Abstract
Objective
To comprehensively evaluate the significance of circular RNAs (circRNAs) as potential diagnostic biomarkers for systemic lupus erythematosus (SLE) via pooled analyses of data from published studies that focussed on the association between circRNAs and SLE.
Methods
The systematic review and meta-analysis protocol was registered in PROSPERO (registration No. CRD42021229383). Relevant studies published before 3 April 2022 were selected to verify the relationship between circRNA expression levels and SLE. Extracted data were analysed using a random-effects model with Meta-DiSc 1.4 and Stata 16 software. Transcription factors related to hsa_circ_0000479 and its parental gene were extracted from the TRCirc and hTFtarget databases, respectively.
Results
A total of 10 studies, involving 438 patients with SLE and 434 controls, were included in the meta-analysis. The pooled sensitivity, specificity, and diagnostic odds ratio of circRNAs in detecting SLE were 0.66 (95% confidence interval [CI] 0.63, 0.70), 0.79 (95% CI 0.76, 0.82), and 10.80 (95% CI 6.58, 17.73), respectively. The area under the summary receiver operating characteristic curve was 0.8366.
Conclusions
Meta-analysis of pooled data indicated a moderate accuracy of circRNAs in diagnosing SLE. The exact diagnostic value of circRNAs and the mechanisms of interaction between circRNAs and their parental genes should be confirmed in further studies.
Introduction
Systemic lupus erythematosus (SLE) is a chronic, complex, and potentially fatal autoimmune disease that is related to disordered innate and adaptive immune systems, which may culminate in excessive activation of immune complexes, biological waste production, and inflammatory damage. 1 The common clinical symptoms include changes in the mucocutaneous system, joints, central nervous system, and haematological system, while some patients may develop lupus nephritis, which is a serious complication of SLE. The diagnosis of SLE is mainly based on the revised American College of Rheumatology (ACR) classification criteria, and delay in diagnosis is associated with increased damage to vital organ systems.2,3 Patients may present with clinical symptoms and immunological abnormalities that indicate SLE but do not meet the classification criteria, with only 10–55% of patients advancing to classified disease. 4 In one study, healthy controls tested positive for antinuclear antibody (ANA) at a titre of ≥1: 80 in 8% of cases, and the prevalence of positive anti-double stranded DNA in patients with SLE was 37%. 5 Additionally, the conventional ANA test lacks specificity, while the sensitivity of the anti-C1q, anti-histone and anti-Sm antibody test is relatively weak. 6 Therefore, there is an urgent need to identify novel reliable biomarkers for use in distinguishing patients with SLE, to help prevent organ damage and mortality.
Numerous circular RNAs (circRNAs) have been identified using the rapidly developing technologies of bioinformatics and high-throughput sequencing. CircRNAs can bind to microRNAs (miRNAs) and act as miRNA sponges, thereby indirectly regulating the expression of genes. 7 Existing evidence demonstrates that circRNAs promote the transcription of parental genes, 8 serve as mRNAs to direct protein synthesis, 9 and regulate protein expression. 10 In addition, circRNAs have been linked to many diseases, such as neurological disorders, 11 cancers, 12 and SLE. Wang et al., 13 summarized the roles of circRNAs in diagnosing SLE, evaluating their therapeutic effect and biological functions in the onset and progression of SLE. However, no comprehensive studies have assessed the diagnostic accuracy of circRNAs in patients with SLE. The present systematic review and meta-analysis aimed to assess the diagnostic accuracy of circRNAs in patients with SLE, and summarize the important transcription factors related to the circRNA parental genes.
Materials and methods
Search strategy
Studies published in English that reported the relationship between circRNAs and SLE were searched in the PubMed, Web of Science, Cochrane Library, Clinical Trials, and EMBASE databases up to 3 April 2022. The search was performed by two independent investigators (LY Wang and MT Lu) using “Lupus erythematosus, systemic” and “RNA, circular” as keywords (illustrated in Supplemental File 1). Search results were initially screened based on title and abstract review. The full text manuscripts of potentially suitable studies were then retrieved and reviewed for compliance with the inclusion criteria defined in the meta-analysis protocol. The protocol and registration information for this meta-analysis were registered in the PROSPERO database (available at https://www.crd.york.ac.uk/PROSPERO; registration No. CRD42021229383).
Literature selection
Following removal of duplicates, two reviewers (LY Wang and WY Li) independently screened and assessed the literature at the title and/or abstract level, and disagreements were resolved by discussion or consensus with a third reviewer (RG Fan), based on the inclusion or exclusion criteria, and the quality of reports. The eligibility criteria for inclusion into this meta-analysis were as follows: (1) SLE was confirmed in all patients using the revised ACR classification criteria; 3 (2) the essential sample size, sensitivity, and specificity or summary receiver operating characteristic (ROC) curve was provided; (3) expression of circRNAs could be detected in the plasma, serum, or peripheral blood mononuclear cells (PBMCs); (4) full text could be obtained; (5) full text was published in English; and (6) the study included healthy or disease controls.
Articles were excluded if they met the following criteria: (1) duplicate data; (2) no available data to construct a 2 × 2 table; (3) patients had malignant tumours or autoimmune diseases; (4) article type was a conference abstract, review, letter, animal study, commentary, editorial, unrelated study, or case report without controls; and/or (5) study lacked available data.
Data extraction and quality assessment
If the available data were not directly reported, data were extracted using GetData Graph Digitizer software, version 2.26 (http://getdata-graph-digitizer.com/). The following characteristics were extracted from included studies: first author, publication year, country, diagnostic criteria, detection method, specimens, sample size, expression of circRNAs, sensitivity, and specificity. Two investigators (LY Wang and WY Li) extracted the values of sensitivity, specificity, and essential sample size, which were used to calculate true-positive, false-positive, true-negative, and false-negative rates. The quality of all included studies was independently assessed by two investigators (RG Fan and SJ Wen) using the Quality Assessment of Diagnostic Accuracy Studies-2 (QUADAS-2) tool, 14 and any disagreement was resolved by discussion. The quality of diagnostic accuracy was assessed using Review Manager software (RevMan, version 5.4; https://training.cochrane.org/online-learning/core-software/revman).
Data synthesis and statistical analyses
Data were analysed using Meta-DiSc (version 1.4; https://meta-disc.software.informer.com/1.4) and Stata (version 16; https://www.stata.com/stata16/) statistical software. Cochran’s Q test and
CircRNAs and their parental genes
Transcription factors regulated by circRNAs were predicted using the TRCirc database (http://www.licpathway.net/TRCirc/view/index), and only the −2 kb/1 kb regions of transcription factors were selected. The human transcription factor (hTF) target database (http://bioinfo.life.hust.edu.cn/hTFtarget#!/) was used to predict transcription factors related to the parental genes of circRNAs. After comprehensive analysis of circRNA-related transcription factors, and those related to the corresponding circRNA parental genes, the overlapping transcription factors were integrated in a network diagram using Cytoscape software, version 7.0 (https://cytoscape.org/).
Results
Characteristics of included studies
A total of 155 articles were identified from the searched databases, and 75 records remained after eliminating duplicates. After reviewing titles and abstracts, two animal studies, three commentaries, three conference abstracts, three editorials, 20 reviews, and 12 unrelated studies were excluded. The full text of 32 studies was then reviewed resulting in removal of 12 studies that lacked available data, nine studies that were irrelevant to SLE and one study that did not meet the eligibility criteria (Figure 1). Finally, 10 studies,18–27 including 15 circRNAs, were finally included in the meta-analysis, with a total of 438 patients and 434 controls (Table 1). All included studies were conducted between 2018 and 2022 in China, and comprised Han Chinese participants. All patients with SLE met the ACR diagnostic criteria. Specimens used for CircRNA analyses were plasma in two studies, PBMCs in six studies, and peripheral blood in the remaining two studies. Patients were compared with healthy controls in nine studies.

Flow diagram of study selection process.
Characteristics of 10 studies included in the current meta-analysis.

ACR, American College of Rheumatology; CircRNA, circular RNA; HC, healthy controls; LN, lupus nephritis; no LN, systemic lupus erythematosus patients without LN; RA, rheumatoid arthritis; qRT–PCR, quantitative reverse transcription–polymerase chain reaction; PBMC, peripheral blood mononuclear cell; PB, peripheral blood; ↑, upregulated; ↓, downregulated.
Quality assessment
The results of the quality assessment of the included studies are shown in supplemental File 2. The majority of the articles selected for this meta-analysis did not show high risks of bias in patient selection, index test usage, reference standard choice, and flow and timing.
Diagnostic accuracy of circRNAs
The overall diagnostic accuracy of circRNAs in detecting SLE is summarized in Figure 2. The summary ROC curve did not have a curvilinear shape (Figure 2a), suggesting that no threshold effect related to the heterogeneity existed in the analysis of the diagnostic threshold using Meta-DiSc, version 1.4. The random-effects model was used because of the high heterogeneity found in the selected studies (

Forest plots for the diagnostic accuracy of circular RNAs in detecting systemic lupus erythematosus, showing: (a) summary receiver operating characteristic curves for all datasets; (b) overall diagnostic odds ratio (OR) and 95% confidence intervals (CIs); (c) pooled sensitivity (with 95% CI) and (d) pooled specificity (with 95% CI).
Sensitivity analyses
Sensitivity was analysed using Stata software, version 16, to find the source of heterogeneity. All studies were found to produce little impact on heterogeneity, and the results of the present meta-analysis were shown to be relatively stable (Figure 3). Therefore, subgroup analysis was conducted according to the number of cases and controls, and no statistically significant difference between subgroups was shown (Supplemental File 4). These results indicate that the sources of heterogeneity might be caused by other factors.

Sensitivity analysis of the individual trials, showing results of influence analysis. Numbered lines correspond to the selected circular RNAs in each of the 10 studies.
Publication bias
Deek's funnel plot asymmetry test revealed the possibility of publication bias for pooled evaluation results (
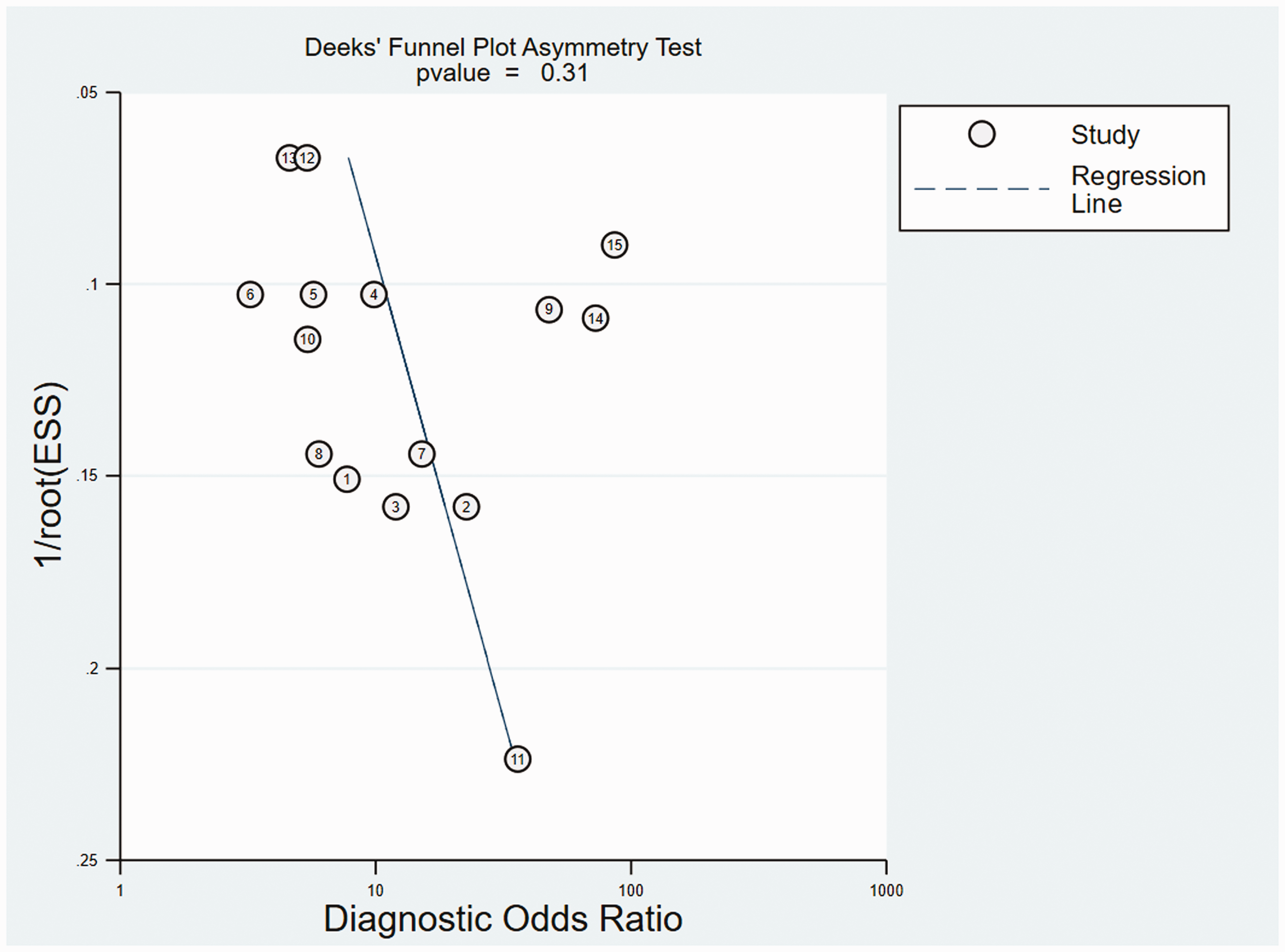
Deek's funnel plot asymmetry test for publication bias, showing possible publication bias in the studies included in the meta-analysis. Numbered circles correspond to the selected circular RNAs in each of the 10 studies.
CircRNAs and their parental genes
Among all included studies, two studies reported hsa_circ_0000479.18,20 The transcription factors regulated by hsa_circ_0000479, and related to the parental gene of hsa_circ_0000479, are summarized in Figure 5. Epithelial stromal interaction 1 (
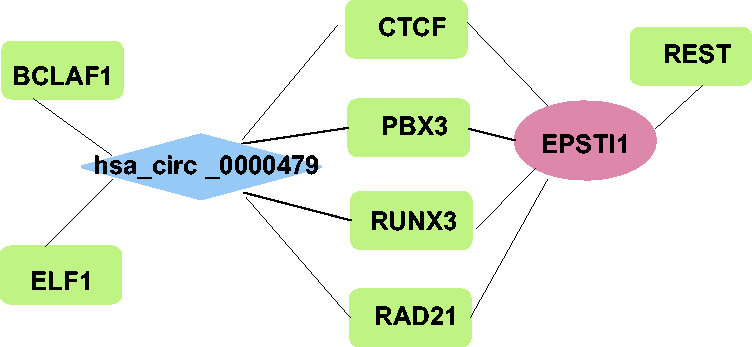
Network diagram showing transcription factors and the parental gene associated with hsa_circ_0000479 circular RNA. BCLAF1, BCL2-associated transcription factor 1; ELF1, E74-like ETS transcription factor 1; CTCF, CCCTC-binding factor; PBX3, PBX homeobox 3; RUNX3, RUNX family transcription factor 3; RAD21, RAD21 cohesin complex component; EPSTI1, epithelial stromal interaction 1.
Discussion
Circular RNA transcripts were first identified in the early 1990s, and have since been characterised regarding their role in immune responses. CircRNAs can induce the activation of cellular immune response pathways, 28 mediate immune regulation via interactions with miRNAs,29,30 and suppress the polarization of M2 macrophages during immune responses. 31 Some studies have reported that circRNAs may serve as potential biomarkers of, and be involved in, inflammatory diseases, such as rheumatoid arthritis, 32 type 2 diabetes mellitus, 33 carcinomas, 34 and SLE. 35 CircRNAs from patients with SLE may contribute to enhanced T-cell apoptosis, 35 regulate the methylation of DNA, 36 and inhibit the effects of miRNAs on genes. 37 CircRNAs and their parental genes have been reported to show aberrant expression in patients with SLE, and to have potent regulatory functions,25,26 and circRNAs are increasingly reported to regulate their parental genes in a variety of ways.8,38 These results suggest that circRNAs participate and play an important role in the immune regulation of SLE. Thus, the present study explored the diagnostic ability of circRNAs in SLE and integrated the transcription factors associated with the parental genes of circRNAs.
The current meta-analysis integrated the results of 10 studies. The pooled results of sensitivity, specificity, LR+, LR–, diagnostic OR, and AUC indicated a moderate accuracy of circRNAs as diagnostic biomarkers for SLE, particularly the summary specificity and AUC values, suggesting that circRNAs may be used as a novel biomarker for SLE diagnosis.
Heterogeneity existed in the data synthesis and analysis, and the results of the sensitivity analysis showed no obvious factors of instability in the present study. Subgroup analysis, conducted according to the number of SLE cases and controls, indicated that the heterogeneity might be caused by other factors. The subgroups divided by specimen groups with insufficient numbers were not suitable for subgroup analysis, which may have resulted in the source of heterogeneity being missed. Patients with lupus nephritis were also included in the present study. The expression of circRNAs in the blood of patients with SLE with renal involvement is higher than that in the blood of patients without renal involvement,22,23 which may have contributed to the presence of heterogeneity and a higher diagnostic accuracy. More studies related to lupus nephritis should be included in future studies. The exact diagnostic value of circRNAs in SLE should be confirmed by future comprehensive studies. There was no statistically significant publication bias in the pooled evaluation results. The results of the literature search did not provide enough data to analyse a particular type of circRNA, therefore, a number of different circRNAs were analysed instead of a single circRNA. The present results might be inaccurate, as some studies were based on a set of circRNAs rather than a single circRNA, however, the present study may provide data to promote the development of future research on certain types of circRNA.
Only two studies focused on hsa_circ_0000479.18,20 The present analysis mainly focused on the relationships of hsa_circ_0000479 with its parent gene,
The results of the present study may be limited by several factors. First, heterogeneity among the studies was reported, which might have affected the interpretation of the results. However, sensitivity analyses and subgroup analysis did not reveal the source of heterogeneity. Possible reasons for heterogeneity may concern the specimen source, disease duration and/or severity, and treatment in patients with SLE. Due to the limited information provided by the included studies, further analysis was not possible. Secondly, a total of 10 articles were included, but only two studies focused on hsa_circ_0000479. Thus, more studies need to be conducted on the same individual circRNA markers, and the technology used for detecting multiple circRNAs needs to be standardized. Thirdly, the number of screened articles was too small to analyse these potential limitations, although there are few studies that have analysed combined rather than single circRNA. Fourthly, all results in the included studies were positive, which may have resulted in publication bias.
In conclusion, to the best of the authors knowledge, the present study is the first to comprehensively evaluate the potential role of circRNAs in diagnosing SLE, and to integrate the transcription factors related to hsa_circ_0000479 and its parental gene. Future research should include a sufficient number of subgroups divided by specimen. To improve the specificity of circRNAs in the diagnosis of SLE, it may be better to combine circRNAs with other criteria, such as antibodies, complements, and clinical manifestations, or find a breakthrough marker among the parental genes of circRNAs.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605221103546 - Supplemental material for Significance of circRNAs as biomarkers for systemic lupus erythematosus: a systematic review and meta-analysis
Supplemental material, sj-pdf-1-imr-10.1177_03000605221103546 for Significance of circRNAs as biomarkers for systemic lupus erythematosus: a systematic review and meta-analysis by Luyuan Wang, Mengting Lu, Wenyu Li, Runge Fan, Sijian Wen, Wen Xiao and Youkun Lin in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_03000605221103546 - Supplemental material for Significance of circRNAs as biomarkers for systemic lupus erythematosus: a systematic review and meta-analysis
Supplemental material, sj-pdf-2-imr-10.1177_03000605221103546 for Significance of circRNAs as biomarkers for systemic lupus erythematosus: a systematic review and meta-analysis by Luyuan Wang, Mengting Lu, Wenyu Li, Runge Fan, Sijian Wen, Wen Xiao and Youkun Lin in Journal of International Medical Research
Supplemental Material
sj-pdf-3-imr-10.1177_03000605221103546 - Supplemental material for Significance of circRNAs as biomarkers for systemic lupus erythematosus: a systematic review and meta-analysis
Supplemental material, sj-pdf-3-imr-10.1177_03000605221103546 for Significance of circRNAs as biomarkers for systemic lupus erythematosus: a systematic review and meta-analysis by Luyuan Wang, Mengting Lu, Wenyu Li, Runge Fan, Sijian Wen, Wen Xiao and Youkun Lin in Journal of International Medical Research
Supplemental Material
sj-pdf-4-imr-10.1177_03000605221103546 - Supplemental material for Significance of circRNAs as biomarkers for systemic lupus erythematosus: a systematic review and meta-analysis
Supplemental material, sj-pdf-4-imr-10.1177_03000605221103546 for Significance of circRNAs as biomarkers for systemic lupus erythematosus: a systematic review and meta-analysis by Luyuan Wang, Mengting Lu, Wenyu Li, Runge Fan, Sijian Wen, Wen Xiao and Youkun Lin in Journal of International Medical Research
Footnotes
Declaration of conflicting interest
The Authors declare that there is no conflict of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the Natural Science Foundation of China (81760561).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
