Abstract
Objectives
Genome-wide association studies have identified FAM13A as a gene linked to chronic obstructive pulmonary disease. Several single-nucleotide polymorphisms—notably, rs7671167, rs2869967, rs2869966, and rs17014601—are associated with chronic obstructive pulmonary disease susceptibility and spirometric indices. This study aimed to investigate the association between FAM13A single-nucleotide polymorphisms and spirometric indices in chronic obstructive pulmonary disease and non-chronic obstructive pulmonary disease patients and assess the correlation of these single-nucleotide polymorphisms with pulmonary ventilation function.
Methods
This study included 160 participants (80 chronic obstructive pulmonary disease patients and 80 controls) who underwent clinical evaluation and spirometry at Can Tho University of Medicine and Pharmacy Hospital from March 2021 to February 2023.
Results
The mean patient age was 66.3 ± 7.9 years. Significant differences in allele frequencies of the studied single-nucleotide polymorphisms were observed between the two groups. TT genotypes at rs7671167 and rs17014601 and CT genotypes at rs7671167, rs2869967, and rs2869966 were associated with significant differences in forced vital capacity, forced expiratory volume in 1 second, and Tiffeneau index.
Conclusion
The CT genotype at rs7671167, rs2869967, and rs2869966 is significantly associated with altered spirometric parameters, suggesting a role in chronic obstructive pulmonary disease–related lung function impairment.
Introduction
The prevalence and mortality of chronic obstructive pulmonary disease (COPD) have been steadily increasing. According to the World Health Organization, approximately 600 million individuals worldwide are affected by COPD. In 1990, COPD was the sixth leading cause of mortality, and it is projected to become the third leading cause by 2030. 1 According to the Asia-Pacific Respiratory Society, the prevalence of COPD in Vietnam was 6.7%, the highest among 12 countries in the region. The increasing prevalence of COPD has contributed to a growing burden on the national healthcare system, underscoring the need for greater attention to COPD management to support social welfare stability. 2 With the rapid advancement of global medicine, the diagnosis and treatment of COPD must adopt a multidimensional approach, emphasizing early detection, prevention, and phenotype-based or targeted therapy. In 2019, the Global Initiative for Chronic Obstructive Lung Disease (GOLD) identified genetics as one of the risk factors for COPD. The interaction between genetic factors and environmental influences necessitates the investigation of genotype associations with COPD risk factors and pulmonary function decline. 1
Family with sequence similarity 13 member A (FAM13A) is a protein-coding gene that interacts with the mediator protein sirtuin-1 to regulate the expression of carnitine palmitoyltransferase 1A (CPT1A). Upregulation of CPT1A promotes fatty acid oxidation, leading to increased production of reactive oxygen species. This oxidative stress induces apoptosis in pulmonary epithelial cells, ultimately contributing to irreversible alveolar damage characteristic of COPD. Numerous studies have emphasized the pivotal role of FAM13A in the pathogenesis of various respiratory diseases, particularly COPD; moreover, single-nucleotide polymorphisms (SNPs) of the FAM13A gene, namely, rs7671167, rs2869967, rs2869966, and rs17014601, have been associated with COPD susceptibility and several spirometric indices used to assess pulmonary ventilation function.2,3 Large-scale statistical analyses with genome-wide association studies have identified FAM13A as one of the genes linked to COPD.4–8 However, data on FAM13A variants in the Vietnamese population with COPD remain limited. Therefore, this study aimed to investigate the association between FAM13A SNPs and selected spirometric indices in individuals with and without COPD as well as the correlation between FAM13A SNPs and pulmonary ventilation characteristics in COPD patients.
Materials and methods
Study population and design
We conducted a cross-sectional study among patients who underwent clinical examination and spirometry testing at the University of Medicine and Pharmacy Hospital, Can Tho, from March 2021 to February 2023. The reporting of this study conforms to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines. 9
Selection criteria for the disease group were patients diagnosed with COPD according to the GOLD criteria and those who consented to participate in the study. 1 COPD diagnosis based on the GOLD criteria was considered in cases presenting with progressively worsening dyspnea over time, particularly during exertion; chronic cough, either intermittent or dry; and chronic sputum production as well as in those with a history of exposure to various risk factors, such as smoking (cigarettes or tobacco), smoke (household smoke, fuel combustion, occupational dust, and organic and inorganic particulates), and toxic fumes and gases. A confirmed COPD diagnosis was established when the postbronchodilator forced expiratory volume in 1 second/forced vital capacity (FEV1/FVC) ratio was <70%. Additionally, the control group consisted of patients presenting with respiratory conditions who were confirmed not to have COPD according to GOLD criteria and those who consented to participate in the study. The control group was matched with the disease group in terms of age and sex.
Exclusion criteria included patients experiencing an acute exacerbation of COPD or those diagnosed with other acute medical conditions.
Sample size
The sample size was calculated using the pilot study method on four polymorphisms. Subsequently, the polymorphism with the largest calculated sample size was selected for the main study. The sample size formula is as follows:
Analysis and determination of the heterozygous CT genotype frequency of rs17014601 were conducted using 30 disease group samples and 30 control group samples to establish the study sample size. The results indicated p1 = 46.7% in the disease group and p2 = 26.7% in the control group, yielding a calculated sample size of n = 80. Consequently, the study was conducted with 80 participants in the disease group and 80 in the control group. To ensure the representativeness of the study population, after obtaining the SNP genotyping results, we performed Hardy–Weinberg equilibrium testing to confirm that the selected sample met the representativeness criteria for genetic variation within the study population.
Study variable and data collection method
The study participants underwent surveys, interviews, medical history assessments, and clinical examinations, providing variables related to anthropometric and clinical characteristics, including: age (years), sex (male or female), height (cm), weight (kg), body mass index (BMI) (kg/m2), pulse rate (beats per minute (bpm)), systolic blood pressure (mmHg), diastolic blood pressure (mmHg), history of tobacco smoke exposure (still smoking or quit less than 5 years), history of pulmonary tuberculosis, and clinical symptoms (cough, sputum production, wheezing, dyspnea, high-pitched rhonchi, low-pitched rhonchi, crackles, moist rales, and decreased breath sounds). 1 Subsequently, the study participants underwent spirometry testing at the Department of Functional Examination of the Can Tho University of Medicine and Pharmacy Hospital using the Ferraris Koko spirometer. This assessment was used to collect spirometric indices both before and after bronchodilator responsiveness testing, including vital capacity (VC) (L, %), FVC (L, %), FEV1 (L, %), peak expiratory flow (PEF) (L, %), FEV1/FVC (%), and forced expiratory flow between 25% and 75% of forced vital capacity (FEF25) (L, %). Following spirometry testing, an evaluation of pulmonary ventilation dysfunction was conducted, and COPD patients were categorized into restrictive ventilatory dysfunction, obstructive ventilatory dysfunction, mixed ventilatory dysfunction, small airway obstruction, and severe airway obstruction subgroups according to the GOLD criteria. 1
Overall, 2–3 mL of peripheral blood was collected from all study participants for SNP analysis of the FAM13A gene. Blood samples were stored at 4°C for 1 week before DNA extraction and subsequently preserved at −30°C for storage and further DNA extraction if needed (polymerase chain reaction extraction protocol is detailed in Appendix 1). Study participants were classified into CC, TT, or CT genotypes at SNPs rs7671167, rs2869967, rs2869966, and rs17014601 of the FAM13A gene. The allele frequencies of C and T for each SNP were determined in the study groups. The association between COPD risk and specific alleles and genotypes of each SNP was analyzed between the disease and control groups. Additionally, correlations between the alleles and genotypes of the studied SNPs and the obstructive ventilatory dysfunction subgroup among COPD patients were assessed according to the GOLD 2019 criteria. Further analyses examined genotype associations of SNPs with different types of ventilatory dysfunction and spirometric indices.
Bias-controlled methods and data analysis
To minimize potential systematic errors in the research process, the following measures were implemented: 1. The questionnaire was piloted to refine its application and was administered by a single individual throughout the data collection phase. 2. All data collection instruments were standardized, uniformly used, and calibrated before each research session. 3. Laboratory tests were conducted by specialized physicians and technicians following validated protocols, with quality control and cross-checking procedures ensuring accuracy and consistency in test results.
Data were entered and analyzed using biomedical statistical methods via SPSS 20.0 software. Continuous quantitative variables were presented as mean ± standard deviation if normally distributed, while non-normally distributed variables were expressed as median or range. Qualitative variables were reported as frequency and percentage. The t-test and analysis of variance were applied to evaluate statistically significant differences in mean values between the two groups. The chi-square test was used for proportion comparisons. The risk of COPD among different genotypes and alleles of SNPs was assessed using odds ratios (OR) with 95% confidence intervals (CIs). The recessive inheritance analysis model for rs7671167, rs2869967, rs2869966, and rs17014601 was CC vs. TT + CT, and the dominant inheritance analysis model was TT vs. CC + CT. Multivariate logistic regression was employed to analyze associations between SNPs, spirometric indices, and ventilatory dysfunction types. Statistical significance was defined as p < 0.05.
Ethical issue
This study was approved by the Biomedical Ethics Committee of the Can Tho University of Medicine and Pharmacy, under certification number 21.550/PCT-HĐĐĐ, dated 30 March 2021. Our study followed the Helsinki Declaration. The research was conducted after obtaining informed consent from all participants. Each participant was thoroughly informed about the study’s objectives and procedures. The medical tests performed were necessary for diagnosis, treatment, and patient monitoring. Patients’ personal information was kept strictly confidential, and deidentified data were used exclusively for research purposes. All collected data have been assessed and approved for use by the review board. Furthermore, no interventions were administered that could affect patients’ health or psychological treatment, ensuring that the study adhered to ethical standards without violations.
Results
Baseline characteristics
The study included 160 participants, comprising 80 individuals in the disease group and 80 in the control group (Figure 1). The overall mean height for both groups was 162.3 ± 5.8 cm, with the disease group exhibiting a higher value than the control group. The mean weight and BMI for the two study groups were 59.3 ± 9.6 kg and 22.5 ± 3.5 kg/m2, respectively, with the disease group showing lower values than the control group (Table 1).
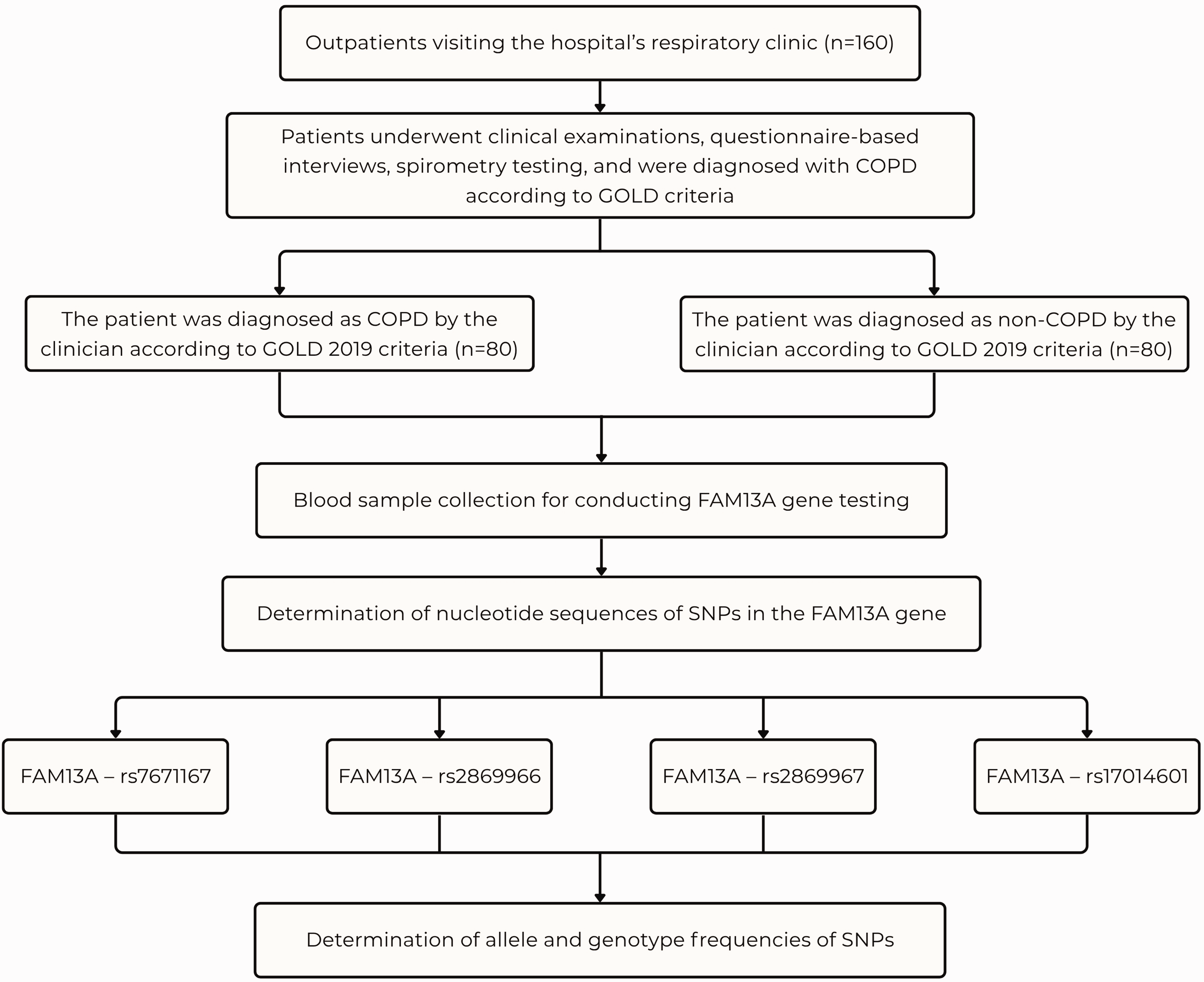
Study population flowchart.
Baseline characteristics of the population.
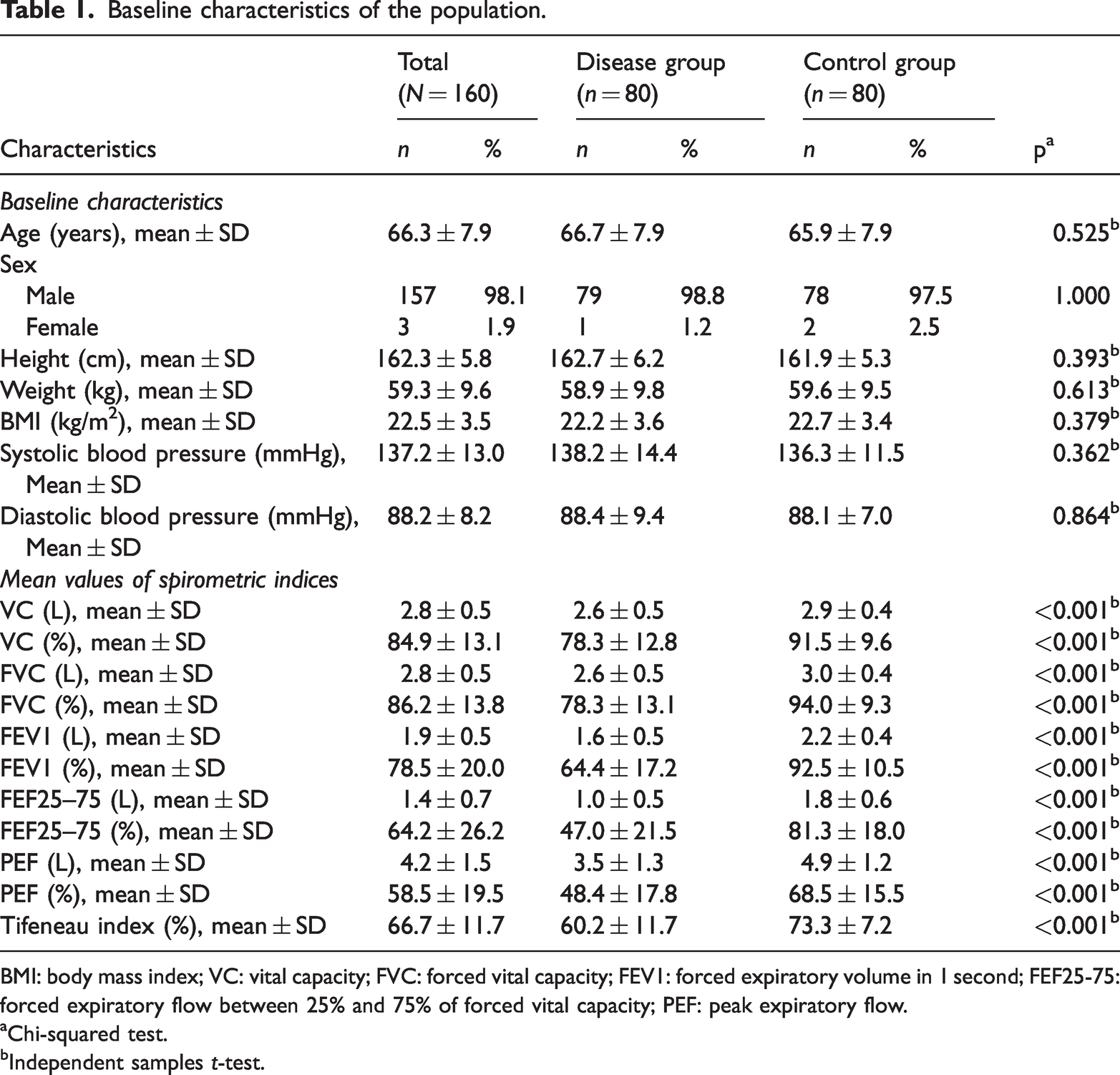
BMI: body mass index; VC: vital capacity; FVC: forced vital capacity; FEV1: forced expiratory volume in 1 second; FEF25-75: forced expiratory flow between 25% and 75% of forced vital capacity; PEF: peak expiratory flow.
Chi-squared test.
Independent samples t-test.
The allele frequencies of C for the SNPs rs7671167, rs2869967, rs2869966, and rs17014601 in the disease and control groups were 45%, 50.6%, 49.4%, and 31.9% and 47.5%, 47.5%, 52.5%, and 21.3%, respectively. The allele frequencies of T for these SNPs in the disease and control groups were 55%, 49.4%, 50.6%, and 68.1% and 52.5%, 52.5%, 47.5%, and 78.8%, respectively. For SNP rs17014601, the distribution of alleles T and C differed significantly between the disease and control groups. Notably, rs17014601 was significantly associated with COPD, with an OR of 2.43 (95% CI: 1.23–4.78, p = 0.010) for CT vs. TT genotype and OR of 2.26 (95% CI: 1.20–4.29, p = 0.012) for the dominant model (Table 2). Regarding SNP rs7671167, the heterozygous CT genotype had the highest prevalence in both groups. Similarly, for SNPs rs2869967 and rs2869966, the CT genotype was the most common in both groups. For SNP rs17014601, the CT genotype was the most prevalent in the disease group (46.3%), whereas the TT genotype was the most frequent in the control group (65%) (Table 2).
Allele and genotype frequencies of the population.
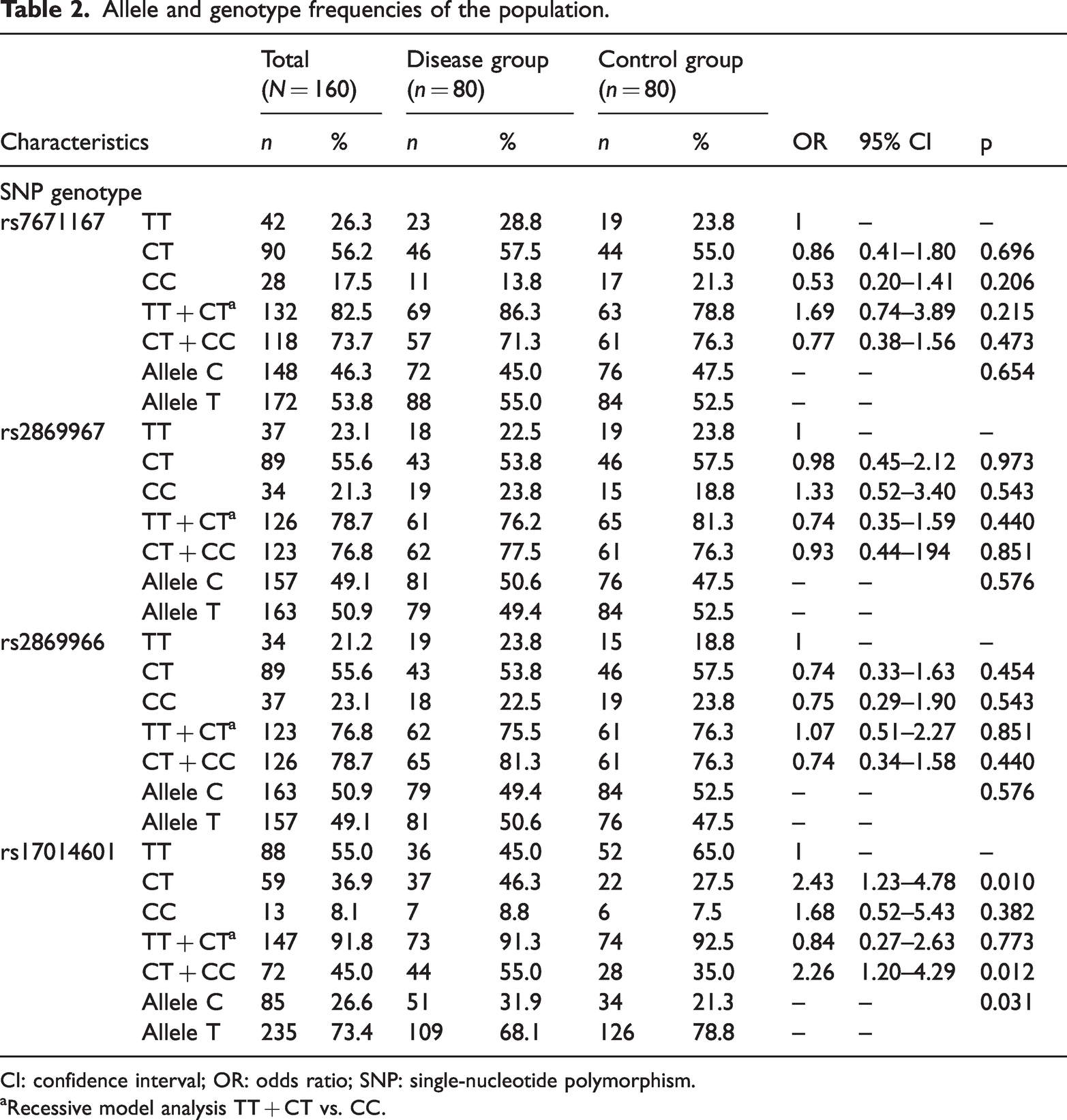
CI: confidence interval; OR: odds ratio; SNP: single-nucleotide polymorphism.
Recessive model analysis TT + CT vs. CC.
Association between SNPs of the FAM13A gene and pulmonary ventilation characteristics in COPD patients
Across the obstruction severity subgroups in COPD patients, the CT genotype exhibited the highest prevalence among the studied SNPs, except for rs17014601, where the TT genotype was most frequent in the moderate and extremely severe groups. No statistically significant differences were observed in the genotype distribution of FAM13A SNPs among COPD patients (Table 3).
Genotypic distribution across obstruction severity subgroups in COPD patients based on GOLD criteria.
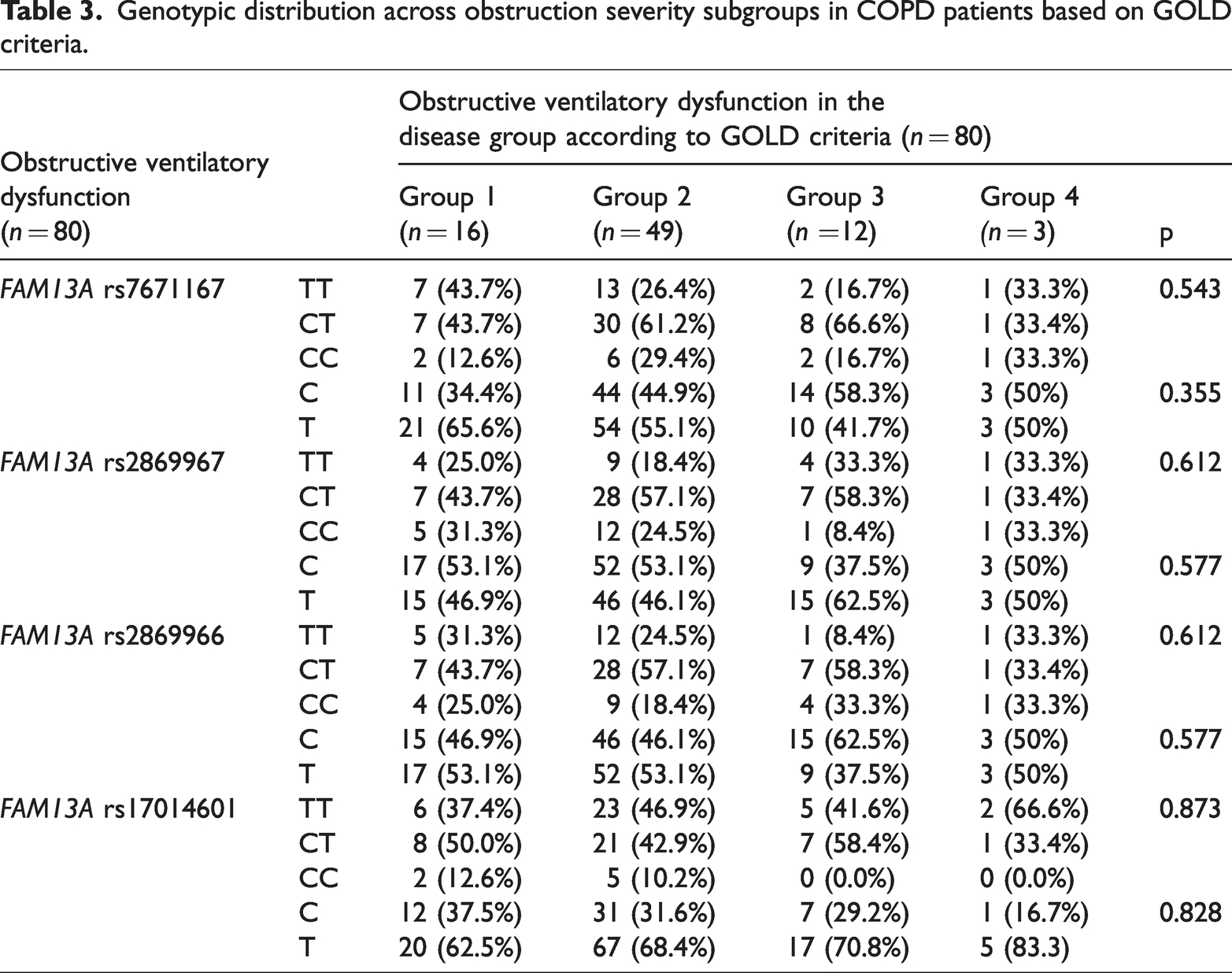
The mean FVC values across genotypes within each SNP in both study groups did not show statistically significant differences. However, a statistically significant difference was observed between the two study groups when analyzing mean FVC values among individuals with TT genotype at SNPs rs7671167 and rs17014601. Similarly, a significant association was identified between the heterozygous CT genotype at SNPs rs7671167, rs2869967, and rs2869966 and mean FVC values in both the disease and control groups (Table 4).
Association between SNP genotypes and mean FVC values.

FVC: forced vital capacity; SNP: single-nucleotide polymorphism.
Analysis within groups.
Analysis between the control and disease groups.
The mean FEV1 values across genotypes within each SNP in both study groups did not show statistically significant differences. However, for rs7671167, a statistically significant difference in mean FEV1 values was observed between the disease and control groups across all genotypes. Similarly, individuals in the disease group carrying the CT genotype of polymorphisms rs2869967 and rs2869966 exhibited significantly lower mean FEV1 values than those in the control group (Table 5). For rs17014601, a statistically significant difference in mean FEV1 values was observed between the two study groups for TT and CT genotypes, with p-values of 0.004 and 0.002, respectively (Table 5).
Association between SNP genotypes and mean FEV1 values.

FEV1: forced expiratory volume in 1 second; SNP: single-nucleotide polymorphism.
aAnalysis within groups.
bAnalysis between the control and disease groups.
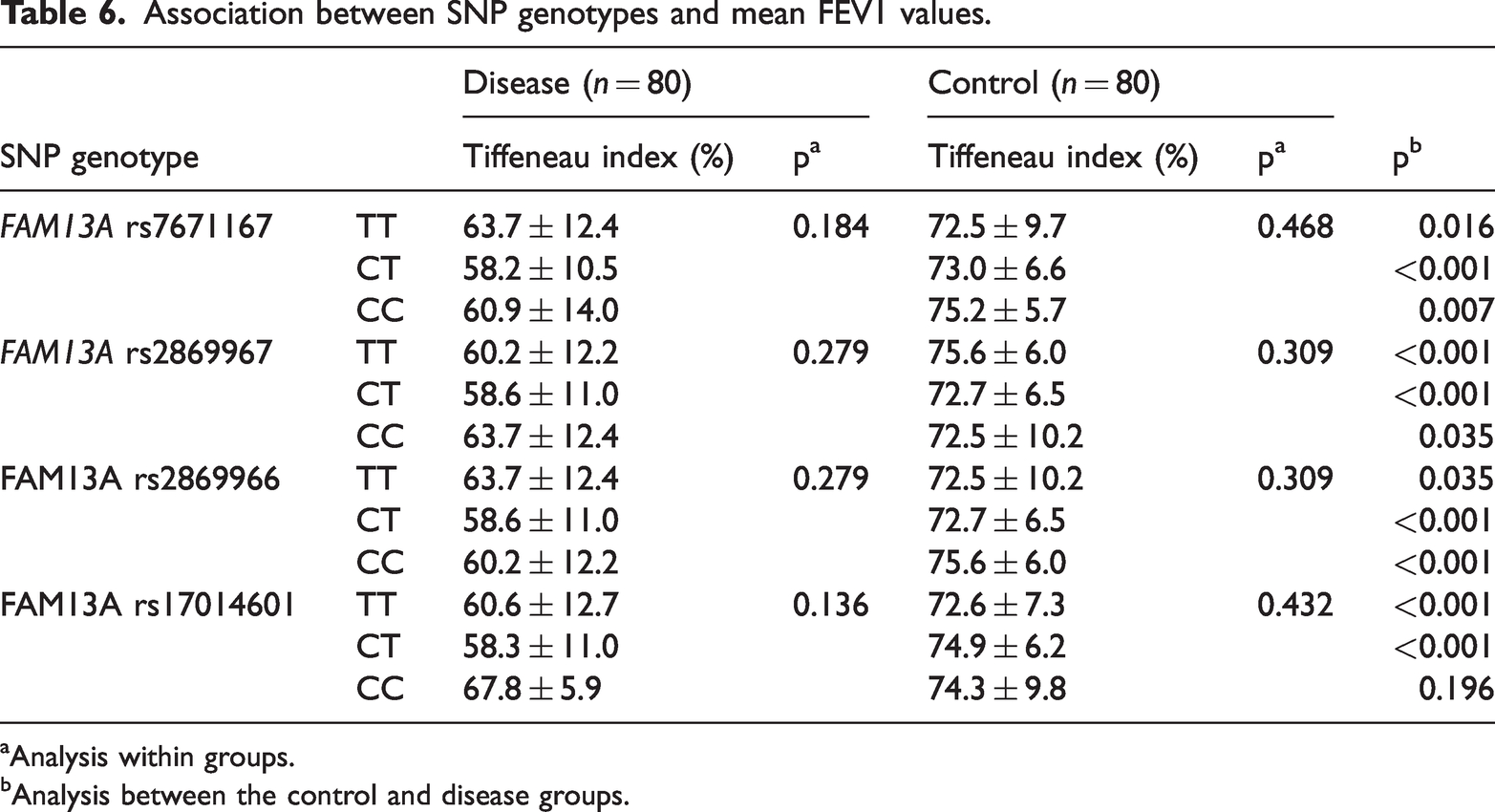
No statistically significant difference was observed in the mean Tiffeneau index values across genotypes for each SNP within any study group. However, the disease group consistently exhibited statistically lower mean Tiffeneau index values than the control group across most SNP genotypes, with the sole exception of the CC genotype of rs17014601 (Table 6).
Association between SNP genotypes and mean FEV1 values.
Analysis within groups.
Analysis between the control and disease groups.
Discussion
Baseline characteristics
Our study participants’ characteristics are consistent with findings reported by Trung in 2021, based on a study of 173 COPD patients. 10 The sex distribution in our study exhibited a notable imbalance between male and female participants, with proportions of 98.1% and 1.9%, respectively. However, this ratio aligns with findings from previous studies conducted at the Can Tho University of Medicine and Pharmacy Hospital, including the research by Pham et al. 11 and Thu et al. 12 The overall mean height and weight of the two study groups were 162.3 ± 5.8 cm and 59.3 ± 9.6 kg, respectively. The disease group exhibited a higher mean height than the control group, whereas the mean weight was lower. Our study anthropometric values are consistent with those of previous studies on Vietnamese populations, particularly BMI values. The overall mean values of heart rate, systolic blood pressure, and diastolic blood pressure among the 160 study participants were 86.2 ± 10.9 bpm, 137.2 ± 13.0 mmHg, and 88.2 ± 8.2 mmHg, respectively. When analyzed by study group, the disease group exhibited higher mean heart rate, systolic blood pressure, and diastolic blood pressure than the control group, although the differences were not statistically significant. In the disease group, the mean systolic blood pressure was 138.2 ± 14.4 mmHg, a value comparable to the findings of Kraen et al., 13 who reported an SBP of 138 ± 18 mmHg. This consistency is reasonable, as both studies were conducted on COPD patients and exhibited similar sex distributions. Additionally, most study participants had a mean age above 60 years, making the observed systolic and diastolic blood pressure values consistent with age-related physiological changes in older individuals.
The values of VC and FVC were equivalent, and both showed a slight reduction below 80% of the predicted values. Additionally, the mean FEV1 value decreased (64.4%), and the Tiffeneau index fell below the 70% threshold. These spirometric characteristics align with the GOLD criteria for COPD diagnosis. In comparison with international studies, the study by Yan et al. 14 reported relatively similar FVC values in the disease and control groups, a finding consistent with our study. According to Ju et al., 15 in COPD patients, the mean values of VC and FVC were both reduced, with FVC decreasing to below 80% and showing a greater reduction than VC. Previous studies have suggested that in cases of severe airway obstruction, excessive alveolar air trapping occurs, leading to an increase in residual volume, which consequently results in a decline in VC and FVC below the 80% threshold. 16
Association between FAM13A SNPs and selected spirometric parameters in patients with and without COPD
According to GOLD criteria, a definitive diagnosis of COPD requires the assessment of chronic airway obstruction to evaluate risk and implement effective management strategies. Therefore, in the disease group, we conducted allele and genotype analysis based on obstruction severity, dividing participants into four subgroups: mild (group 1), moderate (group 2), severe (group 3), and extremely severe (group 4) (Table 3). As shown in Table 3, the frequency of allele T consistently exceeded that of allele C in COPD severity groups 1 and 2 at SNPs rs7671167 and rs17014601. Conversely, in group 3, allele C was more prevalent than allele T at rs7671167 and rs2869966. This indicates a substantial variation in allele C and T distribution across different obstruction severity levels in COPD patients, except for group 4, where the sample size was too small for meaningful analysis. However, the allele frequency distribution across SNPs in COPD subgroups did not exhibit statistically significant differences. We conducted an analysis of genotype frequencies across different levels of airway obstruction in COPD patients and observed that for all three SNPs rs7671167, rs2869967, and rs2869966, the heterozygous CT genotype showed consistently higher frequency than the two homozygous genotypes at all obstruction severity levels. However, in the moderate COPD group (group 2) at rs17014601, the frequency percentages of the TT and CT genotypes were relatively similar at 46.9% and 42.9%, respectively. This differs from the trends observed in other SNPs analyzed at the same obstruction severity level and aligns with the overall genotype frequency distribution among the 80 COPD patients. Thus, across the COPD obstruction severity groups, the CT genotype exhibited the highest frequency among the studied SNPs, except for rs17014601, where the TT genotype was most prevalent in the moderate and extremely severe groups. No statistically significant differences were observed in the genotype distribution of FAM13A SNPs among COPD patients.
Analysis of the heterozygous CT genotype at SNPs rs7671167, rs2869967, and rs2869966 revealed a statistically significant association with mean FVC, Tiffeneau index, and FEV1 values in both the disease and control groups (Tables 4 to 6). This finding supports the observation that pulmonary capacity is reduced in the disease group compared with that in the control group for each genotype in these SNPs, aligning with prior research by Young et al. 17 Similarly, Hancock et al. 7 demonstrated that rs2869967 is one of the SNPs associated with spirometric parameters in COPD patients. 17 Individuals in the disease group carrying the CT genotype of polymorphisms rs2869967 and rs2869966 exhibited significantly lower mean FEV1 values than those in the control group. Similarly, Ziółkowska-Suchanek et al. 18 reported lower mean FEV1 values in COPD patients across all three genotypes CC, CT, and TT. These findings suggest that the decline in FEV1 function within the disease group is associated with certain SNPs of the FAM13A gene. Their study outcome was the statistically significant reduction in mean FEV1/FVC values in the disease group compared with that in the control group across most SNP genotypes, except for the CC genotype of rs17014601, a trend consistent with the findings reported by Young et al. 17 Our findings precisely align with those of Xie et al., 19 particularly concerning the association between rs7671167 and pulmonary function decline in COPD patients. Critically, both studies indicated that the CC genotype at rs7671167 is linked to a more pronounced decrease in FEV1 compared with CT and TT genotypes. This remarkable consistency independently reinforces the evidence regarding rs7671167’s crucial role in COPD progression. Thus, through the analysis of genotype associations within each SNP and selected mean spirometric parameters, we identified statistically significant differences between the disease and control groups for certain genotypes. These findings provide valuable insights for further research on genetic variations and pulmonary function, particularly in COPD patients in Vietnam.
Strengths and weaknesses of the study
Our study followed a specifically designed sample collection process with defined inclusion and exclusion criteria, and the methods were clearly described and reproducible. The novelty of our study is that the outcome provides a reference for the prevalence of FAM13A gene polymorphisms. Therefore, our study data and outcome could be strong evidence for further study in this field. However, our study had a small sample size, mainly due to the limitation of research conditions, and the study was evaluated only in one hospital, which might lead to bias in national and area baseline characteristics. A multicenter study with a larger sample size is required to represent the study population.
Conclusion
Through the study of 160 participants, we observed a statistically significant difference between the disease and control groups when analyzing mean FVC values in individuals with the TT genotype at SNPs rs7671167 and rs17014601. Notably, the heterozygous CT genotype at SNPs rs7671167, rs2869967, and rs2869966 was found to be significantly associated with mean FVC values in both the disease and control groups. For rs7671167, a statistically significant difference in mean FEV1 values was observed across all genotypes between the disease and control groups. Individuals in the disease group carrying the CT genotype of polymorphisms rs2869967 and rs2869966 exhibited significantly lower mean FEV1 values than those in the control group. For rs17014601, a statistically significant difference in mean FEV1 values was observed between the two study groups for the TT and CT genotypes, with p-values of 0.004 and 0.002, respectively.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605251372069 - Supplemental material for Association of FAM13A single-nucleotide polymorphism with spirometry indices and pulmonary ventilation characteristics in patients with chronic obstructive pulmonary disease
Supplemental material, sj-pdf-1-imr-10.1177_03000605251372069 for Association of FAM13A single-nucleotide polymorphism with spirometry indices and pulmonary ventilation characteristics in patients with chronic obstructive pulmonary disease by Binh Huy Nguyen, Nhung Thi Cam Tran, Chuong Quoc Ho, Tam Thai Thanh Tran, Hong Ngoc Phan, De Van Tran, Nga Thi Ngoc Pham, Kien Trung Nguyen and Khanh Hoang Pham in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_03000605251372069 - Supplemental material for Association of FAM13A single-nucleotide polymorphism with spirometry indices and pulmonary ventilation characteristics in patients with chronic obstructive pulmonary disease
Supplemental material, sj-pdf-2-imr-10.1177_03000605251372069 for Association of FAM13A single-nucleotide polymorphism with spirometry indices and pulmonary ventilation characteristics in patients with chronic obstructive pulmonary disease by Binh Huy Nguyen, Nhung Thi Cam Tran, Chuong Quoc Ho, Tam Thai Thanh Tran, Hong Ngoc Phan, De Van Tran, Nga Thi Ngoc Pham, Kien Trung Nguyen and Khanh Hoang Pham in Journal of International Medical Research
Footnotes
Acknowledgments
We want to thank the Can Tho University of Medicine and Pharmacy for creating favorable conditions for this study to be carried out.
Authorship contribution statement
Data availability
Data will be made available on request.
Declaration of conflicting interests
None.
Ethics approval
The study was approved by the Ethics Committee in biomedical research of the Can Tho Univeristy of Medicine and Pharmacy. The study fully complies with the principles outlined in the Declaration of Helsinki, and all patients provided informed consent to participate in the study.
Funding
None.
Supplementary material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

