Abstract
Objective
Circular RNA (circRNA) plays a vital role in the development and progression of malignancies, however, the function of circRNAs in cholangiocarcinoma (CCA) remains unexplored. The aim of this study was to investigate circRNA expression in CCA versus para-cancer tissues, and elucidate any potential associated mechanisms.
Methods
Differential expression of circRNAs between CCA and para-cancer tissue was analysed by microarray hybridization, and validated by real-time quantitative reverse transcription–polymerase chain reaction (qRT–PCR). The downstream pathway was investigated using bioinformatics and qRT–PCR.
Results
Microarray hybridization revealed 10 circRNAs with > 3-fold increased expression versus para-cancer (circRNA_002172, circRNA_002144, circRNA_001588, circRNA_000166, circRNA_000585, circRNA_000167, circRNA_402608, circRNA_006853, circRNA_001589, circRNA_008882), and three circRNAs with > 3-fold decreased expression (circRNA_406083, circRNA_104940, circRNA_006349). CircRNA_000585 was shown by qRT-PCR to be upregulated in tumour versus paired para-cancer tissue from 15 patients with CCA. Bioinformatics analysis revealed a potential pathway comprising circRNA_000585/microRNA-615-5p/angiomotin (AMOT)/Yes associated protein 1 (YAP) in CCA. RT–PCR validation of crucial molecule expression showed downregulation of miR-615-5p, and upregulation of AMOT and YAP in CCA tumours.
Conclusion
Multiple circRNAs are dysregulated in CCA. CircRNA_000585 is upregulated in CCA, and may function by a circRNA_000585/miR-615-5p/AMOT/YAP pathway, which may be a novel CCA pathway.
Introduction
Cholangiocarcinomas (CCAs) are malignant biliary epithelial tumours involving the bile duct tree, that represent the second most common hepatic malignancy. They are categorized as intrahepatic, perihilar, or distal CCA, according to anatomical location, and most patients are diagnosed at advanced stages and cannot undergo resection. 1 For patients with unresectable cholangiocarcinoma, standard systemic therapies are limited, and median overall survival (OS) is below 1 year. 2 Many risk factors and molecules are implicated in cholangiocarcinoma, however, evidence supporting each risk factor and the molecular pathogenesis of CCA remains limited. 3
Circular RNA (circRNA) is a type of RNA with tissue/development-specific expression patterns, generated from exonic and/or intronic sequences by back-splicing. The biological functions of circRNA include regulation of transcription, competition with linear RNA splicing, acting as a microRNA (miRNA) sponge, protein decoy or scaffold, and translation of regulatory peptides. 4 Accumulated evidence shows that circRNA is dysregulated in malignant tumours, and most circRNAs target miRNA as miRNA sponges. The dysregulated circRNAs affect many cellular functions including proliferative signalling, cell migration and invasion, cell apoptosis, and angiogenesis. 4
In CCA, various circRNAs participate in different functions, and may be correlated with prognosis and clinical factors. For example, circRNA CDR1as is highly expressed in tumour tissue compared with adjacent normal tissues, and is associated with tumour, node, metastasis (TNM) stage, lymph node invasion, and postoperative recurrence. 5 Circ-SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 5 (SMARCA5) is inversely correlated with Eastern Cooperative Oncology Group performance score, T and N stages, TNM stage, and abnormal carbohydrate antigen (CA) 19-9 status in CCA. 6 CircRNAs may also be involved in migration, invasion and proliferation of CCA cell lines. For example, an in vitro study revealed that circRNA CDR1as was significantly increased in CCA cell lines, and promoted CCA cell proliferation, migration, and invasion by targeting miR-614. 7 Exosomal mediated circ_0000284 may be a potential biomarker in CCA, as it has shown increased expression in tumour tissues compared with surrounding normal tissues, and this increased expression was associated with enhanced migration, invasion and proliferation in CCA cell lines by targeting lymphocyte antigen-6E. 8 A network analysis identified a potential circ2174-associated genetic pathway in intrahepatic cholangiocarcinoma, whereby circ2174 acts as an miR-149 sponge, which targets octamer-binding transcription factor 2. 9 Circ-0000284 has been shown to promote CCA progression and may be directly transferred from CCA cells to surrounding normal cells via exosomes. 10 Circ_0001649 overexpression was shown to inhibit cell proliferation, migration and invasion by inducing apoptosis in CCA cells in vivo. 11 Finally, circ_0005230 might act as an effective therapeutic target for CCA by sponging miR-1238 and miR-1299. 12
The aim of the present study was to identify the differential expression of circRNAs in CCA, and explore a potential mechanism of circRNAs in CCA.
Patients and methods
Study population
This study included tissue samples collected between July 2019 and December 2019 from consecutive patients with CCA who were being treated at the Second Affiliated Hospital of Nanchang University, Nanchang, China. Diagnosis of CCA was based on pathology of the resected specimen, and patients with oncological or infectious comorbidities were excluded from the study. Demographic and clinical data were collected from patients’ medical records.
Tumour tissue was selected and validated by pathology, and para-cancer tissue was defined as normal tissue situated between 1 and 3 cm from the tumour edge. Selected tissues were snap frozen in liquid nitrogen for 30–60 s, and stored at –80°C prior to use. Total RNA was extracted using Trizol reagent (Invitrogen, Carlsbad, CA, USA) according to a standard protocol, and quantified with a NanoDrop ND-1000 spectrophotometer (ThermoFisher Scientific, Waltham, MA, USA).
The study was approved by the Ethics Committee of the Second Affiliated Hospital of Nanchang University and written informed consent was obtained from all patients who provided samples for the study.
Microarray hybridization
Three CCA specimens and three matched para-cancer specimens from three randomly selected patients with CCA were analysed by microarray hybridization using the Arraystar Human circRNA Array V2 (Arraystar, Rockville, MD, USA), based on Arraystar standard protocols, as previously described. 13 Briefly, total RNA from each sample was digested with Rnase R (Epicentre, Madison, WI, USA) to remove linear RNAs and enrich circRNAs. The enriched circRNAs were then amplified and transcribed into fluorescent cRNA utilizing a random priming method (Arraystar Super RNA Labelling Kit; Arraystar), according to the manufacturer’s instructions. The labelled cRNAs were purified using an RNeasy Mini Kit (Qiagen, Hilden, Germany), and their concentration and specific activity (pmol Cy3/μg cRNA) were measured with the NanoDrop ND-1000. Labelled cRNA (1 µg of each) was then fragmented by adding 5 µl of 10× blocking agent and 1 µl of 25× fragmentation buffer and incubating at 60°C for 30 min. The labelled cRNA mixture was diluted by adding 25 µl of 2× hybridization buffer, and 50 µl of the hybridization solution was transferred into the gasket slide, which was then assembled onto the circRNA expression microarray slide (Arraystar Human circRNA Array V2 [8 × 15 K]). Slides were incubated for 17 h at 65°C in an Agilent hybridization oven (Agilent Technologies, Santa Clara, CA, USA), washed using gene expression wash buffer, then scanned with an Agilent G2505C DNA microarray scanner (Agilent Technologies).
Array image analysis
Acquired array images were analysed with Agilent Feature Extraction software, version 11.0.1.1 (Agilent Technologies), as previously described. 13 Extracted raw data underwent quantile normalization and subsequent data processing using the R software limma package, version 3.25.15 (bioinf.wehi.edu.au/limma). Statistically significant differentially expressed circRNAs between tumour and para-cancer tissue were identified through volcano plot filtering, with fold-change threshold values of 1.5 and statistically significant P values of <0.05 (Student’s t-test). Differentially expressed circRNAs between tumour and para-cancer tissue groups were identified through fold-change filtering. Hierarchical clustering was performed to show distinguishable circRNA expression patterns among the samples. When comparing two groups of profile differences (tumour versus para-cancer tissue), the ‘fold change’ (i.e. the ratio of group averages) between the groups for each circRNA was computed.
Selection and identification of circRNAs
Out of circRNAs with fold changes >3 and P values <0.05, circRNA_000585 was selected for further investigation by real-time quantitative reverse transcription–polymerase chain reaction (qRT–PCR) in paired tissue samples (tumour and para-cancer) from 15 patients with CCA. Briefly, total RNA was extracted from 100 mg of tissue/sample using Trizol reagent (Invitrogen) and reverse transcribed using QuantiTect Reverse Transcription kit (Qiagen), according to the manufacturer’s instructions. The cDNA products were then amplified by real-time qPCR using QuantiTect PCR kit (Qiagen) with the circRNA_000585 and 18S rRNA internal reference primers detailed in Table 1. PCR was performed in an Applied Biosystems QuantStudio™ 5 real-time PCR system (ThermoFisher Scientific) with the following reaction conditions: 40 cycles of denaturation at 95°C for 10 s, annealing at 60°C for 60 s, and elongation at 72°C for 60 s. PCR products normalised to 18S rRNA were quantified using the 2–ΔΔct method.
Primer details for quantitative real-time reverse transcription–polymerase chain reaction of circRNA_000585.

F, forward; R, reverse; bp, base pairs.
Prediction and identification of miRNA
The circRNA/miRNA interaction was predicted with Arraystar's miRNA target prediction software (based on TargetScan and miRanda), and the differentially expressed circRNAs within all comparisons were annotated in detail with circRNA/miRNA interaction information. As miR-615-5p was found to interact with circRNA_000585, miR-615-5p expression levels were evaluated by qRT–PCR in paired tissue samples (tumour and para-cancer) from 15 patients with CCA, as described above. RT reactions were followed by qPCR under the following reaction conditions: 1 cycle of initial denaturation at 95°C for 30 s, 40 cycles of denaturation at 95°C for 5 s, annealing at 60°C for 30 s and elongation at 72°C for 60 s. The miR-615-5p and U6 internal reference primer sequences, annealing temperatures and product lengths are listed in Table 2.
Primer details for quantitative real-time reverse transcription–polymerase chain reaction of microRNA miR-615-5p.

F, forward; R, reverse; GSP, gene specific primer; bp, base pairs.
Prediction and identification of protein/miRNA interactions
MiRNA/protein interactions were searched in the miRNA Pathway Dictionary Database (miRPathDB; https://mpd.bioinf.uni-sb.de/) and revealed that miR-615-5p targets angiomotin (AMOT). Subsequent STRING protein interaction network and function analysis (https://string-db.org/) showed that AMOT interacts with Yes associated protein 1 (YAP); thus, expression levels of AMOT and YAP in paired tissue samples (tumour and para-cancer) from 15 patients with CCA were evaluated by qRT–PCR, as described above, under the following conditions: 40 cycles of denaturation at 95°C for 30 s, annealing at 60°C for 30 s, and elongation at 72°C for 120 s. The AMOT, YAP and 18S rRNA internal reference primer sequences, annealing temperatures and product lengths are listed in Table 3.
Primer details for quantitative real-time reverse transcription–polymerase chain reaction of angiomotin (AMOT) and Yes associated protein 1 (YAP) mRNA.

F, forward; R, reverse; bp, base pairs.
Statistical analyses
Statistical analyses were performed using SPSS software, version 23 (IBM, Armonk, NY, USA). Continuous data are presented as mean ± SD, and categorical data are presented as percentage and number of cases. Paired samples t-test was used to compare different expression levels between cancer and para-cancer tissues, and χ2-test with Yates’ continuity correction or Fisher’s exact test was performed to assess the association between demographic/clinical variables and circRNA_000585 expression in tumour tissue. All calculated P-values were 2-sided and a P value < 0.05 was considered statistically significant.
Results
Tumour and para-cancer tissues from a total of 15 patients were included in the study (seven male and eight female patients; mean age, 60.67 ± 7.52 years [range, 44–71 years]).
CircRNA_000585 is highly expressed in CCA versus para-cancer tissue
Differential expression of circRNAs in CCA versus para-cancer tissue from three patients with CCA is presented as heat maps (Figure 1 and 2). A total of 117 circRNAs were found to be upregulated and 104 circRNAs were downregulated in CCA versus para-cancer tissues, including 10 circRNAs that were >3-fold higher (circRNA_002172, circRNA_002144, circRNA_001588, circRNA_000166, circRNA_000585, circRNA_000167, circRNA_402608, circRNA_006853, circRNA_001589, and circRNA_008882), and three circRNAs that were >3-fold lower (circRNA_406083, circRNA_104940, and circRNA_006349) (Figure 3 and 4). Of these, analyses by qRT–PCR showed that circRNA_000585 was upregulated in tumour tissues versus paired para-cancer tissue specimens from 15 patients with CCA (P = 0.003; Figure 5).

Heatmap of circular (circ)RNA expression levels in cholangiocarcinoma (CCA) tumour specimens (P1–3) and matched para-cancer specimens (C1–3) from three patients with CCA. Each block represents different circRNA expression levels (red represents high expression; green represents low expression).

Fold change in circular (circ)RNA expression levels in cholangiocarcinoma (CCA) tumour specimens and matched para-cancer specimens from three patients with CCA. (A) scatter plot (dots above the upper line represent fold-change >1.5, dots below the lower line represent fold-change >–1.5); and (B) volcano plot of circRNA expression (red dots represent fold-change >1.5 or >–1.5).

Venn diagram showing upregulated circRNAs in cholangiocarcinoma (CCA) tumour specimens versus matched para-cancer specimens from three patients with CCA. Each circle represents different fold change between the two groups.

Venn diagram showing downregulated circRNAs in cholangiocarcinoma (CCA) tumour specimens versus matched para-cancer specimens from three patients with CCA. Each circle represents different fold change between the two groups.

Vertical scatter plot showing elevated circRNA_000585 expression in cholangiocarcinoma (CCA) tumour tissue versus matched para-cancer tissue from 15 patients with CCA (central horizontal line represents mean, upper and lower horizonal lines represent SD; P = 0.003 between groups).
Association between circRNA_000585 expression and clinicopathological characteristics
Associations between circRNA_000585 expression in tumour tissue and clinicopathological or demographic characteristics in patients with cholangiocarcinoma were investigated. No clinicodemographic factors, including age, sex, carcinoembryonic antigen, CA 19-9, tumour location, tumour size, or extrahepatic metastasis were found to be associated with circRNA_000585 expression (Table 4).
Association between circRNA_000585 expression and clinicopathological characteristics in 15 patients with cholangiocarcinoma.

Continuous data presented as mean ± SD.
CEA, carcinoembryonic antigen; CA 19-9, carbohydrate antigen 19-9; I, intrahepatic cholangiocarcinoma; P, perihilar cholangiocarcinoma; D, distal cholangiocarcinoma.
No statistically significance differences in expression between subgroups (all P > 0.05).
miR-615-5p is downregulated in CCA versus para-cancer tissue
Predictions of circRNA/microRNA interactions identified miR-615-5p as a downstream molecule of circRNA_000585. Comparison by qRT–PCR of miR-615-5p expression between tumour and para-cancer tissues, from 15 patients with CCA, showed that miR-615-5p is significantly downregulated in CCA tumour tissues (P < 0.05; Figure 6).

Vertical scatter plot showing decreased miR-615-5p expression in cholangiocarcinoma (CCA) tumour tissue versus matched para-cancer tissue from 15 patients with CCA (central horizontal line represents mean, upper and lower horizonal lines represent SD; P < 0.05).
AMOT/YAP is upregulated in CCA versus para-cancer tissue
Using miRPathDB software, miR-615-5p was found to target AMOT, and STRING protein interaction network and function analysis revealed interactions between AMOT and YAP. Investigation of AMOT and YAP expression by qRT–PCR showed that AMOT and YAP were both upregulated in CCA tumour tissue compared with para-cancer tissue (P = 0.019, Figure 7; and P = 0.0098, Figure 8, AMOT and YAP, respectively).

Vertical scatter plot showing increased angiomotin (AMOT) expression in cholangiocarcinoma (CCA) tumour tissue versus matched para-cancer tissue from 15 patients with CCA (central horizontal line represents mean, upper and lower horizonal lines represent SD; P = 0.019).
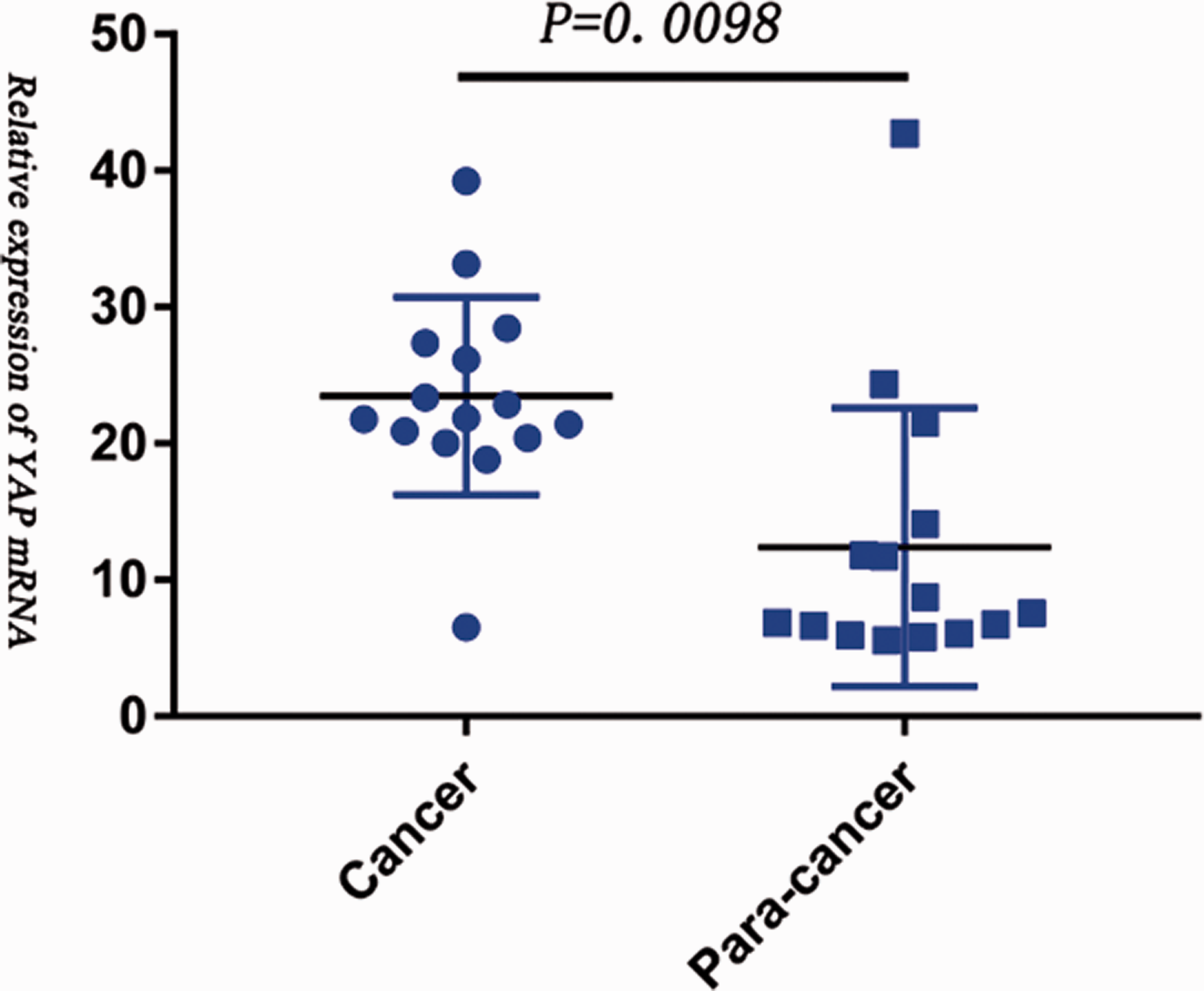
Vertical scatter plot showing increased Yes associated protein 1 (YAP) expression in cholangiocarcinoma (CCA) tumour tissue versus matched para-cancer tissue from 15 patients with CCA (central horizontal line represents mean, upper and lower horizonal lines represent SD; P = 0.0098).
Discussion
CircRNAs have been shown to function in various cancers as microRNA sponges, and regulators of parental gene expression, during transcription, post-transcription, and protein translation. They are highly stable in body fluids, such as plasma and saliva, which makes them ideal potential cancer biomarkers. 14 Dysregulation of circRNAs has been linked with different cancer phenotypes, and altered expression closely correlates with various pathological processes. CircRNAs also play complex roles that affect the malignant behaviours of cancer in the tumour microenvironment. 15
As indicated earlier, circRNAs, including cirRNA CDR1as, circ_0000284, circ2174, circ-SMARCA5, circ_0001649, and circ_0005230, play multiple roles as biomarkers, and in tumour proliferation, migration and invasion.5–12 The present study aimed to explore all cholangiocarcinoma-related circRNAs to reveal circRNA expression that was significantly related to CCA, using high-throughput circRNA microarray and bioinformatics analysis. One hundred and seventeen circRNAs were found to be upregulated and 104 circRNAs were downregulated in CCA, including 10 circRNAs that showed 3-fold higher expression than para-cancer tissue (circRNA_002172, circRNA_002144, circRNA_001588, circRNA_000166, circRNA_000585, circRNA_000167, circRNA_402608, circRNA_006853, circRNA_001589, and circRNA_008882), and three circRNAs that showed 3-fold lower expression than para-cancer tissue (circRNA_406083, circRNA_104940, and circRNA_006349). CircRNA_000585 was then identified by RT–PCR to be upregulated in tumour and paired para-cancer specimens from 15 patients with CCA. CircRNA_00585, also known as hsa_circ_0000515, is a novel circRNA in CCA. The gene is located in chr14:20811305-20811534, with a genomic length of 229 bp (www.circbase.org/cgi-bin/singlerecord.cgi?id=hsa_circ_0000515).
Regulatory networks can be established when circRNAs bind to other molecules forming circRNA–DNA, circRNA–RNA, or circRNA–protein interactions. MiRNA sponging is the most common process for circRNAs, and the circRNA–miRNA–mRNA regulatory axis is related to multiple functions in cell signalling and cancer biogenesis. 16 In the present study, circRNA/microRNA interactions were predicted using software based on TargetScan and miRanda, and the differentially expressed circRNAs within all comparisons were annotated in detail with circRNA/miRNA interaction information. MiR-615-5p was revealed to interact with circRNA_000585, and analysis of miR-615-5p expression by qRT–PCR showed that miR-615-5p is significantly downregulated in CCA compared with para-cancer tissues. MiR-615-5p is highly conserved and involved in embryogenesis, and in the regulation of growth and development during osteogenesis and angiogenesis. MiR-615-5p also participates in regulating cell growth, proliferation and migration, and has been shown to act as a tumour promoter or tumour suppressor, for example, through post transcriptional inhibition of oncogenes involved in essential biological processes, mainly by preventing invasiveness and metastasis, the main causes of tumour reappearance. 17
Using miRPathDB software in the present study, miR-615-5p was found to target AMOT, and STRING protein interaction network and function analysis revealed that AMOT interacts with YAP. Subsequent qRT–PCR showed that AMOT and YAP were upregulated in CCA tissue compared with para-cancer tissue. AMOT has been identified as an extracellular interacting protein that localizes in the membranous, cytoplasmic or nuclear fraction in a cell context-dependent manner, and controls migration, tight junction formation, cell polarity, and angiogenesis. 18 AMOT has also been demonstrated to participate in cancer initiation and progression, and previous studies have focused on the involvement of AMOT in the Hippo–YAP1 pathway. 18 Additionally, AMOT has been shown to promote nuclear translocation of YAP and act as a transcriptional cofactor of the YAP–transcriptional enhanced associate domain (TEAD) complex to facilitate proliferation of biliary epithelial cells and cancer development of the liver, either in response to tissue injury or in the absence of the tumour suppressor Merlin. 19
The AMOT family members promote proliferation and invasion of cancer cells, including breast cancer, osteosarcoma, colon cancer, prostate cancer, head and neck squamous cell carcinoma, cervical cancer, liver cancer, and renal cell cancer. However, in glioblastoma, ovarian cancer, and lung cancer, AMOT inhibits the growth of cancer cells. In addition, there are controversies on the regulation of YAP by AMOT, as AMOT has been shown to promote either the internalization of YAP into the nucleus or the retention of YAP in the cytoplasm of different cell types. 20
Yes-associated proteins are encoded by paralogous genes, which are transcriptional coactivators with 46% amino acid identity. Upregulated activity of YAPs are found in many malignancies, where they promote cell proliferation and overgrowth. 21 Forced YAP expression has been shown to potently promote cell proliferation in vitro, cause organ overgrowth and lead to the occurrence of cancer. 22 YAP also performs other functions, such as induction of epithelial to mesenchymal transition. 23 In addition, YAP/ transcriptional co-activator with PDZ-binding motif (TAZ) has been increasingly recognised to regulate cellular metabolism associated with phenotypic changes, and recent studies suggest that YAP/TAZ-mediated metabolic alterations contribute to metastasis. 24 YAP has also been demonstrated to promote tumour proliferation, angiogenesis and chemoresistance in CCA via TEAD transcription factors. 25 Hyperactivation of YAP/TAZ may promote the development of intrahepatic CCA, and some molecules, such as WW and C2 domain-containing 1 and neurofibromatosis type 2, may suppress CCA development by inhibiting the oncogenic activity of YAP/TAZ. 26
Although the present results are informative, they may be limited by several factors. First, the patient sample size was relatively small, and the low numbers are underpowered to show any statistical significance. Thus, the results need to be confirmed in future, using a larger sample size. The potential pathway implicated in the study should be verified by in vitro and in vivo research. Lastly, further studies are required to reveal the detailed mechanism associated with circRNA_000585 in CCA.
In conclusion, the differential expression of circRNAs in CCA were explored and circRNA_000585 was found to be highly expressed in CCA. A potential mechanism of circRNA_000585 in CCA may involve a circRNA_000585/miR-615-5p/AMOT/YAP pathway, although the present results are limited and further research is required to support these results.
Footnotes
Availability of data and materials
The datasets used and/or analysed in the present study are available from the corresponding author upon reasonable request.
Acknowledgement
We acknowledge the patients who participated in this study.
Author contributions
FMY and LF were responsible for conception and design of the research. FMY and LXX carried out the research and statistical analyses. FMY and LF supervised the paper writing. All authors read and approved the final manuscript.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was supported by Jiangxi Provincial Department of Science and Technology (grant number 20203BBGL73144) and Jiangxi Provincial Education Department (GJJ150141).
