Abstract
Objectives
CCL20 is a chemotactic factor that is involved in immune cell recruitment and cancer progression. However, the role of CCL20 in the prognosis of breast cancer remains unclear. This study analyzed correlations between CCL20 expression and immune infiltration, clinicopathological parameters, and prognosis in breast cancer patients.
Methods
Correlations between CCL20 expression and clinicopathological parameters, prognosis, and immune infiltration in breast cancer were determined using the TIMER, UALCAN, and PrognoScan databases. Furthermore, gene–gene and protein–protein interactions were determined using GeneMANIA and STING network construction, respectively.
Results
CCL20 expression was significantly upregulated in breast cancer and had significant associations with clinicopathological features, including race, sex, age, menopause status, cancer stage, cancer subclass, and nodal metastasis; moreover, patients with higher CCL20 expression exhibited poor prognosis. Meanwhile, CCL20 expression was significantly correlated with the infiltration of immune cells in breast cancer, including monocytes, neutrophils, tumor-associated macrophages, Th1 cells, regulatory T cells, and exhausted T cells. Moreover, the network of CCL20 expression showed the majority genes and proteins were associated with immune reactions.
Conclusions
CCL20 is a prognosis-related biomarker in breast cancer on the basis of its correlation with immune infiltration levels and has potential to also be a therapeutic target.
Introduction
Breast cancer is the most common cancer worldwide and has a 6.6% mortality rate. 1 Breast cancers are classified into different molecular subtypes such as luminal A, luminal B, HER2+, and basal-like/triple-negative. Recently, treatments for breast cancer patients have been improved because of advances in surgery, chemotherapy, and radiotherapy. Unfortunately, the survival time of breast cancer patients remains poor because of chemo-radiation resistance and the high rate of metastasis.2,3 Over the past decades, immune checkpoint inhibitors such as PD-1 and PD-L1 have been identified as exceptional therapeutic targets in breast cancer.4,5 However, it is essential to identify more effective biomarkers and targets for immunotherapy in breast cancer.
C-C motif chemokine ligand 20 (CCL20) is a member of the subfamily of small cytokine CC genes. Cytokines are a family of secreted proteins that are involved in immunoregulatory and inflammatory processes and are an indispensable factor in tumor microenvironment formation.6–8 Chemokines provide a crucial condition for the infiltration of immune cells into the tumor microenvironment.9–11 CCL20 acts as a ligand for the C-C chemokine receptor CCR6, inducing a strong chemotactic response and mobilization of intracellular calcium ions.12–14 CCL20 acts as a chemotactic factor that attracts lymphocytes and neutrophils and is involved in the recruitment of both Th17 cells and regulatory T cells (Tregs) to sites of inflammation.12,15 Recent studies have reported that high CCL20 expression is associated with growth and metastasis of various cancers such as hepatocellular carcinoma, colorectal cancer, pancreatic cancer, and lung cancer.16–18 Furthermore, patients with high CCL20 expression show poorer prognosis. Regarding breast cancer, several studies have shown that CCL20 upregulation significantly promotes migration and invasion of breast cancer cells.19–21 Furthermore, CCL20 overexpression facilitates the proliferation and invasiveness of triple-negative breast cancer (TNBC) cells and accelerates tumor growth in xenograft models. 22 Thus, CCL20 may participate in the pathogenesis of breast cancer by altering immune processes.
In this study, we investigated the expression of CCL20 in breast cancer using the TIMER database and associations between CCL20 expression and clinicopathological parameters and the prognosis of breast cancer patients using the UALCAN and PrognoScan databases, respectively. Furthermore, associations between immune infiltration and CCL20 expression were also examined using the Tumor Immune Estimation Resource (TIMER) database. Finally, gene–gene and protein–protein interaction networks were determined using GeneMANIA and STING network construction, respectively. Our results highlight a potential mechanism through which CCL20 is involved in immune infiltration in breast cancer.
Materials and methods
TIMER database analysis
The TIMER (https://cistrome.shinyapps.io/timer/) database was used to determine CCL20 expression in different cancers through the “Diff Exp” module. The infiltration levels of different immune cells in breast cancer were also analyzed in TIMER. The relationship between CCL20 and immune cell infiltration levels (B cells, CD8+ T cells, CD4+ T cells, neutrophils, macrophages, and dendritic cells) were investigated through the “Gene” module. Additionally, relationships between CCL20 expression and different marker sets for immune cells were assessed through the “Correlation” module.
Gene Expression Profiling Interactive Analysis (GEPIA)
The GEPIA database (http://gepia.cancer-pku.cn/index.html) was used for gene expression analysis on the basis of The Cancer Genome Atlas (TCGA) and GTEx data. CCL20 levels were evaluated from the breast invasive carcinoma (BRCA) datasets. The expression of CCL20 in BRCA and normal adjacent breast tissues was investigated with the option of matching TCGA normal and GTEx data and log2 (TPM+1) for log-scale using the “Expression DIY” module. Four BRCA subtypes were analyzed, including basal-like/triple-negative, HER2+ non-luminal, Luminal A, and Luminal B.
UALCAN database analysis
The UALCAN database (http://ualcan.path.uab.edu/), including transcriptome data from TCGA and MET500 data, was used to investigate CCL20 levels in BRCA tissues. Furthermore, we determined associations between CCL20 and various clinicopathological parameters (race, sex, age, menopause status, cancer stage, cancer subclasses, nodal metastasis status, and TP53 mutation status) of BRCA.
PrognoScan database analysis
The PrognoScan database (http://www.abren.net/PrognoScan/) was used to investigate correlations between CCL20 expression and survival in breast cancer patients. The patients included in these open databases all provided informed consent. Relationships between CCL20 expression and patient prognosis, such as relapse-free survival (RFS) and distant metastasis free survival (DMFS), across a large collection of publicly available cancer microarray datasets can be investigated using PrognoScan. The GSE12276, GSE1379, and GSE9195 datasets were selected for further analysis. Hazard ratios (HRs) with 95% confidence intervals (CIs) and adjusted Cox P-values were calculated.
Analysis of CCL20-interacting genes and proteins
The GeneMANIA database (http://www.genemania.org) was used to construct the CCL20 gene interaction network. The STRING database (https://string-db.org/) was used to construct the protein–protein interaction (PPI) network for CCL20.
Results
CCL20 expression is upregulated in breast cancer
First, we investigated the mRNA expression of CCL20 in human cancers using the TIMER database. Significantly higher expression of CCL20 was found in BRCA, cholangiocarcinoma, colon adenocarcinoma, esophageal carcinoma, head and neck squamous cell carcinoma, kidney chromophobe, liver hepatocellular carcinoma, lung adenocarcinoma, rectum adenocarcinoma, stomach adenocarcinoma, thyroid carcinoma, and uterine corpus endometrial carcinoma compared with the corresponding normal tissues (Figure 1a). Moreover, we found that CCL20 mRNA levels were elevated in the GEPIA and UALCAN databases, especially in the basal-like/triple-negative and HER2+ non-luminal BRCA subtypes (Figure 1b and Figure 2a). Taken together, these findings suggest that CCL20 expression is increased in breast cancer and that CCL20 may play a key role in breast cancer development.

CCL20 mRNA levels in human breast cancer.

Associations between CCL20 expression and clinicopathological parameters and prognosis in breast cancer patients.
Correlations between CCL20 expression and clinical parameters of breast cancer patients
We further determined CCL20 expression among groups of BRCA patients on the basis of different clinical parameters using the UALCAN database. According to tumor stage, CCL20 expression was significantly upregulated in BRCA patients with stage 1, 2, 3, and 4 disease (Figure 2b). Regarding race, significantly higher expression of CCL20 was observed in White and Black BRCA patients, with higher expression also found in Asian patients (Figure 2c). In terms of age and sex, significantly increased CCL20 expression was observed in female patients compared with healthy individuals and male patients (Figure 2d). CCL20 levels were also enhanced in patients from different age groups, including 21 to 40 years, 41 to 60 years, 61 to 80 years, and 81 to 100 years (Figure 2e). Additionally, CCL20 expression was significantly upregulated in pre- and post-menopausal patients (Figure 2f). Consistent with Figure 1b, CCL20 expression was significantly increased in the luminal and triple-negative BRCA subclasses compared with healthy individuals (Figure 2g). With regard to nodal metastasis, CCL20 levels were higher in patients with BRCA classified as N0, N1, N2, and N3 (Figure 2h). Upregulation of CCL20 was found in both TP53-mutant and TP53-wild-type BRCA patients compared with healthy individuals (Figure 2i). Therefore, CCL20 may participate in the growth and metastasis of BRCA.
Poor prognosis in breast cancer patients with higher CCL20 expression
Next, we examined whether breast cancer patients with higher CCL20 expression had poorer survival outcomes using the PrognoScan database. By analyzing the GSE12276 dataset, we found that patients with higher CCL20 expression exhibited significantly lower RFS (HR: 1.09, 95% CI:1.00–1.19, COX P-value: 0.04) (Figure 2j). In the GSE1379 dataset, higher CCL20 expression in BRCA patients was also associated with decreased RFS (HR: 1.54, 95% CI: 0.88–2.71, COX P-value: 0.13) (Figure 2k). Furthermore, RFS and DMFS were analyzed in the GSE9195 dataset, and the results showed decreased RFS (HR: 2.16, 95% CI: 0.60–7.71], COX P-value: 0.24) and DMFS (HR: 2.67, 95% CI: 0.66–10.83, COX P-value: 0.17) in patients with high CCL20 expression than in those with low CCL20 expression (Figure 2l and 2m). These data demonstrate that higher CCL20 expression is associated with poor prognosis in BRCA patients.
Correlation between CCL20 expression and immune infiltration in BRCA
As immune infiltration plays an important role in tumor occurrence and development, we explored the relationship between CCL20 expression and the infiltrating immune cells in breast cancer using the TIMER database. The results showed that CCL20 expression was significantly correlated with tumor purity in BRCA. Additionally, we observed that CCL20 expression was significantly correlated with B cells (r = −0.21, p = 1.99e−11), CD8+ T cells (r = 0.138, p =1.58e−05), CD4+ T cells (r = 0.179, p = 2.36e−08), neutrophils (r = 0.342, p = 1.85e−27), and dendritic cells (r = 0.324, p = 1.07e−24) in BRCA (Figure 3a). Additionally, the relationship between CCL20 expression and different gene marker sets of immune cells were investigated, including monocytes, neutrophils, tumor-associated macrophages (TAMs), T helper 1 (Th1) cells, Tregs, and exhausted T cells (Figure 3b). The results showed significant correlations between CCL20 and levels of specific immune cell markers such as the monocyte marker CD86 (r = 0.377, p = 1.89e−38), the neutrophils markers ITGAM (r = 0.308, p = 1.43e−25) and CCR7 (r = 0.28, p = 2.98e−21), the TAMs markers IL10 (r = 0.305, p = 3.86e−25), CD68 (r = 0.272, p = 3.54e−20), and STAT6 (r = 0.116, p = 1.16e−04), the Th1 markers TBX21 (r = 0.295, p = 1.69e−23), TNF (r = 0.495, p = 5.2e−69), IFNG (r = 0.338, p = 9.26e−31), and STAT1 (r = 0.248, p = 6.52e−17), the Treg markers CCR8 (r = 0.31, p = 7.1e−26) and STAT5B (r = −0.097, p = 1.35e−03), and the T cell exhaustion markers HAVCR2 (r = 0.311, p = 4.26e−26), PDCD1 (r = 0.326, p =1.31e−28), and CTLA4 (r = 0.412, p = 3.26e−46). These findings indicate that CCL20 expression is significantly correlated with immune cell infiltration in breast cancer.
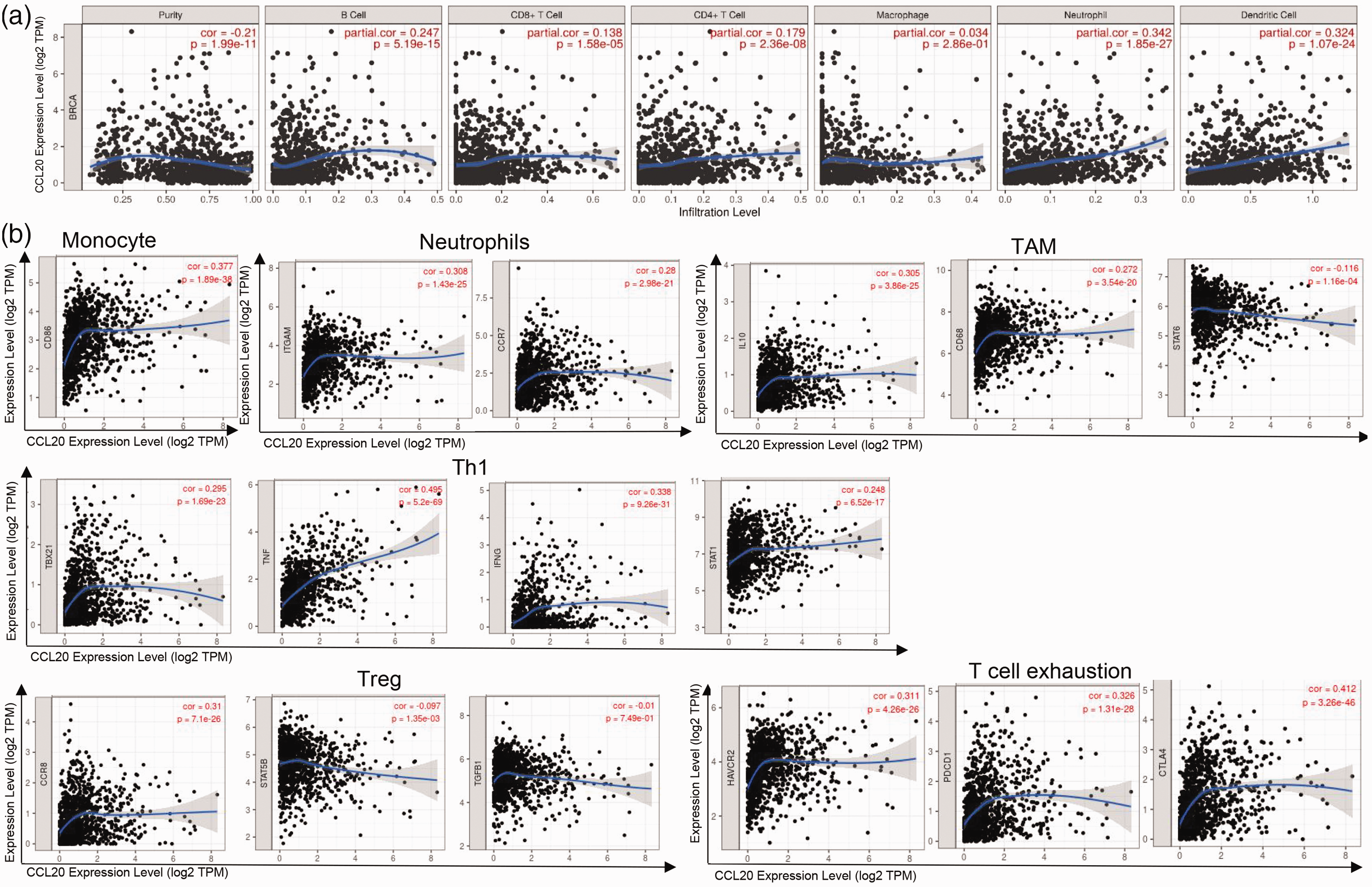
Associations between CCL20 expression and immune infiltration levels in breast cancer patients.
Identification of CCL20-interacting genes and proteins
To examine the gene–gene interactions for CCL20, we constructed an interaction network using the GeneMANIA database. The results showed that the 20 most frequently altered genes closely associated with CCL20 included CCR6, CXCL5, XCL2, CXCL12, CXCL17, and CXCL11 (Figure 4a). These genes participate in immune cell chemotaxis and migration, and thus regulate cellular immune responses. The protein–protein interaction network for CCL20 was constructed using the STRING database and included 11 nodes and 54 edges. The results demonstrated that 10 proteins were correlated with CCL20, including IL6, IL1A, IL1B, and CSF2, which are all involved in modulating inflammatory reactions (Figure 4b).

Interaction analysis of CCL20 at the gene and protein levels.
Discussion
In this study, we found that CCL20 expression was significantly upregulated in breast cancer compared with normal breast tissue through online analysis of the TIMER, UALCAN, and GEPIA databases. Additionally, CCL20 expression was significantly associated with certain clinicopathological characteristics including race, sex, age, menopause status, cancer stage, cancer subclass, and nodal metastasis status. By evaluating the survival outcomes of breast cancer patients from several GSE datasets, we found that patients with higher CCL20 expression had poorer prognoses than those with lower CCL20 expression. Furthermore, CCL20 expression was significantly correlated with markers of infiltrating immune cells in breast cancer, including monocytes, neutrophils, TAMs, Th1, Tregs, and exhausted T cells. Moreover, gene–gene and protein–protein interaction networks for CCL20 were obtained by network construction, and the majority of interacting genes and proteins were associated with immune reactions.
CCL20 is a member of the chemokine family, and thus is involved in the pathological processes of arthritis, psoriasis, and other diseases.23–26 A growing number of studies have shown that high CCL20 expression is associated with diverse cancers. For example, Ding et al. found that CCL20 is upregulated in HCC patients, and that high CCL20 expression is positively correlated with tumor size, tumor number, vascular invasion, tumor differentiation, and recurrence; similar to this study, they also found that patients with higher CCL20 expression exhibited poorer overall survival and recurrence-free survival compared with those with lower CCL20 expression. 16 Moreover, HCC tumor angiogenesis, outgrowth, and distal metastasis were dramatically repressed by blocking CCL20 activity. 27 Studies have also examined the role of CCL20 in breast cancer progression. Upregulation of CCL20 significantly elevates the proliferation and invasion of TNBC cells and facilitates the growth of tumors in xenograft models. Furthermore, CCL20 promotes chemotherapy resistance in breast cancer cells by activating the NF-κB pathway through PKC and p38 MAPK, which subsequently elevates expression of the drug efflux-protein ABCB1, leading to continuous export of taxanes from cells, which is a mechanism of drug resistance in cancer cells. 22 Muscella et al. demonstrated that CCL20 accelerates breast cancer cell migration and invasion and enhances MMP-2 and MMP-9 mRNA and protein expression through the JAK2/STAT3 and PKC-α pathways, respectively. 20 Another study revealed that CCL20 overexpression dramatically enhanced cell invasion and MMP-2/9 secretion in basal-like TNBC cells, which accelerated the bone metastasis of breast cancer. Intraperitoneal injection of anti-CCL20 antibody significantly inhibited the bone metastasis of osteolytic breast cancer cells in mice. 28 Thus, treatments targeting CCL20 or its downstream signaling pathways may improve outcomes in breast cancer.
Notably, CCR6 was the gene most significantly correlated with CCL20 on the basis of our constructed network. Previous evidence has shown that the chemokine binding receptors primarily expressed on endothelial cells, immune cells, and some tumor cells are G protein-coupled receptors with seven transmembrane regions. Notably, CCR6 is the only binding receptor for CCL20,29,30 and is mainly expressed on B cells, memory and effector T cells, Th17, Tregs, and immature dendritic cells.31–34 High CCR6 expression is found in inflammatory bowel disease, rheumatoid arthritis, and other pathological autoimmune diseases.25,35 Importantly, CCR6 is upregulated in many cancers, where it promotes tumor development. For example, CCR6 expression is increased in colon cancer tissues compared with normal adjacent tissues, which facilitates the lymph node metastasis and distant metastasis of colon cancer. 36 Additionally, high levels of CCR6 have been observed in gastric cancer tissues and are significantly correlated with recurrence and poor overall survival. 37 CCR6 is elevated in HCC patients and is associated with tumor metastasis and poor prognosis. Moreover, CCR6 expression is positively correlated with CCL20 levels in HCC tissues. 27 Regarding breast cancer, elevated CCR6 expression has been found in primary breast cancer cells, and CCR6 activates diverse signaling kinases downstream if CCL20 such as AKT, JNK, and NF-kB, resulting in the proliferation and migration of breast cancer cells. 19 Thus, the CCL20/CCR6 axis is a prospective therapeutic target for breast cancer treatment.
According to our results, CCL20 mainly interacts with inflammatory cytokines and chemokines. The GeneMANIA database contains different modes of action among proteins, such as physical interactions, co-expression, co-localization, genetic interactions, pathways, and shared protein domains. The STRING database contains information on more than 5,000 species, more than 20 million proteins, and more than 3 billion interactions. These protein interactions include direct physical effects and co-expression correlations. Therefore, different predictive modes can lead to different results. Furthermore, different literatures may be included in the two databases, leading to slight differences in the predicted results. However, the two databases both showed that CCL20 interacted with chemokines and inflammatory factors, indicating that CCL20 might mediate the infiltration and inflammatory response of immune cells through inflammatory factors and chemokines.
Immunotherapy with inhibitors of the immune checkpoint, especially PD-L1/PD-1, has been an exceptional addition to the arsenal of treatments for solid tumors. As the most common cancer diagnosed in women worldwide, identifying the efficacy of immunotherapy for breast cancer is very important. Zhang et al. demonstrated that D-mannose promotes immunotherapy and radiotherapy in TNBC by degrading PD-L1. Mechanically, D-mannose can activate AMPK to phosphorylate PD-L1 at S195, leading to abnormal glycosylation and proteasomal degradation of PD-L1, which enhances T cell activation and T cell-mediated killing of tumor cells. Notably, the combination of D-mannose with anti-PD-1 therapy significantly repressed tumor growth and extended the lifespan of mice injected with breast cancer cells. 38 Shen et al. indicated that pharmacological disruption of the MTDH-SND1 complex by stabilizing Tap1/2 mRNA dramatically elevated tumor antigen presentation, synergizing with anti-PD-1 therapy in metastatic breast cancer. 39 Additionally, CTLA-4 is another negative regulator of T cell-mediated immunity. Evidence has shown that tumor cells of TNBC patients express CTLA-4 in different cell compartments, and this increased expression of CTLA-4 in patients may be a key mediator of immune escape. 40 Consistent with these studies, we also found that upregulation of CCL20 was positively correlated with markers of T cell exhaustion including PD-L1, PD-1, and CTLA-4, suggesting that CCL20 expression in breast cancer patients may promote immune escape.
However, our results need further validation by both in vivo and in vitro experiments to confirm the biological function of CCL20 in the pathogenesis of breast cancer. Additionally, suitable population data are needed to investigate the correlation between CCL20 and immunotherapy response.
Conclusion
This study indicated that CCL20 expression was significantly upregulated in breast cancer and significantly associated with different clinicopathological parameters, for example, patients with higher CCL20 expression exhibited poorer prognoses. Furthermore, CCL20 expression was significantly correlated with immune cell infiltration in breast cancer, including monocytes, neutrophils, TAMs, Th1 cells, Tregs, and exhausted T cells. Therefore, CCL20 is a prognosis-related biomarker for breast cancer that has great potential to be a prospective therapeutic target.
Footnotes
Acknowledgments
We acknowledge all the public databases involved in this study for providing their platforms as well as their contributors for uploading meaningful datasets. Users can download relevant data for free for research and publish relevant articles. Our study is based on open-source data, so there are no ethical issues or other conflicts of interest.
Author contributions
Hongyan Zhai conceived and designed the study and performed the analysis. Na Zhang, Dan Mo, and Tingting Qin prepared figures and the manuscript. All authors were involved in editing the manuscript and final approval of the submitted and published versions.
Data availability statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
