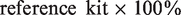Abstract
Objective
To assess the detection performance of the hematopoietic stem cell enumeration kit developed by BD Biosciences.
Methods
Cord blood samples were prepared using a hematopoietic stem cell enumeration kit developed by BD Biosciences and Stem-Kit reagents from Beckman Coulter. CD34+ cells were enumerated using a BD FACSCanto instrument and FACSDiva software.
Results
A total of 519 samples were analyzed in this study. The hematopoietic stem cell enumeration kit developed by BD Biosciences yielded absolute counts of CD34-positive cells that were on average 8.7% lower than Beckman Coulter Stem-Kit reagents (range: −5.7% to−14.7%). The BD Biosciences kit yielded relative counts that were on average 9.9% higher compared with Beckman Coulter Stem-Kit reagents (range: −2.1% to +13.8%). The intraclass correlation coefficients for absolute and relative counts of CD34-positive cells were 0.9967 (95% confidence interval [CI]: 0.9961–0.9972) and 0.9512 (95% CI: 0.9423–0.9587) for the BD Biosciences and Beckman Coulter kits, respectively.
Conclusions
The hematopoietic stem cell enumeration kit developed by BD Biosciences can be used to enumerate CD34-positive stem cells from cord blood samples.
Introduction
Cord blood-derived cells are alternative sources of hematopoietic stem cells (HSCs) for either allogeneic or autologous transplantation. Advantages of cord blood-derived cells include an expanded donor pool, lower risk to donors, higher proliferative capacity, and reduced histocompatibility.1–3 CD34+ cell content is a widely used predictive factor for accessing engraftment and survival following umbilical cord blood transplantation.4–6 CD34 is expressed on HSCs and is considered a reliable marker for stem cells with regenerative capacity. 7 Despite the major limitation of lower HSCs, 8 with a single cord blood unit containing approximately 10% fewer stem cells than bone marrow, the number of CD34+ cells obtained per unit volume is similar. 9 Moreover, CD34+ cell count is reported to correlate with the total nucleated cell dose, which is crucial for optimal clinical transplantation outcomes. 10 , 11 Although lower doses can also be successful, 8 most studies have reported that the number of viable CD34-positive HSCs is correlated with velocity of engraftment.9–11
Existing techniques for the enumeration of CD34-positive HSCs are typically dual-platform methods, in which total white blood cells are enumerated by an automated hematology analyzer and CD34+ cell frequency is assessed using flow cytometry. The main disadvantage of these methods is the combination of the inherent errors of the two methods; these errors can be variable among laboratories, impairing standardization. Therefore, single-platform methods are now preferred because they allow for the direct enumeration of CD34-positive HSCs 12 and their use in China is supported by current guidelines. 13 Available single-platform methods are based on the guidelines from the International Society of Hematotherapy and Graft Engineering (ISHAGE) and use sequential gating steps.12,14,15 The current guidelines recommend use of commercially available reagents from Beckman-Coulter (Brea, CA, USA);12,14,15 thus, these reagents should be considered as the gold standard for the evaluation of novel methods. Viability evaluation via addition of a dye (e.g., with the percentage of live and dead cells at each passage determined using the trypan blue exclusion test12,14–16) is recommended, especially for previously frozen samples. 17 BD Biosciences (San Jose, CA, USA) has developed a Trucount-based single-platform HSC enumeration kit based on the ISHAGE guidelines 15 that uses the 7-aminoactinomycin D (7-AAD) viability dye. This single-platform method does not require washing after lysing red blood cells and does not require the use of a hemocytometer to count white blood cells; therefore, the system error is small and it is considered the preferred method for CD34+ cell enumeration. 12
Previous studies have not examined the reliability of the BD™ stem cell enumeration kit for cord blood in China. Cord blood stem cells represent an important source of HSCs for transplantation. 18 Therefore, this study aimed to compare the detection performance of the HSC enumeration kit (flow cytometry) developed by BD with that of the Stem-Kit reagents from Beckman Coulter, which has already been granted CE certification.
Materials and methods
Study design and samples
This was an open-label, multicenter, parallel, controlled, registered technical trial. Umbilical cord blood samples were collected at a cord blood bank (Shanghai Stem Cell Technology Co., Ltd., Shanghai, China) from April to August 2016 and CD34+ HSCs were enumerated at three hospitals: the Shanghai Tenth People’s Hospital (SH10), Shanghai General Hospital (SHGH), and Shanghai Jiaotong University School of Medicine affiliated to Shanghai Children Medical Center (SCGC). The study was approved by the Ethics Committees of all three participating hospitals (Shanghai Children’s Medical Center, Shanghai JiaoTong University School of Medicine, SCMCIRB-L201404-05; Shanghai Tenth People’s Hospital of Tongji University, SHSY-IEC-3.0/15-48/02; and Shanghai General Hospital, Shanghai JiaoTong University School of Medicine, 2014-SJ-056-X161). The requirement for individual consent was waived by all three committees because the study used anonymized samples remaining following routine clinical work.
Fresh cord blood and cryopreserved cord blood were tested in the study (both from the cord blood bank). The inclusion criteria were: (1) sufficient volume (≥600 µL) or volume after dilution, if applicable; (2) white blood cell count <30 × 103/µL or count after dilution with low-temperature phosphate-buffered saline (PBS) (2–8°C) containing 0.5% bovine serum albumin (BSA), if necessary; and (3) fresh samples or anti-coagulated using ethylenediaminetetraacetic acid (EDTA), heparin, citrate phosphate dextrose, and/or acid citrate dextrose solution formula A. The exclusion criteria were: (1) coagulation or hemolysis; and (2) incorrect collection, storage, or processing of the sample.
CD34-positive stem cell enumeration kits
The primary goal of this study was to evaluate the CD34+ HSC enumeration kit (flow cytometry) developed by BD Biosciences [BD stem cell enumeration (SCE) kit]. BD SCE reagents include BD Stem Cell Reagent [CD45-fluorescein isothiocyanate (FITC)/CD34-phycoerythrin (PE)], 7-AAD, 10× ammonium chloride lysing solution, and 50 BD Trucount tubes.
The reference kit was the BCI Stem-Kit™ reagents from Beckman-Coulter (BCI Stem-Kit). Stem-Kit reagents include a CD45-FITC/CD34-PE ready-to-use reagent; CD45-FITC/IsoClonic Control-PE reagent (a mixture of unconjugated and PE-conjugated monoclonal antibody clone 581); Stem-Count Fluorospheres containing a dye with a fluorescence emission range of 525 to 700 nm; 7-AAD; and NH4Cl lysing solution.
Measurement processes
On each day, fresh or thawed cord blood samples were selected according to the inclusion and exclusion criteria. Fresh samples had to be measured within 24 hour after collection, while thawed samples were measured within 1 hour after thawing in each laboratory at the three hospitals. All samples were stored at 2–8°C. Before staining, cells were counted in samples by microscopy or using an automatic blood cell counter. Samples with white blood cell counts >30.0 × 103/µL were diluted using low-temperature PBS (2–8°C) containing 0.5% BSA before staining. Samples with granules or lipemia were diluted and filtered before staining even if the white blood cell count was <30.0 × 103/µL.
For the BD SCE kit, acquisition/analysis was performed using BD FACSCanto clinical software, version V2.4 or higher and BD SCE module, version V1.0.1 or higher on a BD FACSCanto II cytometer. For the BCI Stem kit, acquisition/analysis was performed using BD FACSDiva software, version V7.0 or higher and BCI FACSDiva experimental template, version V1.6 or higher on a BD FACSCanto II cytometer.
Quality control
Before the start of the study, all personnel involved at the clinical research unit were trained in the use of the flow cytometer (BD FACSCanto II) to ensure that all three hospitals were gating the same way using the same software template. FACSCanto clinical software and seven-color setup beads were used to calibrate the flow cytometer. After successful calibration (all parameters in the calibration report showing “Pass”), the cytometer was optimized for the BD Stem Cell software, BD SCE kit, and BD stem cell control (BD SCC) staining using 7-AAD. The same parameters for setup and optimization were used for FACSDiva software.
The BD SCC was stained/lysed using the “low-quality content” and “high-quality content” reagents provided with the BD SCE kit. The BD Stem Cell software from the BD FACSCanto II flow cytometer was used to enumerate CD34+ cells in the BD SCC after staining/lysing. If the results (absolute count of CD34+ cells and percentage of CD34+ cells among CD45+ cells) were in the target value range, quality was considered acceptable.
BCI Stem-Trol quality control cells were added to one sample of “normal” complete blood (EDTA anticoagulation) to obtain the BCI quality control sample, which was then subjected to staining/lysing using the reference kit. The BCI FACS Diva mode in the BD FACSDiva software was used for enumeration. If the measurement results (normalized absolute counts of the quality control material) were in the target value range, the quality was considered acceptable.
Data collection
The absolute count of viable CD45+/CD34+ double-positive HSCs, as well as the percentage of viable CD34+ cells among viable CD45+ cells (relative counts), were obtained.
Sample size
The kits used in this study were type III in vitro diagnostic reagents. According to the Technical Guidelines for Clinical Trials of In Vitro Diagnostic Reagents and Review Guidelines for the Registration of Testing Reagents Specifically for Flow Cytometry issued by the China Food and Drug Administration, at least 500 assessable cord blood samples should be included in the study.
Statistical methods
SAS 9.4 (SAS Institute, Cary, NY, USA) was used for statistical analysis. A per-protocol set analysis was conducted in this study. For all continuous variables, a normality test was first conducted using the Kolmogorov–Smirnov test. Normally distributed data were described as means ± standard deviations, while non-normally distributed data with were described as medians and ranges. Qualitative data were described as frequencies and percentages.
The absolute and relative errors between the results from the study and reference kits were calculated according to the following equations.
The mean absolute and relative errors of each indicator, as well as the 95% confidence intervals (95% CIs), were calculated.
For each indicator, scatterplots of the results were generated. Correlation and regression analyses were conducted for the two kits, and the determination coefficient (R2), regression equation, intercept, slope, and 95% CIs were calculated. The interclass correlation coefficient (ICC) was used for consistency analysis.
For the Bland–Altman analysis, the difference between each pair of results was calculated using the equation: error = results of using test reagent – results of using predict reagent. Qualitative results of indicators were described as above. The reference range was calculated according to the 95% interval (P2.5 to P97.5) of healthy subjects (according to the t distribution and u distribution). For all assessable samples, regression tests showed that R2 was >0.90 and the 95% CI of the slope was between 0.85 and 1.15. Values of P < 0.05 were considered statistically significant.
Results
From April to August 2016, 557 samples from the three hospitals were eligible. After applying the exclusion criteria, 519 samples were included in the study: 297 from the SCMC, 116 from the SH10, and 106 from the SHGH. Among all samples, 43.6% were fresh and 56.4% were thawed, while 53.4% were from male donors and 46.2% were from female donors (Appendix Table 1).
Error analysis
The BD SCE kit yielded lower absolute CD34+ cell counts than the BCI Stem kit (mean: −8.7%) for all samples, ranging from −5.7% (SHGH) to −14.7% (SCMC). The BD SCE kit yielded higher relative CD34+ cell counts than the BCI Stem kit (mean: +9.9%) for all samples, ranging from −2.1% (SH10) to +13.8% (SCMC) (Appendix Table 2).
ICC consistency analysis
To assess the value of the BD SCE kit, CD34+ cells were enumerated using the BD SCE kit and the BCI Stem kit and ICCs were calculated. The ICC for absolute counts of CD34+ cells was 0.9967 (95% CI: 0.9961–0.9972) for all samples and ranged from 0.9960 (95% CI: 0.9942–0.9972) (SH10) to 0.9971 (95% CI: 0.9958–0.9981) (SHGH). The ICC for relative counts of CD34+ cells was 0.9512 (95% CI: 0.9423–0.9587) for all samples and ranged from 0.8949 (95% ranging CI: 0.8519–0.9259) (SH10) to 0.9688 (95% CI: 0.9545–0.9787) (SHGH; Appendix Table 3). Measurement of CD34+ cells in cord blood using the BD™ stem cell enumeration kit was highly consistent at all three hospitals (all ICCs >0.95; Figure 1).

Examples of gating protocols. (a) BD SCE kit, acquisition/analysis using BD FACSCanto clinical software; (b) BCI Stem kit, acquisition/analysis using BD FACSDiva software.
Regression analysis
To assess the correlation between values measured using the BD SCE kit and the BCI Stem kit, a regression analysis was performed. The R2 value for absolute counts of CD34+ cells comparing the two kits was 0.99, with a regression equation of y = −0.38 + 0.98x (Figure 2). This suggested that measurements of absolute counts of CD34-positive cells using the two methods were highly linear and nearly equivalent. The R2 for relative counts of CD34-positive cells comparing the two kits was 0.91, with a regression equation of y=0.03 + 0.95x (Figure 3). Thus, measurements of relative counts of CD34-positive cells using the two methods was also highly linear and nearly equivalent. Appendix Figures 1–3 show the individual results for the three hospitals.

Linear regression between absolute counts of CD34+ cells in cord blood determined using the BD SCE kit and the BCI Stem kit. Measurements of absolute counts of CD34-positive cells using the two methods were highly linear and nearly equivalent.

Linear regression between relative counts of CD34+ cells in cord blood determined using the BD SCE kit and those from the BCI Stem kit. Measurements of the relative counts of CD34-positive cells between the two methods were highly linear and nearly equivalent.

Bland–Altman plot of absolute counts of CD34+ cells in cord blood showing agreement between the BD SCE kit and the BCI Stem kit. The two measurement methods had high agreement for absolute counts of CD34-positive cells.

Bland–Altman plot of relative counts of CD34+ cells in cord blood showing agreement between the BD SCE kit and the BCI Stem kit. The two measurements methods had high agreement for relative counts of CD34-positive cells.
Bland–Altman analysis
To assess agreement between the BD SCE kit and the BCI Stem kit, a Bland–Altman analysis was performed. As shown in Figure 4, most differences in absolute counts of CD34-positive cells between the two kits fell within the mean value ± 1.96 standard deviations. Thus, the two measurement methods had high agreement for absolute counts of CD34-positive cells. As shown in Figure 5, most differences in relative counts of CD34-positive cells between the two kits fell within the mean value ± 1.96 standard deviations. As shown in Appendix Figures 1–3, the individual results for the three hospitals demonstrated that the two measurement methods had high agreement for relative counts of CD34-positive cells.
Reference ranges
Using the BD SCE kit, the reference range (P2.5 to P97.5) for absolute viable CD34-positive cells was 3.50 to 66.10 cells/µL (median: 16.81 cells/µL) and the reference range (P2.5 to P97.5) for relative counts of CD34-positive cells was 0.09 to 0.99 (median: 0.34) (Appendix Table 4).
Discussion
Enumeration of CD34-positive stem cells is critical for autologous and allogeneic stem cell transplantation protocols. 12 Available methods have disadvantages such as low reproducibility and reliability as well as lack of evaluation of cell viability. Therefore, this study aimed to assess the detection performance of the HSC enumeration kit developed by BD Biosciences in comparison with that of Stem-Kit reagents from Beckman Coulter. The latter kit has already been granted CE certification.
Despite differences observed in the absolute and relative counts of CD34-positive cells between the BD SCE kit and the BCI Stem kit, the correlations between the two kits were linear for the entire range of detected cells (R2 > 0.95 for absolute counts of CD34+ cells). Some differences were observed among the three hospitals (minimum R2 value 0.8 for relative counts of CD34+ cells at Shanghai Tenth People’s Hospital). These findings may be explained by differences in staff technical experience and quality control that are not clinically relevant. In addition, the BD SCE kit does not require bead pipetting, which could potentially reduce errors19–21 and result in slightly higher total counts. To the best of our knowledge, this is the first study to compare the results of CD34-positive cell enumeration in cord blood using these two kits.
Our results are supported by those of Dauber et al., 20 who compared the BD SCE kit and the BCI Stem kit using peripheral blood, bone marrow, and cord blood. Furthermore, Omana-Zapata et al. 21 and Preti et al. 22 evaluated the BD SCE kit using the FACSCalibur and FACSCanto II flow cytometry systems and reported similar performance for the two systems. Similar results were reported by Massin et al. 23 using the FACSCanto and FACSCanto II flow cytometry systems.
The present study did not specifically examine cell viability differences using fresh vs. thawed samples between the two kits. Dauber et al. 20 reported that fresh and thawed cells had similar CD34-positive cell frequencies, independently of sample origin and isolation kit used. Of course, gating strategy can lead to significant variability in results between hospitals, and only the ISHAGE protocol has yielded reproducible results. 24 In the present study, all three hospitals used the same ISHAGE-based protocol, and the results were consistent among all three. Aliquoting samples and beads is another major source of variation. 16 The BD SCE kit includes lyophilized pellets to remove this source of aliquoting error.
Most transplantation protocols require a minimum number of CD34+ cells/kg for successful engraftment.25,26 Therefore, accurate enumeration of viable CD34-positive cells is critical for treatment success. Our results strongly suggest that the BD SCE kit can be used to enumerate CD34-positive stem cells from cord blood samples. The performance of the BD SCE kit was comparable with that of the CE-certified Stem-Kit reagents from Beckman Coulter.
The present study had several limitations. Because only cord blood samples were tested, and only three centers were included, we could not evaluate the performance of these kits for cord blood compared with other sample types. In addition, the kits were not tested across multiple flow cytometry platforms, which might also have resulted in different performance.
Conclusion
The BD SCE kit yielded results that were consistent with the reference CE-approved BCI Stem kit.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605211055384 - Supplemental material for Enumeration of viable CD34+ cells in cord blood using a novel stem cell enumeration kit
Supplemental material, sj-pdf-1-imr-10.1177_03000605211055384 for Enumeration of viable CD34+ cells in cord blood using a novel stem cell enumeration kit by Sheng Huang, Kangli Xu, Shanshan Xu, Shijun Yu, Yan Miao and Juan Wang in Journal of International Medical Research
Footnotes
Acknowledgements
The authors acknowledge the help of Jerry Zhang, Jiahui Shi, Lucy Sun, and Namei Hu from Beckton Dickinson for coordination of the study and reviewing the study report. We also acknowledge the contributions of Lei Zhang from Kun Tuo in designing the statistical analysis plan and completing the final statistical analysis report. Help from the laboratory technicians of the Shanghai Cord Blood Bank in providing and de-identifying specimens is gratefully acknowledged.
Author contributions
S.H., K-L.X.: designed and performed the study, acquired and analyzed data, and co-wrote the paper. S-S.X.: performed the study and acquired data. S-J.Y.: provided flow-cytometry analyses and critical revision. Y.M.: supervised the study. J.W.: designed the study and co-wrote the paper. All authors read and approved the final manuscript.
Declaration of conflicting interest
Shijun Yu is employed by BD Biosciences, which funded this research. Four employees of BD Biosciences coordinated the study and reviewed the manuscript. The other authors have no conflicts of interest to declare.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and publication of this article: This study was funded by BD Biosciences (San Jose, CA, USA) and the Shanghai “Rising Stars of Medical Talent” Youth Development Program – Clinical Laboratory Practitioner Program.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.