Abstract
Objectives
Commensal bacteria in the nasal cavity may act as opportunistic pathogens that cause infections under certain conditions. Screening for commensal bacteria in the nasal cavity may aid in understanding their roles in microbiota balance and preventing potential infections.
Methods
Nasal samples were collected from healthy preclinical medical students and used to inoculate various bacterial culture media, by means of the WaspLab microbiology automated system. Bacterial colonies were then identified by matrix-assisted laser desorption ionization-time of flight mass spectrometry. Antibiotic resistance phenotypes of
Results
In total, 549 bacterial strains were isolated from 161 participants. These strains included the following genera:
Conclusions
A diverse group of nasal commensal bacteria inhabited our population of healthy volunteers. These data can improve comprehension of the potential roles of these nasal commensal bacteria in regulating microbiota balance and promoting or mitigating potential future infections.
Keywords
Introduction
Commensal bacteria in the human nasal cavity are important members of the normal flora that contribute to maintenance of normal physiological and immune functions. 1 , 2 These diverse commensal species possess immunoregulatory functions; they also inhibit or eliminate incoming pathogens. However, normal microbial flora may harbor opportunistic pathogens that cause infection in states of immune dysfunction or nasal microbial dysbiosis. Common bacterial infections are frequently associated with nasal colonization by a genetically homologous strain. 3 , 4 Previous studies have suggested that some microbial interactions can affect human nasal colonization by opportunistic pathogens. 1 However, the interactions that mediate pathogenicity involving nasal commensal bacteria remain poorly understood; the increasing resistance of these bacteria to most available antibiotics poses a substantial public health problem. 3
Medical students represent an important subset of healthy individuals in a given population, such that studies of their nasal bacteria might improve recognition of important commensal species that regulate microbiota balance. Furthermore, medical students represent an important group within the context of healthcare staff, who might spread bacteria to other community members or susceptible patients.
12
Medical students could participate in greater awareness regarding hospital-acquired infections. Therefore, analyses of prevalence in medical students are needed to assess their carriage statuses before clinical rotations. This study evaluated the nasal carriage rates of cultivable commensal bacteria in healthy preclinical medical students in China, including
Materials and methods
Study Population
Samples were collected from healthy, third-year, preclinical medical students from the Clinical Medicine Science Department of the Fourth Military Medical University, with approval by the ethics committee of Xijing Hospital, Fourth Military Medical University (approval number: B670201422215). All participants provided written informed consent prior to participation in the study. All experiments were performed in accordance with relevant guidelines and regulations. Participants were excluded if they met any of the following criteria: systemic or local infectious disease within the past 1 month; administration of antibiotic drugs within the past 1 month; a history of travel history in the past 2 months; and/or a history of chronic respiratory disease (e.g., chronic obstructive pulmonary disease or bronchiectasis), immunocompromised state, autoimmune disease, chronic granulomatous disease, diabetes, or tumors.
Specimen collection and transportation
Nasal specimens were collected for bacterial isolation using the flocked eSwab transport system (Copan, Brescia, Italy). One moistened swab was introduced into both anterior nares approximately 2 cm into the nasal passage, gently rubbed across the nasal mucous membranes, and rotated five times (both clockwise and counterclockwise) to acquire squamous epithelial cells. All samples were immediately stored in eSwab Liquid Amies preservation medium and transported to the laboratory for processing, or stored at 4°C until further analysis.
Specimen processing and bacterial isolation
All swabs were loaded into the WASPLab laboratory automation systems (Copan) for plating on 5% sheep blood agar, chocolate agar, and MacConkey agar plates (Autobio, Zhengzhou, China). The inoculated plates were moved by a conveyor belt to a digital imager, where images were obtained at time point 0; they were then moved into the WASPLab incubator and incubated at 35°C with 5% CO2 for 16 hours, in accordance with the laboratory’s standard operating procedures. Upon reaching the end of incubation, a second image of plates was collected and used for manual assessment of bacterial growth. Plates with no bacterial growth after 16 hours of incubation were incubated for an additional 24 hours (40 hours total) and a third image was collected for manual assessment of bacterial growth. All isolates were identified by matrix-assisted laser desorption ionization–time of flight mass spectrometry.
Matrix-assisted laser desorption ionization-time of flight mass spectrometry identification
Plate preparation, mass spectrum generation, and processing were performed in accordance with previously described standards
13
for the Vitek MS system (bioMérieux, Marcy l'Etoile, France). Briefly, a 1-mL loop was used to apply a portion of a single colony to a disposable polypropylene target slide (bioMérieux). One milliliter of a-cyano-4-hydroxycinnamic acid matrix solution (3.1% wt/vol; bioMérieux) was immediately applied to the isolate and allowed to dry at room temperature, prior to mass spectrometric analysis. All isolates were analyzed with the Vitek MS in linear positive-ion mode using the Vitek MS Acquisition Station software version 2.0 (bioMérieux). The similarities of samples with reference isolates in the MS-ID database (within Vitek MS Acquisition Station software) was then calculated and expressed as an optimized quantitative value (confidence value). Newly passaged isolates of
Antimicrobial susceptibility testing for S. aureus
Following identification by matrix-assisted laser desorption ionization-time of flight mass spectrometry,
Results
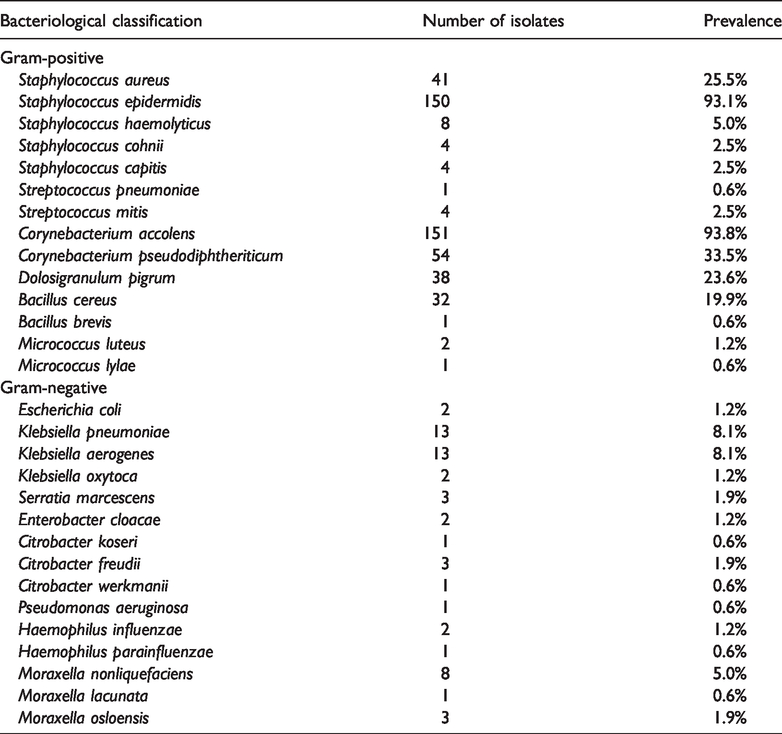
In total, 161 participants were enrolled in this study; the mean participant age was 20.36 ± 1.13 years and most participants were men (140, 87.0%). Table 1 shows the overall bacteriological profile of nasal specimens collected from participants in this study. Bacteriological analysis identified 549 cultivable bacterial strains from nasal samples, including the following genera:
Prevalences of cultivable bacteria in nasal specimens from medical students in China.
Overall, there were 72 co-occurrence patterns of commensal bacteria in nasal specimens cultured from the participants in this study; the number of bacterial species in a single sample varied between two and six. The compositions and prevalences of the top five co-occurrence patterns of cultivable nasal bacteria are listed in Table 2; representative photographs of these patterns of nasal commensal bacteria are shown in Figure 1. The dominant co-occurrence pattern of cultivable nasal bacteria consisted of
Top five major co-occurrence patterns of cultivable bacteria in nasal specimens from medical students in China.


Representative photographs of the top five co-occurrence patterns of commensal bacteria in nasal specimens collected from medical students in China, grown on appropriate culture medium. (a)
The susceptibility patterns of
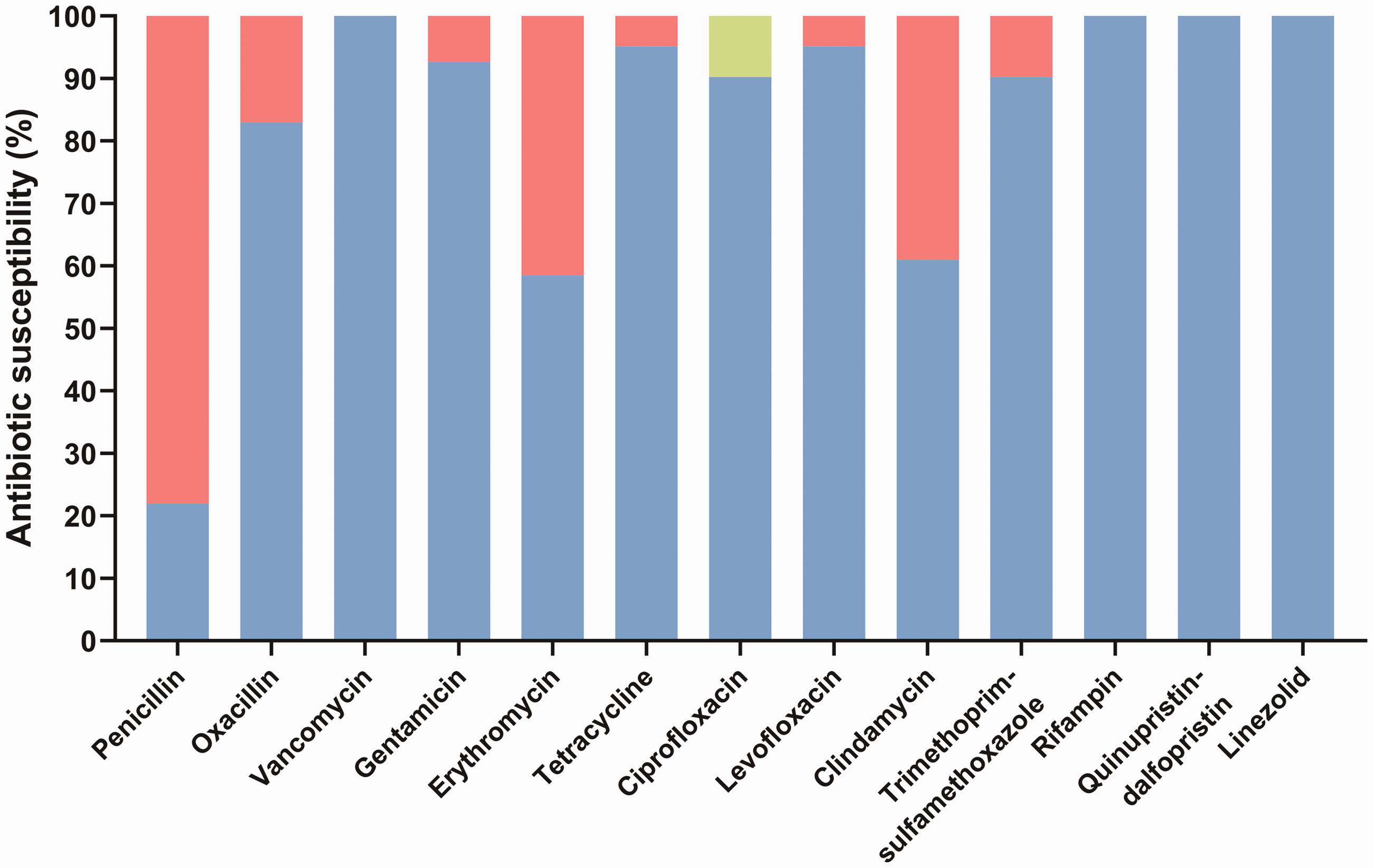
Antimicrobial susceptibilities of
Discussion
Early studies of the human microbiota indicated that commensal bacteria in the healthy respiratory tract comprise species from the following genera (including some potential pathogens):
Most nasal commensal bacteria function as mutualists, such that they alter the local host microenvironment in a manner that can help to protect against colonization by (and proliferation of) pathobionts, such as
Although commensal bacteria have been shown to exhibit beneficial effects, there is evidence that these bacteria can also serve as opportunistic pathogens.
Notably, most catheter-related, device-related, and prosthetic joint infections are caused by CoNS, especially by
An important limitation of this study was that it relied upon a traditional culture method, rather than metagenomic sequencing; thus, the results could not demonstrate overall microbial species diversity, because the nasal cavity harbors a diverse population of uncultivable commensal microbiota. 2 Another important limitation was the small sample size, because eligible individuals were limited to a predefined student population. Further investigations are necessary to gain insight into the mechanisms involved in interspecies interactions that regulate microbiota balance. Moreover, additional data are needed to develop effective microbiota-based approaches for control of bacterial pathogens.
In conclusion, we demonstrated the prevalences and compositions of commensal microbiota in nasal samples from preclinical medical students; our results indicated considerable diversity in the healthy nasal commensal community in this population of healthy individuals. These findings can improve general knowledge concerning the potential roles of these nasal commensal bacteria in regulation of microbiota balance, as well as the risks of future infections that involve these species. The results highlight the need for enhanced awareness among medical students regarding their roles in the transmission of pathogens and prevention of relevant infections.
Footnotes
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by grants from Disciplinary Research Startup Program of Xijing Hospital in Fourth Military Medical University (XJZT19ML23 and XJZT18ML54).
