Abstract
Objective
Long non-coding RNAs (lncRNAs) play a crucial role in non-small cell lung cancer (NSCLC). This study aimed to investigate the novel biomarker, lncRNA RP11-58O9.2, in patients with NSCLC.
Methods
RP11-58O9.2 expression in NSCLC cells and tissues was detected by reverse transcription-quantitative polymerase chain reaction. Patient survival was analyzed in relation to RP11-58O9.2 expression levels. RP11-58O9.2 expression was knocked down and endogenous expression was verified in two NSCLC cell lines. Cell proliferation was then assessed by Cell Counting Kit-8 and colony-formation assays, and cell invasion and migration were assessed by Transwell and wound-healing assays, respectively. In vivo experiments were performed in mice, and the combination of RP11-58O9.2 and miR-6749-3p was predicted by miRanda.
Results
RP11-58O9.2 was highly expressed in NSCLC cell lines and tissues, and was associated with advanced stage, lymphatic metastasis, and differentiation group. High RP11-58O9.2 levels were also associated with shorter survival. RP11-58O9.2 knockdown inhibited the proliferation, invasion, and migration of lung cancer cells, and tumor growth in mouse xenografts in vivo. RP11-58O9.2 may target and regulate miR-6749-3p.
Conclusions
LncRNA RP11-58O9.2 is associated with NSCLC prognosis and promotes lung cancer progression. Further studies are needed to investigate the mechanisms and the regulatory association between RP11-58O9.2 and miR-6749-3p.
Keywords
Introduction
Lung cancer (LC) is a common cancer worldwide, and its mortality rate remains high. 1 LC comprises non-small cell LC (NSCLC) and small cell LC. NSCLC accounts for approximately 85% to 90% of all LCs, and can be divided into three categories: adenocarcinoma, squamous cell carcinoma, and large cell carcinoma. 2 In addition to traditional non-surgical methods, biological targeted therapy has become a popular treatment for LC in the past decade. Nevertheless, the 5-year survival rate of NSCLC remains low, because of atypical early symptoms in some patients, difficult diagnosis, and easy recurrence after treatment.3,4 Studies are therefore needed to identify novel molecular biomarkers and therapies to facilitate personalized treatment and predict prognosis.
Long non-coding RNAs (lncRNAs) are RNAs longer than 200 nucleotides, which function in multi-level regulatory gene expression but do not encode proteins. 5 LncRNAs usually exist in the nucleus or cytoplasm, and have a wide range of biological and cellular functions.6–8 LncRNAs have recently been shown to act as crucial regulators in gene expression networks by controlling nuclear structure and transcription and modulating mRNA stability in the cytoplasm, and by translation and post-transformation modification.9–11 For example, lncRNA-RAB11B-AS1 can promote LC development by regulating RAB11B expression, 12 while lncRNA-SNHG7 can increase FAIM2 expression, accelerating apoptosis and inhibiting proliferation. 13 However, our understanding of the vast competing endogenous RNA (ceRNA) network between lncRNAs and microRNAs in the human body is currently poor. Further studies are therefore needed to clarify the roles of lncRNAs.
A previous study showed that lncRNA-RP11-58O9.2 was upregulated in esophageal cancer and regulated more than 20 target genes, and was associated with multiple miRNAs, including miR-6749-3p. 14 However, information on the association between RP11-58O9.2 and NSCLC development is lacking. This study thus aimed to determine the association between lncRNA-RP11-58O9.2 and NSCLC and the ceRNA mechanisms of RP11-58O9.2.
Methods
NSCLC patients and tissues
Paired NSCLC and normal tissue samples were collected from patients at the Second Affiliated Hospital of Harbin Medical University. The inclusion criteria were patients with a first pathological diagnosis of NSCLC lung cancer without treatment. Patients with severe heart, lung, kidney, or other system dysfunction, other malignant tumors, or unclear consciousness were excluded. None of the patients diagnosed with NSCLC had received any treatment prior to surgery for diagnosis. All patients or their family members provided signed informed consent before specimen collection. The tissues were stored at −80°C until further use. This study was approved by the ethics committee of the Second Affiliated Hospital of Harbin Medical University of the Joint Logistics Team (Harbin, China; Approval number: KY2018-228; Date of approval: 28.02.2018) and was carried out according to the guidelines approved by the Second Affiliated Hospital of Harbin Medical University. We also compared the expression difference of RP11-58O9.2 between normal lung tissue and lung cancer by searching the gene database (http://gepia.cancer-pku.cn/detail.php?gene=RP11-58O9.2).
Cell culture and transfection
Three NSCLC cell lines (H1299, H1975 and A549) and one bronchial epithelial cell line (16HBE) were obtained from the American Type Culture Collection and grown in Roswell Park Memorial Institute (RPMI) 1640 medium containing 10% fetal bovine serum (FBS), streptomycin (100 μg/mL), and penicillin (100 U/mL) at 37°C with 5% CO2. Two NSCLC cell lines (H1299 and A549) were selected for transfection. Before transfection, the well-growing cell lines were seeded into 6-well plates and cultured until they reached approximately 90% confluence. Lentiviruses used for RP11-58O9.2 knockdown and negative control vectors were engineered and produced by Genechem Co., Ltd. (Shanghai, China), and the short hairpin RNA (shRNA) was marked with enhanced green fluorescent protein (EGFP). The transfection mixture containing lentivirus vectors carrying negative control and RP11-58O9.2 shRNAs were added to the culture plate for transfection. After 6 to 8 hours, the transfection solution was replaced with fresh RPMI 1640 medium containing serum for further culture. After puromycin screening, we obtained stable RP11-58O9.2-knockdown cell lines to verify knockdown efficiency. The shRNA sequences were as follows: forward, 5′-GCGATCATTTCTGCTGCATTT-3′, and reverse, 5′-AAATGCAGCAGAAATGATCGC-3′.
Reverse transcription quantitative polymerase chain reaction (RT-qPCR)
Total RNA was extracted from NSCLC cancer tissues, adjacent tissue specimens, and three NSCLC cell lines using TRIzol reagent. Total RNA was reverse-transcribed into cDNA using an all-in-one first-strand cDNA synthesis kit (Seven Biotechnology Co., Ltd., Beijing, China), according to the manufacturer’s instructions. PCR was carried out using a SYBR Green RT-qPCR reagent kit (Seven Biotechnology Co., Ltd.). The primer sequences are listed in Table 1. Data were normalized to glyceraldehyde 3-phosphate dehydrogenase expression and further normalized to the negative control. The data were analyzed using the 2−ΔΔCt method.
Primers used in this study.

RP11-58O9.2, lncRNA RP11-58O9.2; GAPDH, glyceraldehyde-3-phosphate dehydrogenase.
Cell Counting Kit-8 assay (CCK-8)
H1299 and A549 cells, each including a shRP11-58O9.2 and a negative control group, were subjected to CCK-8 assays to assess cell proliferation. In total, 100 μL of medium containing 2000 cells was added to 96-well plates for 24, 48, 72, and 96 hours of culture, after which 10 μL of CCK-8 reagent (Seven Biotechnology Co., Ltd.) was added to each well and incubated for another 2 hours. Finally, the absorbance at 450 nm was evaluated using a microplate reader.
Colony-formation assay
H1299 and A549 cells containing shRP11-58O9.2 and negative controls were seeded into 6-well culture plates at 800 cells/well. Three parallel samples were used to ensure the accuracy of the results. Samples were cultured for 2 weeks in RPMI 1640 containing 10% FBS. After discarding the medium, the colonies were fixed with methanol, washed with phosphate-buffered saline to remove the methanol, and dyed with crystal violet for 30 minutes. Finally, the number of colonies were calculated using ImageJ (US National Institute of Health, Bethesda, MD, USA).
Cell wound-healing assay
The shRP11-58O9.2 and negative control NSCLC cells were seeded in 6-well plates and incubated in RPMI 1640 containing 10% FBS for 24 hours. The cell monolayers were then scratched using a 200-μL sterile plastic pipette tip, the cell medium was changed to RPMI 1640 without FBS, and the cells were incubated for a further 24 hours. Wound healing was monitored using a phase contrast microscope, and images were processed using ImageJ. These experimental steps were repeated at least three times, and appropriate results were selected for statistical analysis.
Transwell assay
Cell invasion analysis was performed using a 24-well Transwell chamber and Matrigel-coated membrane (ACRO Biosystems Co., Ltd., Beijing, China). NSCLC cells (1 × 105) in RPMI 1640 without FBS were added to the upper chamber, and RPMI 1640 with FBS was placed in the lower chamber. After 24 hours, the cells in the upper chamber were removed using a cotton swab and the cells that invaded the lower surface of the filter were fixed with 4% paraformaldehyde, stained with crystal violet, and counted. These steps were repeated three times using two NSCLC cell lines (H1299 and A549).
In vivo experiment
For the tumor growth study, ten 5 to 6-week-old BALB/c mice (Vital River Laboratories, Beijing, China) were divided into two groups: an experimental group (n = 5) injected with A549 cells transfected with shRP11-58O9.2 lentiviruses, and a negative control group (n = 5) injected with cells transfected with control lentiviruses. Tumor size was examined every week and tumor volume was calculated as: length × width 2 × 0.5. After 6 weeks, the tumors were weighed. The experiment was performed in compliance with authenticated animal protocols of the Ethics Review Committee of Harbin Medical University.
Prediction of microRNA (miRNA) and ceRNA networks
The miRanda tool (parameters: - sc 140, - en -20, - restrict) was used to predict the existence of binding sites between miRNAs and lncRNAs, and possible miRNA-related mRNAs were obtained by searching the miRDB database. Finally, mRNAs with higher scores were screened out.
Statistical analyses
SPSS version 21.0 (IBM Corp., Armonk, NY, USA) was used for statistical analyses. Correlations between RP11-58O9.2 and clinicopathological parameters were analyzed by χ2 and Student’s t-tests and survival analysis was carried out by log-rank test. Overall survival (OS) was evaluated by the Kaplan–Meier method, and multivariate Cox proportional hazards analysis was performed with survival as the dependent variable. P < 0.05 was considered significant.
Results
Patients
Tissues were obtained from 78 patients with NSCLC, including 31 patients with advanced-stage (III–IV) NSCLC and 47 with stage I–II NSCLC. Twenty-six patients had lymph node metastases and 52 had no lymph node metastases. The patients characteristics are shown in Table 2.
Relationship between RP11-5809.2 expression level and clinicopathological factors in patients with non-small cell lung cancer.

LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma.
RP11-58O9.2 was highly expressed in NSCLC cell lines and tissues
According to the result of the genetic database (http://gepia.cancer-pku.cn/detail.php?gene=RP11-58O9.2), RP11-58O9.2 expression was significantly upregulated in NSCLC cases compared with normal tissues (Figure 1a). RP11-58O9.2 expression in the 78 LC and paired adjacent tissues was then confirmed by RT-qPCR, and RP11-58O9.2 expression was significantly higher in NSCLC tissues than in adjacent tissues (P < 0.001) (Figure 1b). In addition, RP11-58O9.2 was more highly expressed in A549, H1299, and H1975 NSCLC cells than in the bronchial epithelial cell line 16HBE (P < 0.001) (Figure 1c).

RP11-58O9.2 expression and prognosis of non-small cell lung cancer (NSCLC). (a) Box plots of expression in tumor and normal samples from GEPIA. (b) RP11-58O9.2 expression in 78 pairs of NSCLC tissue and adjacent non-tumorous tissue was validated by reverse transcription-quantitative polymerase chain reaction (***P < 0.001). (c) Expression levels of RP11-58O9.2 were higher in NSCLC cells than in normal cell lines. (d) Associations between RP11-58O9.2 levels and clinicopathological factors. (e–g) Expression of RP11-58O9.2 in 78 patients with NSCLC. (h) High expression levels of RP11-58O9.2 in NSCLC were significantly associated with poor overall survival based on Kaplan–Meier analysis. *P < 0.05, **P < 0.01, ***P < 0.001. LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma.
LncRNA RP11-58O9.2 and clinicopathological factors
The correlations between RP11-5809.2 expression and clinicopathological parameters in patients with NSCLC are shown in Table 2. RP11-5809.2 expression was defined as high or low using its median expression level in all samples as the cut-off value. High RP11-5809.2 expression was associated with higher clinical stage (P < 0.001) compared with low RP11-5809.2 expression, suggesting that RP11-58O9.2 expression was related to tumor development. Lymph node metastasis was also more common in patients with higher RP11-5809.2 expression levels compared with patients with lower expression levels (P < 0.001). However, RP11-5809.2 expression was not significantly correlated with age, sex, tumor size, differentiation, smoking, or histology. RT-qPCR analysis also revealed that RP11-5809.2 expression levels were associated with clinical stage (P < 0.001), lymph node metastasis (P < 0.001), and degree of differentiation (P = 0.0018) (Figure 1d–g). The different results for degree of differentiation may have been caused by an insufficient sample size.
RP11-58O9.2 expression levels and prognosis in patients with NSCLC
The correlation between RP11-5809.2 expression and OS in patients with NSCLC was evaluated by Kaplan–Meier analysis and log-rank test. Kaplan–Meier survival analysis demonstrated that patients with low RP11-5809.2 expression had longer survival than those with high expression (χ2 = 18.966, P < 0.001; Figure 1h). We also explored the association between RP11-5809.2 and prognosis by Cox regression analysis (Table 3). Univariate analysis revealed that clinical stage, lymph node metastasis, RP11-5809.2 expression (all P < 0.001), and the degree of differentiation (P = 0.007) were significantly associated with OS. The significant risk factors in univariate analysis were further analyzed by multivariate Cox analysis, which identified RP11-5809.2 level, lymphatic metastasis, and clinical stage as risk factors for NSCLC prognosis.
Cox analysis of prognostic parameters in patients with non-small cell lung cancer.

CI, confidence interval; HR, hazard ratio.
RP11-58O9.2 knockdown inhibited NSCLC cell progression
To explore the role of RP11-5809.2 in NSCLC, we knocked down RP11-5809.2 expression in A549 and H1299 cells using a lentiviral system (Figure 2a, b) and verified the knockdown efficiency by RT-qPCR (Figure 2c). We then evaluated the proliferation, invasion, and migration of NSCLC cells by CCK-8, colony-formation, Matrigel invasion, and wound-healing assays, respectively. CCK-8 (Figure 2d, e) and colony-formation (Figure 2f, g) assays showed that RP11-5809.2 knockdown significantly inhibited cancer cell proliferation, while wound-healing and Matrigel invasion assays showed that knockdown also reduced cell migration (Figure 2h–k) and invasion (Figure 3a, b), compared with the negative control group.

RP11-58O9.2 knockdown inhibited the proliferation and migration of lung cancer cells. (a, b) Expression of enhanced green fluorescent protein in transfected cells assessed by fluorescence and light microscopy. (c) Transfection and knockdown efficiency were verified by reverse transcription-quantitative polymerase chain reaction. (d, e) Cell proliferation determined by CCK8 assay after transfection. (f, g) Colony-formation assay. (h–k) Cell wound-healing assay.

RP11-58O9.2 knockdown inhibited the invasion of lung cancer cells and tumor growth in mice xenografts in in vivo experiments. (a, b) Transwell assay to detect cell invasion. (c–e) Tumor tissues were isolated from mice 6 weeks after cell injection, and tumor weight and volume were measured and calculated
RP11-5809.2 knockdown inhibited cancer cell growth in vivo
Ten mice were injected subcutaneously with A549 cells to investigate the effects of RP11-5809.2 on A549 cell progression in vivo. Tumor formation was monitored weekly. All mice developed tumors, but tumors in mice injected with RP11-5809.2 lentivirus-shRNA-EGFP were smaller and weighed less than those in the negative control group (Figure 3c–e).
ceRNA network regulated by RP11-58O9.2 in tumors
Using the miRanda tool, we confirmed that RP11-58O9.2 included a theoretical binding site for miR-6749-3p (Figure 4a). Examination of the miRDB database suggested that miR-6749-3p was possibly associated with and had binding sites for multiple mRNAs (Figure 4b). Of these mRNAs, TNRC6B, ORC5, and CDK19 have previously been confirmed to be associated with the progression of lung cancer. 14
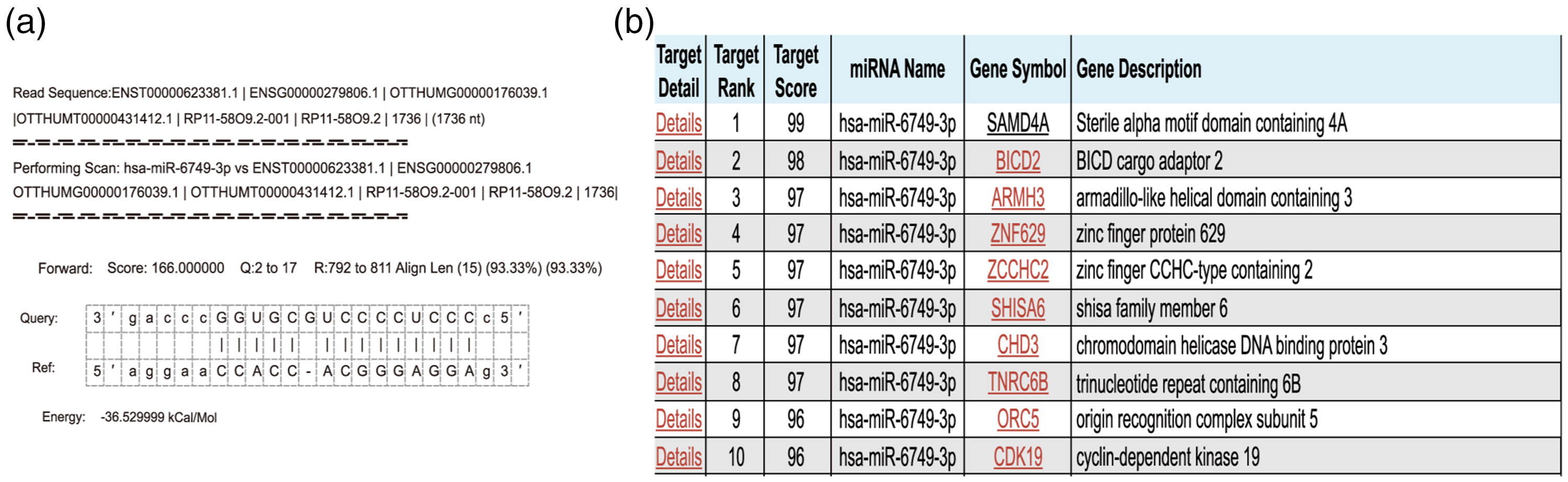
The ceRNA network regulated by RP11-58O9.2 in tumors. (a) Binding sites between miRNAs and lncRNA from miRanda. (b) mRNAs associated with miR-6749-3p from the miRDB website.
Discussion
In this study, we examined the novel biomarker lncRNA RP11-58O9.2 and its association with NSCLC. LC has high morbidity and mortality rates worldwide, and despite new therapeutic measures, the 5-year survival rate of patients with advanced LC in China was still only approximately 16% in 2022.15–19 OS and LC-specific survival rates also decrease significantly with increasing LC stage, indicating that the stage at initial diagnosis is a crucial factor affecting LC prognosis.20,21 The development of molecularly targeted agents and immune checkpoint inhibitors has provided better treatment options for patients with NSCLC; however, drug resistance represents a new challenge for cancer therapy. 22 LncRNAs are correlated with resistance to chemotherapy and targeted therapy, 23 and thus have potential applications as biomarkers for drug sensitivity and clinical prognosis of NSCLC. In addition, antagonists and/or mimics of lncRNA drug targets can be developed and used together with the chemotherapy or targeted drugs to enhance their effectiveness.24–26
A previous study found that lncRNA RP11-5809.2 was associated with restenosis caused by benign hyperplasia in patients with esophageal cancer, 14 but its role in LC remains unknown. We therefore explored the association between lncRNA RP11-5809.2 and NSCLC development. We showed that RP11-58O9.2 expression differed between NSCLC and normal tissues, and confirmed that lncRNA RP11-5809.2 was highly expressed in NSCLC tissues and cell lines by RT-qPCR. We also showed that patients with stage III–IV NSCLC and/or lymphatic metastasis had higher RP11-5809.2 levels than those with stage I–II NSCLC and/or no lymphatic metastasis, and that patients with high RP11-5809.2 expression had shorter survival than those with low expression. These results suggest that RP11-5809.2 may be a molecular marker for evaluating prognosis in patients with NSCLC. The results of in vitro and in vivo experiments revealed that RP11-5809.2 knockdown inhibited the proliferation, invasion, and migration of NSCLC cells and suppressed tumor growth in mice, further indicating that RP11-58O9.2 is associated with NSCLC prognosis and promotes NSCLC progression through certain mechanisms. Notably, multivariate analysis showed that the degree of differentiation had no significant correlation with RP11-5809.2 expression levels (P = 0.709), but this might have been related to the insufficient sample size.
There have been no previous reports on the mechanisms of RP11-58O9.2 in LC, and we therefore predicted the binding sites between miRNAs and RP11-58O9.2 lncRNA using miRanda, and used the miRDB website to obtain its related mRNAs. RP11-58O9.2 was predicted to act as a molecular sponge for miR-6749-3p. Regarding miR-6749-3p-regulated mRNAs, we selected the genes with the highest scores and found that target genes, including SAMD4A, BICD2, ARMH3, and ZNF629, may be associated with miR-6749-3p, while TNRC6B, ORC5, and CDK19 have been confirmed to be associated with the progression of lung cancer in previous reports.14,27,28
However, this study had some limitations. We did not verify the ceRNA network of RP11-58O9.2. This study described the biological function of RP11-58O9.2 and clarified its relationship with tumor progression; however, we did not conduct in-depth research on the mechanism of action, and further studies are needed to clarify this.
In conclusion, lncRNA RP11-58O9.2 is highly expressed in NSCLC and is associated with cancer prognosis and LC progression. RP11-58O9.2 may thus act as a novel biomarker for the diagnosis and treatment of LC. Although we predicted the ceRNA network, its specific mechanism of action remains unclear, and further studies are needed to verify these results.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605231206295 - Supplemental material for LncRNA RP11-58O9.2 predicts poor prognosis and promotes progression of non-small cell lung cancer
Supplemental material, sj-pdf-1-imr-10.1177_03000605231206295 for LncRNA RP11-58O9.2 predicts poor prognosis and promotes progression of non-small cell lung cancer by Xuan Miao, Wen Xi and Yongxia Bao in Journal of International Medical Research
Footnotes
Acknowledgements
We would like to express our gratitude to the laboratory supervisor for their attentive guidance. We would also like to thank the students who provided us with assistance, including Guanyu Li, Han Wang, and Feng Zhou. We also greatly appreciate the assistance of Jing Yin in data analysis.
Author contributions
Yongxia Bao designed and supervised the project. Xuan Miao performed most of the experiments. Wen Xi jointly conducted data analysis and article correction. All the authors read and approved the manuscript.
Data availability statement
The authors declare that the data in this article are available.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This work was supported by the Natural Science Foundation of Heilongjiang Province of China (Grant No.: LH2020H060).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
