Abstract
Objective
Endometrial cancer (EC) is one of the most prevalent cancers in women. Long non-coding RNAs (lncRNAs) are potential diagnostic biomarkers in patients with EC.
Methods
We obtained clinical information and transcriptome data for 552 patients with EC from The Cancer Genome Atlas database. Cuproptosis-associated lncRNAs were obtained through Pearson’s correlation analysis. Univariate and multivariate Cox regression analyses were applied and a signature predicting overall survival (OS) among patients with EC was constructed. We also analyzed the tumor immune microenvironment and drug sensitivity. The results were validated by quantitative real time-polymerase chain reaction, and 5-ethynyl-2′-deoxyuridine and wound-healing assays.
Results
Seven cuproptosis-associated lncRNAs related to prognosis were screened out and a signature was constructed. OS was significantly superior in the low-risk group. In addition, patients in the low-risk group had more CD8+ T cell infiltration, a stronger type II interferon response, and greater cisplatin sensitivity. Expression levels of some of the lncRNAs were significantly increased by cuproptosis. Furthermore, silencing of lncRNA AC084117.1 significantly inhibited the proliferation and migration of EC cells.
Conclusion
We constructed a seven cuproptosis-associated lncRNA signature to predict the prognosis of patients with EC with good predictive power.
Keywords
Introduction
Endometrial cancer (EC) is one of the most prevalent cancers of the female reproductive system, with about 41,700 new cases worldwide in 2020, and showing an increasing trend.1,2 The incidence rate of EC in developed countries far exceeds that of cervical cancer, making it the most common gynecological malignant tumor and the second most common female tumor. 3 The main treatment is currently surgery, possibly supplemented by radiotherapy, chemotherapy, hormone therapy, or immunotherapy, according to its histology, stage and molecular classification.4,5 Notably however, there has been no significant improvement in the survival of patients with EC since the mid-1970s. 3
Long non-coding RNAs (lncRNAs) are functional RNA molecules with transcripts longer than 200 bp, which cannot be translated into proteins.6,7 Many researches have shown relationships between lncRNAs and tumor occurrence and development, metastasis, drug resistance, the immune microenvironment, diagnosis, and prognosis, and have identified them as potential therapeutic targets and diagnostic biomarkers.7–12 Several studies have constructed gene signatures based on lncRNAs to predict the prognosis of cancer patients. For instance, Pan et al. constructed a lncRNA signature based on cancer-associated fibroblasts, which could predict the prognosis of patients with colorectal cancer, as well as other pan-cancers. 13 Lang et al. constructed a fatty acid metabolism-related nine-lncRNA signature to predict the overall survival (OS) of patients with cervical cancer, and as an independent prognostic factor. 14 In addition, an N7-methylguanine-related lncRNA signature had reliable prognostic predictive value in EC. 15
Copper is an essential trace element necessary for maintaining normal life activities, and plays a significant role in various physiological functions, such as mitochondrial respiration, antioxidant defense, and autophagy.16,17 Numerous studies have also confirmed that an imbalance of copper homeostasis is related to tumor initiation and progression. 18 For example, depletion of mitochondrial copper suppressed triple-negative breast cancer, 19 and patients with Wilson’s disease may have an increased incidence rate of hepatocellular carcinoma. 20 Previous studies also found that serum copper levels were imbalanced in patients with EC, and its levels were related to the depth of myometrial invasion. 21 Moreover, recent studies identified a novel mode of cell death, referred to as, cuproptosis, which is markedly distinct from the known mechanisms of apoptosis, pyroptosis, necroptosis, and ferroptosis. 22 Cuproptosis is induced by the accumulation of copper in mitochondria, targeting lipoacylated tricarboxylic acid cycle proteins to induce cell death. 23 Research on cuproptosis in EC, however, remains very limited.
In this study, by combining cuproptosis with lncRNAs, we constructed a signature to estimate the prognosis of patients with EC. We also explored the tumor microenvironment and drug sensitivity, and verified our findings by polymerase chain reaction (PCR), 5-ethynyl-2′-deoxyuridine (EdU) assay, and wound-healing assay. The results of this study may help to evaluate the prognosis of patients with EC from the perspective of cuproptosis, and aid the development of promising therapeutic methods.
Materials and methods
Data collection and identification of cuproptosis-associated lncRNAs
Clinical information and transcriptome data for patients with EC were collected from The Cancer Genome Atlas (TCGA) database, and transcriptome data were normalized to log2 (fragments per kilobase of transcript per million mapped reads+1). Only patients with OS ≥90 days were included in the analysis. We also carried out a literature search to identify cuproptosis-associated genes. We then evaluated the correlation between the cuproptosis-associated genes and corresponding lncRNAs by Pearson’s correlation analysis with a correlation coefficient >0.6 and P < 0.001. All the data analyzed were from a publicly available database and there was thus no need for ethical approval.
Construction and verification of prognostic signature according to cuproptosis-associated lncRNAs
We randomly divided the patients into training and validation sets in a ratio of 1:1 (Figure 1). The training set was used to construct a prognostic signature, which was then verified in the validation set and the whole set. We identified prognosis-related cuproptosis-associated lncRNAs by univariate Cox regression analysis, followed by LASSO Cox regression to minimize overfitting, and multivariate Cox analysis to construct a cuproptosis-associated lncRNA-based prognostic signature. The formula used was: risk score = ∑i (βi × expression of lncRNA i), where βi is the coefficient of lncRNA. EC patients in all three sets were then separated into high- and low-risk groups on the basis of the median risk score in the training set. Kaplan–Meier (K-M) survival curves and the log-rank test were used to compare survival differences. Moreover, the time-dependent receiver operating characteristic (ROC) curves and area under the curve (AUC) values were determined to estimate the accuracy of the signature.

Flow chart of bioinformatics analysis. EC, endometrial cancer; TCGA, The Cancer Genome Atlas; lncRNA, long non-coding RNA; EdU, 5-ethynyl-2′-deoxyuridine; PCR, polymerase chain reaction.
Identification of independence of prognostic signature
K-M curves were created to describe the predictive power of the prognostic signature in relation to different clinical characteristics, including age, stage, grade, surgical approach, and disease type. Univariate and multivariate Cox regression analyses were conducted to analyze the correlations between the clinical parameters and risk-score model, and ROC curves were applied to evaluate the exactness of the predictions.
Differences in tumor immune microenvironment
We carried out principal component analysis (PCA) to explore the differences between the high- and low-risk groups in terms of the expression profiles of all the studied genes, cuproptosis-associated genes, cuproptosis-associated lncRNAs, and prognosis-related cuproptosis-associated lncRNAs using the scatterplot3d package in R. We compared the immune checkpoint blockade responses between the high- and low-risk groups using the “ggpubr + ggplot2” package and displayed them by box plots. We then compared the immune function and infiltrating immune cells between the two groups using heatmaps and radar maps. We obtained somatic mutation data from TCGA database and compared the tumor mutation burden (TMB) using the “maftools” package. We also compared tumor immune dysfunction and exclusion between the two groups. Finally, we compared the IC50 values of different drugs in the two groups using the R package pRRophetic.
Cell lines and reagents
Human Ishikawa and HEC-1B EC cells were purchased from the Cell Bank of the Chinese Academy of Science (Shanghai, China) and cultured in Dulbecco's Modified Eagle Medium or Minimum Essential Medium, respectively, containing 10% fetal bovine serum (Sigma, St. Louis, MO, USA), 100 µg/mL streptomycin, and 100 µg/mL penicillin in a humidified incubator at 37°C with 5% CO2. The potent copper ionophore elesclomol was purchased from MedChemExpress (Shanghai, China), and copper(II) chloride was purchased from Sangon Biotech (Shanghai, China).
Quantitative real time (RT-q)PCR
Total RNA was extracted from Ishikawa and HEC-1B cells after treatment with TRIzol® reagent (Haoke Biotechnology, Hangzhou, China), and reverse transcription was performed using a Primescript RT Kit (Haoke Biotechnology) according to the manufacturer’s protocol. qRT-PCR was conducted by SYBR-Green Premix (Vazyme, Nanjing, China) with specific PCR primers (Supplementary Table 1) to detect lncRNA expression. Glyceraldehyde 3-phosphate dehydrogenase was used as an internal control, and the 2−ΔΔCT method was used to standardize relative lncRNA expression values.
Wound-healing assay
EC cells were incubated in 6-well plates for 24 hours and then treated with AC084117.1 small interfering RNA (siRNA, si-AC084117.1), negative control (NC) siRNA (si-NC), or nothing for 2 or 3 days until they reached 90% confluence. Scratch wounds were then created using a 200-µL pipette tip, the cells were washed three times with phosphate-buffered saline, and photos were taken immediately. The cells were then cultured in serum-free medium for a further 48 hours and photographed again.
EdU assay
Cell proliferation was determined using an EdU assay kit (Beyotime Biotechnology, Shanghai, China). About 1 × 105 cells were incubated in 24-well plates for 24 hours and then treated with si-AC084117.1, si-NC, or nothing for 2 days, followed by incubation with 10 μM EdU at 37°C for 2 hours. The cells were then fixed with 4% paraformaldehyde for 15 minutes, incubated with phosphate-buffered saline containing 0.3% Triton X-100 for 10 minutes, and incubated with Click reaction solution in the dark at room temperature for 30 minutes. Finally, the cells were stained with 1 × Hoechst 33342 (Beyotime Biotechnology, Shanghai, China) for 10 minutes and images were captured using an Olympus BX51 immunofluorescence microscope (Olympus, Tokyo, Japan).
Statistical analysis
All statistical analyses were carried out using R version 3.6.1 and GraphPad Prism 9 software. Experimental results were expressed as mean ± standard error of mean, and statistical significance was determined by the unpaired t-test. A value of P < 0.05 was considered significant. We carried out LASSO regression and multivariate Cox regression analyses to identify the relevant genes.
Results
Patients and genes
Clinical information and transcriptome data for 552 patients with EC were collected from TCGA, and 501 patients with OS ≥ 90 days were included in the analysis. These patients were divided into a training set (251 patients) and a validation set (250 patients). Clinical information for these patients, including age, stage, grade, surgical approach, disease type, and OS are presented in Table 1.
Clinical information for patients with endometrial cancer.

NA, not available.
Nineteen cuproptosis-associated genes were identified from the literature: NFE2L2, NLRP3, ATP7B, ATP7A, SLC31A1, FDX1, LIAS, LIPT1, LIPT2, DLD, DLAT, PDHA1, PDHB, MTF1, GLS, CDKN2A, DBT, GCSH, and DLST. 24
Identification and validation of a cuproptosis-associated lncRNA signature
First, we identified a total of 927 cuproptosis-associated lncRNAs in EC patients (Supplementary Table 2). Using univariate Cox regression analysis, we screened out 22 prognosis-related cuproptosis-associated lncRNAs with P < 0.05 (Figure 2a). We then conducted LASSO (Figure 2b, c) and multivariate Cox analyses to further determine the relevant genes, and finally obtained seven prognosis-related cuproptosis-associated lncRNAs, AC084117.1, AC090617.5, FARSA-AS1, AC011479.2, Z99572.1, AC004466.2, and AC008966.2 (Figure 2d), which were all associated with glutaminase (GLS) (Figure 2e). Using the above formula, we established the following prognostic model: risk score = 0.3039 ×expression of AC084117.1 − 1.2129 ×expression of AC090617.5 + 0.4903 ×expression of FARSA-AS1 + 0.5194 × expression of AC011479.2 + 1.5936 × expression of Z99572.1 +0.6218 × expression of AC004466.2 − 0.9087 ×expression of AC008966.2. Based on the median risk score in the training set, we separated the EC patients in the training, validation, and whole sets into high- and low-risk groups (Figure 3a–c), and showed the survival status of each EC patient (Figure 3d–f). Differences in expression of the cuproptosis-associated lncRNAs were visualized using heatmaps, which showed that expression levels of risk-type cuproptosis-associated lncRNAs (AC084117.1, FARSA-AS1, AC011479.2, Z99572.1, and AC004466.2) were significantly higher and levels of protective-type lncRNAs (AC090617.5 and AC008966.2) were significantly lower (Figure 3g–i) in the high-risk group. K-M curves revealed that the prognosis in the low-risk group was superior to that in the high-risk group in all three sets (Figure 3j–l), and the AUC values for 1-, 3-, and 5-year OS ranged from 0.621 to 0.837 (Figure 3m–o), indicating that the signature had excellent prognostic predictive power in patients with EC.

Identification of prognostic-related cuproptosis-associated long non-coding RNAs (lncRNAs) in endometrial cancer (EC). (a) Forest plot of prognosis-related cuproptosis-associated lncRNAs by univariate Cox analysis. (b) LASSO coefficient profiles of candidate cuproptosis-associated lncRNAs. (c) Selection of optimal penalty parameter for LASSO algorithm. (d) Information on seven prognostic-related cuproptosis-associated lncRNAs by multivariate Cox analysis and (e) Correlation between seven prognostic-related lncRNAs and cuproptosis-associated genes.

Cuproptosis-associated long non-coding RNA (lncRNA)-based signature predicted overall survival (OS) in patients with endometrial cancer (EC) in the training, validation, and whole sets. (a–c) Risk score distribution. (d–f) Survival status and survival times. (g–i) Heatmaps of expression profiles of seven cuproptosis-associated lncRNAs. (j–l) Kaplan–Meier survival curves and (m–o) Receiver operating characteristic curve analysis of 1-, 3-, and 5-year survival. AUC, area under the curve.
Cuproptosis-associated lncRNA-based prognostic signature was an independent prognostic indicator
We grouped EC patients according to various clinical factors to explore the correlations between these factors and the lncRNA signature. The prognosis of patients in the high-risk group was poorer than that in the low-risk group, except for patients with cystic, mucinous, and serous neoplasms (Figure 4a–e). We further estimated the prognostic value of the signature using univariate and multivariate Cox analyses, and showed that the lncRNA signature was an independent prognostic indicator (P < 0.001) (Figure 4f, g). Three-year ROC curves also showed an AUC of 0.744, which was better than for any other clinical characteristic, indicating its excellent predictive power (Figure 4h).

Cuproptosis-associated long non-coding RNA (lncRNA)-based risk score model was an independent prognostic indicator. Kaplan–Meier survival curves of prognostic risk score model based on different clinical characteristics: (a) age, (b) stage, (c) grade, (d) surgical approach, and (e) disease type. (f) Univariate Cox analysis. (g) Multivariate Cox analysis and (h) Receiver operating characteristic curve analysis of predictive power of risk score model compared with clinical parameters. AUC, area under the curve.
We also carried out PCA to explore the differences between the high- and low-risk groups in four expression profiles including all the studied genes (Figure 5a), 19 cuproptosis-associated genes (Figure 5b), 927 cuproptosis-associated lncRNAs (Figure 5c), and the seven prognosis-related cuproptosis-associated lncRNAs (Figure 5d). The seven prognosis-related lncRNAs had the strongest ability to differentiate between these two groups.

Comparison of tumor immune microenvironment between high- and low-risk groups. (a) Principal component analysis of all studied genes, (b) cuproptosis-associated genes, (c) cuproptosis-associated long non-coding RNAs (lncRNAs), and (d) prognosis-related cuproptosis-associated lncRNAs. (e) Box plots of immune checkpoints. (f) Heatmap of immune function. (g) Radar map of infiltration of immune cells. Waterfall diagram of somatic mutation status in patients with endometrial cancer in (h) high- and (i) low-risk groups. (j) Tumor mutation burden (TMB) in high- and low-risk groups. (k) Kaplan–Meier (K-M) survival curves in high- and low-TMB groups. (l) K-M survival curves of prognostic risk score model combined with different TMB. (m) Tumor immune dysfunction and exclusion between high- and low-risk groups. (n) Drug correlation and (o) sensitivity analysis of cisplatin. CCT018159, cisplatin.
Tumor immune microenvironment in high- and low-risk groups
We compared the tumor immune microenvironment between the high- and low-risk groups. Most immune checkpoints were highly expressed in the high-risk group (Figure 5e), which partly explained the poorer prognosis. Regarding immune function, the type I and type II interferon responses differed significantly between the two groups in the heatmap (Figure 5f), while the radar map showed greater infiltration of CD8+ T cells in the low-risk group (P < 0.001) (Figure 5g).
We revealed the somatic mutation statuses of EC patients in the high- and low-risk groups using a waterfall diagram, and found that PTEN had the highest mutation rate in both groups (Figure 5h, i). TMB is an indirect indicator reflecting the ability of tumors to produce neoantigens, and can predict the efficacy of immunotherapy for tumors. TMB was significantly higher in patients in the low-risk group (P = 0.018) (Figure 5j), consistent with the better survival of patients with high TMB (Figure 5k). Combining TMB and risk score showed that patients with a high TMB and low-risk had the optimal survival outcome (Figure 5l); however, there was no discrepancy in TIDE between the groups (Figure 5m). Finally, regarding drug sensitivity, the IC50 value of cisplatin, as a commonly used chemotherapy agent in patients with EC, was much lower in the low-risk group (Figure 5n, o).
Validation of correlation between lncRNAs and copper
To prove the correlation between the signature lncRNAs and cuproptosis, we treated human Ishikawa and HEC-1B cells with 2 µM copper chloride, 200 nM elesclomol, or a combination of both for 24 hours. Expression levels of the seven lncRNAs were all increased by cuproptosis, and most of the differences were significant (Figure 6), with AC084117.1 expression showing the greatest increase following both monotherapy and combination therapy (Figure 6a), while expression levels of AC00896.2 showed no significant increase in Ishikawa cells (Figure 6g).

Expression levels of seven long non-coding RNAs after induction of cuproptosis.
Silencing AC084117.1 inhibited the proliferation and migration of EC cells
According to the PCR results, we selected AC084117.1 for further research and developed si-AC084117.1-1, si-AC084117.1-2, si-AC084117.1-3, and si-NC. si-AC084117.1-2 had the greatest silencing effect and was selected for subsequent wound-healing and EdU assays. Silencing of AC084117.1 significantly inhibited the proliferation and migration of EC cells (Figure 7).
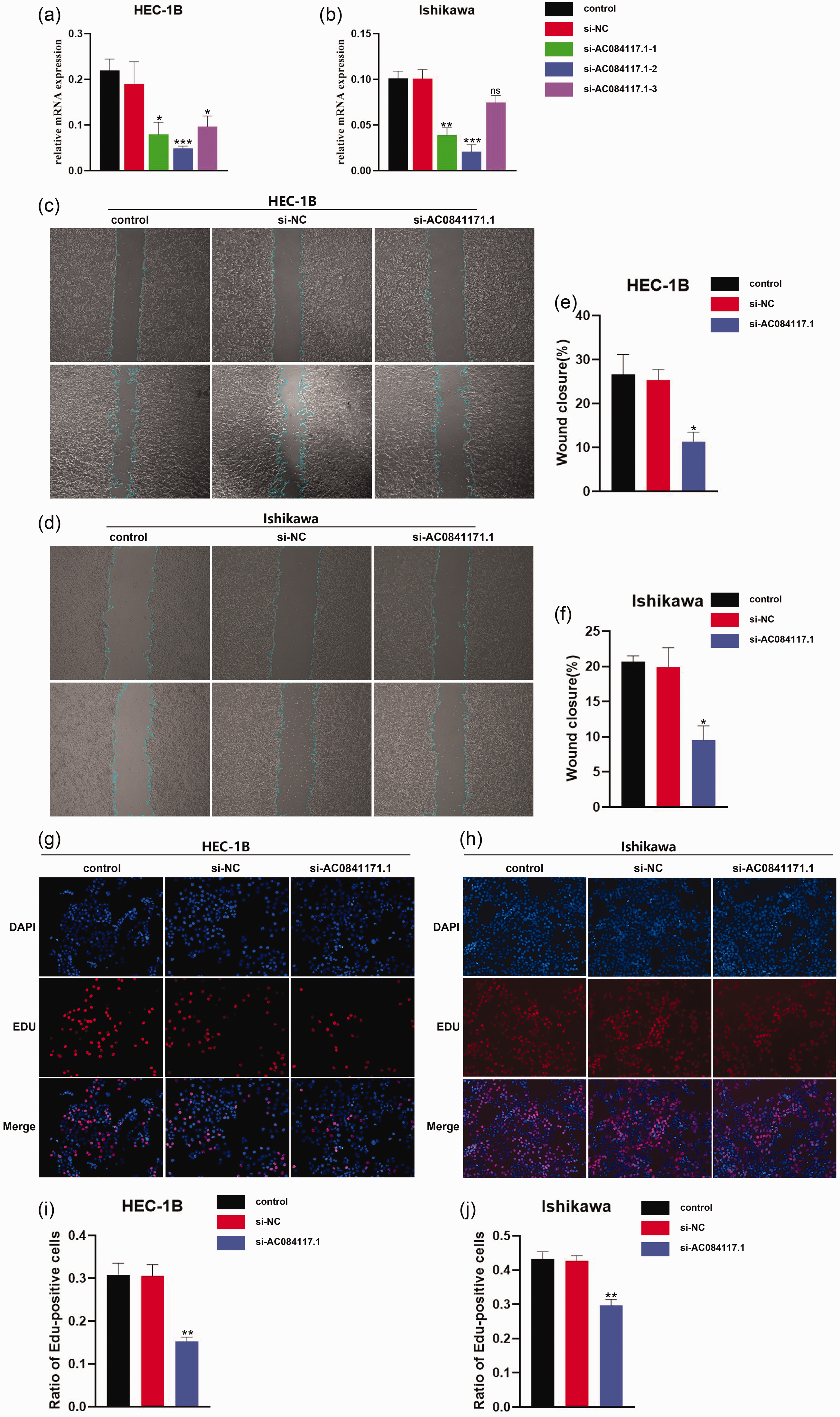
Silencing of AC084117.1 inhibited the proliferation and migration of endometrial cancer (EC) cells. (a, b) Comparison of silencing effects of si-AC084117.1-1, si-AC084117.1-2, si-AC084117.1-3, and si-NC by polymerase chain reaction in HEC-1B and Ishikawa cells. (c–f) Results of wound-healing assay to detect effect of si-AC084117.1 on migration of EC cells and (g–j) Results of 5-ethynyl-2′-deoxyuridine assay to detect effect of si-AC084117.1 on proliferation of EC cells. NC, negative control.
Discussion
EC is the sixth most prevalent cancer among females worldwide. Its incidence rates have increased steadily in recent decades, seriously endangering women's health and fecundity.2,3 There is thus an urgent need to identify valid biomarkers to predict and improve the prognosis of patients with EC. Recent research found that a signature involving cuproptosis-associated lncRNAs was an effective indicator for predicting the prognosis of tumor patients; however, it has been explored in many cancers, but not in EC. For example, Xu et al. demonstrated a 10-cuproptosis-associated lncRNA prognostic signature to predict OS in patients with colon adenocarcinoma, providing promising insights into drug selection, 25 while a cuproptosis-associated lncRNA prognostic signature was also used to evaluate the prognosis and immunotherapy response in patients with clear cell renal cell carcinoma. 26 Cuproptosis-associated lncRNA prognostic signatures have also been constructed in osteosarcoma, 27 cutaneous melanoma, 28 head and neck squamous cell carcinoma, 29 hepatocellular carcinoma, 30 and soft tissue sarcoma, 24 but not in EC.
In this study, we constructed a seven-lncRNA signature based on cuproptosis to comprehensively analyze EC patients, including in terms of survival, tumor immune microenvironment, and drug sensitivity. We initially identified 19 cuproptosis-associated genes from previous studies and screened out 927 cuproptosis-associated lncRNA by Pearson’s correlation analysis. We finally obtained seven cuproptosis-associated lncRNAs following univariate and multivariate Cox analyses, and constructed a prognostic signature. OS was notably worse in EC patients in the high-risk group. In addition, we further verified the correlation between these seven lncRNAs and cuproptosis by PCR, showing that expression levels of most of the seven lncRNAs were up-regulated after induction of cuproptosis.
Our analysis showed all seven lncRNAs were associated with GLS. GLS is the initial enzyme in glutamine metabolism, converting glutamine to glutamate, which plays a critical role in the metabolism, growth, and proliferation of cancer cells and is an important tumor metabolic hallmark.31,32 The selective GLS1 inhibitorCB-839 has been clinically tested in multiple cancers, including triple-negative breast cancer, lung cancer, renal cancer, and colorectal cancer, 33 and some studies have also explored the role of GLS in EC. For example, Zhou et al. found that estrogen promoted the growth of EC cells by inhibiting autophagy and increasing GLS levels through up-regulating c-Myc, 34 while another study found that GLS1 expression was elevated in patients with EC progression, associated with a poor prognosis. Furthermore, treatment with the GLS1-targeting compound 968 inhibited the proliferation and metastasis of EC cells via Akt/mammalian target of rapamycin/S6 signaling pathway inhibition. 35 However, there have been no related reports of cuproptosis and GLS in patients with EC. Notably, using TCGA database, we confirmed that patients with EC and high GLS expression had poorer survival outcomes (Supplementary Figure 1).
Regarding the tumor immune microenvironment, immune checkpoint, immune function, and immune cell infiltration differed between the low- and high-risk groups. Type II IFN (known as IFN-γ in humans) has functions in immune stimulation and immune regulation, enhances the antigen-presentation ability of dendritic cells and macrophages, and promotes immunologic recognition and apoptosis of tumor cells by up-regulating the expression of MHC-I and STAT-1-associated cyclin-dependent kinase, thus playing an important role in anti-tumor immunity.36–38 Our results showed that the type II IFN response was more intense in the low-risk group, which might contribute to their better survival outcome. Furthermore, cytotoxic CD8+ T cells are considered as a pivotal factor in eliminating tumor cells and can provide long-term protective immunity. 39 In the current study, the expression of CD8+ T cells was higher in the low-risk group. These results suggest that cuproptosis might affect the prognosis of patients with EC by regulating immunity.
Finally, we also discovered that patients with EC in the low-risk group had higher sensitivity to cisplatin. Platinum-based combination chemotherapy is currently an important adjuvant treatment for EC,5,40,41 and phase III clinical trials found that the combination of doxorubicin and cisplatin effectively prolonged progression-free survival compared with doxorubicin alone in patients with advanced EC. 42 In addition, chemotherapy with doxorubicin-cisplatin improved progression-free and OS compared with whole-abdominal irradiation. 43 The superior prognosis of low-risk patients with EC may thus be partly due to their greater sensitivity to cisplatin.
This study had some limitations. First, the cuproptosis-associated lncRNA signature should be validated independently to make it more reliable, but there was no external validation cohort with both clinical and transcriptome data. Second, it would be better to validate the seven cuproptosis-associated lncRNAs in a clinical setting; however, the lack of specimens from patients with EC meant that the experimental part in this study was mainly carried out using human EC cells.
Conclusion
We identified seven cuproptosis-associated lncRNAs in patients with EC and constructed a signature to predict the prognosis of these patients. We also analyzed this signature in relation to the tumor immune microenvironment and TMB, and sensitivity to cisplatin, as a common chemotherapeutic drug in EC. These findings may offer novel insights into the mechanisms of cuproptosis and help to improve the survival outcome of patients with EC.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605231213435 - Supplemental material for Identification and validation of a seven cuproptosis-associated lncRNA signature to predict the prognosis of endometrial cancer
Supplemental material, sj-pdf-1-imr-10.1177_03000605231213435 for Identification and validation of a seven cuproptosis-associated lncRNA signature to predict the prognosis of endometrial cancer by Luya Cai, Limeng Cai, Lin Zhou, Yating Zhao and Jianhua Qian in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_03000605231213435 - Supplemental material for Identification and validation of a seven cuproptosis-associated lncRNA signature to predict the prognosis of endometrial cancer
Supplemental material, sj-pdf-2-imr-10.1177_03000605231213435 for Identification and validation of a seven cuproptosis-associated lncRNA signature to predict the prognosis of endometrial cancer by Luya Cai, Limeng Cai, Lin Zhou, Yating Zhao and Jianhua Qian in Journal of International Medical Research
Supplemental Material
sj-pdf-3-imr-10.1177_03000605231213435 - Supplemental material for Identification and validation of a seven cuproptosis-associated lncRNA signature to predict the prognosis of endometrial cancer
Supplemental material, sj-pdf-3-imr-10.1177_03000605231213435 for Identification and validation of a seven cuproptosis-associated lncRNA signature to predict the prognosis of endometrial cancer by Luya Cai, Limeng Cai, Lin Zhou, Yating Zhao and Jianhua Qian in Journal of International Medical Research
Footnotes
Author contributions
JHQ conceived and designed the study. LYC and LMC drafted the manuscript and analyzed the data. LZ and YTZ handled the picture and article format. All authors have read and approved the final published manuscript.
Data availability statement
Declaration of conflicting interests
The authors declare that there is no conflict of interest.
Funding
This work was supported by the National Natural Science Foundation of China (82071665) and the Key Research and Development Program of Zhejiang Province, China (2020C03116).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
