavailable in a usable size or resolution. The increasing occurrence of egregious diagnostic errors in papers published in major journals has caused great concern. For example, neoplasms that investigators claim to have found in new mutant mouse lines are often hyperplasias, dysplasias, cystic lesions, or other degenerative conditions; furthermore, too many publications base the entire evidence for phenotypes on erroneous pathological diagnosis. 7
Submitting Images to Pathbase
Individual users may sign up to submit their images to Pathbase, using the Submit Images link at the bottom of the index page. The user is given a series of menus and input boxes to add the details of the image, strain, diagnosis, and so on. Clicking on Upload allows users to browse their machine, select the image, and upload directly to Pathbase. The images are then subject to curation and checking before being made publicly available. At the moment, the majority of images being submitted are TIFs and JPGs, but it is possible to upload WSI images in Aperio or Hamamatsu format using the same system. Because of image sizes and users' available bandwidth, this may be inefficient. Users should contact Pathbase to arrange for a batch upload facility using FTP.
Integration With Other Projects and Databases
Pathbase has established links with the Jackson Aging Center, 3,31 the Mouse Phenome Database, 4 and the Mouse Tumor Biology Database 13 to curate and host images from ongoing studies of age-related lesions and normal tissue variation from 31 inbred strains of mice. Quantitative data for the other systems-based parameters measured are loaded into the Mouse Phenome Database, and the two datasets (ie, the image data set and the quantitative scores for all the diagnoses) are integrated between the Mouse Phenome Database and Pathbase. Similar integration has been recently established with the European Radiobiology Archives 28 and with the Northwestern University Janus radiobiology database (http://janus.northwestern.edu/janus2/), 14 which has coded 50,000 individual mouse records to MPATH to link the two datasets.
Mouse Anatomy and Pathology Ontologies
Pathologists are trained to use language carefully. Over the centuries, since Rudolf Virchow
30
and others formalized pathology as a medical specialty, language has evolved to encompass new discoveries and technologies but also deal with synonyms, spelling variations, and so forth. These variables are exacerbated by different training sites, training methods, and institutional specialties. Databases depend on standardized terminologies, cross-references for synonyms, and spelling. To address these generic problems, the bioinformatics community has, in recent years, produced complex term hierarchies—ontologies—describing various areas of knowledge (gene properties, anatomies, etc) where the terms are linked by relationships (eg,
MA was developed to standardize anatomical terms. 10 MA has a formal ontological structure built on the kind of framework contained in, for example, an anatomy reference book 15 but is open and under constant development and refinement. This is a dynamic process, and, as such, MA is regularly updated as users provide input to curators. A textbook on comparative microscopic anatomy of the mouse and human is currently being written (P. Treuting, corresponding editor [personal communication]), and similar ones on embryology are now available. 12 These will provide further extension of comparative anatomical nomenclature.
Pathology nomenclature for the mouse has been captured in the form of an ontology called MPATH, as built with the expertise of the Mouse Pathology Ontology Consortium, a group of 20 DVM and MD pathologists and biologists who regularly meet to review and update the ontology. MPATH can be browsed online with the click/expand hierarchy browser; it can be searched online with free text expressions; and it can be downloaded as an OBO-format human-readable flat file from the Pathbase site (http://eulep.pdn.cam.ac.uk/Pathology_Ontology/mpath1/mpath_obo_2002_07_26.txt) or from the OBO Foundry repository (http://www.obofoundry.org/). The downloaded OBO flat-file version may be imported into ontology-aware software or other databases.
MPATH is the only ontology currently available to describe the full range of mouse histopathology in a formally structured way, with subsumption hierarchies allowing accurate inference. It is segmented into aspects of pathology that are familiar to traditionally trained pathologists. The most current release is fully defined and contains terms for all the major classes (594 to date) of pathological lesions and processes relevant to the mouse. Many tissue responses are common to multiple anatomical sites, and as far as possible, the redundancy of specifying a response in multiple tissues is avoided by the curatorial creation of cross products with an appropriate anatomy ontology such as MA, the mouse adult anatomy. The use of cross products prevents the combinatorial explosion that causes ontology bloat in poorly structured ontologies—the inclusion in the ontology of all possible precomposed variations of instances of an entity. For details of the semantic issues surrounding the description of mammalian phenotypes, see Hancock et al. 9 Other controlled vocabularies for rodent pathology (eg, that being derived from the recent INHAND [International Harmonization of Nomenclature and Diagnostic criteria] initiative of the international societies of toxicopathology 29 ) have inappropriate content and structure for studies of mutant mice—particularly, the kind of phenotyping studies used in functional genomics. And whereas it is vital to ensure that MPATH can cover the terms in such glossaries, they do not have the computational reasoning power of ontologies needed for informatics.
The definitions used in MPATH range from the simple format found in
Conclusions
Biology has become Big Science, and with the advent of the large-scale systematic mutagenesis and phenotyping programs worldwide, the place hitherto occupied by high energy physics has been filled by the new biology. As part of this, pathology as a discipline has a crucial role in advancing our understanding of gene function in health and disease, and pathologists are now key players in this global endeavor. This responsibility presents challenges—those of training, modernization, and standardization—but most of all, it requires coordination to make the most of efforts distributed worldwide. We need new resources, and we need a new worldview in veterinary pathology if we are to play the important role that we have been assigned. Learning to use the new resources and informatics is a challenge to which the profession must be sure to rise.
Footnotes
Dr. Sundberg has a research contract with the Procter and Gamble Company that is totally unrelated to this project.
This work was supported by grants from the European Commission (contract Nos. QLRI-1999-00320, PATHBASE; LSHG-CT-2006-037188, EUMODIC), the North American Hair Research Society, the National Institutes of Health (CA089713, RR17436, AR49288), and the Ellison Medical Foundation.
