Abstract
The most commonly used methods for pathogen detection and identification in oral mucosal infectious diseases are DNA or RNA quantitative polymerase chain reaction detection, bacterial or fungal cultures, and immunohistochemical analysis. These traditional methods are time-consuming and can only detect one specific targeted pathogen at a time. An efficient and sensitive method with higher species richness is urgently needed. Metagenomic next-generation sequencing (mNGS) is a new method of pathogen detection with high efficiency and sensitivity. In this case report, mNGS was used to identify the pathogens in oral mucosal tissues of a patient with complex oral mucosal infections and oral leukoplakia. Candida albicans, human gamma herpesvirus 4, and many other pathogens were identified using this method. For complex oral mucosal infections, mNGS is a more efficient and sensitive approach that can replace conventional detection methods.
Keywords
Introduction
The oral cavity has the second largest and most diverse microbiota, harboring over 700 species of bacteria. 1 The soft tissues of the oral mucosa are complex habitats where microbes colonize. Like systemic infections, oral mucosal infectious diseases are usually caused by bacteria, viruses, and fungi such as Porphyromonas gingivalis, herpes virus, and oral Candida, respectively. For viral-induced oral mucosal infectious diseases, the most common diagnostic method is quantitative polymerase chain reaction (qPCR) detection of viral DNA or RNA, which is highly efficient and sensitive. The limitation of this method is that only specifically targeted viruses can be detected, and the species richness is limited. For oral mucosal infectious diseases caused by bacteria and fungi, bacterial or fungal cultures are the most common diagnostic method. However, the time of culture can be relatively long, generally between 48 and 72 hours, and it can also be hard to distinguish different species from one another. 2 A method with high efficiency, sensitivity, and species richness is urgently needed to detect pathogens for oral mucosal infectious diseases.
Metagenomic next-generation sequencing (mNGS) is a new detection method with high efficiency and sensitivity that involves high-throughput sequencing; additionally, the turnaround time of mNGS is shorter compared with other detection methods. Pathogen detection and identification by mNGS is emerging in clinical practice. Only a small amount of DNA needs to be extracted from a sample to simultaneously detect and identify pathogens using mNGS. 3 Moreover, mNGS is not dependent on culture so it is more sensitive than conventional culture methods. 3 mNGS has already been successfully applied in several clinical trials to diagnose infections, and the results have shown that mNGS provides more precise and faster pathogen detection and identification. 4 However, after a review of the literature, mNGS has rarely been used to diagnose oral mucosal infectious diseases.
In this case report, we detail a patient who had undergone s/p allogeneic bone marrow transplantation and been taking immunosuppressants for a long time. Therefore, the patient was susceptible to oral mucosal infectious diseases. For safe and effective treatment, we used mNGS to identify the pathogens in oral mucosal tissues of the patient. We then discuss these results, which are of great significance to the treatment of patients with oral mucosal infectious diseases.
Case report
A 60-year-old male had been suffering from a rough feeling in the gingiva of his left maxillary posterior teeth and left buccal mucosa for 1 year. No tobacco-chewing or smoking habits were reported by the patient. The basic condition of this patient was also complex. In his medical history, the patient reported leukemia 13 years ago and had undergone s/p allogeneic bone marrow transplant in the same year, which led him to regularly take immunosuppressants (tab methylprednisolone, 4 mg orally per day) for several years. Suffering from insomnia and anxiety, the patient also takes zolpidem tartrate orally (10 mg at bedtime).
Clinical examination of the mouth on the initial visit was as follows: the mouth opening was only 27 mm, there were painless white verrucous plaque lesions with multiple peaks on the gingiva and vestibulum oris mucosal located in the left maxillary first molar, and the mucosa of the lesion area was fragile and prone to bleeding (Figure 1a). There was a white lesion with flat or slightly wrinkled surface located in the left buccal mucosa with an ulcer of approximately 3 mm in diameter (Figure 1b, c). According to the clinical symptoms, the patient was clinically diagnosed with oral leukoplakia (OLK) with infection. Incisional biopsy of the diseased mucosa was performed for a definitive diagnosis.

Photo and biopsy of a case of complex oral mucosal infection in a 60-year-old male. (a) Painless white verrucous plaque lesion with multiple peaks on the gingiva and vestibulum mucosal located in the left maxillary first molar. (b) White lesion with flat or slightly wrinkled surface located in the left buccal mucosa and (c) An ulcer of approximately 3 mm in diameter occurred in the left buccal mucosa.
The biopsy tissue was divided into two parts. One part was fixed in 10% formalin solution, and the other was stored at −80°C until DNA and RNA extraction. After embedding, the sample fixed in 10% formalin solution was used for hematoxylin–eosin staining for histopathology. The other tissue sample was used for mNGS. Briefly, 0.5 mL of BioSpec beads (0.5-mm diameter, ZIRCONIA/SILICA Cat. No. 11079105z, Thermo Fisher Scientific, Waltham, MA, USA) were added into the tissue sample. After vigorous agitation for 30 minutes, the sample was ready for DNA extraction. RNA was reverse transcribed and synthesized to double-stranded complementary DNA using the SuperScript II Reverse Transcription Kit (18064-014, Invitrogen, Waltham, MA, USA). After second-strand DNA synthesis, DNA libraries were constructed. Agilent 2100 (Santa Clara, CA, USA) was used for quality control, and DNA libraries were then sequenced using the BGISEQ-100 platform. Next, clean reads were mapped to a human reference database using Burrows–Wheeler Alignment (version 0.7.10). The mapped data were then processed for advanced data analysis.
Histopathological examination revealed squamous epithelial hyperplasia, hyperkeratosis, and inflammatory cell infiltrated into the connective tissue (Figure 2). Subsequently, a histopathological diagnosis of OLK was made. Further pathogenic microorganism DNA/RNA high-throughput genetic sequencing (PMseq)-DNA detection took approximately 36 hours, and the results showed overlapping infections with Candida albicans and Human gamma herpesvirus 4 (EBV). Other bacteria such as Rothia dentocariosa, Prevotella denticola, and Veillonella parvula, were also detected. The PMseq-DNA detection results are shown in Figure 3.
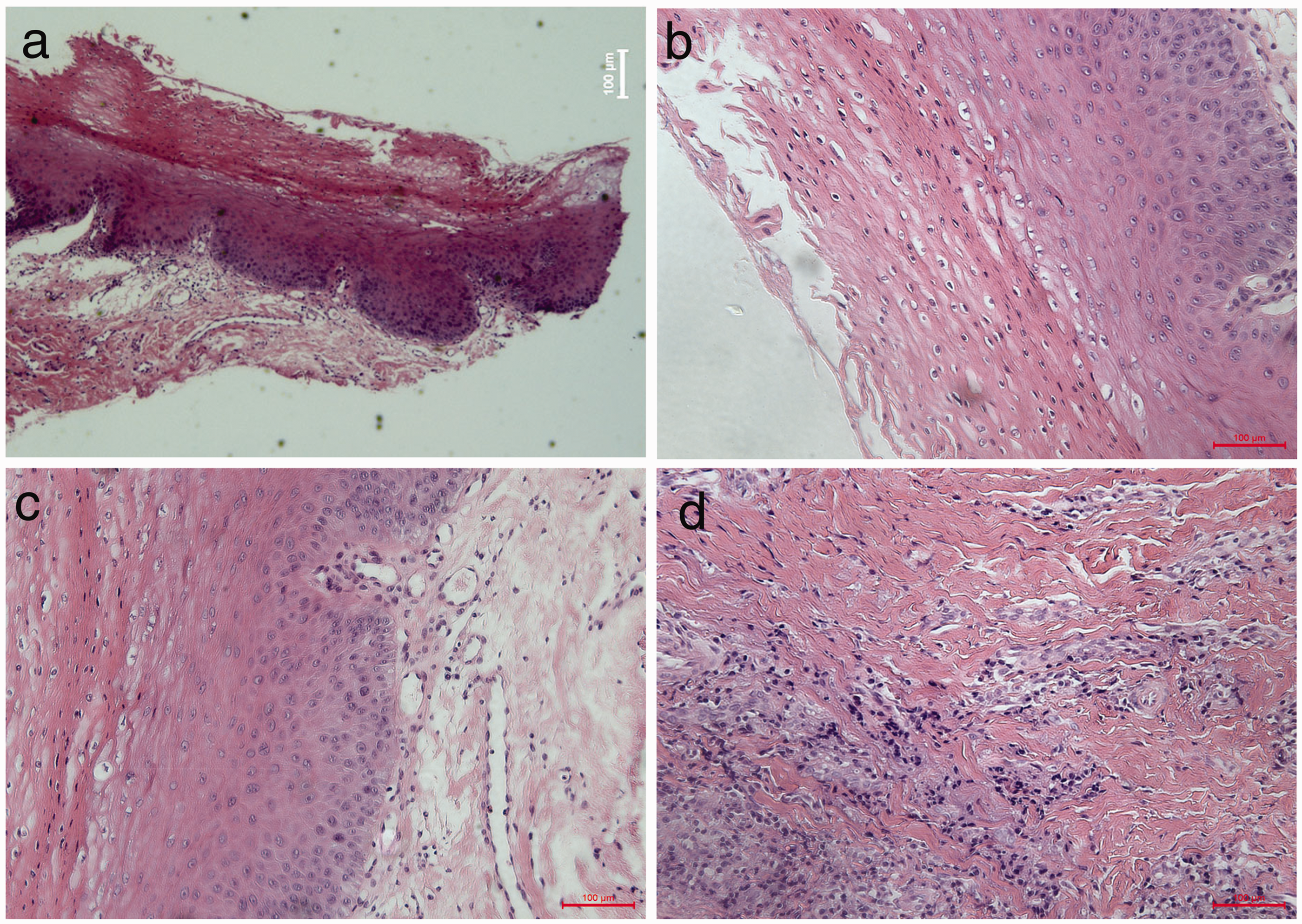
Hematoxylin–eosin staining for histopathology. (a) Squamous epithelial hyperplasia and hyperkeratosis. (b) Hyperkeratosis and parakeratosis. (c) Hyperplasia of the granular and spinous layers and (d) Inflammatory cell infiltration into the connective tissue.

Pathogenic microorganism DNA/RNA high-throughput genetic sequencing (PMseq)-DNA detection. PMseq-DNA detection was used to analyze the microbial nucleic acid sequences in the samples and identify the microorganisms by comparing them with the nucleic acid sequences of the existing microbiota in the database. (a) Candida albicans, (b) Human gamma herpes virus 4, (c) Rothia dentocariosa, (d) Prevotella enoeca, (e) Veillonella parvula, (f) Neisseria sicca and (g) Actinomyces gerencseriae.
On the basis of the PMseq-DNA detection results, 200 mg oral fluconazole was prescribed on the first day, and then 100 mg on the second day, for a total of 7 days. Acyclovir (200 mg orally) was prescribed 5 times per day for 7 days. Additionally, 2% sodium bicarbonate solution was used as mouthwash after eating. The follow-up visit was 2 months later, at which time the ulcer in the left buccal mucosa showed complete response after treatment and the surface of the white lesions had become flatter (Figure 4).

Photo of the case of complex oral mucosal infection 2 months after treatment. The white lesions in the left buccal mucosa had faded, and the ulcer had completely healed.
The reporting of this study conforms to the CARE guidelines. 5
Discussion
In this case, we used mNGS to detect the pathogens in the oral mucosal tissues of a patient with complex oral mucosal infection and OLK who was susceptible to oral mucosal infectious diseases due to s/p allogeneic bone marrow transplantation and use of immunosuppressants. The turnaround time of mNGS was 36 hours, which is shorter than other detection methods. mNGS detects pathogenic microorganisms in infected samples, not the tissue itself and can simultaneously detect almost all bacteria, viruses, and fungi. Compared with traditional detection methods, mNGS has higher efficiency, sensitivity, and species richness. For complex oral mucosal infections, mNGS can replace conventional detection methods.
OLK is a common and potentially malignant oral disorder that has a high malignant transformation rate of between 0.13% and 34% into oral squamous cell carcinoma. 6 The etiology of OLK is complex, and infection is considered to be one of the major pathogenic factors of OLK. Candida infection has been found in 34% of OLK, and Candida invasion is a significant risk factor for malignant transformation in OLK. EBV infection 7 is also strongly associated with the malignant transformation of OLK. The oral microbial environment is complex. Early detection and inhibiting the growth of pathogens in the oral cavity is of great significance for the treatment and inhibition of malignant transformation in OLK.
Candida infection may cause damage to the oral mucosa, such as atrophy, hyperplasia, and dysplasia. 8 Candida can be divided into many different species, such as Candida albicans, Candida tropical, and Candida glabrata. 9 Candida culture is the most commonly used method in the clinical diagnosis of oral Candida. The most frequently used primary isolation medium for Candida culture is Sabouraud Dextrose Agar (SDA). 10 However, the time of culture is relatively long, generally between 48 and 72 hours, and it is hard to distinguish different species from one another. 2 Pathogen detection and identification using mNGS can only take approximately 36 hours or less, and different species of Candida can be accurately distinguished. For patients with HIV infection, on corticosteroids or antibiotic therapies, and in post-transplant immunosuppression have higher infection rates for oral candidiasis. 11 In this case, the patient had been regularly taking immunosuppressants for 5 years, which is a predisposing systemic factor for pathogenic Candida. Polyene and azole are effective treatments for Candida albicans infection. Studies have found that fluconazole is superior to other drugs, showing a statistically significant improvement in its therapeutic effect for oral candidiasis.12,13 Candida albicans prefers an acidic environment. Using 2% sodium bicarbonate solution to maintain an alkaline environment within the oral cavity can inhibit the growth of Candida albicans. Inhibiting the growth of Candida albicans is of great significance for the treatment and inhibition of malignant transformation in OLK.
EBV is a human herpes virus and an opportunistic pathogen in immunocompromised patients. 14 EBV is associated with a large number of infectious diseases such as Hodgkin's lymphoma, Burkitt's lymphoma, and nasopharyngeal carcinoma. 15 Moreover, EBV infection is an important risk factor for malignant transformation in OLK. 16 Accordingly, anti-viral therapy is of great importance. Acyclovir is a commonly used drug to inhibit EBV, which is used as an oral treatment in this case. Besides, EBV infection requires regular follow-up.
Rothia dentocariosa, Prevotella denticola, and Actinomyces gerencseriae were also identified at the same time. Rothia dentocariosa is a part of the normal flora of the human oral cavity and pharynx. 17 Prevotella denticola and Actinomyces gerencseriae are often regarded as principal causes of periodontitis.18–20
In conclusion, we applied mNGS to pathogen detection and identification for a patient with complex oral mucosal infection. The results revealed that mNGS had higher efficiency, sensitivity and species richness compared with traditional detection methods. Thus, mNGS could significantly improve the treatment of oral mucosal infectious diseases.
Footnotes
Acknowledgement
We would like to thank Patho-Genesis (Shenzhen, China) for assistance with metagenomic next-generation sequencing services.
Author contributions
YH Song performed the data analysis and wrote the manuscript. FJ Song performed hematoxylin–eosin staining for histopathology. YH Song, FJ Song, and S Chen contributed to data analysis and manuscript preparation. ZF Song and SF Liu contributed to the conception of the study and helped perform data analysis with constructive discussions. YH Song and FJ Song contributed equally to this work.
Data availability
All data are available from the corresponding author by request.
Declaration of conflicting interests
The authors declare no competing interests.
Ethics statement and informed consent
The patient provided informed consent for treatment at the initial visit. The patient was well informed of all purposes that might be used for research, and consent for publish was signed by the patient.
Funding
The authors disclosed receipt (pending publication) of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Shanghai (22ZR1454200) and Shanghai Stomatological Hospital Talent Project (SSDC-2019-RC01) to Shangfeng Liu.
