Abstract
Objective
To explore the association between Niemann–Pick C1-like 1 gene (
Methods
Using propensity score matching, 490 T2DM patients and 490 matched controls were recruited from 13 communities in Guangxi, China.
Results
The rs2073547 genotype distribution differed significantly among patient groups. Low-density lipoprotein cholesterol levels were similar among different rs2073547 genotypes and alleles. The rs2073547 AG genotype was significantly more prevalent in patients with T2DM. After adjusting for risk or protective factors for diabetes, AG and GG+AG genotypes of rs2073547 were associated with significantly increased risks of T2DM. Compared with the AA genotype, the AG genotype was associated with a significantly higher risk of T2DM in participants with gamma-glutamyl transpeptidase (GGT) <45 U/L, systolic blood pressure (SBP) ≥140 mmHg, or triglyceride <1.70 mmol/L. In participants with GGT <45 U/L or SBP ≥140 mmHg, the GG+AG genotype was associated with a significantly higher T2DM risk versus the AA genotype.
Conclusions
The rs2073547 polymorphism of
Keywords
Introduction
Globally, the number of patients with diabetes mellitus (DM) reached 451 million in 2017 and this is expected to increase to 693 million by 2045.
1
The prevalence of DM has increased rapidly in China during the past few decades.
2
Dyslipidemia is a common metabolic disorder that is also increasing in prevalence worldwide, and was recently estimated to be 42.84% in middle-aged and older Chinese individuals.
3
Adverse consequences of dyslipidemia that seriously threaten the health of patients include atherosclerosis,
4
coronary heart disease, and stroke.
5
Therefore, lipid-lowering drugs, especially those reducing low-density lipoprotein cholesterol (LDL-C), are widely prescribed. The most common lipid-lowering drugs are statins, which reduce LDL-C concentrations by inhibiting the 3-hydroxy-3-methylglutaryl-coenzyme A reductase gene (
Advances in technology have enabled genome-wide association studies (GWAS) to identify more than 100 risk variants associated with T2DM.8,9 Studies of mutations in genes encoding drug targets have been useful in the prediction of both drug efficacy and adverse effects.10,11 For instance, common single nucleotide polymorphisms (SNPs) in
The Niemann–Pick C1-like 1 gene (
Materials and Methods
Study participants
A total of 786 patients with T2DM and 1015 controls without T2DM were recruited consecutively between January 2011 and September 2012 from 13 communities in Nanning, Guangxi, southern China. All participants met the following requirements: (a) age ≥40 years; (b) resident in Nanning for ≥5 years; and (c) not receiving ezetimibe treatment. T2DM was diagnosed according to the World Health Organization diagnostic criteria published in 1999; individuals with type 1 DM, gestational DM, and other types of DM were excluded from the study. Propensity score matching (PSM) based on age, gender, ethnicity (Han and minorities including Zhuang, Miao, Yao, Molao, Buyi, Dai, Dong, Gaoshan, Hui, Zang, Maonan, and Tujia), smoking status, drinking status, and hours of exercise per week were used to control for these potential confounders. 18 For the final analysis, 490 T2DM patients and 490 matched controls were selected. The ethics committee of the First Affiliated Hospital of Guangxi Medical University approved the study. All participants provided written informed consent before the collection of any data or samples.
Data collection
All participants completed an epidemiological questionnaire that included sociodemographic characteristics, personal history, family history, and other lifestyle habits. Trained personnel obtained anthropometric data such as height, weight, systolic blood pressure (SBP), diastolic blood pressure (DBP), heart rate, and waist circumference (WC), as well as peripheral blood samples from the participants. Serum levels of triglycerides (TG), total cholesterol (TC), high-density lipoprotein cholesterol, LDL-C, aspartate aminotransferase, alanine aminotransferase, and gamma-glutamyl transpeptidase (GGT) were measured at the First Affiliated Hospital of Guangxi Medical University. Lipid profiles were measured using the Architect C16000 autoanalyser (Abbott Diagnostics, Des Plaines, IL, USA) and blood glucose was measured using the glucose oxidase method. 19 Some continuous data were changed into binary data according to clinical significance and reference range.
DNA isolation and genotyping
Genomic DNA from all participants was manually isolated from peripheral blood using a DNA extraction kit (Tiangen Biotech, Beijing, China) according to the manufacturer’s instructions.
Post-hoc calculation of sample size
The sample size for the study was calculated according to the web-based program https://ihg.gsf.de/cgi-bin/hw/power2.pl to determine whether the study would be adequately powered, based on methods described previously.20,21 Assuming a disease prevalence of 0.1, a high-risk allele frequency of 0.05, and an alpha (type 1 error) of 0.05, the total sample size required for a power of 0.98 was calculated to be 830 for a multiplicative model, 888 for an additive model, and 955 for a dominant model. Therefore, the total sample size of 980 matched participants (490 per group) was sufficient.
Statistical analysis
Comparisons of variables between patients with T2DM and controls were carried out using the Student’s t-test or paired sample t-test for continuous variables and the chi-squared test or paired chi-squared test for categorical variables. Conditional logistic regression analysis was used to identify factors associated with T2DM. To analyze genotype distributions, the Hardy–Weinberg equilibrium (HWE) for each SNP was tested using the paired chi-squared test with one degree of freedom. One-way analysis of variance and the Student’s t-test were used to investigate associations between genotypes, SNP alleles, and LDL-C levels. Associations between SNP genotypes and T2DM were analyzed using conditional logistic regression under different genetic models (additive, dominant, and recessive) to adjust for potential confounders. Stratified analyses according to the important factors were performed using unconditional logistic regression. Odds ratios (ORs) and 95% confidence intervals (95% CIs) were used to evaluate the association strength between T2DM and controls. PSM and all analyses were conducted using SPSS 23.0 software (IBM Corp., Armonk, NY, USA). A two-tailed
Results
Baseline information
A total of 786 patients with T2DM and 1015 controls without T2DM were initially enrolled. Of these, PSM selected 490 patients with T2DM and 490 matched controls. Baseline characteristics of study participants are presented in Table 1. After PSM, there were no significant differences in age, gender, ethnicity, smoking status, drinking status, or the number of exercise hours between the two groups (Table 1). However, patients with T2DM had significantly higher SBP (≥140 mmHg), DBP (≥90 mmHg), WC (men ≥90 cm, women ≥85 cm), TG (≥1.7 mmol/L), GGT (≥45 U/L), and a higher occurrence of DM family history than controls (
Comparisons of general characteristics between patients with type 2 diabetes mellitus and controls.
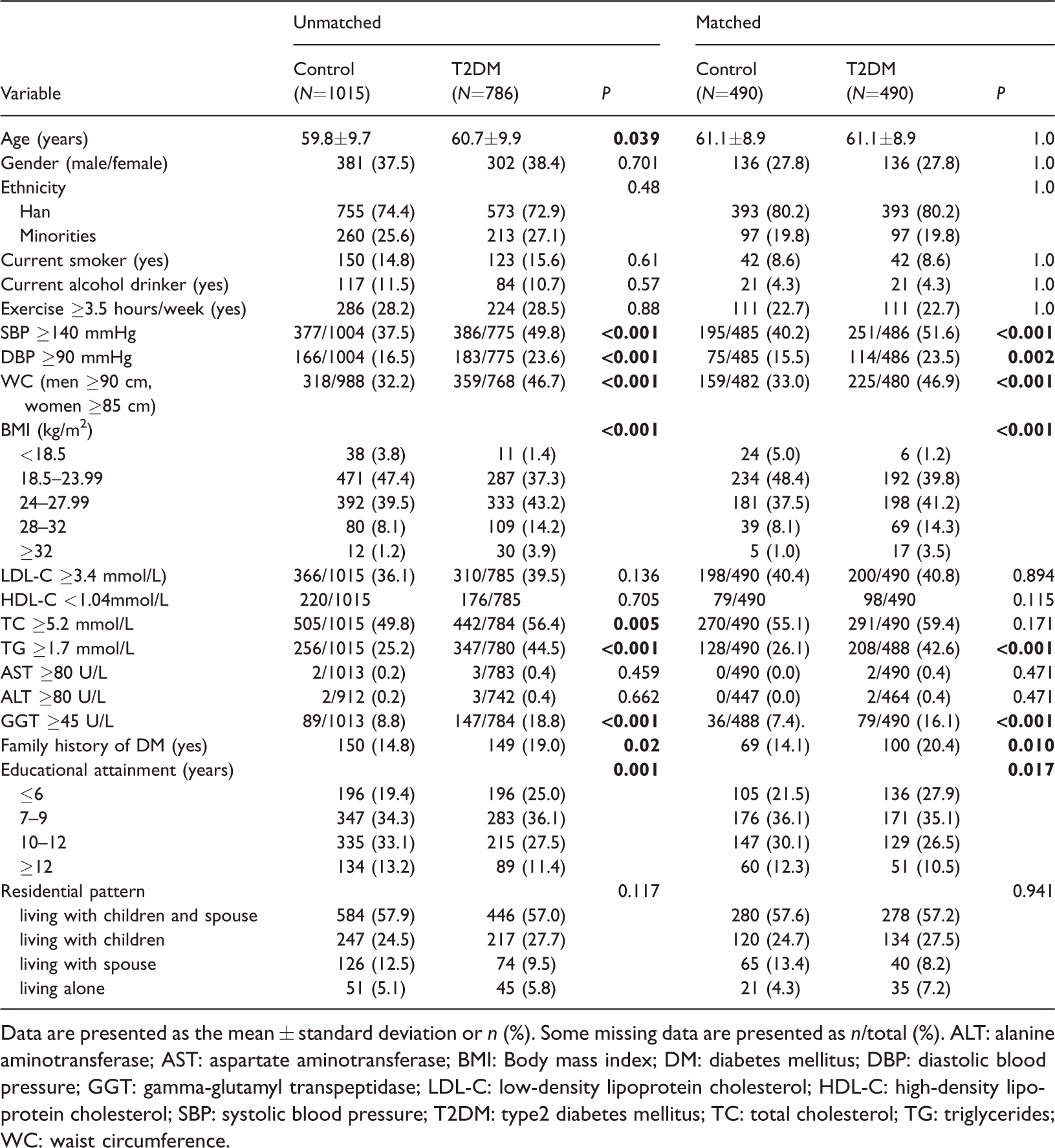
Data are presented as the mean ± standard deviation or
Factors associated with T2DM
Factors associated with T2DM in univariate analysis were identified by conditional logistic regression analysis. This revealed that GGT ≥45 U/L (OR: 1.927, 95% CI: 1.188–3.126), SBP ≥140 mmHg (OR: 1.539, 95% CI: 1.120–2.115), TG ≥1.7 mmol/L (OR: 1.738, 95% CI: 1.268–2.381), and a family history of DM (OR: 1.927, 95% CI: 1.188–3.126) were independently associated with the presence of T2DM (each
Conditional logistic regression analysis of the clinical factors associated with type 2 diabetes mellitus.

95%CI: 95% confidence interval; OR: odds ratio; BMI: Body mass index; DM: diabetes mellitus; DBP: diastolic blood pressure; GGT: gamma-glutamyl transpeptidase; SBP: systolic blood pressure; TG: triglycerides; WC: waist circumference.
Comparison of genotype distributions and allelic frequencies between T2DM and matched control groups
Genotype distributions and allelic frequencies of rs2073547 and rs217386 SNPs in
Genotype distributions, allelic frequencies, and Hardy–Weinberg equilibrium analysis of the two single nucleotide polymorphisms.

Associations between genotypes, alleles of rs2073547, and LDL-C levels
The association between rs2073547 genotypes, alleles, and serum LDL-C levels was investigated (Table 4). No significant differences in LDL-C levels were detected among different genotypes or alleles of rs2073547 in the two groups or total population.
Association between genotypes, alleles of rs2073547, and LDL-C levels.

LDL-C: low-density lipoprotein cholesterol; T2DM: type2 diabetes mellitus.
Associations between genotypes, alleles of rs2073547, and T2DM
We evaluated the association between
Associations between rs2073547 and type 2 diabetes mellitus.
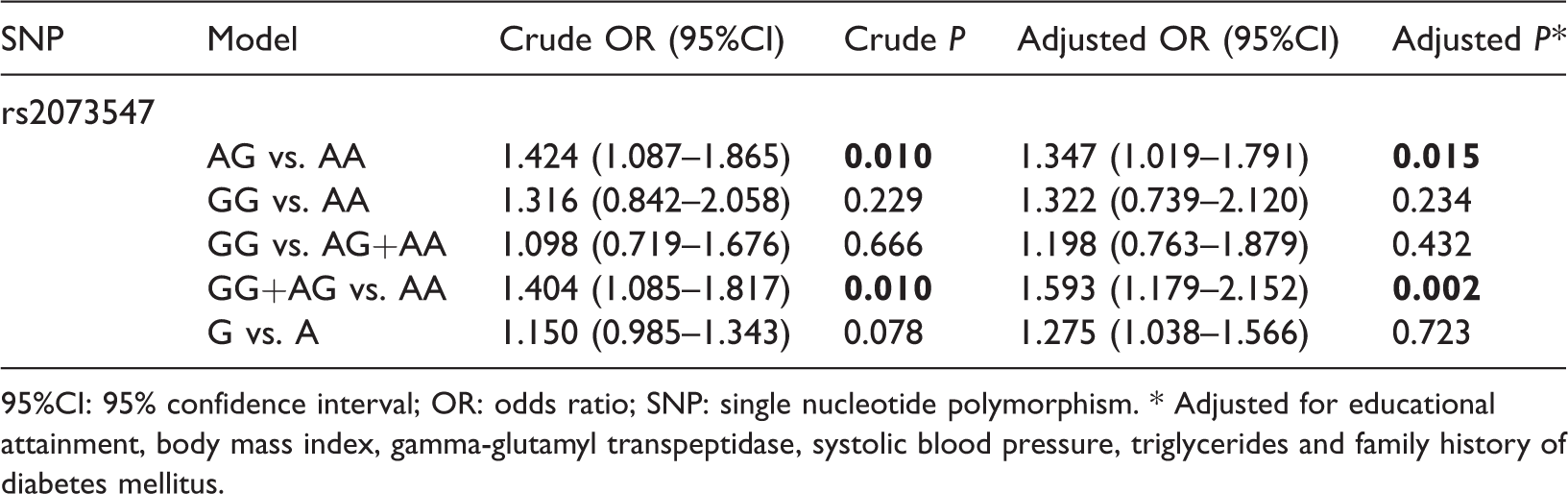
95%CI: 95% confidence interval; OR: odds ratio; SNP: single nucleotide polymorphism. * Adjusted for educational attainment, body mass index, gamma-glutamyl transpeptidase, systolic blood pressure, triglycerides and family history of diabetes mellitus.
Stratified analysis of the association between rs2073547 genotypes and T2DM
We also carried out an analysis of the association of rs2073547 genotypes and T2DM stratified by BMI, GGT, SBP, and TG using different inheritance models. As shown in Table 6, the AG genotype was associated with a significantly greater risk of T2DM than the AA genotype (GGT <45 U/L group: OR: 1.408, 95% CI: 1.060–1.871,
Stratified analysis of rs2073547 by BMI, GGT, SBT, and TG.

95% CI: 95% confidence interval; OR: odds ratio; BMI: body mass index; GGT: glutamyl transpeptidase; SBP: systolic blood pressure; TG: triglycerides.
Discussion
T2DM is a global health problem, and its complex pathogenesis is not yet fully understood. However, GWAS have identified several genetic variants that help explain some of the individual variations in T2DM susceptibility.8,22 Because multiple genetic and environmental factors affect T2DM incidence, a combination of PSM and multivariate logistic regression analysis were adopted in this study to minimize the confounding effects of clinical factors known to be associated with T2DM. Previous studies suggested that elevated TG,
23
GGT,
24
BMI,
25
and a family history of DM
26
are associated with T2DM risk. Although less well studied, elevated SBP may also be a risk factor for T2DM.
27
Borrell et al. reported that educational attainment was inversely associated with the prevalence of DM among certain racial/ethnic groups.
28
In the present study, similar results were obtained in a community-based population. To more clearly elucidate the relationship between
NPC1L1 is associated with cholesterol metabolism, and
In the present study, we investigated the association between rs2073547 and rs217386 genotypes, rs2073547 alleles, and serum LDL-C levels. LDL-C levels were found not to be associated with rs2073547, indicating that the rs2073547 polymorphism of
To our knowledge, our study is the first to investigate the association between
Nevertheless, our findings may provide a new insight into ezetimibe-based monotherapy or combination therapy in Chinese patients with dyslipidemia. Although the reduction of cardiovascular events by LDL-C-lowering drugs is believed to be beneficial despite increased risks of new-onset DM, our results suggest more focus on personalized and precision therapies is warranted to avoid some of the adverse effects of these drugs.
There are a number of limitations associated with our study. It had a cross-sectional design and was a single-center study, so the generalizability of the data to the entire Chinese population remains unknown. Moreover, only two
In conclusion, our study suggests a possible role for the
Footnotes
Acknowledgments
We thank all T2DM patients and control individuals who participated in this study.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Ethical approval and informed consent
The ethics committee of the First Affiliated Hospital of Guangxi Medical University approved the study. All participants provided written informed consent before the collection of any data or samples.
Funding
This work was supported by grants 2016YFC0901200 and 2016YFC0901205 from the Ministry of Science and Technology.
