Abstract
Objective
To address challenges such as blurred boundaries and irregular shapes in brain tumor magnetic resonance imaging scans, we developed a lightweight detection framework to enhance automated diagnosis and meet real-time clinical requirements.
Methods
We proposed an improved You Only Look Once version 12 (YOLOv12n)-based model featuring three modules. First, the Attention-based C2f with Frequency-domain Feed-Forward Network (A2C2f-DFFN) module was incorporated into the backbone network; it combined an attention mechanism with a frequency-domain feedforward network to enhance global context modeling and detailed feature reconstruction. Second, the C2f with Token Statistics Self-Attention and Dynamic Tanh (C2TSSA-DYT) module was employed in the feature fusion neck; it utilized statistical self-attention and a dynamic Tanh activation function to improve robustness in complex backgrounds. Finally, the dynamic upsampling operator was adopted in the feature reconstruction stage; it dynamically generated sampling weights to effectively prevent boundary blurring and detail loss.
Results
On the Kaggle brain tumor dataset, our method achieved 93.2% precision, 88.4% recall, and 94.1% mean average precision at IoU threshold 0.5 (
Conclusion
The enhanced YOLOv12n framework proposed in this study achieved good balance between accuracy and efficiency in brain tumor detection tasks, demonstrating strong robustness, which makes it suitable for use in clinical computer-aided diagnosis systems.
Keywords
Introduction
Brain tumors pose an increasing threat to global health, with the World Health Organization and the Global Burden of Disease study reporting a steady increase in global incidence. 1 These tumors are characterized by complex pathology and high recurrence rates, making early and accurate diagnosis essential for effective treatment planning and improved patient prognosis. 2 Magnetic resonance imaging (MRI), known for its high soft-tissue contrast and noninvasive nature, has become the primary modality for brain tumor detection and diagnosis. 3 However, tumor manifestations on MRI are highly heterogeneous, often exhibiting blurred boundaries, irregular shapes, and significant size variations, which complicate radiological assessment.
Traditional diagnosis relies heavily on visual interpretation by experienced radiologists, a process that is not only time-consuming and labor-intensive but also subject to considerable interobserver variability. 4 This approach increasingly fails to meet the growing clinical demand for efficient and highly accurate diagnostics. Consequently, computer-aided diagnosis (CAD) systems for automated brain tumor detection have gained significant attention. 5 Advancements in deep learning, particularly deep convolutional neural networks (CNNs),6,7 have enabled the development of end-to-end trainable models capable of automatically extracting multilevel tumor features, thereby improving detection accuracy and robustness. Among object detection frameworks, the You Only Look Once (YOLO) series8–15 has gained prominence due to its efficient architecture and rapid inference speed, making it well-suited for medical image analysis.
However, existing algorithms face several critical challenges when deployed in resource-constrained clinical environments. A significant gap remains in developing a detection framework that can simultaneously achieve high accuracy, real-time efficiency, and strong robustness to specific MRI characteristics. First, from a lightweight design perspective, models such as MK-YOLOv8 16 and STAR-YOLO 17 often underperform when handling complex tumor morphologies, as they typically compromise on the feature extraction capacity. Second, regarding feature representation, many methods lack specialized modules to enhance discriminability at blurred boundaries and fine anatomical details. For example, although Gayathiri et al. 18 employed sophisticated preprocessing and autoencoders, their approach was not optimized for real-time detection. Similarly, Yang et al. 19 and Pandey et al.20,21 incorporated attention and multiscale fusion mechanisms; however these approaches often came at the expense of increased complexity or limited generalization. Third, in terms of robustness and generalization, methods that rely predominantly on expert-annotated data or single-domain datasets20,22 frequently exhibit scalability issues.
In recent years, attention-based architectures, particularly the transformer and its variants, have demonstrated considerable potential in medical image analysis. For example, Swin Transformer–based models have been successfully applied to brain tumor segmentation, leveraging hierarchical self-attention to capture long-range dependencies across scales. 23 Similarly, Pan et al. 24 integrated channel and spatial attention modules into CNNs to selectively emphasize informative features. However, the deployment of such methods in real-time, lightweight detection systems is often constrained by their substantial computational overhead. Moreover, existing attention mechanisms are generally not explicitly designed to address the specific challenges presented by brain MRI. Therefore, there is an urgent need to develop more efficient and specialized attention mechanisms that can be seamlessly integrated into lightweight detectors such as YOLO.
To address these limitations, we proposed an improved lightweight brain tumor detection framework based on You Only Look Once version 12 (YOLOv12n), with a focus on enhancing feature modeling, interaction, and reconstruction capabilities without compromising efficiency. The main contributions of this work are as follows:
The Attention-based C2f with Frequency-domain Feed-Forward Network (A2C2f-DFFN) module was proposed via the integration of attention mechanisms with frequency-domain feedforward networks to enhance global context modeling and detailed feature preservation, effectively improving discriminative capability in cases with blurred tumor boundaries. The C2f with Token Statistics Self-Attention and Dynamic Tanh (C2TSSA-DYT) module was designed by employing statistical self-attention mechanisms and dynamic activation functions to strengthen global dependency modeling while maintaining the local advantages of convolution, thereby enhancing robustness in complex backgrounds. The dynamic upsampling (DySample) operator was adopted, utilizing a dynamic weight generation mechanism to alleviate feature detail loss resulting from the use of conventional upsampling methods, significantly improving the localization accuracy of small tumors.
Related work
Efficient multiscale attention
The Efficient Multi-scale Attention (EMA) module 25 utilizes a dual-branch design to capture channel and spatial dependencies. The convolutional branch applies group-wise convolution and Softmax to generate channel-refined features, whereas the coordinate pooling branch performs horizontal and vertical average pooling to produce spatial attention features. The outputs of both branches are fused through summation and refined via sigmoid activation and weight modulation. This architecture effectively integrates channel and spatial attention mechanisms and maintains computational efficiency.
Content-Aware ReAssembly of Features (CARAFE) module
The CARAFE 26 upsampling operator achieves feature map upscaling through two core modules. In the kernel prediction module, the input feature map undergoes channel compression and content encoding to generate and normalize upsampling kernels. Subsequently, in the feature reassembly module, the predicted kernels perform dot product operations with local regions of the input feature map, ultimately producing an output feature map with resolution increased by a factor of sigma (σ). Through its content-aware kernel prediction mechanism, this operator significantly enhances feature preservation during the upsampling process.
Methods
This study proposed an improved architecture based on YOLOv12n to enhance detection capabilities in brain tumor MRI scans through three core modules. As presented in Figure 1, in the backbone network, the A2C2f-DFFN module integrated channel attention with a dual feedforward network in the frequency domain, enhancing the extraction of salient tumor features and suppressing background interference. In the feature fusion neck, the C2TSSA-DYT module employed statistical self-attention to model inter-channel dependencies and used dynamic Tanh (DyT) activation to adaptively adjust activation thresholds, improving robustness in detecting tumors with varying intensities and irregular morphologies. During the upsampling stage, the DySample operator dynamically generated sampling weights, effectively mitigating feature detail loss and boundary blurring. This architecture maintained a lightweight design and significantly improved detection performance in challenging scenarios involving blurred boundaries, small lesions, and complex backgrounds.

A schematic of the enhanced YOLOv12n framework. The proposed A2C2f-DFFN, C2TSSA-DYT, and DySample modules were incorporated into the backbone, neck, and upsampling paths, respectively, enabling robust feature representation, interaction, and reconstruction for superior accuracy-efficient brain tumor detection. YOLOv12n: You Only Look Once version 12; DySample: dynamic upsampling; A2C2f-DFFN: Attention-based C2f with Frequency-domain Feed-Forward Network; C2TSSA-DYT: C2f with Token Statistics Self-Attention and Dynamic Tanh.
A2C2f-DFFN module
Although YOLOv12n exhibits notable real-time capability and detection accuracy in brain tumor detection tasks, 27 its performance is often constrained by insufficient feature representation and limited discriminative power, particularly when confronted with blurred tumor boundaries. To mitigate this limitation, we introduced the A2C2f-DFFN module, which enhances multiscale feature modeling and detail restoration.
As depicted in Figure 1, the module processes input features through the following pipeline. First, features underwent an initial convolutional layer for preliminary spatial feature extraction and dimensional adjustment, establishing a foundation for subsequent processing. The resulting features were then propagated into a core structure composed of stacked Ablock-DFFN units. Within this structure, the Ablock incorporated an attention mechanism to capture long-range dependencies and effectively integrated global contextual information. Concurrently, the DFFN 28 module employed a gating mechanism to selectively preserve and combine low- and high-frequency components. Low-frequency signals helped restore global structures and contours, whereas high-frequency components enhanced fine details and edge information. By decomposing and recombining features in the frequency domain, the DFFN effectively balanced structural coherence with detail recovery, resulting in significant improvements in deblurring performance and visual quality. Subsequently, the outputs of the core module were further refined via an additional convolutional layer to integrate discriminative representations. A residual connection was also incorporated, directly adding the original input features to the convolved outputs. This cross-layer feature reuse not only mitigated gradient vanishing during deep network training but also accelerated convergence and improves numerical stability. The final output of the module was obtained through this additive fusion operation.
C2TSSA-DYT module
Although the A2C2f-DFFN module enhances the model’s ability to capture tumor boundaries and fine-grained details through improved feature extraction, its ability to model global dependencies and multiscale interactions in complex backgrounds remains limited. To address this limitation, we introduced the C2TSSA-DYT module, which strengthens the model’s adaptability to global spatial relationships and dynamic feature distributions, thereby further refining the detection robustness of brain tumor MRI.
As depicted in Figure 1, the proposed module comprised three key stages: convolutional feature extraction, dual-branch modeling, and feature fusion. The input features were first transformed by a convolutional layer and subsequently divided into two streams. One stream was processed by a series of TSSA-DYT units, in which the token statistics self-attention (TSSA) 29 mechanism captured global dependencies using token-level statistics, substantially reducing computational overhead and maintaining long-range contextual modeling. The subsequent DyT 30 activation incorporated an input-adaptive factor that modulated the shape of the Tanh function, enhancing nonlinear representational capacity. The second stream served as a residual bypass, preserving local structural features. The outputs of both branches were concatenated along the channel dimension and subsequently compressed and fused through a convolutional layer to generate the final output. This design effectively integrated global context and retained the local representational advantages of convolution, significantly improving feature discrimination and robustness in complex brain tumor detection scenarios.
DySample operator
In the feature reconstruction stage of brain tumor detection, the choice of upsampling operator critically influences detection accuracy and computational efficiency. Although commonly used methods such as nearest-neighbor and bilinear interpolation offer high computational efficiency, they often result in the loss of fine details and boundary blurring. Deconvolution can restore some high-frequency information; however, it tends to introduce checkerboard artifacts, compromising feature consistency. To address these limitations, we adopted the DySample operator, which dynamically generates interpolation weights adapted to feature distribution across different spatial locations. This approach effectively preserves boundary structures and detailed information and maintains lightweight computation, thereby significantly enhancing resolution recovery and localization accuracy in brain tumor detection tasks.
As demonstrated in Figure 2, DySample
31
mainly consisted of a sampling point generator and a grid sampling module. Given an input feature map

A schematic of the DySample operator. DySample: dynamic upsampling.
As demonstrated in Figure 3, the sampling point generator produced a sampling set

Architecture of the DySample point generator. DySample: dynamic upsampling.
Experiment
Dataset
The medical imaging dataset used in this study was obtained from the publicly available “Medical Image Dataset: Brain Tumor Detection” on Kaggle. 32 It consists of MRI scans of three common brain tumor types: meningioma, glioma, and pituitary tumor (Figure 4). All images were anonymized in accordance with medical privacy guidelines. During data preprocessing, rigorous screening was performed to exclude low-quality images (e.g. blurred, improperly exposed, and those lacking anatomical structures). The final curated dataset comprised 2451 training and 613 testing images. Data partitioning maintained class balance and sample representativeness. Moreover, the dataset’s diversity in imaging conditions, tumor locations, and visual characteristics provided a reliable foundation for validating method robustness.
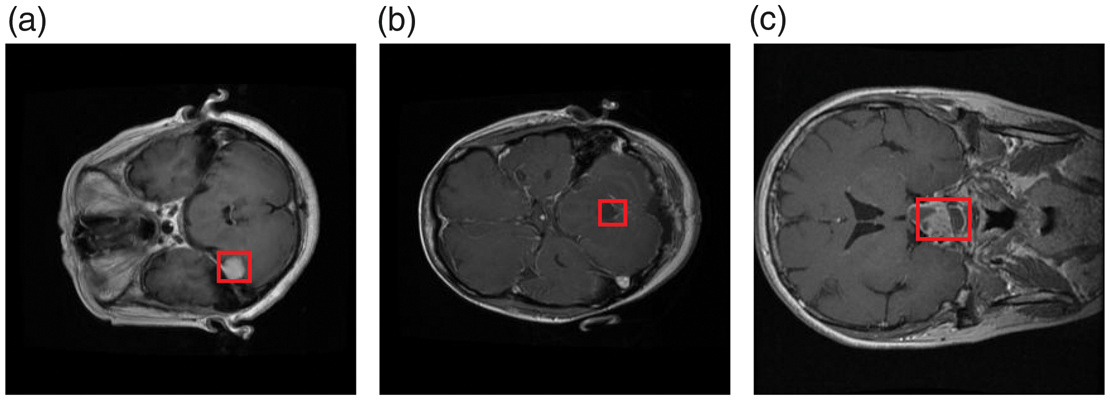
Representative MRI samples from the dataset depicting three tumor categories: (a) meningioma, (b) glioma, and (c) pituitary tumor. MRI: magnetic resonance imaging.
Discussion on dataset selection and limitations
Although the selected Kaggle dataset serves as a widely recognized benchmark for fair algorithmic comparison, we acknowledge several limitations regarding potential biases and generalizability. First, the dataset lacked detailed metadata related to patient demographics (e.g. age, sex, and ethnicity) and acquisition protocols (e.g. MRI machine vendors and magnetic field strengths). This absence of information may have introduced implicit demographic or acquisition biases, potentially affecting the model’s fairness across diverse patient populations. Second, public datasets often undergo preselection, in which “textbook-quality” cases may be overrepresented compared with the noisy, subtle, or artifact-prone images encountered in routine clinical practice. Consequently, although our results demonstrated strong theoretical performance, the model’s generalizability to real-world, multicenter clinical environments requires further validation using external, heterogeneous cohorts to ensure robustness against domain shifts.
Experimental environment and hyperparameter configuration
Experiments were conducted on a high-performance computing platform using an NVIDIA RTX 4090 graphics processing unit (GPU; 24 GB VRAM). The models were trained using PyTorch 1.10.0 and CUDA 11.3, with input images resized to 640 × 640 pixels. Training employed a batch size of 64 over 200 epochs, using an initial learning rate of 0.01, momentum of 0.937, and weight decay of 0.0005. This configuration ensured stable convergence and maintained training efficiency.
Evaluation metrics
To align with our clinical objectives, we employed specific metrics. Recall (R) was prioritized to minimize missed diagnoses for patient safety, whereas precision (P) was measured to ensure diagnostic reliability. The mean average precision at IoU threshold 0.5 (mAP@0.5) served as the comprehensive indicator of accuracy. Furthermore, frames per second, Giga Floating-point Operations Per Second (GFLOPs), and parameters were utilized to objectively measure the model’s computational efficiency and architectural complexity.
In this study, model performance was evaluated using F1-score, P, R, average precision (AP), and mean average precision (mAP) as the primary metrics. In addition, the number of parameters (Parameters) was reported to assess the computational complexity of different models. The corresponding formulations are expressed as follows:
Experimental analyses
Algorithm comparison results
To evaluate the proposed method, we compared it with leading lightweight detectors, including YOLOv8n–v12n, RT-DETR-r18,
33
YOLOv5m-ESA,
34
and SCC-YOLO.
35
As demonstrated in Table 1, our method achieved state-of-the-art accuracy with 93.2% P, 88.4% R, and 94.1%
Comparison of results obtained from multiple models.

FPS: frames per second; GFLOPs: Giga Floating-point Operations Per Second; mAP@0.5: mean average precision at IoU threshold 0.5; YOLOv12n: You Only Look Once version 12.
Table 2 presents a comparison of AP of different detectors across three tumor types. Although all the models performed well for meningioma, our method achieved competitive results, slightly inferior to those of RT-DETR-r18 but surpassing those of most YOLO variants. Notably, in the more challenging glioma category, our approach attained the highest AP, outperforming RT-DETR-r18 by 3.2% and YOLOv11n by 1.3%. Similarly, for pituitary tumors, our model exhibited the best performance, exceeding YOLOv8n and RT-DETR-r18 by 2.5% and 1.6%, respectively. These results validate the robustness and generalizability of our method, particularly in detecting complex tumor types with data constraints. Table 3 presents a comparison of P achieved by different detectors across the three tumor categories. Our method achieved the highest P in all classes, reaching 97.1% for meningioma, 84.9% for glioma, and 97.6% for pituitary tumors. Notably, in the challenging glioma category, it surpassed YOLOv8n and RT-DETR-r18 by 2.7% and 0.6%, respectively. These results validate the consistent superiority and diagnostic reliability of our approach, particularly in P-sensitive medical detection tasks. Table 4 presents a comparison of R rates across the three tumor categories. Our method achieved the highest R for glioma and pituitary tumor cases and maintained competitive performance for meningioma cases. It surpassed the existing models by 1.3%–1.8% in glioma cases and 2.3%–3.3% in pituitary tumor cases, demonstrating enhanced sensitivity and robustness, particularly in challenging detection scenarios.
Comparison of the average precision (AP, %) of multiple algorithms.

YOLOv12n: You Only Look Once version 12.
Comparison of the accuracy of multiple algorithms (precision (P), %).
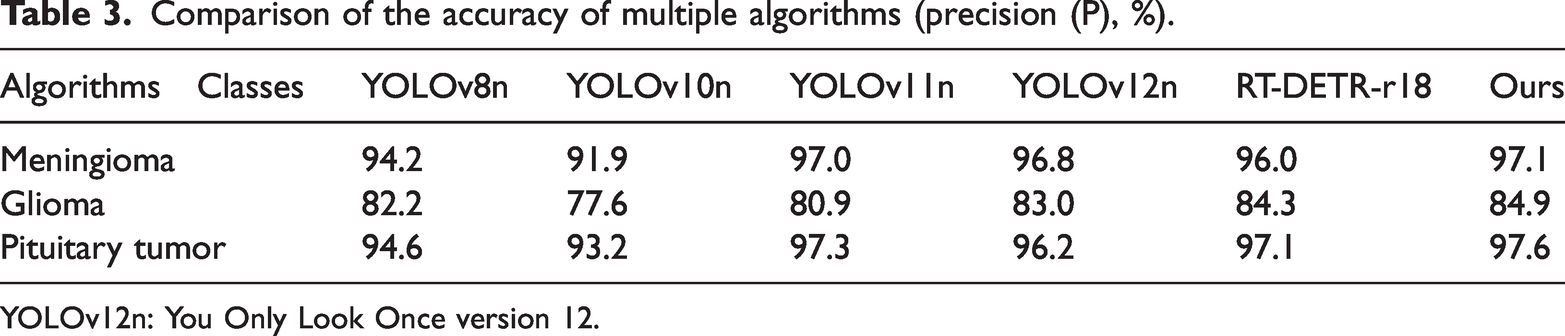
YOLOv12n: You Only Look Once version 12.
Comparisons of various algorithms in terms of recall (R, %) rate.

YOLOv12n: You Only Look Once version 12.
Visualization of results
To visually benchmark our method against baselines (YOLOv8n–v12n and RT-DETR-r18), we performed the following multidimensional analysis:
Confusion matrices. These visualized class-wise accuracy and background misclassification rates across all models (Figure 5).
Normalized confusion matrices for (a) YOLOv8n, (b) YOLOv10n, (c) YOLOv11n, (d) YOLOv12n, (e) RT-DETR-r18, and (f) ours. YOLOv12n: You Only Look Once version 12. F1-confidence curves. These demonstrated prediction stability and robustness under varying confidence thresholds (Figure 6).
F1-Confidence curves: (a) YOLOv8n, (b) YOLOv10n, (c) YOLOv11n, (d) YOLOv12n, (e) RT-DETR-r18, and (f) ours. YOLOv12n: You Only Look Once version 12. Qualitative visualization. These intuitively demonstrated our method’s superiority in handling blurred boundaries and small lesions compared with those of baselines (Figure 7).
Visual comparison of detection results on sample MRI scans: (a) YOLOv8n, (b) YOLOv10n, (c) YOLOv11n, (d) YOLOv12n, (e) RT-DETR-r18, and (f) ours. MRI: magnetic resonance imaging; YOLOv12n: You Only Look Once version 12. Continued.




To comprehensively evaluate classification performance, normalized confusion matrices were generated on the test set (Figure 5). These matrices visually illustrated per-class accuracy and misclassification trends, supplementing conventional metrics such as P and R. All models exhibited strong performance for well-defined tumor categories such as meningioma and pituitary tumors, with R rates consistently >0.95. The proposed method achieved the highest R for meningioma cases at 0.99, followed by RT-DETR-r18 at 0.98, and YOLO-series models ranged between 0.95 and 0.96. Substantial performance differences emerged in the challenging glioma category. YOLOv8n and YOLOv10n exhibited limited capability. Although YOLOv11n and YOLOv12n improved R to 0.86 and 0.84, respectively, background confusion remained notable. RT-DETR-r18 reached 0.83 R with 0.17 background confusion. The proposed method outperformed all others, achieving 0.88 R with only 0.12 background confusion, approximately halving the error rate of the weakest models. For pituitary tumors, all models exhibited comparable performances, with the proposed method attaining 0.96 R and minimal background confusion. Overall, the proposed method demonstrated superior robustness, significantly reducing missed detections and aligning with the quantitative results (Tables 2 to 4), thereby confirming its enhanced discriminative capability in complex diagnostic scenarios.
To evaluate prediction reliability, we analyzed the F1-score against the confidence threshold (Figure 6), which reflects the P–R trade-off and the model’s capacity for high-confidence true positives. Comparative analysis revealed notable performance differences. For meningioma and pituitary tumor cases, all models exhibited strong performances, indicating that these well-defined tumors were easier to identify. In contrast, glioma remained the most challenging category, with F1-scores typically ranging from 0.7 to 0.8. YOLOv8n and YOLOv10n exhibited relatively weak performances, with YOLOv10n achieving a peak F1-score of only 0.86 and requiring lower confidence thresholds, suggesting limited robustness. YOLOv11n and YOLOv12n demonstrated clear improvement with smoother curves across confidence ranges. RT-DETR-r18 matched YOLOv11n’s peak but declined sharply at high confidence, indicating reduced stability. The proposed method achieved superior overall performance, attaining the highest F1-score with an optimal confidence threshold of 0.744. This enabled the maintenance of excellent performance at higher confidence levels and minimized the false detection risk. Our method showed particular improvement in glioma detection, effectively elevating and stabilizing the overall curve. These results demonstrate the superiority of the proposed method in terms of accuracy, robustness, and its ability to handle challenging categories, making it highly suitable for medical imaging tasks that require high reliability.
Figure 7 demonstrates the practical advantages of our method through comparative visualization of sample MRI scans. Compared with contemporary models, our approach consistently produced bounding boxes with higher confidence and superior anatomical alignment. Visual comparisons revealed clear performance differences. YOLOv8n coarsely localized the meningioma but showed low confidence and misalignment in ambiguous cases. YOLOv10n exhibited slightly better performance but remained sensitive to ill-defined margins, with the confidence level dropping to 0.67 in challenging gliomas. YOLOv11n demonstrated stable perfrmance in glioma cases but struggled with pituitary tumor cases, often deviating from the true boundaries. YOLOv12n exhibited multitarget recognition ability; however, it demonstrated inconsistent confidence levels, scoring only 0.46 for pituitary tumor cases. RT-DETR-r18 outperformed most YOLO variants, achieving confidence levels of 0.89 and 0.86 for meningioma and glioma cases, respectively; however, its performance for small pituitary tumors remained poor. In contrast, the proposed method achieved the highest confidence levels across all categories (0.93, 0.90, and 0.90) and produced bounding boxes that align closely with lesion boundaries. It almost eliminated false positives and negatives, demonstrating strong robustness particularly in detecting small tumors within complex anatomical contexts. These results confirm the practical superiority of our approach in clinical detection accuracy and reliability.
Discussion
In recent studies on MRI-based brain tumor detection, the central focus has been on improving the detection accuracy and preserving computational efficiency. The YOLOv11n architecture, in particular, has shown considerable promise in clinical imaging applications. Chen et al.
36
enhanced YOLOv11n with GhostConv, Online Convolutional Re-parameterization (OREPA), and EMA mechanisms, whereas Han et al.
37
integrated Shuffle3D and dual-channel attention modules. Both studies reported increased mAP and R with reduced computational overhead. In a similar direction, our model prioritizes R stability and robustness, especially for small or low-contrast tumors, addressing key factors in reducing missed diagnoses in clinical practice. Integrated detection–segmentation frameworks represent another significant advancement. Saranya and Praveena
38
combined YOLOv5 with a fully convolutional network for real-time cascade segmentation, whereas Chourib
39
and Xiong et al.
40
extended YOLOv11n using transfer learning and boundary-sensitive mechanisms, achieving
Conclusion
This study presented an enhanced brain tumor MRI scan detection framework based on YOLOv12n. By integrating the A2C2f-DFFN and C2TSSA-DYT modules along with the DySample operator, the proposed model systematically enhanced its capabilities in feature modeling, interaction, and reconstruction. Our results demonstrated that the method outperformed the original YOLOv12n and several state-of-the-art lightweight models in terms of P, R, and
Nevertheless, this study has certain limitations. For instance, detection performance under extremely low-contrast conditions requires further improvement, and the current work is confined to two-dimensional (2D) MRI scans without exploration of three-dimensional (3D) or multimodal data. Future research should focus on extending the model to 3D and multimodal scenarios, thereby enhancing its robustness under complex imaging conditions and promoting its practical application and validation within clinical workflows.
Footnotes
Acknowledgments
The authors would like to acknowledge the creators of the public Kaggle datasets 32 used in this study. We also acknowledge the use of AI-powered language editing tools to enhance the readability and clarity of the manuscript.
Author contributions
Ronghui Zheng: Conceptualization, methodology, data collection, formal analysis, writing-original draft, and software use. Shanshan Cai: Validation, project administration, data curation, visualization, writing-review & editing, and supervision.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.








