Abstract
Objectives
Chemokines have been suggested to play significant roles in the progression of malignant cancers. This study aimed to identify the chemokines related to malignant progression in thyroid carcinoma.
Methods
The mRNA expression levels of 52 chemokines were compared between differentiated thyroid cancer (DTC) samples and normal thyroid tissues from The Cancer Genome Atlas database; survival analysis was then performed on the basis of differentially expressed chemokines. A retrospective study was conducted on the level of differentially expressed chemokines in 76 DTC patients. Functional pathway analysis was performed to explore chemokine-related regulatory mechanisms.
Results
We identified 20 chemokines with differentially expressed mRNA levels through publicly available data. High levels of CCL22 and CCL26 were found to be related with metastasis in clinical DTC samples. High levels of CCL22 were found to be significantly related to poor prognosis in DTC patients. Pathway analyses revealed that cytokines might affect cancer progression through cytokine-cytokine receptor and cytokine-interleukin interactions.
Conclusions
CCL22 and CCL26 could serve as prognostic biomarkers in thyroid carcinoma.
Introduction
The incidence of thyroid carcinoma has steadily increased over the past few decades. 1 According to the latest statistics, there were approximately 567,000 new cases of thyroid carcinoma worldwide in 2018, ranking fifth in incidence among all cancers. 2 The most prevalent type of thyroid carcinoma is differentiated thyroid carcinoma (DTC), which accounts for approximately 90% of cases. Surgery, 131I therapy, and thyroid stimulating hormone suppression therapy have resulted in favorable prognoses for DTC patients, as reflected by 5-year survival rates of approximately 90%. 3 However, approximately 10% to 20% of DTC patients experience local recurrence and/or distant metastases within 10 years after the initial operation, 4 and the disease-specific mortality of this population can reach 70%. 5 Therefore, exploring proper diagnostic methods to identify distant metastasis is crucial to appropriately tailoring treatment in clinical practice. 6 The level of serum thyroglobulin (Tg) is one of the most common indicators for DTC patients who received 131I therapy in long-term follow-up. 7 However, the presence of antithyroglobulin antibody reduces the accuracy of Tg detection. Prognostic predictions for such patients with more advanced or high-risk disease remains a difficult problem that perplexes clinicians. Therefore, exploring potential biomarkers to identify the patients with a high risk of local recurrence and/or distant metastases is crucial for appropriate follow-up and to formulate treatment plans in clinical practice.
Chemokines are a group of small proteins that elicit physiological and immune responses by recruiting specific cell populations to sites of infection or malignancy. Specifically, chemokines play a pivotal role in tumor progression, angiogenesis, metastasis, and prognosis by affecting the leukocyte migration, inflammatory responses, lymphocyte homing and immune responses. 8 Several chemokine-chemokine receptor axes including CXCR1/2-CXCL1-3/5-8, CXCR3-CXCL9/10/11, CXCR4-CXCL12, and CXCL5/CXCR2, have been confirmed to have effects on protumor or antitumor immune effector cells in lung cancer, hepatocellular carcinoma, and gastric cancer tumors.9–12 Hence, studying chemokines can provide a novel understanding of the mechanisms of tumor development, disease evaluation, and even treatment strategies.
The majority of studies on chemokines in cancer have focused on patients with primary tumors, and chemokine information has seldom been reported in thyroid cancer patients with metastasis. To explore chemokine alterations in metastatic DTC, we performed a retrospective study in DTC patients using experimental exploration and bioinformatics analysis, with the aim of demonstrating that chemokine expression can provide significant information about disease progression.
Materials and Methods
Datasets and screening differentially expressed genes (DEGs)
The mRNA expression levels of 52 chemokines in thyroid carcinoma were downloaded from The Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov/). The search strategy was as follows: primary site (thyroid), data category (transcriptome profiling), project ID (TCGA-THCA), experimental strategy (RNA-Seq), and workflow type (HTSeq counts). The other filters were set as default. Practical Extraction and Reporting Language (Perl) was used to extract the sample information, generate the mRNA expression matrix, and annotate gene symbols. Data from a cohort of 567 cases (509 thyroid carcinoma samples and 58 normal samples) were obtained from TCGA. The EdgeR package from Bioconductor was used to screen DEGs between thyroid carcinoma and normal tissues. An adjusted P-value <0.001 and |log fold change|>1 were set as the cutoff criteria for DEGs. Considering the detection conditions and research interests, 52 chemokines were screened including CCL1, CCL2, CCL3, CCL4, CCL5, CCL7, CCL8, CCL11, CCL13, CCL14, CCL15, CCL16, CCL17, CCL19, CCL21, CCL22, CCL23, CCL24, CCL25, CCL26, CCL28, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL8, CXCL9, CXCL10, CXCL11, CXCL12, CXCL13, CXCL16, CX3CL1, IL2, IL4, IL5, IL6, IL7, IL10, IL13, IL15, IL16, IL18, IL19, IL20, IL26, IL27, IL32, IL34, TNF, and MIF.
Subjects and detection methods
This study was approved by the Ethics Committee of Shanghai Tenth People’s Hospital (SHSY-IEC-KY-4.0/16–18/01) and conducted in accordance with the guidelines provided by the World Medical Association and the Helsinki Declaration. Written informed consent was obtained from all study subjects.
DTC patients who underwent total thyroidectomy between January 2017 and June 2017 were recruited from the Thyroid Disease Center Nuclear Medicine Subcenter of Shanghai Tenth People's Hospital. Ultimately, 76 patients were included in this study, and we have de-identified all patient details. The reporting of this study conforms to STROBE guidelines. 13 The exclusion criteria for this study included: (1) patients with any chronic or infectious diseases, (2) patient who received surgery in the last 6 months, and (3) patients with other histories of malignancy.
The diagnostic criteria for tumor metastasis were: (i) pathology of postoperative and/or biopsy tissue suggested metastasis, and/or (2) therapeutic 131I whole body scan, single-photon emission computed tomography (SPECT/CT), and/or positron emission tomography/computed tomography (PET/CT) showed distant metastases (iodine uptake foci were localized to lungs, bones, and/or other soft tissues in accordance with characteristic anatomical imaging findings such as lung nodules and/or lytic bone lesions).
Venous blood was taken from DTC patients to detect the expression level of chemokines before thyroidectomy. The serum was processed in accordance with recommendations for the detection methodology. Briefly, 781 μL of standard buffer and 250 μL of standard diluent were added to the bottle as the sample dilution buffer (SBD). Next, 50 μL of sample, 60 μL of SBD, and 20 μL of serum were added to a 96-well detection plate after even dilution. All serum procedures were performed on ice. The human multifactor detection kit (171-AK99MR2 Bio-Plex Pro Human Chemokine Assays) and the liquid-phase microbead suspension protein chip were provided by Shanghai Huaying Biomedical Technology Co., Ltd. (Bio-Plex Magpix System; Shanghai, China) and used to test the level of chemokines. We used the 5-parameter mode (5PL) recommended by Bio–Rad to detect standards by fluorescence emission. The concentration of each chemokine was calculated according to the standard curve (concentrations are reported as pg/mL) after the deviation point was automatically corrected and the effective point was fit.
Statistical and survival analysis
Normally distributed data are expressed as mean±standard deviation, and non-normally distributed data are expressed as median (interquartile range). In accordance with the normality and homogeneity of variance of the data, t tests or rank-sum tests were used for comparisons between two groups, and single-factor ANOVA or rank-sum tests were used for comparisons between multiple groups. We used SPSS 20.0 software (IBM Corp., Armonk, NY, USA) for statistical analysis and GraphPad Prism 8.0 (GraphPad Software, Inc., San Diego, CA, USA) for data visualization. Receiver operating characteristic (ROC) curve analysis was performed to evaluate the specificity and sensitivity of key genes. Area under the curve (AUC) and P values were then calculated. Whether the expression levels of key chemokine genes were associated with overall survival was investigated using Kaplan–Meier plotter (http://www.kmplot.com/). P < 0.05 was considered statistically significant.
Evaluating expression levels with clinical characteristics
UALCAN (http://ualcan.path.uab.edu.) is an interactive web-portal that can perform in-depth analyses of gene expression data. 14 UALCAN provides a protein expression analysis option using data from the Clinical Proteomic Tumor Analysis Consortium (CPTAC) Confirmatory/Discovery dataset. We evaluated the protein expression of key genes in thyroid carcinoma, with P < 0.05 considered statistically significant.
Constructing the protein-protein interaction (PPI) network and co-expression analyses
The Search Tool for Retrieval of Interacting Genes (STRING, version 10.5, https://string-db.org/) database is an online tool that was used to construct the PPI network of key chemokine genes and to analyze functional interactions between the gene products. 15 An interaction with a combined score of >0.4 was considered significant (species: Homo sapiens). Protein co-expression analyses were achieved using GeneMANIA (http://www.genemania.org), a website that provides information about protein co-expression and colocalization.
Functional enrichment analysis
The Kyoto Encyclopedia of Genes and Genomes (KEGG) is a database that provides functional associations of gene and proteins. Gene ontology (GO) analysis provides a standardized description of gene products in terms of molecular function (MF), biological process (BP), and cellular component (CC). KEGG and GO functional enrichment analyses were conducted using the online tool DAVID (DAVID, https://david.ncifcrf.gov/) and visualized using the R package. P < 0.05 was considered statistically significant.
Results
Identification of DEGs in thyroid carcinoma
The expression levels of 52 chemokines in 567 samples are shown in a heatmap (Figure 1a). Twenty DEGs of chemokines were identified between thyroid carcinoma and normal samples, and the clustering volcano is shown in Figure 1b. There were 14 upregulated chemokines in tumor samples including CCL11, CXCL6, CCL13, CCL17, CXCL5, CCL26, CXCL8, CCL1, CCL7, CXCL1, CX3CL1, CCL15, CCL22, and CXCL16 (Figure 1c). Six chemokine genes were downregulated in tumor samples including CCL21, CCL19, IL2, IL6, CXCL13, and CCL2 (Figure 1d).

Expression of chemokines in a public database and screening for differentially expressed chemokines. (a) Clustering heatmap of 52 expressed chemokine mRNAs in 567 thyroid samples. The ordinate axis indicates the chemokine names. (b) Volcano plot showing the chemokine mRNAs. Red represents high expression, and blue represents low expression and (c) The 14 upregulated chemokine mRNAs in thyroid tumor tissues were CCL11, CXCL6, CCL13, CCL17, CXCL5, CCL26, CXCL8, CCL1, CCL7, CXCL1, CX3CL1, CCL15, CCL22, and CXCL16. (d) The six downregulated chemokine mRNAs in thyroid tumor tissues were CCL21, CCL19, IL2, IL6, CXCL13, and CCL2.
Clinical characteristics of the patients
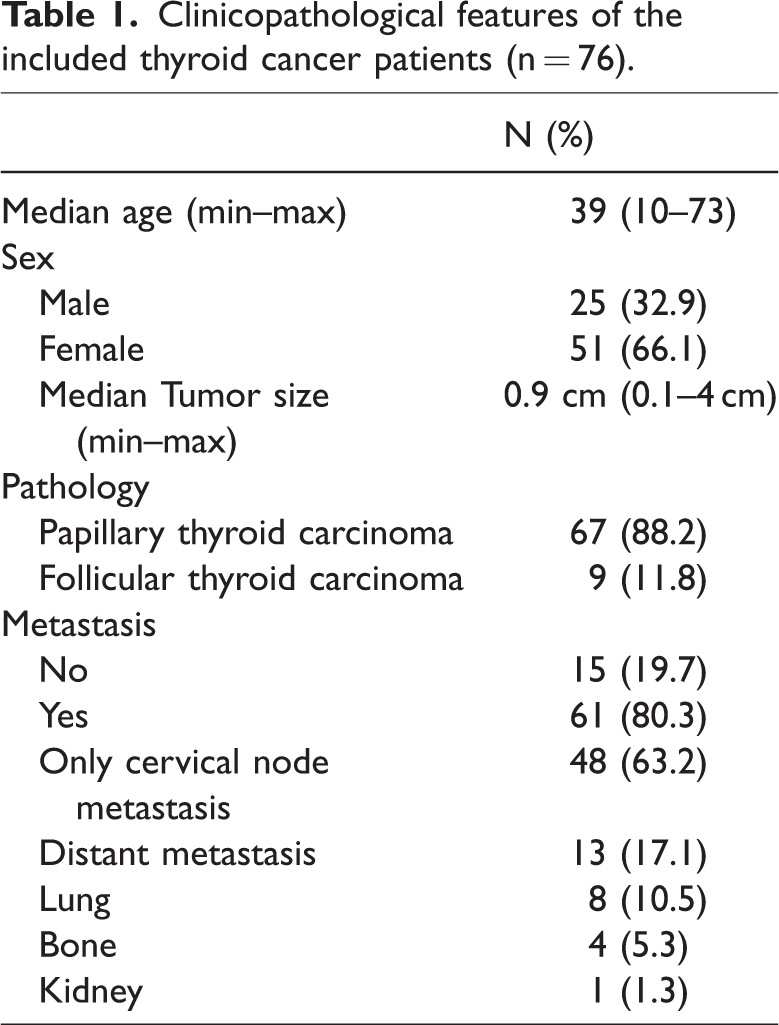
This study included 76 DTC patients, with a mean age of 39 years (range: 10–73), and the sex ratio (female:male) was 25:51. There were 48 patients with cervical lymph node metastasis only and 13 with distant metastasis. Basic clinical characteristics of the included patients are shown in Table 1. In the distant metastasis group, eight patients were diagnosed with lung metastases, four with bone metastases, and one with kidney metastases.
Clinicopathological features of the included thyroid cancer patients (n = 76).

Expression levels of DEGs and identification of key genes
Table 2 shows protein expression levels of the 20 DEGs preoperatively identified in DTC patients between the metastatic and non-metastatic groups. CCL22 expression was higher in the metastatic group (n = 61, 1148.24 (885.57, 1477.73) pg/mL) than in the non-metastatic group (n = 15, 909.05 (673.25, 1125.42) pg/mL), U = 258, P = 0.042, Figure 2a). Compared with the non-metastatic group, CCL22 expression was increased in the lymph node metastasis (n = 48, 1064.26 (894.364, 1355.28) pg/mL) and distant metastasis (n = 13, 1332.32 (908.305, 1608.24) pg/mL) groups, although no difference was found among the three groups (U = 4.805, P = 0.091, Figure 2b). CCL26 expression was higher in the metastatic group (n = 61, 25.82 ± 6.30 pg/mL) than in the non-metastatic group (n = 15, 21.76 ± 5.71 pg/mL, P = 0.030, t = 2.22, Figure 2c). CCL26 expression gradually increased in the non-metastatic group, only lymph node metastasis group (n = 48, 21.94 ± 5.60 pg/mL), and distant metastasis group (n = 13, 21.18 ± 6.22 pg/mL); however, no difference was found among the three groups (F = 2.797, P = 0.068, Figure 2d). According to ROC curve analysis, the AUCs of CCL22, CCL26, and the combination to predict metastasis in DTC were 0.678 (P = 0.045), 0.718 (P = 0.014), and 0.780 (P = 0.002), respectively (Figure 2e).
The expression levels of 20 chemokines in 76 differentiated thyroid cancer patients.

*Comparison between the metastatic and non-metastatic groups, P < 0.05. Normally distributed data are represented by mean ± standard deviation, and non-normally distributed data are represented by the median (interquartile range).
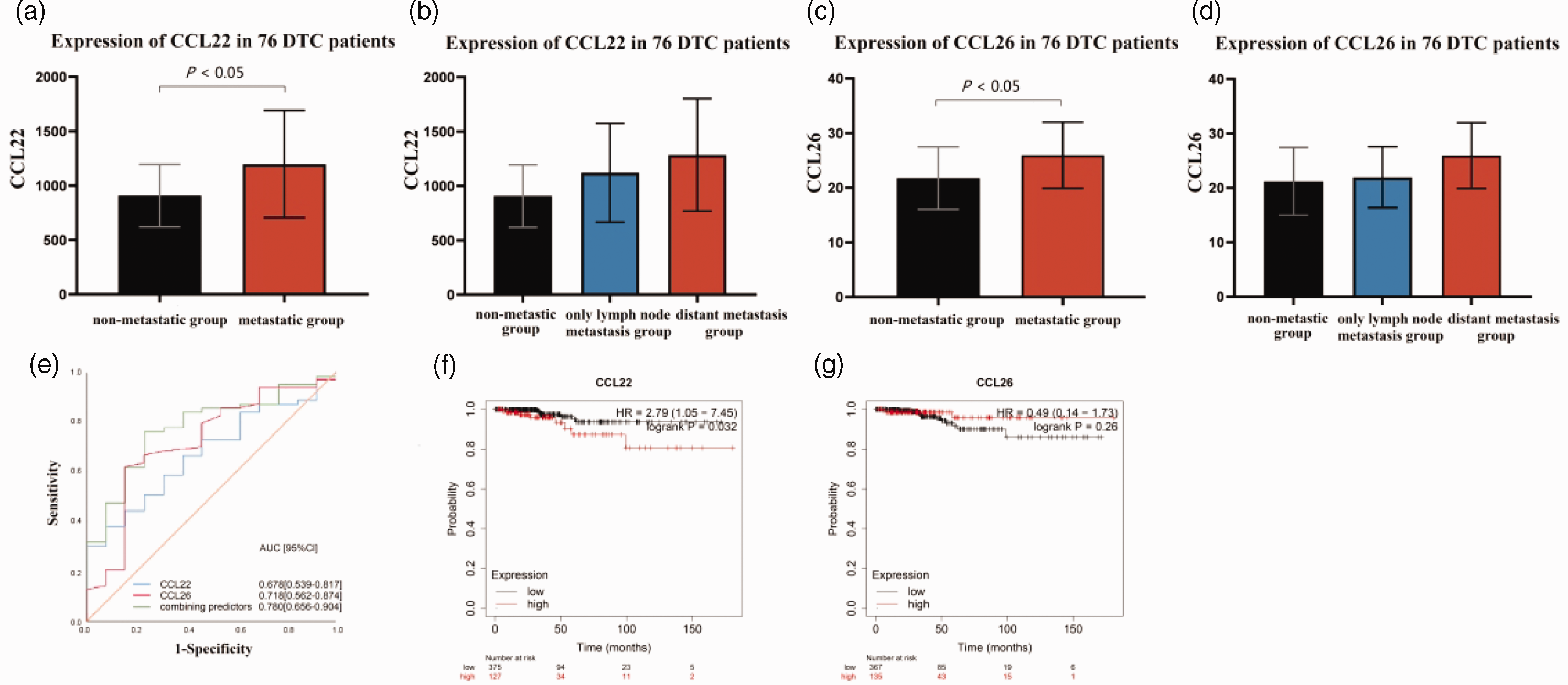
CCL22 and CCL26 expression in 76 DTC patients stratified by the presence or absence of metastatic lesions. (a) CCL22 expression in non-metastatic and metastatic differentiated thyroid cancer (DTC) patients. (b) CCL22 expression in DTC patients without metastases, with only lymph node metastasis, and with distant metastases. (c) CCL26 expression in non-metastatic and metastatic DTC patients. (d) CCL26 expression in DTC patients without metastases, with only lymph node metastasis, and with distant metastases. (e) Combined prognostic value of CCL22 and CCL26 in 76 DTC patients. (f) The prognostic value of CCL22 in thyroid carcinoma patients from The Kaplan–Meier Plotter database. (g) The prognostic value of CCL26 in thyroid carcinoma patients from The Kaplan–Meier Plotter database.
Kaplan–Meier analysis was then used to predict the prognostic value of CCL22 and CCL26. The results demonstrated that higher expression of CCL22 was associated with worse overall survival for thyroid cancer patients (hazard ratio [HR]: 2.79, 95% confidence interval [CI]: 1.05–7.45, log-rank P = 0.032, Figure 2f and 2g). In contrast, lower CCL26 expression may be associated with worse overall survival in thyroid carcinoma patients, although no significant difference was observed (HR: 0.49, 95% CI: 0.14–1.73, log-rank P = 0.260).The relationship between CCL26 expression and prognosis in thyroid cancer patients needs to be confirmed by more studies.
Clinical subgroup analysis
Subgroup analyses on the basis of tumor histology, age, disease stage, and tumor grade showed that the transcription levels of CCL22 were significantly higher in thyroid carcinoma patients than in healthy controls. The expression levels of CCL22 and CCL26 were different among different histological subtypes (Figure 3a and 3e). Tall thyroid papillary carcinoma showed the highest expression levels of CCL22 and CCL26. DTC patients of different ages showed higher CCL22 and CCL26 expression than healthy controls (Figure 3b and 3f).

CCL22 and CCL26 transcript expression levels in subgroups of patients with thyroid carcinoma, stratified by histology, sex, age, and other criteria (UALCAN). (a) Relative CCL22 expression in thyroid cancers of different histological grades. (b) Relative CCL22 expression in healthy controls and thyroid carcinoma patients aged 21–40, 41–60, 61–80, or 81–100 years. (c) Relative CCL22 expression in healthy controls and in thyroid carcinoma patients with or without nodal metastasis. (d) Relative CCL22 expression in healthy controls and stage I to stage IV thyroid carcinoma patients. (e) Relative CCL22 expression in thyroid cancers of different histological grades. (f) Relative CCL22 expression in healthy controls and thyroid carcinoma patients aged 21–40, 41–60, 61–80, or 81–100 years. (g) Relative CCL22 expression in healthy controls and thyroid carcinoma patients with or without nodal metastasis and (h) Relative CCL22 expression in healthy controls and stage I to stage IV thyroid carcinoma patients. Data are expressed as mean ± standard error; *P < 0.05; **P < 0.01; ***P < 0.001.
It was noteworthy that the expression of CCL22 in patients with lymph node metastasis was higher than in patients without lymph node metastasis (Figure 3c). CCL22 expression in stage 1 patients was significantly different from in patients with stage 2, stage 3, and stage 4 disease (Figure 3d). Notably, CCL22 expression was higher in advanced stages (stages 3 and 4). Similarly, the presence of lymph node metastasis and different individual cancer stages had an effect on the expression of CCL26 (Figure 3g and 3h). Thus, the expression levels of CCL22 and CCL26 in thyroid carcinoma were different between the metastatic and non-metastatic groups and may serve as potential diagnostic indicators for thyroid carcinoma progression.
PPI network construction and co-expression analyses
As expected, 11 nodes and 52 edges were obtained in the PPI network of CCL22 (Figure 4a). Network construction revealed 10 proteins that were significantly related to the expression of CCL22 including CCR7, CCR4, IL13, CXCR3, CCR8, IL4, CCR2, STAT3, IL10, and CCR1 (Figure 4a). There were 11 nodes and 43 edges in the PPI network of CCL26, and the proteins related to its expression included CCR1, CX3CR1, CXCR3, CCR3, CCR2, CCR8, IL13, IL4, STAT6, and SLC12A2 (Figure 4b). Text mining, co-expression, and protein homology were the main connections between experimentally and curated databases. GeneMANIA was used to validate the co-expression genes of key genes. The results demonstrated that PF4V1, CCL2, CCL8, CCL11, XCL1, CXCL12, CXCL14, CXCL10, CCL21, PPBP, CXCL9, CCL28, CXCL11, XCL2, CCL13, CXCL17, CCL17, and ACKR2 were correlated with CCL22 and CCL26 (Figure 4c).

(a) The protein-protein interaction (PPI) network significantly coregulated with CCL22 (CCR7, CCR4, IL13, CXCR3, CCR8, IL4, CCR2, CCR1, STAT3, and IL10). (b) The PPI network significantly coregulated with CCL26 (CCR1, CX3CR1, CXCR3, CCR3, CCR2, CCR8, IL13, IL4, STAT6, and SLC12A2). (c) GeneMANIA was used to validate the genes co-expressed with CCL22 and CCL26 and (d–g) Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses of the co-expressed genes with CCL22 and CCL26. P-values are displayed on the x-axis and GO functional enrichment and KEGG pathways are shown on the y-axis.
Functional networks in thyroid carcinoma
To further investigate the specific function of these chemokine genes, the key genes and their co-expressed genes were uploaded to the online tool DAVID. KEGG analysis revealed that the key genes were predominantly involved in the chemokine signaling pathway, cytokine-cytokine receptor interaction, NF-kappa B signaling pathway, and Toll-like receptor signaling pathway with their co-expressed genes (Figure 4d). GO analysis found that in terms of cellular components, the key genes were enriched in the extracellular space, extracellular region, and cell and external sides of the plasma membrane (Figure 4e). When analyzed in terms of biological processes, the key genes were associated with chemokine-mediated signaling pathways, chemotaxis, inflammatory responses, immune responses, and monocyte chemotaxis (Figure 4f). Regarding molecular function, the key genes were mainly enriched in chemokine activity, CCR chemokine receptor binding, CXCR3 chemokine receptor binding, heparin binding, chemokine receptor binding, CXCR chemokine receptor binding, and receptor binding (Figure 4g).
Discussion
This study used chemokine profiling to reveal the chemokine alterations that are related to DTC metastasis. We identified 14 upregulated genes and six downregulated genes in DTC samples compared with normal thyroid tissues using bioinformatics and identified CCL22 and CCL26 as key genes that are related to distant metastasis through experimental results. Our findings showed that CCL22 and CCL26 were highly expressed in the serum of DTC patients with distant metastasis, suggesting that high CCL22 expression is significantly related to poor prognosis in DTC by integrated analysis.
CCL22 is known as a macrophage-derived chemokine that is expressed by macrophages, dendritic cells, B cells, and T cells. CCL22 is constitutively expressed under homeostatic conditions and inducible upon inflammation.16,17 High CCL22 expression has been observed in various types of cancer. For instance, it has been found that CCL22 leads to reduced antitumor immunity in glioma. 18 CCL22 may also be an independent prognostic predictor in breast cancer. 19 These findings may help to demonstrate the possibility of CCL22 being a biomarker for predicting progression in thyroid cancer.
CCL22 might also be a promising treatment target for malignancies. A recent study showed that vaccines targeting CCL22 can induce the infiltration of CD8+ T cells and M1 macrophages into the tumor microenvironment, thereby enhancing anti-tumor ability in vivo. 20 Our findings suggest that the chemokine-receptor interaction is a key pathway and that CCR4 is an important interacting protein with CCL22. CCR4 is a transmembrane protein that acts as the receptor for CCL22, which is expressed by activated T helper 2 (Th2), regulatory T cells (Tregs), and memory CD4+ T cells. 21 A previous study reported that the CCL22/CCR4 axis can recruit regulatory T cells in tumors, leading to tumor immune escape. 22 This suggests a potential regulatory role for CCL22 in the immune escape of thyroid cancer that might explain the regulatory mechanism of elevated CCL22 in the serum of patients with distant metastases.
CCL26, also called eotaxin-3, macrophage inflammatory protein 4-alpha or thymic stroma chemokine-1, is another chemokine that we experimentally identified to be associated with distant metastasis, although statistical significance was not reached to suggest that it is associated with thyroid cancer prognosis. CCL26 is the chemoattractant for eosinophils and basophils and is produced by monocyte-derived macrophages, endothelial cells, epithelial cells, and smooth muscle cells. 23 As CCR3 and CX3CR1 have been demonstrated to be functional ligands for CCL26, CCL26 could recruit CX3CR1+ myeloid-derived suppressor cells (MDSCs) to tumor tissues. 24
Kawano et al. suggested that high levels of CCL26 in osteosarcoma were associated with increased motility and invasion in vitro. Meanwhile, they also showed an increased number of lung metastases in a lung metastasis model in nude mice in which recombinant CCL26 was added. 25 Another study demonstrated that CCL26 induces tumor-associated macrophage infiltration by binding to the CCL26 receptor in colorectal cancer. 26 Furthermore, hypoxia is a common feature of solid tumors, and CCL26 can be activated by hypoxia-inducible factors. 27 It has been found that chronic hypoxia not only increases CCL26 expression in ovarian cancer but also increases MDSC recruitment to hepatocellular carcinoma by increasing CCL26 expression. 28 Finally, we found increased CCL26 expression in the metastatic group, which is consistent with other studies.
The overall density of tumor-associated macrophages was significantly increased in PTC tumors compared with in benign thyroid disease, 29 indicating that lymphocytes were subjected to chemotaxis during tumor growth. Upregulated CCL22 expression in the tumor microenvironment might account for the increased proportion of Tregs, which are ideal cells for tumors to suppress spontaneous and therapy-induced T cell antitumor immunity and promote immune evasion. 30 Another way in which CCL22 may promote the occurrence and progression of malignancies is by inducing the accumulation of CCL22-dependent Th2 cells in the tumor microenvironment, resulting in an imbalance between Th1 and Th2 cells.31,32 Evidence suggests that chemokines and immune cells have a great influence on the tumor microenvironment through chemokine signaling and chemotaxis responses.
There are some limitations to this retrospective study. First, the CCL22 and CCL26 expression levels detected in our study reflect the preoperative DTC status, and the relationships between the trend of CCL22 and CCL26 expression and disease status during the 3-year course is unknown. Second, because the cohort of patients for protein profiling was of a relatively limited sample size, the patients were grouped and compared according to whether they had distant metastatic lesions. Third, studies of protumor and antitumor chemokine–chemokine receptor signaling pathways may be more meaningful for clinical practice than a single chemokine or chemokine receptor, and further mechanistic in vitro and in vivo experiments will be helpful for verification. These points will be addressed in follow-up studies.
Conclusion
This study found that two key genes, CCL22 and CCL26, were highly expressed in metastatic DTC patients compared with the non-metastatic group at both the transcriptional and protein levels. As potential prognostic target genes, CCL22 and CCL26 may regulate inflammatory cells and chemokines in the tumor microenvironment of thyroid carcinoma. These proposed chemokine-based prognostic markers not only provide promising biomarkers for thyroid carcinoma but also provide new insights into monitoring long-term treatment strategies.
Footnotes
Acknowledgements
With many thanks to my supervisors, Zhongwei Lv and Dan Li, for their guidance during this research.
Author contributions
Junyu Tong designed the statistical analysis and drafted this manuscript. Wen Jiang and Xiaoying Zhang performed the experiments. Ru Wang, Tingting Qiao, Yingchun Song, and Dingwei Gao contributed to data collection. Xiaqing Yu, Zhongwei Lv, and Dan Li conceived the project and reviewed the manuscript. All authors approved the final version of the manuscript. Junyu Tong, Wen Jiang, and Xiaoying Zhang contributed equally and share co-first authorship. Xiaqing Yu, Zhongwei Lv, and Dan Li contributed equally as corresponding authors.
Data availability statement
Some of the data supporting the findings of this study are available within public databases.
Declaration of conflicting interests
The authors declare that there is no conflict of interest that could be perceived as prejudicing the impartiality of this article.
Funding
The authors disclosed receipt (pending publication) of the following financial support for the research, authorship, and/or publication of this article: This study was sponsored by the Natural Science Foundation of China (82071964), Shanghai Leading Talent Program sponsored by Shanghai Human Resources and Social Security Bureau (2019), Key discipline construction project of the three-year action plan of Shanghai public health system by Shanghai Municipal Health Commission (GWV-10.1-XK09), the Natural Science Foundation of Shanghai (21ZR1449600), the Science and Technology Commission of Shanghai Municipality (22YF1433600) and Shanghai Tenth People's Hospital, Tongji University School of Medicine (2021SYPDRC064).
