Abstract
Microbiota and its contribution to brain function and diseases has become a hot topic in neuroscience. We discuss the emerging role of commensal bacteria in the course of stroke. Further, we review potential pitfalls in microbiota research and their impact on how we interpret the available evidence, emerging results, and on how we design future studies.
All disease begins in the gut —attributed to Hippocrates. (ca. 400 BC)
Microbiota and the brain
Humans and bacteria coevolved throughout millions of years and have established, alongside host-pathogen relation, a tight commensal alliance. Human host niches are colonized by several bacterial populations – microbiota, with a huge collection of genes – themicrobiome. Each body site has its specific microbiota composition and interindividual differences between the same body habitat are smaller than intraindividual variance, between different body sites. 4 The largest and most complex bacterial community resides within the gastrointestinal system, particularly the large intestine, and comprises trillions of bacteria. 5 Gut microbiota is protective for the organism through competition with pathogens. By participating in formation of the intestinal barrier and development of the immune system, it implements structural functions. Finally, it is crucially involved in metabolism, fermenting indigested food, producing vitamins, and deactivating xenobiotics. 6
Recent evidence from different fields of medicine suggests that microbiota plays an important role in many diseases, including those of the nervous system. 7 The microbial population of the gut has been found to engage in an intense signaling with the central nervous system (CNS) via neural, immunological, and direct humoral signaling pathways and hence has been incorporated into the concept of bidirectional brain–gut signaling. 8 Indeed, CNS alters the intestinal microenvironment by regulating gut motility and secretion, as well as mucosal immune responses via the enteric nervous system and the neuronal-glial-epithelial unit.9–11 Bacteria react to catecholamines or host hormones and produce neurotransmitters and neuromodulators or their precursors.12–14 Moreover, gut microorganisms influence the enteric nervous system and might stimulate afferent signaling from the gut to the brain via the vagus nerve. 15 In a recent study, gut microbiota was identified as a regulator of host serotonin production. 16 If not tightly controlled by the host, microbiota and bacterial components (e.g. LPS) threaten to harm or even kill the host. Not surprisingly then, there is a very close interaction of gut microbiota and the host immune system, with important consequences for systemic immunity.17–19 Consequently, mice raised and kept germ free (GF) appear not to develop a normal immune system, 20 which needs to be kept in mind when working with such mice in animal models of disease (see below).
Studies in GF mice have shown that gut microbiota is important for the development and proper function of the host CNS. This in turn has important consequences for modeling brain diseases in GF mice (see below). For example, GF animals have a leaky blood brain barrier (BBB), 21 as well as an impaired microglia structure and function. The latter findings were replicated in animals after depletion of microbiota with antibiotics. 22 Further, gut microbiota possibly influences brain chemistry and behavior, culminating in the claim that psychological phenotypes (e.g. anxiety) could be transferred between individuals via fecal transplantion.23–28
Animal and human studies have suggested involvement of microbiota in the pathogenesis of autism,29,30 experimental autoimmune encephalomyelitis,31,32 neuromyelitis optica, 33 Guillain-Barré syndrome, 34 and Parkinson. 35 Neuropsychiatric diseases related with alterations in the brain–gut axis include depression,34,36 anxiety,36,37 and pain disorders.38,39 In the context of stroke, it is also important to mention that gut microbiota is linked to metabolic profiles of symptomatic atherosclerosis.40–44
Microbiota and acute stroke
In Figure 1, we summarize a speculative framework for brain–gut communication after acute brain lesion, such as a stroke. We synthesize in it recent insights into the interplay between a brain lesion and the immune and autonomic nervous system (ANS), as well as accumulating knowledge about the interaction of the gut associated lymphatic system (GALT) with the gut microbiota. Briefly, via the ANS and/or the hypothalamus-pituitary-adrenal glands axis (HPA) a brain lesion can affect intestinal function, including immunity. In experimental stroke, T and B cell numbers are significantly reduced in Peyer's patches already 24 h after stroke onset.
45
Increased intestinal permeability may promote bacterial translocation to extraintestinal organs, blood stream, or lymphatic fluid, engaging local and systemic immunity and potentially leading to organ infection (e.g. pneumonia) or even sepsis.
46
General concept of brain–gut microbiota interactions after central nervous system lesion.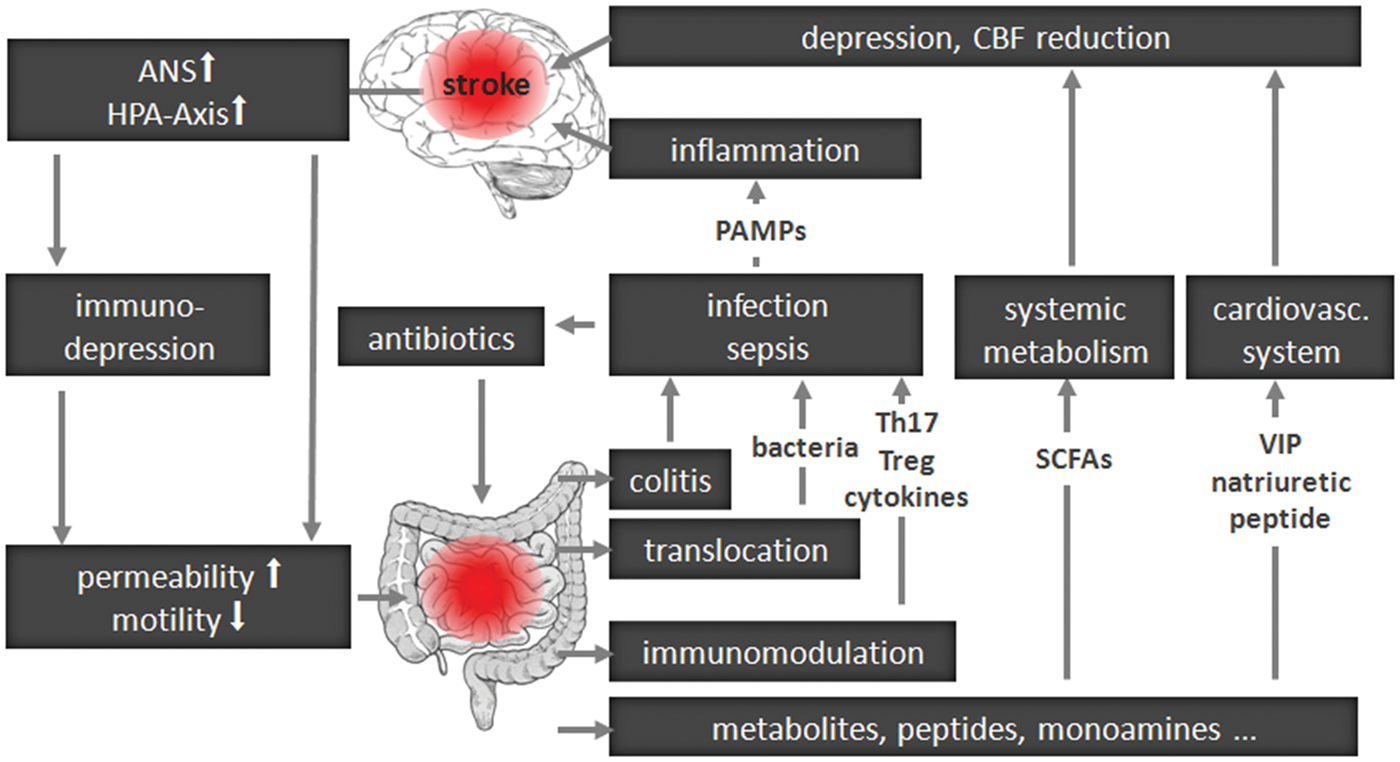
Brain function can be modulated by metabolic signaling via short chain fatty acids (SCFAs), products of bacterial fermentation in the intestine, or intestinal hormones such as ghrelin or leptin reaching the CNS. Further, an active interplay of commensal bacteria and GALT can modulate systemic immunity, impacting the outcome after brain lesion. Injury to the BBB following stroke allows brain-infiltration of immune cells, reactive to CNS antigens. Concurrent with changes in BBB integrity, microbiota may bias the immune system by inducing, instructing, and shifting immune cell populations (such as regulatory T-cells or IL-17 producing Th17 cells).20,47,48 As post-stroke infections are common complications, 49 stroke patients are often treated with a potent combination of antibiotics, 50 which might lead to a dramatic alteration in the composition of gut microbiota, 51 adding a further level of complexity.
Following in the footsteps of researchers in other fields, many groups worldwide have now started to study the impact of gut microbiota on stroke outcome in experimental models, and to characterize microbiota composition changes after stroke in humans. Surprisingly, the existing knowledge, based on brain–gut interaction after stroke, is rather limited. Tascillar et al. reported bacterial translocation to extraintestinal structures in a mouse middle cerebral artery occlusion model (MCAo). 52 Caso et al. found that in rats, stress before MCAo fostered bacterial translocation to diverse organs. 53 Swidsinski et al. observed transient colonic inflammation and abrupt disappearance of several bacterial groups after stroke in humans and animals after acute brain injury. 54
In summary, it is plausible that gut microbiota plays a role in shaping the outcome after acute cerebral ischemia. However, as we anticipate a certain “hype” regarding the role of gut-brain interactions after stroke, we would like to draw attention to some of the issues that have led to an overselling of the microbiome or microbiomania. 55
Potential pitfalls in microbiota studies
Key issues in investigating the role of gut microbiome for stroke outcome.
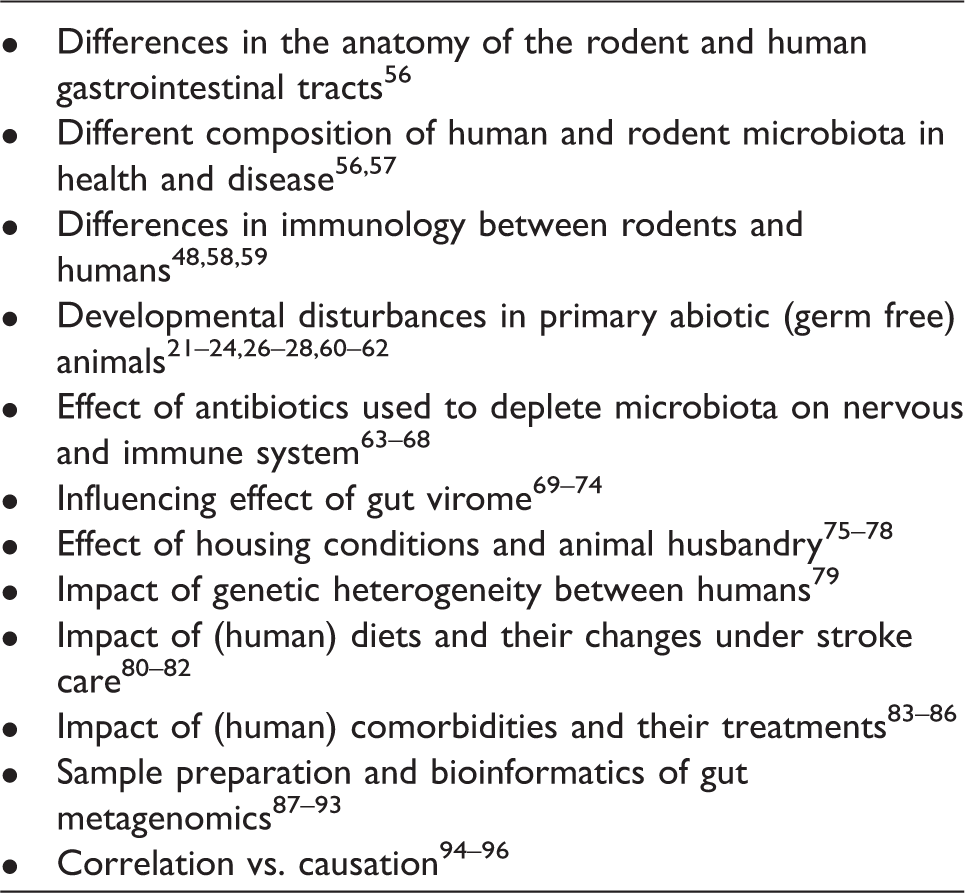

Mouse vs. man
Despite similarities, there are several important differences between rodents and humans in gastrointestinal anatomy and function, many of them driven by differences in diet. For example, the caecum of mice plays a very important role in fermentation of digested food, whereas in humans, it has no known function. Goblet and Paneth cells are distributed differently in rodent and man, and rodents do not have haustra in the colon. 56 These differences are not substantial, but may have important implications for the region of the gut from which feces are sampled.
Since rodents are coprophagic and in the laboratory live on a specialized diet, it is not surprising that microbiota composition differs substantially from that of humans. Coprophagy and cohousing may lead to a homogenization of microbiota between individual animals. 56 Housing conditions, 75 familial transmission, 76 caging, 77 vendor, as well as genotype77,78 strongly influence the composition of intestinal microbiota in rodents. Although both in mice and man the most abundant phyla are Firmicutes and Bacteroidetes, at the genus level more than 80% of the bacterial sequences found in mice are not found in humans. 57
The intestinal immune system maintains the balance between combating pathogens and tolerance for commensal microbiota and food antigens. Dendritic cells (DC) are centrally involved in the generation of gut homing and differentiation of T cells, and thereby in determination of immunogenic or tolerogenic T-cell responses.58,59 Available data suggest similar properties for murine and human intestinal immunity. However, information from humans is scarce, and marked differences between the human and the mouse intestinal immune system have already been described. For example, γδ T cells are found significantly less frequently in the intraepithelial compartment of humans than in that of mice.48,58,59 Moreover, mouse, but not human B cells express TLR4. Therefore, mice, but not humans demonstrate polyclonal B cell responses for antibacterial defense, independently of T-cells. 48 This might have relevance for the secretion of immunoglobulin A (IgA), which is known to protect the gut epithelium from luminal antigens and contributes to host-microbe symbiosis. 97
Sampling, DNA extraction, and metagenomics
The spatial organization of microbiota along the gut and across the gut lumen complicates microbiome analysis. All this makes sampling location and technique (e.g. aerobic vs. anaerobic; feces vs. biopsy) critical contributors to the results. In addition to the variance introduced by sampling, discrepancies between studies resulting from different DNA extraction methods have to be considered.87,88 The standardization in microbiome research, as proposed for animals 89 and for humans, 90 is urgently needed.
Largely advances in metagenomics drove the recent surge in interest and knowledge on the role of commensal microorganisms in health and disease. Bacterial 16S rRNA sequencing or, more recently, shotgun sequencing techniques have enabled the quantification of known and unknown microorganisms at species-level resolution and revealed a tremendous microbial diversity that was not captured previously by cultivation-based methods. Many studies find metagenomic differences between healthy or diseased individuals (including experimental animals), or cohorts at risk. However, interpretation of switches in the taxonomic and functional composition of the microbiome is often limited by low sample numbers and missing replications, missing normalization to genome size, as well as heterogeneous sample processing, variability in amplification efficiency and bias by copy number variation, among others.91–93
Additionally, the gut contains not only bacteria but also viruses such as plant-derived viruses, giant viruses, and bacteriophages. 69 Bacteriophages have a high host specificity and impact microbial activity. The human gut phageome is estimated at 10 15 bacteriophages. The role of the gut virome for health and disease remains opaque. 70 Bacteriophages are considered to shape the bacterial community in the gut 71 and to modify host immune responses, e.g. by changing bacterial PAMPs and supporting the mucosal barrier of the host.71,72 Thus, phage-viral-bacterial host dynamics in the gut also need to be considered in human health and diseases. 73 Very recent data suggest that even the gut virome is altered in patients suffering from ischemic bowel disease, showing disease-specific patterns in ulcerative colitis as compared with Crohn's disease. 74 The role of the gut virome in human stroke or animal models has not been investigated so far.
Rodent models to study functional role of microbiota
Mouse models using animals raised in germ-free isolators – “GF mice” – are the workhorse of experimental microbiome research. When interpreting data from GF animals, however, one has to consider that they may deviate from normal physiology in several important ways.60,61
GF mice have underdeveloped immune structures (Peyer’s patches, mesenteric lymph nodes, and splenic white pulp) and differ from conventionally colonized mice in the abundance of several immune cell populations, such as proinflammatory invariant NK cells (iNK),20,62 IgA-producing plasma cells, and lamina propria CD4+ cells. Serum of GF mice contains fewer immunoglobulins, in particular IgG. 60
The morphology and physiology of the gastrointestinal tract is altered in GF animals. They have an enlarged caecum, a reduced overall intestinal surface area, longer and thinner intestinal villi and an increased gastrointestinal transit time. 62 Interestingly, absence of gut microbiota influences the development not only of the gastrointestinal system but also of other organs. GF mice have higher bone mineral density than conventionally colonized mice and a different metabolic status. 62 As discussed above, recent data show that the development and function of the CNS of GF animals is also affected by the absence of microbiota. Besides a “leaky” BBB 21 and altered microglia morphology and function, 22 they differ from conventionally colonized mice in levels of neurotransmitters, synaptogenesis, 26 and behavioral phenotypes.23,24,26–28
To circumvent the developmental deficiencies that result from raising a rodent in an environment lacking viral, bacterial, or parasitic agents, severe reductions of microbial diversity and abundance can be induced by combination therapies with antibiotics.63–65,98,99 However, antibiotics may have modulatory effects on the immune system66,67 and protective as well as detrimental effects on the CNS.67,68 In addition, antibiotic treatment starting in early adolescence may lead to disorders found in GF animals, such as behavioral changes (reduced anxiety and memory deficits) or altered levels of neuromodulators. 65
Other models include selective colonization with specific strains or colonization with human microbiota. Selective colonization, however, still remains an artificial system, not taking into account the effects of complex community interactions within the microbiome. In the case of “humanized” mice, it is still not possible to fully mimic the host-microbiota interplay due to differences in the composition of normal mouse and human microbiota and differences in the immune system. 89
Human studies
The ultimate goals in microbiota research are studies in humans linking disease susceptibility or pathology to changes in the composition of commensal bacteria. Not surprisingly, human microbiota research also suffers from idiosyncratic limitations. Substantial genetic variation between individuals, quite diverse living conditions and nutritional habits result in major heterogeneity of microbiota composition. Further, microbiota is affected by co-morbidities and their treatments, and depends on the age of the host. 100 Different drugs taken, and in particular administration of antibiotics affect human bacterial communities.83–86 For example, Dethlefsen et al. reported restoration of the microbiome composition only 4 weeks after the end of ciprofloxacin treatment. However, shifts in several taxa were detected even 6 months after antibiotic therapy. 51 The individual adult human microbiome seems to be relatively stable, 101 but nonetheless there are studies reporting high variability over time. 102 An important consideration, in particular with respect to studies in stroke patients, is our own experience that patients who are existentially threatened by a CNS disease may object to microbial sampling in observational studies (GUTSTROKE study, NCT02008604). In addition, no information on pre-stroke microbiota composition is available; the majority of patients experience a change of nutrition and receive antibiotics or other drugs in acute stroke care, which impact the immune system and microbial communities. As the effects of stress, life style change, and a potential influence of hospital bacterial strains have to be taken into account, complex combinations of controls are necessary (such as patients with non-CNS emergencies, TIA patients, healthy individuals who have lived with the patient, etc.). Box 1 summarizes critical issues when investigating the role of gut microbiome for stroke outcome.
In conclusion, the recent availability of powerful and inexpensive sequencing methods that allow the characterization of individual metagenomes and the use of GF animals and experimental fecal transplant approaches have led to a surge in insights into the role of gut microbiota in health and disease. In neuroscience, commensal microbiota has been incorporated into the concept of brain–gut signaling. Several groups have provided experimental evidence that intestinal microbiota is involved in the development of neurological diseases. Although as yet, little is known about the role of gut microbiota in stroke, it is highly plausible that microbiota affect outcome after stroke, and a number of groups worldwide are pursuing research into this relevant topic. However, we should learn from other fields of biomedicine, in which microbiota research is more advanced, to avoid overconfidence and overenthusiasm in interpreting our results. Cognizant of the many pitfalls and limitations of the emerging field of gut microbiome research, we will successfully elucidate the highly complex interactions of the trillions of foreign organisms we harbor in our guts with our immune and nervous systems and drawing from this knowledge we will be able to develop novel therapeutic strategies against CNS disorders.
Footnotes
Funding
The authors received financial support from the German Research Foundation (Exc 257) and the Federal Ministry of Education and Research (01 EO 08 01). KW received PhD-scholarship from the International Max Planck Research School for Infectious Diseases and Immunology (IMPRS-IDI) and the Sonnenfeld-Stiftung.
Acknowledgments
The authors would like to thank Ms. Catharine Aubel for proofreading the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
