Abstract
Objective
The ataxia telangiectasia mutated (
Methods
We retrieved and identified the available case–control studies that met the inclusion criteria from the PubMed, Web of Science, and Embase databases. Odds ratio (OR) and 95% confidence intervals (CIs) were used to measure the association between rs1801516 and cancer risk. Additionally, we performed sensitivity, subgroup, and publication bias analyses.
Results
After inclusion criteria were met, the meta-analysis included 29 studies, with 9,453 cancer patients (cases) and 14,646 controls. No association was found between rs1801516 and cancer risk (pooled OR = 0.911; 95% CI, 0.740–1.123). Concordantly, no association was found between rs1801516 and cancer risk after subgroup analysis by source of controls, cancer type, or ethnicity, which confirmed the finding of the dominant model that this SNP is not involved in the occurrence of cancer.
Conclusions
Through this meta-analysis, we found no association between rs1801516 and cancer occurrence as a risk factor. These data provide useful information for future case–control studies on cancer etiology.
Introduction
Cancer occurrence is increasing because of an aging population, the prevalence of smoking, physical inactivity, and other lifestyle factors. 1 Cancer is a cellular abnormality initiated by uncontrolled growth caused by an accumulation of damage or mutations in the genetic material due to hereditary and environmental factors, which causes cells to evade the many signals that control cell growth and death. 2 Genetic factors have a greater effect on cancer initiation than do lifestyle or environmental factors. 3 Many candidate genes and variations have been identified that contribute to cancer susceptibility.
The ataxia telangiectasia mutated (
Variations in the
When there is considerable variation in the results of studies on topics that have been studied extensively, meta-analysis can be used to identify a common effect.
16
Such an analysis was conducted by Gao et al. (2010)
17
to assess whether the
Materials and Methods
Reporting for this meta-analysis followed the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines. All data in this study were from published studies and did not involve patients directly. Therefore, ethics committee approval and informed consent were not required.
Identification of relevant studies
We performed a literature search of three online literature databases (PubMed, Web of Science, and Embase) to screen and identify available studies to be included in the meta-analysis. The keywords used were “
The inclusion criteria were as follows: (1) the studies were designed as case–control; (2) the cases in the identified studies were cancer patients; and (3) the studies reported the frequencies of
Data analysis
Two investigators (Yueting Li and Pengxu Shi) independently extracted the data from each eligible publication, including the last name of the first author, the year of publication, the geographic region, the genotyping method, the sample size, and the genotyping results for both cases and controls. Data pertaining to patient ethnicity, control source, and cancer type were extracted with a view to determining the contributions of underlying characteristics to the study findings.
Statistical analysis
The Hardy–Weinberg equilibrium (HWE) of control genotypes was calculated using a χ2 test. The strength of the association of rs1801516 and cancer was evaluated using odds ratios (ORs) and 95% confidence intervals (CIs). To calculate the pooled estimates of the ORs and 95% CIs among the studies, a random effects model was used to resolve inter-study heterogeneity. 20 A random effects model evaluates the likely effect size across different cancers and takes heterogeneity across studies into account. It is different from a fixed effects model, which evaluates the most likely effect size from multiple studies by hypothesizing that they are sampled with the same cancer or from a single population, but the model can be biased by high heterogeneity across studies.
Three genetic models (allele, dominant, and recessive) were used to measure the overall pooled ORs. As described in the previous study, we compared OR1 (AA vs. aa), OR2 (Aa vs. aa), and OR3 (AA vs. Aa), where A is the risk allele. 16 We selected the most appropriate genetic model using a previous study.21,22
We evaluated the degree of inter-study heterogeneity using a Q statistic,23,24 where
By means of sensitivity analysis, we evaluated whether a single study could influence the pooled effect size. Specifically, we omitted each study from the meta-analysis in turn and subsequently evaluated whether any significant alterations were made to the pooled effect size.
Publication bias was investigated by using funnel plots generated for each study, in which the standard error of log(OR) was plotted against the log(OR). Possible publication bias was determined when the plot was asymmetric, in which case an Egger test was used to determine degree of asymmetry, with
All statistical calculations were performed using Stata version 10.0 (Stata Corp., College Station, TX, USA).
In silico analysis
To predict the potential association between rs1801516 and the expression of
Results
We searched online literature databases and identified relevant articles for inclusion. The literature screening flowchart is shown in Figure 1. According to the established inclusion criteria, 29 publications were ultimately screened and included in our meta-analysis.13,14,26–52 These 29 case–control studies collectively contained 9,453 patients with cancer (cases) and 14,646 unaffected participants (controls). Individuals with different ethnic backgrounds were included (e.g., Caucasian, Hispanic, African and Asian, which included Arab, East Asian, and Sinhalese). Table 1 summarizes the main characteristics of the included studies, and Table 2 summarizes the genotype and allele frequencies of rs1801516 SNP and HWE in controls. Of the 29 studies, four deviated significantly from HWE.13,42,46,47

Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) search flow diagram.
Baseline characteristics of qualified studies in this meta-analysis.

Distribution of genotype and allele frequencies of the

Heterogeneity detection and pooled analysis
The association between the rs1801516 polymorphism and cancer risk was evaluated using pooled ORs (with 95% CIs) under the dominant, recessive, homozygous codominant, heterozygous codominant, and allele genetic models (Figure 2, Table 3). We selected the dominant model to perform the pooled analysis according to our model selection strategy.21,53 The pooled results of the dominant model indicated that there was no association between the rs1801516 polymorphism and cancer risk (OR, 0.911; 95% CI, 0.740–1.123). After removing the four studies that deviated significantly from HWE, there was no obvious change in the pooled result (OR, 0.914; 95% CI, 0.725–1.152). Subgroup analysis was then performed, which also failed to detect any association between rs1801516 and cancer risk in any ethnicity (Table 4). Moreover, no association was observed in the subgroup analysis between rs1801516 and cancer by source of controls (hospital-based and population-based). Additionally, we performed a subgroup analysis by type of cancer, again finding no association between rs1801516 and the occurrence of breast cancer, prostate cancer, rectal cancer, leukemia, bladder cancer, colorectal cancer, renal cancer, head and neck cancer, cervical cancer, melanoma, thyroid cancer, ovarian cancer, or lung cancer (Table 4).

Forest plot of the association between the rs1801516 polymorphism of the
Summarized ORs with 95% CIs for the association of

Stratified analysis of the association of

Sensitivity analysis
We next sought to determine the contribution of individual studies to the pooled results via sensitivity analysis. To do this, we removed each study from the analysis in turn and determined pooled ORs. We detected no significant changes between each of these analyses and the overall results of the meta-analysis, indicating that none of the included studies significantly altered the overall results. Therefore, our meta-analysis results are stable and reliable.
Publication bias
Publication bias was assessed by generating and analyzing a funnel plot (Figure 3); no significant effects of publication bias were detected by Egger’s test (Table 3).
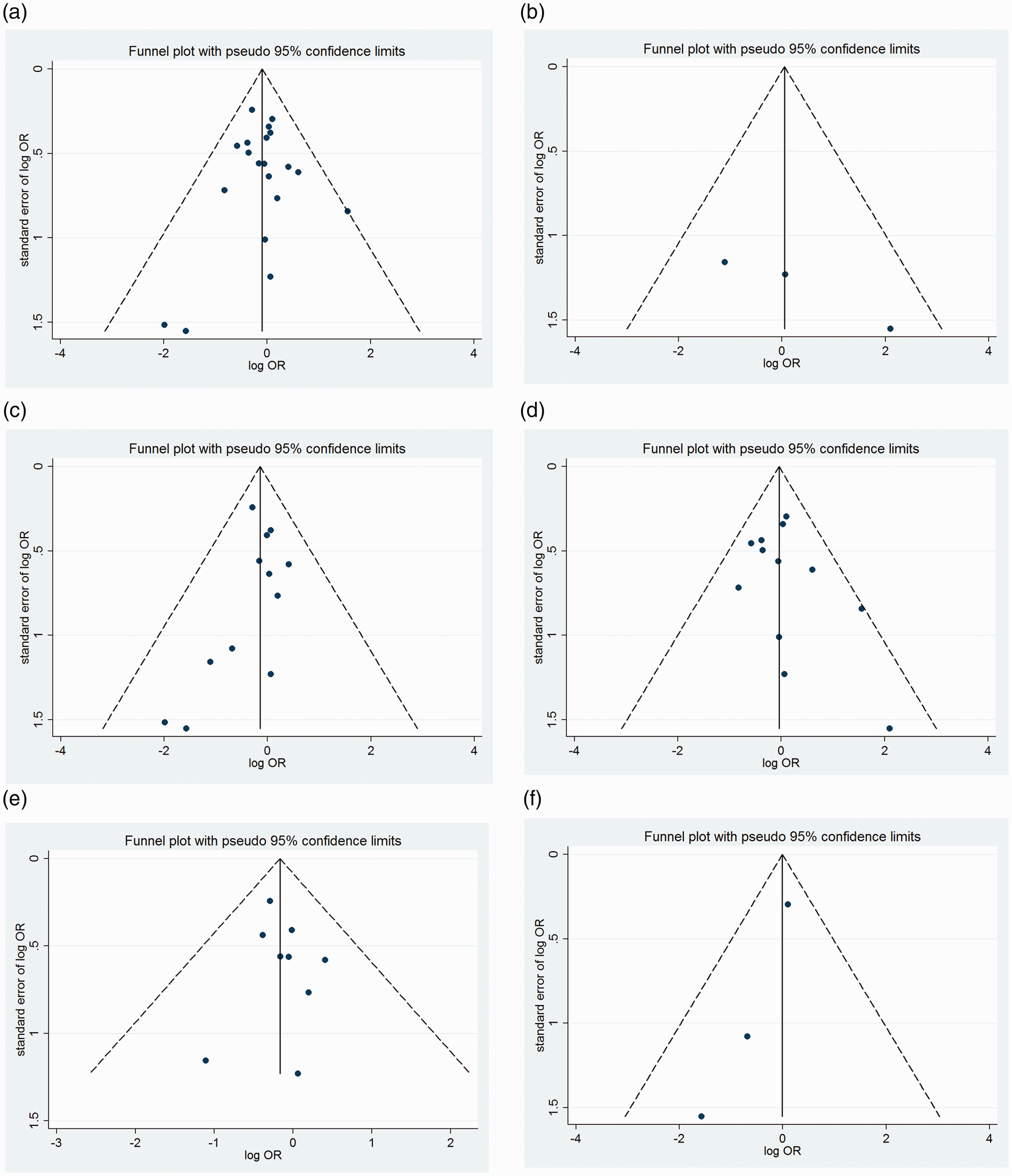
Funnel plot analysis depicting publication bias in the association between the rs1801516 polymorphism of the
In silico analysis
Using eQTL analysis, we found that, compared with the G allele, the A allele of the rs1801516 locus did not alter expression of

In silico analysis of ATM expression related to rs1801516 polymorphism. ATM, ataxia telangiectasia mutated. The box shows the average value and the line indicates the standard error.
Discussion
We explored the underlying relationship between the rs1801516 SNP of the
Previously, a putative association between rs1801516 and the occurrence of cancer was analyzed in two meta-analyses.17,18 These studies found that this SNP in
Our ability to conclusively define stable effects by subgroup, however, was limited by the relatively small sample sizes included in the subgroup analysis. Only one study reported an association between rs1801516 and cancer risk in Arab, East Asian, and Sinhalese ethnicities. Moreover, except for breast and thyroid cancers, the association of any other cancer type was reported in only one study. Thus, because of the limited sample size, the results of the association between rs1801516 and cancer risk in these subgroups are not comprehensive.
A prior meta-analysis of individual patient data showed that
There are some potential limitations to our current analysis. First, we did not analyze gene–gene interactions or epigenetics, which are factors that can influence cancer. Smoking, physical inactivity, and emotional state are also involved in the occurrence of cancer. Second, just one SNP in the
Conclusion
In this study, we showed that there was no association between the rs1801516 SNP in the
Footnotes
Authors’ contributions
Daqing Jiang conceived and designed the study. Yueting Li and Pengxu Shi were responsible for collecting the data, performing statistical analyses, and preparing the manuscript. All authors contributed to drafting the manuscript, and all read and approved the final version.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
