Abstract
Chromogenic immunohistochemistry (IHC) is an important molecular localization assay in biomedical research and nonclinical drug development, enabling the visualization of specific epitopes within tissues. The methodology is widely used in drug target selection, risk assessment, understanding disease biology, and characterizing histopathological findings in nonclinical studies. The Scientific and Regulatory Policy Committee of the Society of Toxicologic Pathology formed a working group to compile essential information on chromogenic IHC assays not performed in compliance with Good Laboratory Practice (GLP) from nonclinical studies, using relevant literature and the Working Group members’ collective expertise. In this “Points to Consider” article, emphasis is placed on factors influencing IHC data quality, including sample selection, general assay considerations, data generation and interpretation, and effective reporting. The Working Group members deliberated extensively on pertinent topics, aiming to provide specific and practical guidance for pathologists, histologists, and allied scientists engaged in chromogenic IHC assays. While refraining from an exhaustive exploration of the intricate technical details associated with chromogenic IHC, this article offers insights to enhance the accuracy, credibility, and reproducibility of chromogenic IHC, thereby facilitating informed decision-making in the nonclinical development of biomedical products.
Keywords
This “Points to Consider” article is a product of a working group of the Scientific and Regulatory Policy Committee (SRPC) of the Society of Toxicologic Pathology (STP). It has been reviewed and approved by the SRPC and Executive Committee of the STP, but it does not represent a formal Best Practice recommendation of the Society; rather, it is intended to provide key “Points to Consider” in designing nonclinical studies or interpreting data from nonclinical toxicity and safety studies intended to support regulatory submissions. This article has been reviewed and endorsed by the British Society of Toxicological Pathology (BSTP) and European Society of Toxicologic Pathology (ESTP). The points expressed in this document are those of the authors and do not reflect the views or policies of their employing institutions. Readers of
Introduction
Molecular localization studies performed on cell preparations, tissue sections, and sometimes intact organs or organisms are core biomedical research assays supporting basic biological discovery and biomedical product development. In particular, immunohistochemistry (IHC) is an essential tool in modern pathology practice. Briefly, IHC uses highly specific antibodies and linked chemical reactions to visualize the distribution of specific molecules (antigens) at the light microscopic or ultrastructural levels (Figure 1). All IHC assays are based on incubating the specimen (where the terms “specimen,” “biospecimen,” and “sample” are used interchangeably within this article) with a primary antibody (monoclonal or polyclonal) that specifically recognizes and durably binds an antigen of interest (if it is present). Detection of bound primary antibodies is accomplished by noting the position of a colored product (chromogenic IHC); light emission (fluorescent IHC or immunofluorescent histochemistry); or size-defined, electron-dense metal particle (ultrastructural IHC). What sets IHC apart from other molecular detection methodologies is that IHC “staining” allows visualization of proteins or sometimes other molecules (e.g., DNA, RNA, polysaccharides) in the context of their locations inside of or associated with specific cells or tissue features. With proper assay optimization and antibody selection, IHC is a highly sensitive and specific detection technique. 106
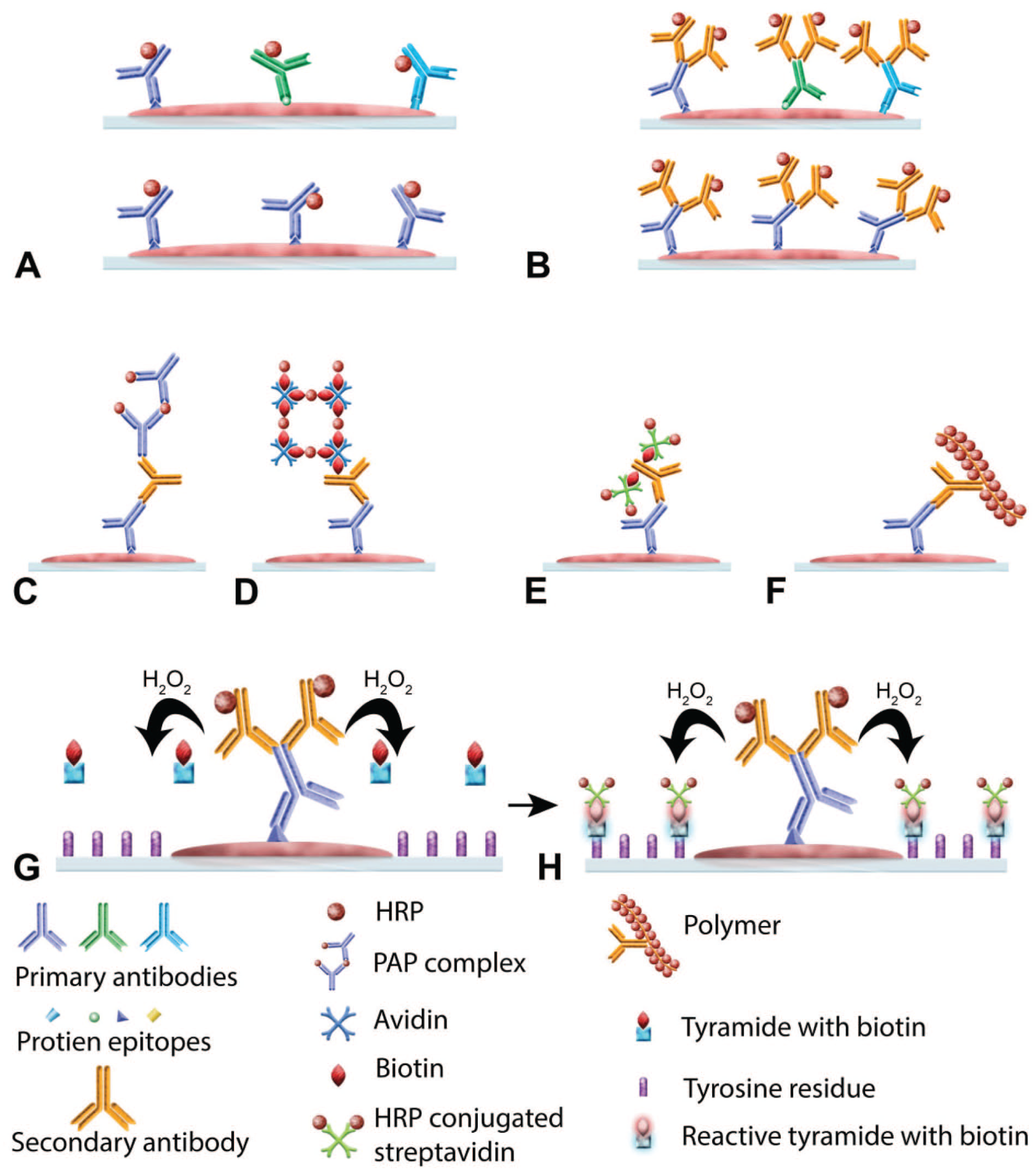
Basics of immunohistochemistry. (A) Illustration of the direct method of detecting a protein by immunohistochemistry where a polyclonal (upper) or monoclonal (lower) antibodies tagged to an enzyme such as horseradish peroxidase (HRP) are used. (B-F) Illustration of various indirect methods of detecting a protein by immunohistochemistry. A simple indirect method is shown in (B). In this method, primary antibody (polyclonal—top and monoclonal—bottom) is unconjugated and secondary antibody is conjugated to HRP or alkaline phosphatase. (C) Illustration of the peroxidase-antiperoxidase (PAP) method. First, unconjugated primary and secondary antibodies are added to the reaction, followed by the addition of PAP complex. (D) Illustration of the avidin-biotin complex (ABC) method. After the addition of unconjugated primary antibody, biotinylated secondary antibody is added, followed by the addition of HRP-conjugated avidin. (E) Illustration of the labeled streptavidin-biotin method is shown. Unconjugated primary antibody, biotinylated secondary antibody, and HRP-conjugated streptavidin are added sequentially. (F) Polymer-based method is demonstrated. After adding unconjugated primary antibody, secondary antibody and enzyme-conjugated polymer are added to the reaction. (G and H) Illustration of the catalyzed signal amplification method (also known as tyramide signal amplification). After the addition of unconjugated primary antibody and HRP-conjugated secondary antibody, biotinylated tyramide and hydrogen peroxide are added to the reaction. In the presence of hydrogen peroxide, HRP converts biotinylated tyramide to a reactive molecule, which binds to the tyrosine residues present in the proximity of the protein of interest. This is followed by the addition of HRP-conjugated streptavidin. Modified from Janardhan. 64
Despite the seemingly conceptual simplicity of IHC, generation of high-quality data by IHC can be challenging for many reasons. In this regard, key factors include preanalytical variables, reagent selection, assay conditions, visual interpretation, and approaches to data analysis.72,84,105 Nonetheless, molecular localization studies using IHC performed on tissue sections and cell lines are a core tool supporting basic biomedical research and nonclinical product development. For example, the comparison of “normal” (control) and/or diseased human to animal tissue is often undertaken to understand disease biology, assess target prevalence, identify putative tissue responses suggesting a potential liability for product development, verify the relevance of animal models, and classify observed safety signals. The IHC can be used to characterize findings in nonclinical safety studies (including discrimination of toxic changes from incidental background findings) or to demonstrate pharmacodynamic (PD) effects. At times, IHC is used to visualize the test article itself (e.g., antisense oligonucleotides, proteins, small molecules, stem cells).
Given the importance of IHC as a research tool, the Society of Toxicologic Pathology (STP) charged its Scientific and Regulatory Policy Committee (SRPC) with convening an IHC Working Group to develop “Points to Consider” for using chromogenic IHC assays in nonclinical product development. The decision to limit the charge to light microscopic chromogenic IHC of formalin-fixed, paraffin-embedded (FFPE) samples reflects the preponderance of this assay for conventional nonclinical safety studies where long-term retention of study materials requires a stable detection system; in general, chromogen signals persist longer than fluorescent signals. The methodology has also been shown to be easier to standardize across labs and is amendable to automation. Specific topics to address in the charge included sample collection and processing, data generation and interpretation by visual scoring (also commonly referred to as manual scoring) rather than automated digital analysis, and effective communication of methods and results. The IHC Working Group included 10 experienced toxicologic pathologists who were members of the STP and/or several allied global organizations of pathology: British Society of Toxicological Pathology (BSTP), European Society of Toxicologic Pathology (ESTP), International Academy of Toxicologic Pathology (IATP), American College of Veterinary Pathologists (ACVP), and/or Digital Pathology Association (DPA). In order to devise specific and useful points to consider, the IHC Working Group members first generated a detailed outline of potentially relevant topics and key discussion points, after which they met regularly to share their experiences and discuss aspects of likely value to the target audiences (i.e., pathologists, histologists, and allied scientists). The Working Group’s deliberations focused on IHC assays in which Good Laboratory Practice (GLP) compliance is not necessary as such non-GLP methods are more common in both the discovery and safety assessment settings. Information on chromogenic IHC applications in GLP-compliant nonclinical studies (e.g., tissue cross-reactivity studies for antibody therapeutics) may be gleaned elsewhere.10,19,79,82
A comprehensive review of the basic principles and technical aspects of chromogenic IHC is beyond the scope of this article. Interested readers may explore such details in many published manuscripts36,50,52,58,64,65,72,76,84,93,105,107,119,121,123,125,132,133 and books.18,32,81 Therefore, the current SRPC “Points to Consider” article is intended to describe principal considerations related to sample collection and processing as well as data generation, interpretation, and reporting for non-GLP light microscopic chromogenic IHC assays as typically used to drive decision-making in nonclinical development of biomedical products. Technical details of IHC assay design are mentioned but are limited to those features that substantially influence data generation and interpretation.
Biospecimen Selection and Informatics
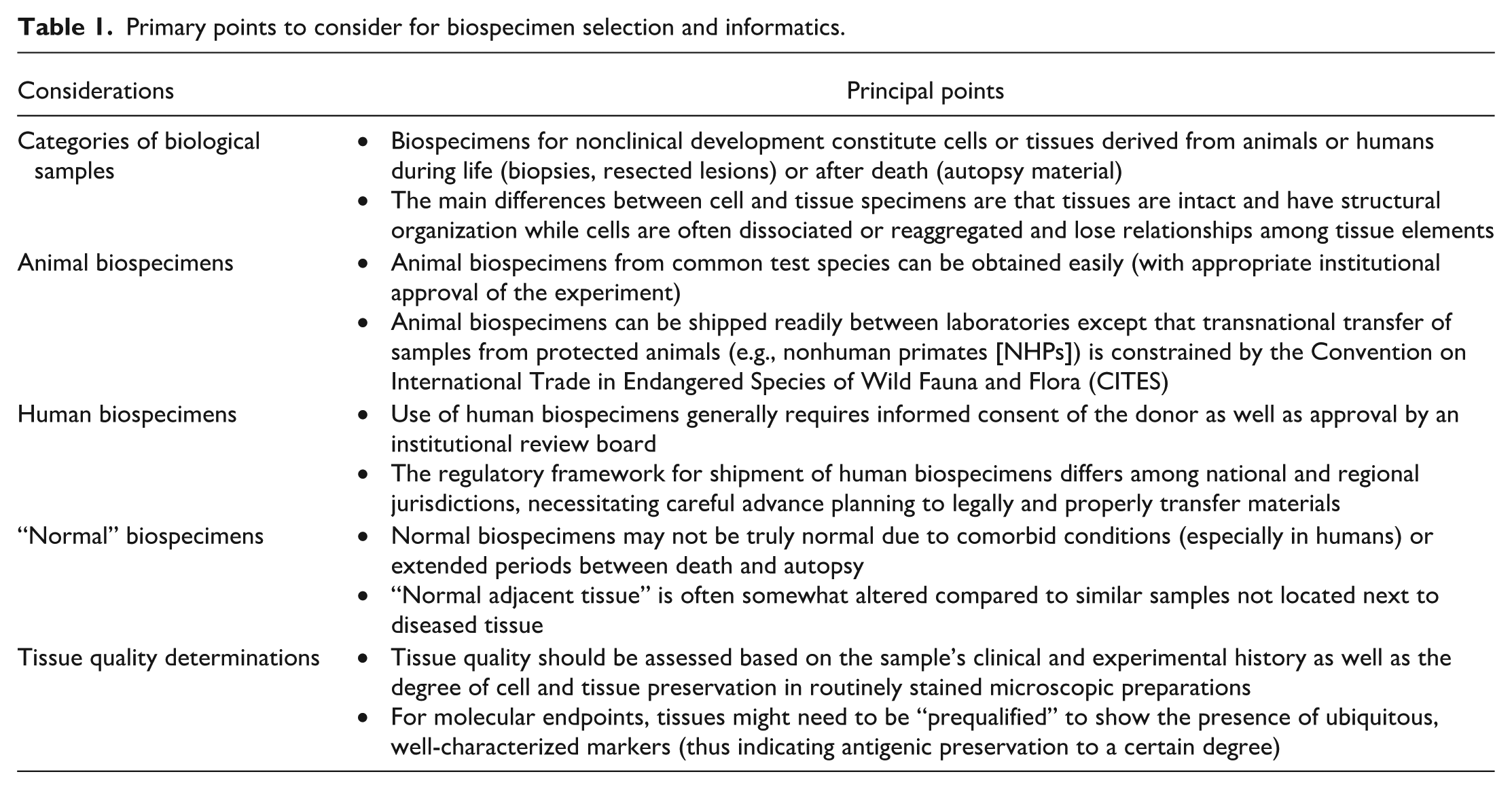
The success of molecular localization studies using IHC (or other molecular pathology assays) is determined in large part by attributes of the biospecimen (cell or tissue sample) and the accurate collection of relevant metadata (information describing various attributes of a sample) associated with the specimen. Depending on the study objectives and molecular target(s), the appropriate sample type is impacted by many factors (Table 1). This section describes parameters that influence the choice of samples for IHC assays in non-GLP nonclinical studies.
Primary points to consider for biospecimen selection and informatics.

Biospecimen Sourcing
Healthy and diseased cells and tissues from both humans and animals are essential research resources that impact development programs for virtually all biomedical products. Such samples may be obtained from living (by biopsy or surgical resection) or newly deceased (by autopsy/necropsy) individuals only when needed or may be procured from sample archives (e.g., biorepositories [“tissue banks”]). In general, chromogenic IHC is most successful when sample preservation is initiated as soon as possible after collection, although the intrinsic stability or fragility of the antigen to be detected influences this trend.
Biospecimen categories
For nonclinical studies, samples slated for chromogenic IHC are cells or tissues. Conventional cell preparations are commonly dissociated (e.g., two-dimensional [2D] cell cultures) or derived (e.g., three-dimensional [3D] cell cultures, microphysiological systems, organoids) and may be isolated from freshly harvested, unfixed tissue (i.e., primary preparations) or may be derived by serial passage of immortalized cell lines.47,67 In contrast, tissues generally comprise representative pieces of an organ (e.g., conventional tissue sections, tissue microarrays, tissue slices); organ-like structure (e.g., organoids); whole organs (e.g., micromass cultures, whole-mount organ preparations); or entire organisms (e.g., fish, whole rodent embryos).46,47 The main differences between cells and tissues are that (1) 2D and 3D structural organization remains intact in tissues and (2) any cell attributes depending on proximity to other cell types or specific tissue features (e.g., barriers or chemical gradients) are less likely to be disrupted in tissues compared to cell preparations.
Animal biospecimens
Relevant animal biospecimens are comparatively simple to obtain as long as animal care and use regulations are followed in designing and conducting nonclinical studies. In many instances, healthy animals for many test species (mouse, rat, rabbit, dog, minipig, nonhuman primate [NHP], and sheep for biomedical product development) including various rodent stocks and strains are used in nonclinical studies, and tissues are harvested at the end of the experiment. Alternatively, isolated tissues (flash-frozen or fixed) from a specific test species may be ordered from vendors. Animal models of disease (whether spontaneous or induced) provide sources of abnormal cells and/or tissues even though the clinical and/or structural manifestations of the disease may vary depending on such factors as the animal’s genotype, genetic background, sex, and age. Because nonclinical studies with animals are typically prospective, the quality (in terms of cell, molecular, and tissue integrity) is generally high for both control and diseased tissues as samples may be harvested and preserved within minutes after death, thus minimizing the extent of postmortem cell and tissue dissolution due to the action of endogenous cell enzymes (i.e., autolysis) and/or exogenous enzymes released by leukocytes or microbes (i.e., heterolysis). Transferring animal biospecimens among facilities is straightforward except for NHPs, where additional documentation is necessary for international shipments. The US Department of the Interior, based on the globally accepted Convention on International Trade in Endangered Species (CITES), regulates the shipment of NHP biospecimens (including cells and tissue sections) between research facilities in different countries. 128
Human biospecimens
Global demand for research with human biospecimens continues to grow while sample availability is constrained. 11 Centralized oversight and regulatory guidance are lacking, and no international consensus exists regarding what constitutes a human biological sample. 30 Thus, caution is warranted in transferring human biospecimens between international sites because regulatory requirements may differ among national and regional jurisdictions. The direct and indirect impact of incorrectly sourcing human samples raises many questions for various stakeholders and may introduce significant barriers that slow research progress as scientists seek to manage this risk. Specialized commercial vendors are available that provide samples with regulatory compliance. Nevertheless, institutions should have review processes in place to ensure that local government guidelines are met and that proper informed consent has been obtained for the intended biospecimen use.
Sources of human tissue include remnant clinical biopsies, resected tissues or organs (including “normal adjacent tissue” submitted for diagnosis), and autopsy material. Clinical biopsies are often small and may limit the number of tissue sections which may be obtained from remnant clinical material. Remnant clinical samples that would otherwise be slated for disposal are frequently retained for biomedical research. Autopsy material may be the only means to obtain some nondiseased human tissue types (e.g., brain, heart). Such specimens may be accompanied by clinical information but are coded so that the identity of the individual subject cannot be ascertained readily by investigators; generally, organizations that manage biorepositories must have established policies and procedures that prevent the release of personal information when human samples are shipped out for research. Use of remnant clinical samples requires informed consent of the donor or patient as well as approval in advance by an Institutional Review Board (IRB). Since informed consent may be withdrawn later in most locales, physical tracking of tissue blocks and slides may be required. Additional information may be found in the US National Cancer Institute’s publication describing best practices for utilizing biospecimen resources. 129
“Normal” biospecimens
In general, normal cells and tissues are easily obtained from animals in control groups of prospective nonclinical studies. For human tissues, “normal” connotes samples from healthy individuals (i.e., those lacking unintended comorbid [concurrent medical] conditions). However, because of the way these biospecimens are sourced, “normal” human tissues may not be truly normal. Control material obtained at autopsy (including “normal adjacent tissue” surrounding disease loci and tissues from apparently healthy donors) and remnant clinical samples may be aberrant as many patients may have comorbidities such as microbial infections or long-standing health issues such as atherosclerosis, cancer, diabetes, nonalcoholic steatohepatitis, obesity, or many other ailments. Moreover, many patients may have been or are being treated for these underlying conditions by pharmacologic means or have a history of prior or current exposure to a plethora of exogenous entities (e.g., alcohol, cannabis, pesticides, solvents, tobacco) or environmental factors (e.g., microplastics, radiation). Comorbidities and chemical exposures may not be available or well-documented in the donor’s clinical history. For samples collected at autopsy, terminal morbidities including cachexia, disseminated intravascular coagulation, hypoperfusion, pyrexia, or sepsis may substantially impact gene and protein expression from the molecular signature characteristic of a physiologically normal state.34,95,101,124 For example, terminal sepsis may increase the expression of various proinflammatory biomarkers as well as result in injury to key organs such as the kidney, liver, and lung. These alterations may not be apparent on routine histopathological evaluation but may impact the expression of molecular targets assessed by IHC techniques. Agonal duration (i.e., the time spent in passing from life to death) may also impact gene and protein expression patterns, with the changes varying with the time required for transition. The Hardy scale has been developed based on the duration of this dying process (Table 2) as a means of estimating the potential impact of perimortem factors on molecular expression. 124 For these reasons, a broad label such as “normal” may be less useful for human specimens compared to a relative definition of the tissue quality. Instead, the concepts “control” tissue, “comparator” tissue, or “experimental control tissue” may be more appropriate when describing the status of human material. Such comparator human tissues may be broadly characterized with respect to their suitability for various research purposes based on information regarding the medical history of the donor, cell and tissue histomorphology, and/or molecular profiling.
Tissue quality determinations
Assessment of the acceptability of animal and human biospecimens for use in nonclinical research (including molecular assays like chromogenic IHC) involves a tiered review of sample quality. The first step is to examine the clinical or experimental history of a specimen (including attributes like an individual’s demographic data, disease status, comorbid conditions, chemical exposures, and time between death and sample preservation). For chromogenic IHC, the next step is a microscopic evaluation of conventionally stained material (e.g., Romanowsky-stained cytology preparations or hematoxylin and eosin [H&E]-stained tissue sections) by a pathologist. Evaluation of the conventionally stained specimen confirms the identity and presence of desired cell and/or tissue features while checking the presence, extent, and uniformity of preservation. In conjunction with the clinical history, the veracity of the original diagnosis may be verified generally (e.g., confirm that a neoplasm in a specimen is a carcinoma) or specifically (i.e., a pathologist may affirm the diagnostic classification of the carcinoma). Subsequently, the likely integrity of antigens in biospecimens may be prequalified (confirmed in advance) by processing samples to demonstrate either ubiquitous and well-characterized markers such as CD31 (expressed in blood vessels), Ki67 (evident in proliferating cells), or vimentin (present in many nonepithelial cells), thereby confirming global preservation of the sample, or to verify specific retention of antigenic targets known to be sensitive to preanalytical conditions (e.g., phosphorylated proteins like histones and tyrosine kinases, which degrade if the time between sample collection and tissue fixation is prolonged). Although the preservation of one marker does not guarantee the preservation of another marker, such an approach can be adapted to include robust and more labile markers.
Biospecimen Metadata
Collection, storage, and retrieval of detailed sample metadata are essential for meaningful generation and interpretation of molecular data from animal and human biospecimens. Metadata for biospecimens includes many categories of information regarding characteristics related to health status and sample quality (Table 3). Demographic information typically includes the individual’s species, genetic background (e.g., stock, strain, breed, ethnicity, and known mutations), sex, and age. The clinical history details the primary disease (if identified) or presentation, any comorbid conditions, treatments, and concurrent exposures (when known) to other potential causes of disease (e.g., chemicals, pathogens). The experimental history records parameters assigned to or shared by individual members of particular research groups; examples include test article identity, dose, duration, and route as well as principal environmental and husbandry conditions (e.g., diet, humidity, light cycle, season, temperature). Preanalytical factors (e.g., fixative type, fixation duration, time between sample collection and preservation) can modulate gene and protein expression. Finally, documentation of appropriate research oversight (e.g., Institutional Animal Care and Use Committee [IACUC] or IRB approvals for animal and human samples, respectively) and, where warranted, the presence of the patient’s informed consent may be appended to the metadata.
Metadata considerations for animal and human biospecimens.
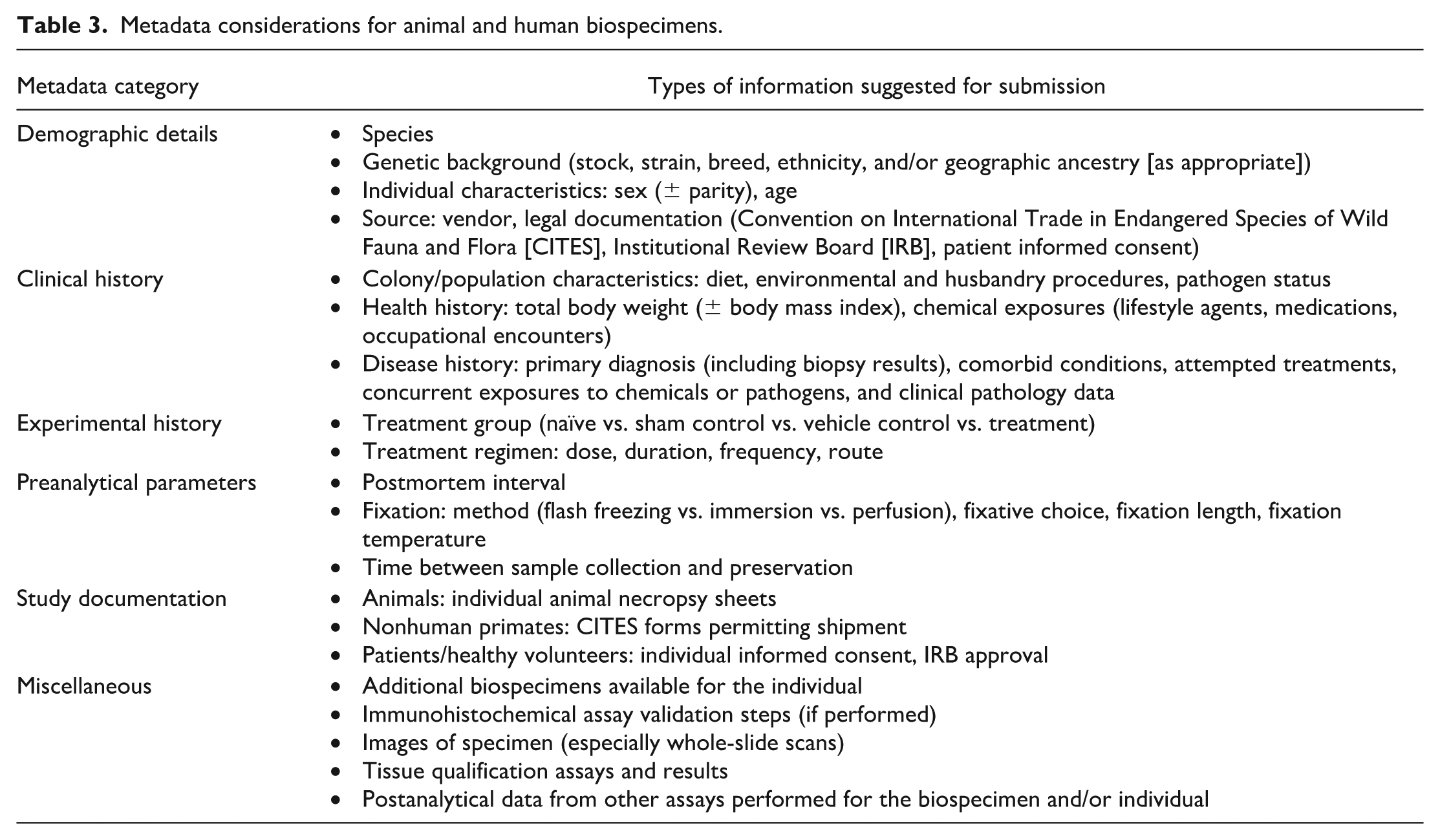
Suggestions for metadata that will improve research outcomes when using human biospecimens may be found in the Biospecimen Reporting for Improved Study Quality (BRISQ) recommendations. 94 Although intended as guidelines to enhance the content of scientific publications, the BRISQ elements are applicable to animal specimens as well. An algorithm for chromogenic IHC of tissues to demonstrate membrane, cytoplasmic, and nuclear markers with and without antigen retrieval has been developed to qualify human-derived formalin-fixed, paraffin-embedded (FFPE) samples. 41 Results of such tissue qualification assessments provide valuable metadata for archival specimens to facilitate their use to address future research questions. Taken together, consideration of such information is essential when interpreting chromogenic IHC data for nonclinical biospecimens.
Knowledge of the time between sample collection and preservation is particularly important in IHC data interpretation since diminished circulation (leading to glucose depletion and hypoxia) initiates antigen degradation. Depending on the sample type, the time component is usually recorded as either “postmortem interval” (i.e., period between death and sample preservation) or “time to preservation” (i.e., period from biopsy extraction or lesion resection, sometimes referred to as “ischemia time”) to denote the time elapsed between halting the blood supply to the specimen and cessation of metabolic pathways leading to cell and tissue degradation. Autolysis related to delayed specimen preservation leads to disintegration of nucleic acids and proteins (and thus many antigens in cells and tissues) and thus may fundamentally alter apparent molecular localization in chromogenic IHC. Generally, less autolysis is observed with biopsy and resected organs compared to autopsy material, but the time between collection and preservation may still be several hours for resected organs that are set aside during lengthy surgical procedures. Importantly, cells survive for extended periods after an organism dies, and cells expire at different rates depending on their location within the body. Autolysis and/or heterolysis are rapid in cells that are less tolerant of oxygen deficits (e.g., bone marrow) or that are exposed to large quantities of endogenous (e.g., exocrine pancreas, gastric mucosa) or exogenous (e.g., intestinal mucosa, sites of inflammation) enzymes, while cells in other tissues (e.g., cardiac and skeletal muscle) are more resistant. For a given tissue, different cell populations may exhibit divergent responses to delayed fixation. In the brain, neuronal gene expression profiles are reduced rapidly while astrocytes and microglial cells exhibit time-dependent increases in gene expression that continue for at least 24 hours after organism death. 33 In addition, delayed fixation can have a significant impact on protein phosphorylation status. These changes to the phosphoproteome distribution are unpredictable and cannot be assigned to individual proteins. 53
Taken together, integration of all available metadata is essential when interpreting chromogenic IHC data from tissue specimens. Researchers should develop robust procedures for collecting metadata if samples are procured in their own institutions and should insist on comprehensive metadata when acquiring materials from biospecimen repositories.
Antibody and Assay Considerations
In general, IHC is based on incubation of a biospecimen in a solution containing an antigen-specific primary antibody under conditions that allow the antibody to form a stable attachment to the target antigen. The effectiveness of IHC assays depends on properties inherent in both the antibodies and the assays (Table 4).
Primary points to consider for antibodies and assays.
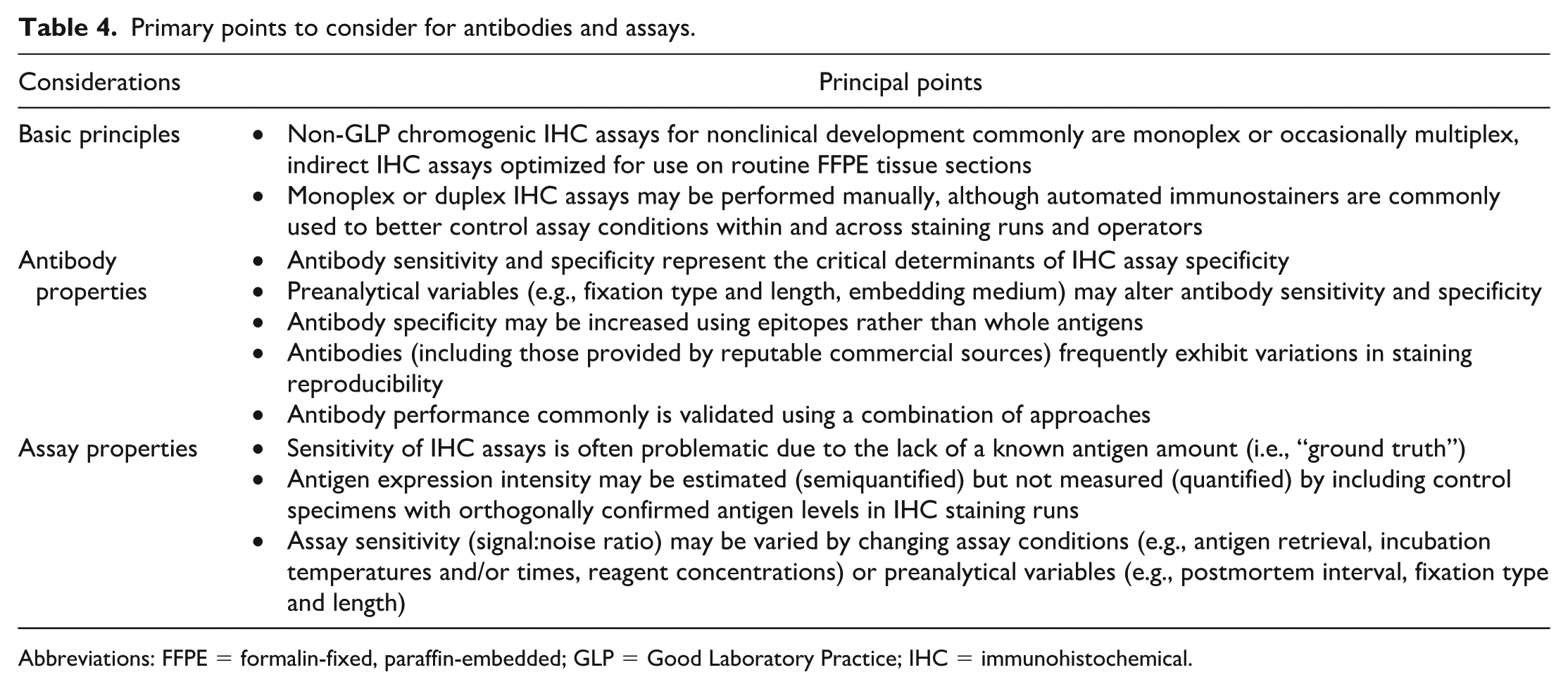
Abbreviations: FFPE = formalin-fixed, paraffin-embedded; GLP = Good Laboratory Practice; IHC = immunohistochemical.
Two detection systems may be used to reveal sites of primary antibody binding: chromogenic IHC, where deposition of a colored product is evaluated by bright-field microscopy, and immunofluorescent IHC, where light emissions are examined by fluorescence microscopy. Depending on the question, IHC (chromogenic or immunofluorescent) may involve monoplex assays (Figure 2A), in which a single primary antibody is applied, or multiplex assays in which 2 or more distinct primary antibodies are added (Figure 2B and 2C). This article emphasizes chromogenic IHC, as this approach has many advantages and relatively few disadvantages for non-GLP nonclinical studies. These advantages and disadvantages are presented in Table 5. Sample protocols for specific target antigens may be perused online.5,37,62,130 Where considered helpful by the authors, characteristics of immunofluorescent IHC are described briefly to permit comparison with chromogenic IHC assays.
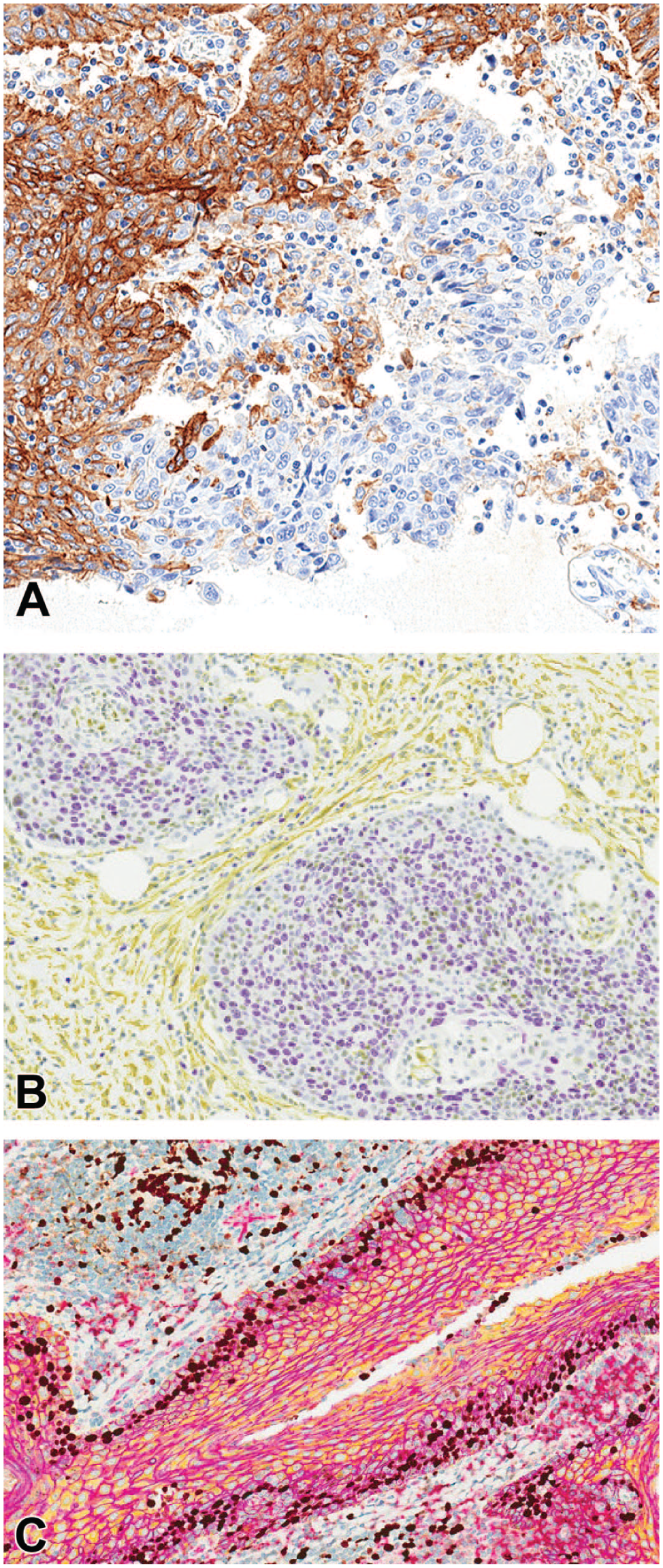
(A) Example of chromogenic monoplex staining: pulmonary carcinoma (non-small cell lung cancer [NSCLC]) from a human patient demonstrating extensive but not universal expression of programmed death-ligand 1 (PD-L1), the molecule that binds the immune checkpoint receptor programmed cell death protein 1 (PD-1). (B) Examples of chromogenic duplex staining: Ki67 and smooth muscle actin (SMA). Dividing cell nuclei stained with Ki67 marked with purple chromogen and smooth myocytes stained with anti-SMA marked with yellow chromogen. (C) Examples of chromogenic multiplex staining: CD31 marked in red, E-cadherin marked in pink, cytokeratin marked in yellow, and Ki67 marked in brown. Modified from Bolon 2017 12 , and Price. 104
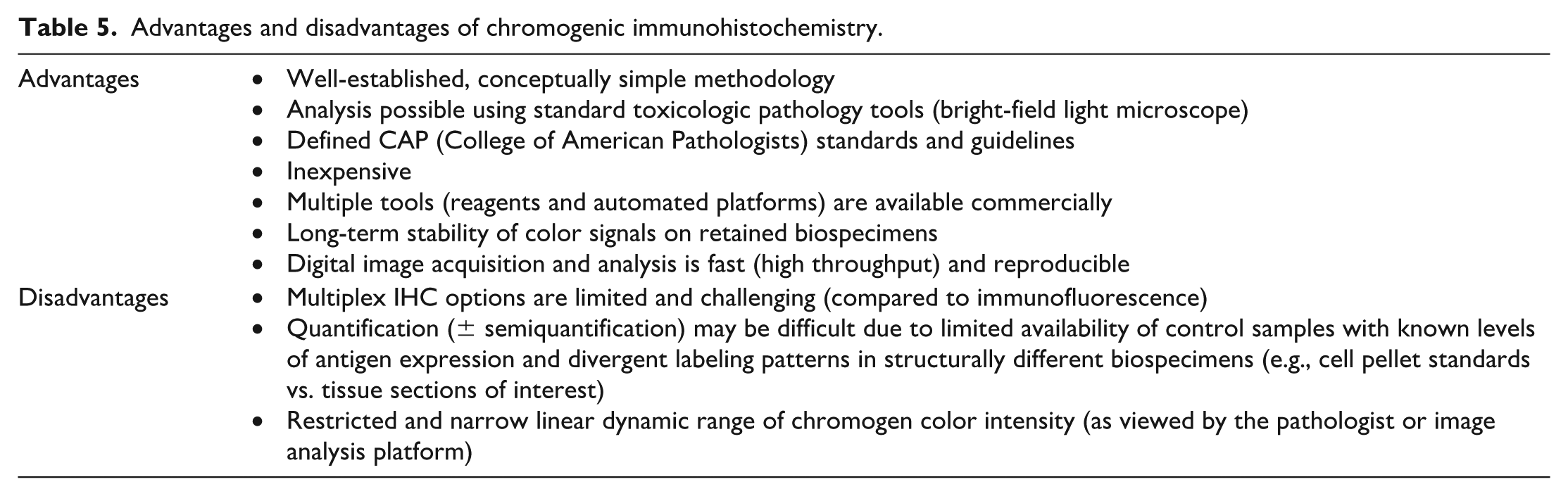
Advantages and disadvantages of chromogenic immunohistochemistry.
Basic Principles
IHC (chromogenic or immunofluorescent) involves binding of primary antibodies to antigens. Chromogenic IHC signals are detected and amplified using a conjugated enzyme to convert a non-colored substrate to a colored precipitate. The most common enzymes are horseradish peroxidase (HRP) and alkaline phosphatase (AP). The enzyme may be conjugated to the primary antibody (“direct IHC”) or linked to another reagent (“indirect IHC”) for amplification. This reagent is typically a secondary antibody (which binds to the primary antibody) conjugated to a linker molecule, while the enzyme is conjugated to a molecule (sometimes but not always an antibody) that binds the linker. The most widely used substrate for chromogenic IHC assays is 3,3’-diaminobenzidine tetrachloride (DAB), which forms a brown substrate. Other chromogens have been developed for these IHC enzymes to produce precipitates of different colors, including the HRP substrates 3-amino-9-ethylcarbazole (AEC, red precipitate) and 3,3’,5,5’ tetramethylbenzidine dihydrochloride (TMB, blue) as well as the AP substrates 5-bromo-4-chloro-3-indolyl phosphate/nitro-blue tetrazolium (BCIP/NBT, blue), BCIP/tetra-nitro-blue tetrazolium (BCIP/TNBT, purple), and Fast Red (red); other choices include agents producing green, teal, or yellow precipitates. Typical chromogenic IHC assays apply a hematoxylin counterstain, and the stained specimen is coverslipped using a suitable mounting medium.
In situations where the outcome of chromogenic IHC might present interpretive challenges, immunofluorescent IHC offers an alternative means for antigen localization. Immunofluorescent IHC is often selected where endogenous materials also used to label IHC reagents (e.g., biotin) might interfere with detection of a chromogenic signal and where very low antigen expression falls below the level of detection for chromogenic IHC staining. If signal quantification is needed, immunofluorescent IHC is sometimes preferred as the signal saturation from immunofluorescent IHC assays is more easily controlled than that from chromogenic IHC assays. Potential disadvantages of immunofluorescent IHC assays include autofluorescence (e.g., blood vessel walls [collagen], erythrocytes, and neurons); more limited ability to colocalize antigens with structural features since the dark field necessary for immunofluorescent microscopy obscures other cellular/tissue details; a greater need to standardize exposure times for signal quantification; and more challenging, time-consuming, and expensive multiplex optimization (Table 5). The impact of preanalytical variables on the antigen that may affect the value of signal quantification needs to be thoroughly explored prior to signal quantification, independent of the staining (chromogenic or fluorescent) modality. If immunofluorescent IHC is used, one should be aware of the fade rate of the chosen fluorophore and analyze or scan specimens prior to signal fading.
Signal detection systems incorporating different substrates can be used in multiplex IHC assays to simultaneously stain for several molecules in the same biospecimen. These multiplex IHC assays usually rely on sequential staining and signal detection (chromogen deposition) followed by heat denaturation of the bound primary and secondary antibodies to allow for a second assay to be run and detected with a separate set of antibodies and a different color chromogen. Monoplex IHC and sometimes duplex IHC assays may be performed manually, but many IHC assays including nearly all 3-plex and above assays rely on automated histostainers to afford better standardization and documentation of the many steps and their conditions (e.g., incubation length and temperature, reagents) involved in successfully performing such assays.25,26 Recommended practices addressing such considerations have been published recently by the Society for Immunotherapy of Cancer (SITC) Pathology Task Force. 118
Several key concepts must be well understood when considering the design of chromogenic IHC assays as well as the reliability and relevance of chromogenic IHC data. Definitions for these core terms and the relationships among them are presented in Table 6. The remainder of this section offers further details relevant to the successful conduct of chromogenic IHC during non-GLP nonclinical studies.
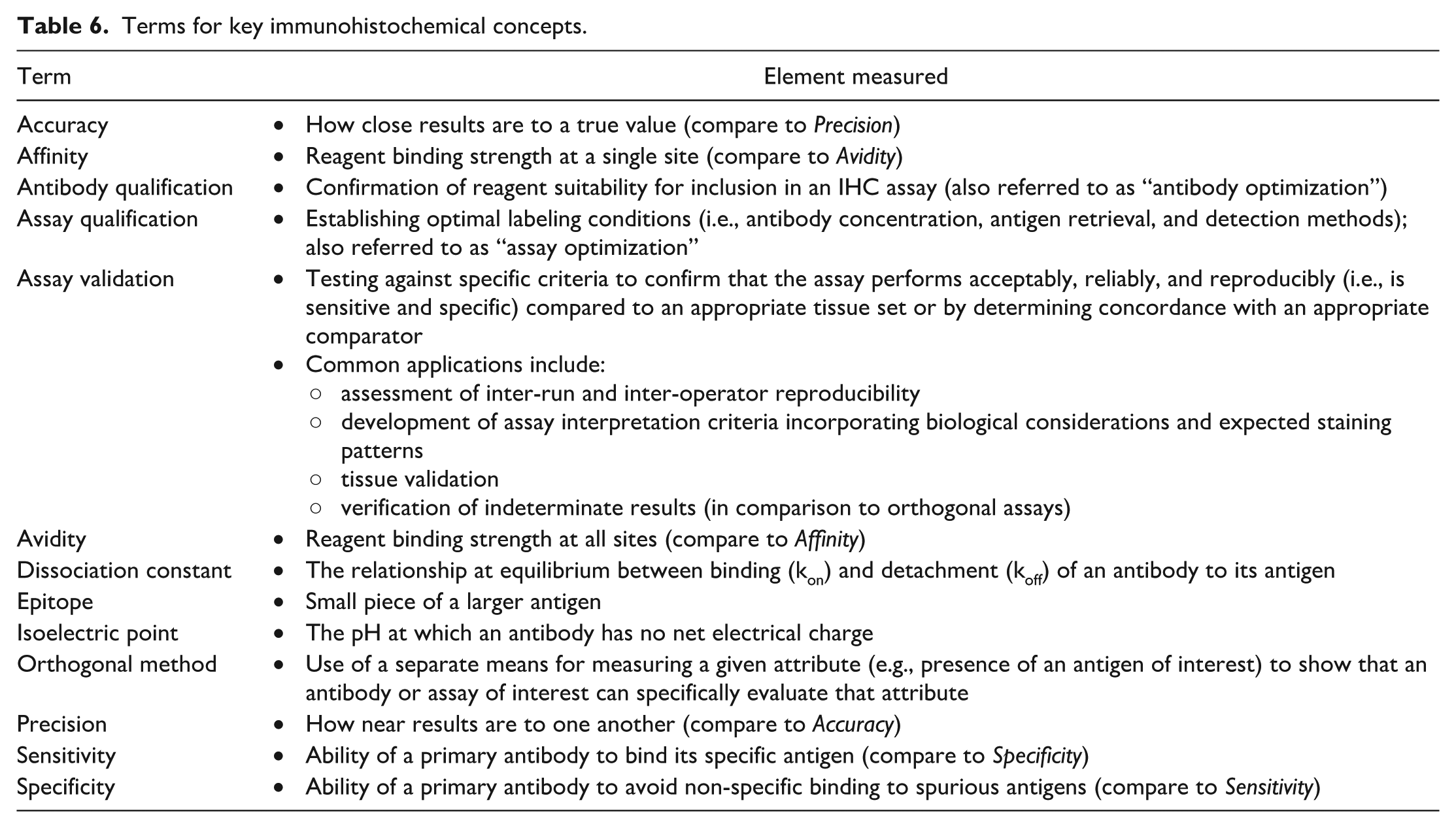
Terms for key immunohistochemical concepts.
Antibody Properties
The success of IHC assays depends on many properties inherent to the antibody reagents. The principal parameters in this regard are antibody sensitivity (i.e., correct recognition and binding of a target antigen), antibody specificity (i.e., minimal unintended binding to spurious antigens), and antibody affinity (i.e., the binding strength between an antibody and antigen).
Antibody sensitivity and specificity
Of all elements in an IHC assay, antibody sensitivity and specificity represent the critical determinants of IHC assay specificity. Antibody sensitivity most often means the ability of a highly diluted antibody to detect an antigen, while antibody specificity generally means that the antibody only detects its target antigen of interest. 125 Antibody sensitivity and specificity are dictated by the complementarity-determining region (CDR) of the variable (Fab) region of the antibody. In general, antibody reagents are generated by immunizing animals (commonly mice, rats, rabbits, or goats but sometimes donkeys or chickens) with an antigen of interest, allowing them to develop an antigen-specific humoral immune response to the antigen, and then by either affinity-purifying polyclonal antibodies or using hybridoma or cloning processes to generate monoclonal antibodies. Immunization to produce antibody reagents is often done with minimally processed antigens that retain their native conformations. Thus, the antigen used to produce an antibody may exhibit substantial differences from the same fixed and embedded antigen, which can be structurally warped. The alterations in antigen structure created by chemical conjugation during fixation and/or dehydration and temperature fluctuations experienced during embedding can reduce antibody specificity by both blunting specific antibody binding to a target antigen and introducing spurious neoantigens that permit nonspecific antibody attachment. Recently, antibody specificity has been boosted by selecting epitopes (i.e., pieces of a larger antigen) as the molecule against which a monoclonal antibody is raised. The smaller size and linear conformation of epitopes lowers the chance that their chemical structure will be altered by fixation and processing.
Antibody affinity
Antibodies have differing affinities for their target antigens, which determines their dissociation constants (i.e., equilibrium between antibody binding and release). As with any chemical reaction, the affinities of antibodies are affected by both extrinsic and intrinsic factors. For example, extrinsic parameters associated with the reaction conditions include incubation temperatures and times as well as reagent concentrations; these same factors also impact the 3D conformation of the antigens in the biospecimens. Factors intrinsic to the antibody include the chemical composition (i.e., amino acid makeup) and isoelectric point (i.e., the pH at which all charged amino acids in the antibody yield no net electrical charge for the entire molecule). For these reasons, affinity varies depending on the reagents and conditions. Therefore, an antibody may produce good specific binding (CDR-mediated binding to intended target) in some circumstances and non-specific results (CDR-mediated binding to cross-reactive [unintended] target and/or non-CDR-mediated [background] binding) under others, while a suboptimal antibody may yield acceptable results in some instances.
Optimizing antibody performance
Antibodies, including those provided by generally reputable commercial sources, often show reproducibility issues. 7 This challenge holds for different lots of a particular antibody product produced by a given vendor and also among equivalent antibodies produced by different vendors. Our collective experience and a critical reading of the literature indicate that many irreproducible IHC assays start with the false premise that the antibody is inherently specific and sensitive, thus precluding the need to optimize the assay by determining the best conditions for antibody use. Therefore, the quality of antibody reagents needs to be evaluated and optimized during IHC assay development. In particular, antibody specificity needs to be confirmed diligently.
Multiple approaches may be used to assess the sensitivity and specificity of primary antibodies. Five standard strategies have been proposed to validate antibodies, including (1) reduced expression or (2) increased expression of the target antigen in control cell lines, (3) antibody comparisons, (4) immunoprecipitation, and (5) orthogonal confirmation.58,83,127 In general, these strategies have to be performed under a particular set of assay conditions,44,49,58,127 and thus antibodies will need to be reassessed if the assay conditions are modified. Examples of possible applications are listed here:
Test the antibody on control cell lines expressing molecules that exhibit high homology to the target antigen. A common practice in this regard is to use cells that do not express the target antigen but instead have been transfected to express different members of the same molecular family that do not bear the target antigen.
Compare staining by the antibody being assessed with staining produced by an independent primary antibody that binds the same antigen but at a different epitope.
Test the antibody on sufficient biospecimens (normal and/or diseased, as warranted) of high quality for which the expression pattern of the target antigen of interest is known. Staining in cells or tissues not known to express the antigen of interest may raise concerns with respect to the specificity of the antibody.
Test the antibody against multiple biospecimens that are expected to express the antigen of interest. This multi-tissue screening gives insight into possible variability of antibody/assay performance across samples that have been harvested, fixed, and handled under various conditions.
Perform a Western blot or immunoprecipitation with the antibody to demonstrate that the antibody is recognizing a protein with the right molecular weight. However, antibodies may recognize non-linear epitopes, and Western blots should be done using either denaturing or nondenaturing conditions depending on the epitope recognized by the antibody. In addition, certain proteins may have different states of glycosylation and proteolytic cleaved states that may yield products of different molecular weights on Western blot, indicating that blot results need to be interpreted with caution.
The specificity of the primary antibody is demonstrated over time as more tests are performed to increase the weight of evidence. The ability to successfully undertake these approaches is often limited by the technical resources of the IHC laboratory.
Assay Properties
Attributes of IHC assays besides the primary antibody also influence assay success. In this respect, sensitivity is a particularly key parameter when seeking to validate assay performance. Assay sensitivity usually describes the ability of a developed IHC assay to detect very low levels of target antigen expression, while assay specificity represents the signal-to-noise ratio of the IHC assay when applied to a biospecimen.
Assay sensitivity
The sensitivity of an IHC assay has been historically difficult to define, mostly due to the lack of a known antigen amount (i.e., “ground truth”) in the biospecimen of interest. This situation is complicated because amplification leading to chromogen precipitation at sites of antibody binding to antigen is usually considered to be qualitative and not quantitative, with the IHC staining intensity following a sigmoid curve with a threshold needed for minimal detection and a plateau where the signal is saturated. Between these extremes is a narrow range where deposition of the colored product is linear. An approximation (semiquantitative) of relative antigen quantity may be estimated based on IHC signal intensity (i.e., the shade of the colored deposit, where darker colors imply more antigen) for biospecimens confirmed to possess given amounts of a target antigen using orthogonal methods. For example, well-characterized cell lines with known levels of protein expression, as assessed by flow cytometry or mass spectrometry, can be condensed into pellets by centrifugation and processed into FFPE “tissue” blocks where each pellet exhibits a different staining intensity based on their specific quantity of antigen; sections of several pellets (where each pellet expresses different antigen amounts) can be included in staining runs to provide an intensity scale for estimating antigen quantity in the sections of test tissues. Preanalytical variables can also impact target antigen, which needs to be explored as part of the assessment if an assay is suitable for meaningful signal quantification. Scanned photomicrographs of the IHC-stained sections can be evaluated by digital image analysis to measure the optical density of chromogenic deposits.109,121,132 Care must be taken in interpretation since staining intensity in cultured (isolated) cells does not always correlate with protein levels in tissue sections displaying similar staining intensity. Mass spectrometry can be used as an alternative orthogonal method to confirm antigen expression.
Assay sensitivity can be modified by altering conditions in one or more of the IHC assay steps. Common choices in this regard include changing incubation temperatures and/or times and/or the reagent concentrations. During pilot runs for IHC assay development and optimization, such adjustments are made deliberately to yield a suitable signal-to-noise ratio (where “signal” reflects the specific and “noise” represents the nonspecific binding of the antibody). The optimal sensitivity of an IHC assay will depend on the nature of the antigen. For example, conditions for an assay devised to identify a cell type-specific antigen with fairly constant expression (e.g., the CD3 receptor for T lymphocytes) will be optimized to demonstrate clearly visible positive staining in cells of the target population while producing little to no background (noise) staining in other cell types or extracellular substances. In contrast, for an IHC assay intended to detect a target antigen with variable expression from cell to cell (and where the level of expression is an important characteristic for understanding the biology of the target), the assay will be optimized to achieve a sensitivity that addresses the scientific question using a broader range of color shades, using orthogonal data to define the most suitable conditions for performing the assay.
Assay specificity
Acceptable assay specificity typically is evaluated by including one or more biospecimens that are known to exhibit substantial nonspecific antibody binding. Nonspecific binding leads to variably intense and often widespread background staining of cell and/or tissue components such as cytoplasm and extracellular matrix (Figure 3). Because they frequently demonstrate nonspecific staining, common specimens for assessing assay specificity include brain (neurons and white matter), kidney (epithelial cells), liver (hepatocytes), smooth muscle (myocytes), and stomach (glandular mucosa). Assay specificity is confirmed by a strong signal associated with antibody binding to sites with known expression of the target antigen accompanied by limited or no background staining elsewhere. 125
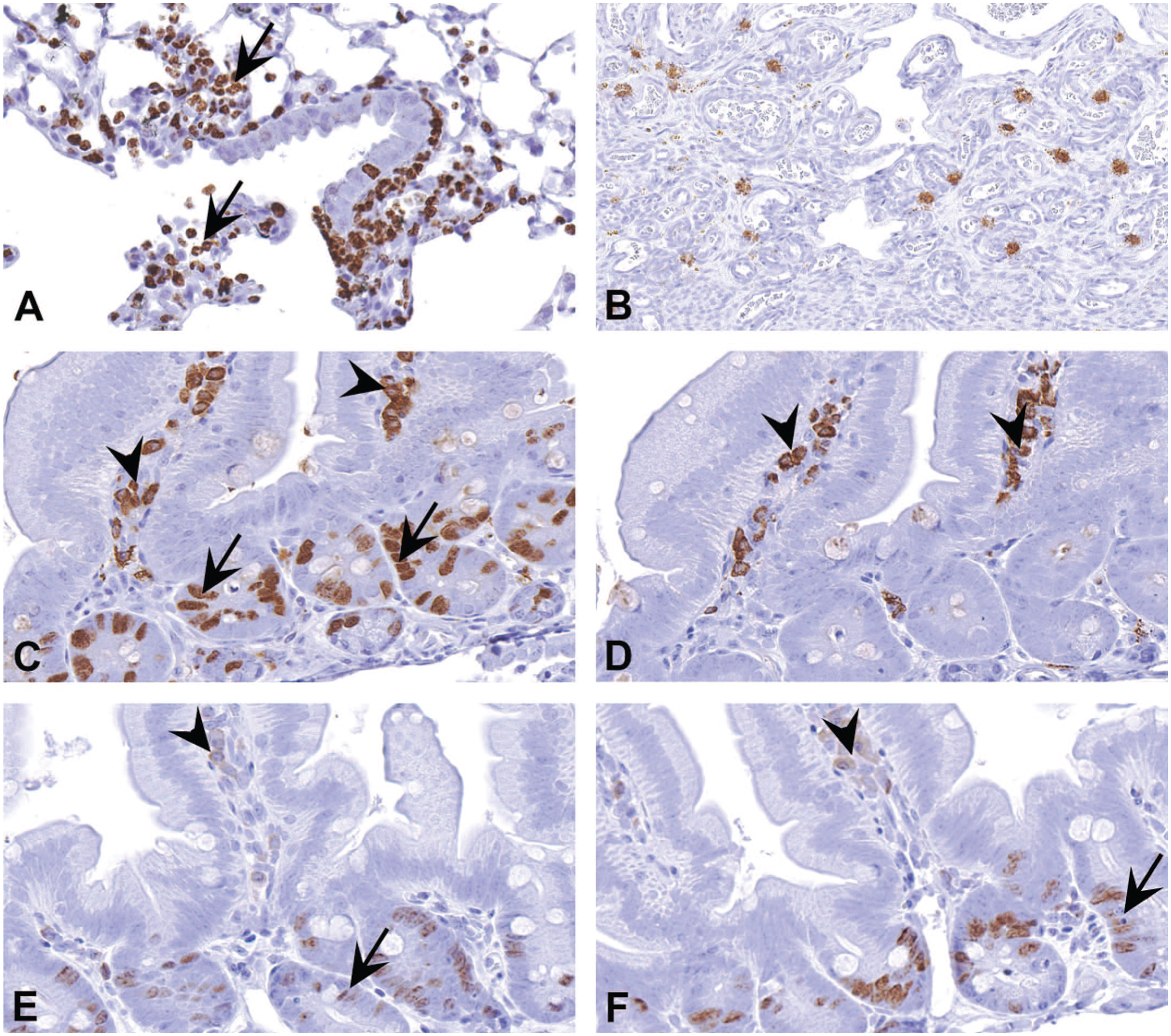
Nonspecific staining in a variety of cell types. (A) Nonspecific staining of neutrophils (arrows) by Ki67 antibody in a section of mouse lung. (B) Nonspecific staining of mast cells in a section of rat uterus. (C) Section of mouse intestine stained with a bromodeoxyuridine (BrdU) mouse monoclonal antibody. The nuclear staining in the crypt epithelial cells is specific (arrows). However, cytoplasmic staining in the plasma cells (arrowheads) in the lamina propria is nonspecific. (D) A section of the same intestine in (C) stained by omitting primary antibody. Notice cytoplasmic staining in plasma cells (arrowheads) even in the absence of primary BrdU antibody. This results from using mouse monoclonal antibody on mouse tissues. Various methods have been described to overcome this issue. (E) and (F) provide two such examples. In (E) and (F), sections of intestine are stained for BrdU using a mouse-on-mouse horseradish peroxidase (HRP) polymer and a mouse-on-mouse HRP kit, respectively. In both, there is reduction in the nonspecific cytoplasmic staining of plasma cells (arrowheads). However, the staining for BrdU is also substantially reduced (arrows). Such a reduction can be an issue when the antigen of interest is not abundant in tissue. Modified from Janardhan. 64
Optimizing assay performance
Guidelines have been published for qualification/optimizing and validating IHC assays.44,49,50,126 These resources may be consulted to obtain a more detailed explication of the considerations discussed below.
Briefly, improvements in assay sensitivity may be attempted by altering some or many assay conditions. Common options include antigen retrieval (e.g., through the use of tissue-dissolving reagents [enzymes] or heat to expose antigens); head-to-head comparisons of different primary antibodies; modifications of assay conditions (incubation temperature and time); adjustments to reagent concentration; and selection of the amplification steps and detection systems. In many cases, two or several of these options are altered in tandem to achieve optimal performance. In assay validation, the method used to validate the assay depends on its intended use. Clinical assays used for diagnostic purposes and to make treatment decisions require rigorous testing of many factors and their effects on assay performance, sensitivity, and specificity for healthy and diseased tissues. Chief among these factors are the effects of preanalytical factors (e.g., postmortem interval [length of tissue ischemia/time to preservation], fixative type and length, and antigen stability in precut slides); analytical reproducibility (e.g., intra-run, inter-run, inter-instrument, and inter-operator); and diagnostic reproducibility among pathologists (if a semiquantitative score is the expected readout for the assay). In contrast, IHC assays developed for nonclinical applications (especially if non-GLP) permit some flexibility in setting the performance norms since the intent is to explore descriptive and hypothesis-driven research objectives rather than make diagnostic and treatment decisions for human patients.
Staining Optimization
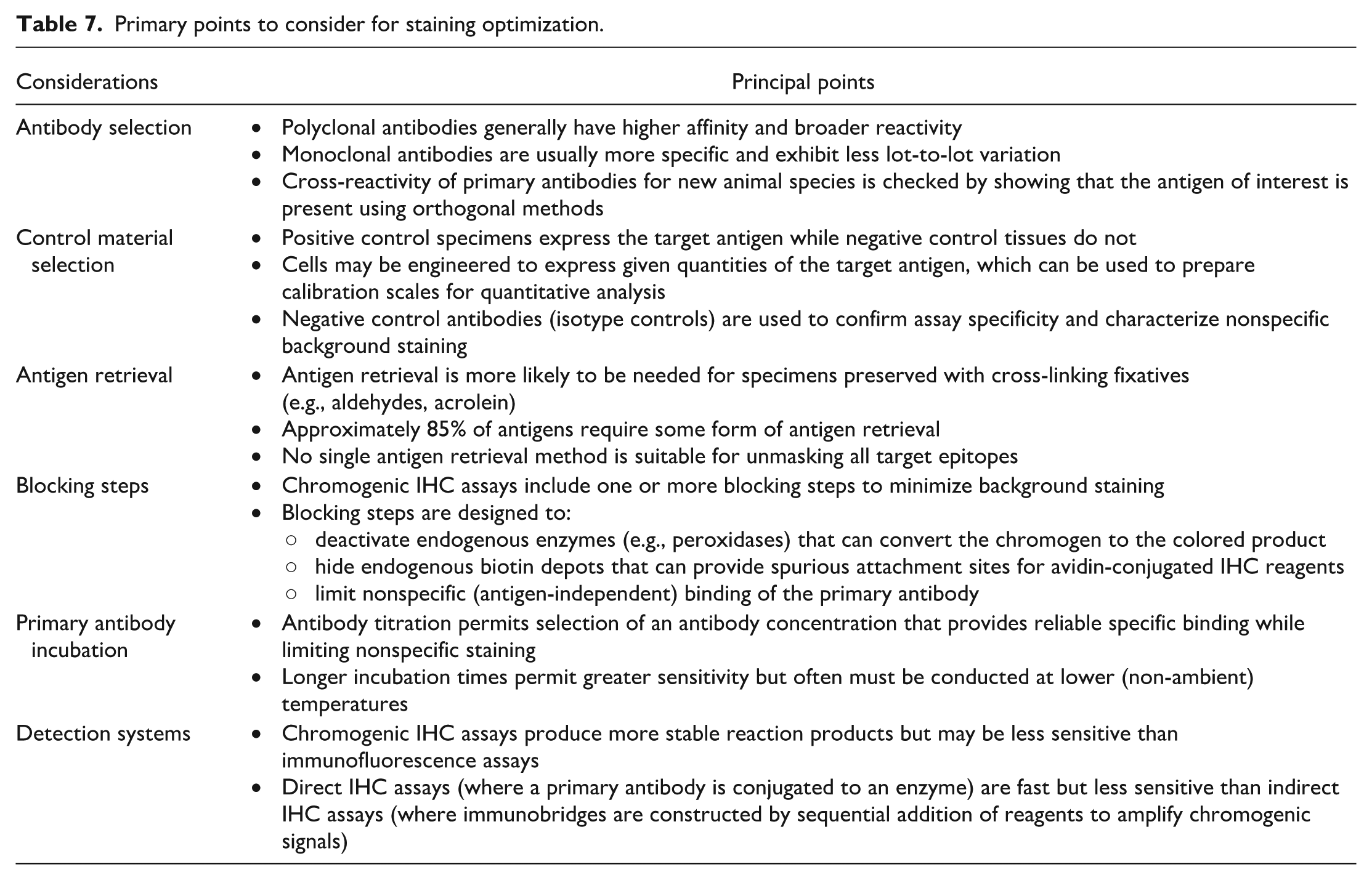
IHC provides a powerful experimental pathology tool for two purposes. First, IHC can visualize phenotypic details of cells or tissues, pathologic findings, mechanisms of action, or downstream effects of various treatments in cells or tissues. Second, IHC may enable semiquantification or quantification of different cell types and/or molecular expression levels. To be a useful tool for either purpose, IHC assays must be optimized for many variables. The objectives of staining optimization are to ensure antibody sensitivity and specificity while maximizing assay sensitivity and specificity. This section briefly covers the many variables that should be optimized when developing complex, multi-step IHC assays (Table 7).
Primary points to consider for staining optimization.
Selection of Appropriate Antigens of Interest (“Markers”)
Marker selection is a critical step in developing an IHC assay. Careful consideration should be given to whether a chosen marker (antigen of interest) will provide a meaningful answer to the scientific question being asked. For example, evaluation of apoptosis should utilize a primary antibody that binds cleaved caspase 3, a specific marker that drives programmed cell death, rather than an antibody recognizing caspase 3, the widely expressed uncleaved (inactive) protein. 15 Thorough understanding of known expression patterns of a chosen marker is important for interpreting IHC staining. Is the chosen marker expressed in the species of interest? For example, toll-like receptors (TLR) that are an important part of the innate immune system, such as TLR11, TLR12, and TLR3, are expressed in mouse but not human cells. 69 Is the chosen marker specific to a cell type of interest? In demonstrating lymphocytes, anti-CD3 will detect all T lymphocytes, while anti-CD4 and anti-CD8 recognize T-helper and T-cytotoxic/suppressor subsets of lymphocytes as well as a population of macrophages, respectively. 6 Is the chosen marker specific for a certain cell type while exhibiting a region-specific expression pattern? For example, cytokeratin 7 is uniformly expressed in transitional epithelium of the renal pelvis and ureter but is heterogeneously expressed in urinary bladder. 117 The biology of the cell and molecular targets of interest—whether there are different isoforms, phosphorylation states, or interacting proteins—might affect IHC results. For instance, various isoforms of the AKT (Ak strain transforming) protein kinase family like AKT1, AKT2, and AKT3 have differential expression in normal tissue and are associated with different processes that drive tumorigenesis. Thus, depending on the question being asked, the AKT isoform of interest will dictate selection of the appropriate primary antibody to use in IHC assays. 60
Selection of the Detection (Primary) Antibody
Several factors influence the selection of a primary antibody to be used for an IHC assay. The key parameters are the type of antibody (polyclonal or monoclonal) and the antibody source (commercial vendor vs. proprietary [developed and produced by the user]).
Reagent categories
Polyclonal antibody preparations consist of multiple antibodies, each specific for a different epitope on the target antigen of interest. Polyclonal reagents are produced by immunization of a host animal followed by harvesting of serum to concentrate the antibodies that recognize the antigen of interest. Polyclonal antibodies generally have higher assay sensitivity and broader reactivity (ability to recognize different isoforms of a target antigen) compared to monoclonal antibodies, but polyclonal reagents might also be less specific due to the variety of different epitopes recognized and the presence of other antibodies in the preparation that may bind non-target proteins.39,55,93 This potential specificity problem can be reduced by affinity purification of the polyclonal antibodies against the target antigen of interest. Finally, lot-to-lot variation of polyclonal antibodies with respect to their ability to recognize the target antigen should be carefully considered. If the polyclonal antibody will be used in an IHC assay that will be repeated many times and/or over a long period of time, multiple identical or equivalent lots of the polyclonal reagent might need to be procured and stored to maintain stability of the staining pattern. 98
Each monoclonal antibody recognizes a single epitope of the target antigen. Monoclonal reagents may be produced by classical immunization or recombinant methods in various species.73,107,131 The majority of commercially available monoclonal antibodies are produced in mice, rats, or rabbits. While monoclonal antibodies are generally more specific and have less lot-to-lot variation compared to polyclonal antibodies, off-target cross-reactivity is still possible since some epitopes are shared by multiple proteins.107,131
The choice of polyclonal vs. monoclonal antibody should be driven by the sensitivity and specificity needed as well as whether the chosen antibody can be used for IHC under the conditions to be used for tissue processing. The binding specificity of the selected antibody for its target antigen should be well demonstrated using the control materials during IHC assay optimization and confirmed using orthogonal methods.
Species considerations in antibody selection
In some cases, the source species of the antibody and the biospecimen are key factors in primary antibody selection. The reason is that additional complications arise when IHC assays use a primary antibody sourced from the same species from which biospecimens have been collected (e.g., mouse antibody on mouse tissue, rabbit antibody on rabbit tissue), usually in the form of higher nonspecific background staining. In such instances, a labeled primary antibody or a precomplexing method (e.g., “mouse on mouse” kits) may be used to minimize background staining16,45 since this method involves precomplexing the mouse antibody with antimouse IgG secondary antibody, followed by using mouse IgG to block unbound secondary antibody. Labeled antibodies are also helpful in signal amplification (which enhances assay sensitivity) and simplifying immunofluorescent multiplex assays (by using labels that fluoresce in different colors).
Reagent considerations in antibody selection
Many labels may be applied to primary antibodies for IHC assays. Examples include linkers such as biotin or digoxigenin and epitope tags such as FLAG or STREP for chromogenic IHC or fluorophores like fluorescein isothiocyanate (FITC) or rhodamine for fluorescent IHC. When using labeled antibodies, appropriate blocking steps (e.g., avidin/biotin blocks for biotinylated antibodies, detection reagents depending on the type of label present) should be incorporated when warranted into the IHC assay.17,64 If the antibody is being labeled by the user, characterization of the labeled product should also be performed including calculation of the labeling index (mols of label/mols of antibody) and demonstration that the label does not interfere with target binding. Such information is also necessary so that similarly labeled isotype control antibodies can be generated.
Of the thousands of commercially available primary (and secondary) antibodies, many will have characterization data including protein concentration, known species cross-reactivity, preferred antigen retrieval methods, best concentration/dilution for use, and the ability to use the reagent in procedures besides IHC assays (e.g., enzyme-linked immunosorbent assay [ELISA] and/or Western blots), among others.107,120 This information can be useful for beginning IHC assay optimization. However, the information provided by the vendor should not be relied on without confirmation.7,35 If no antibody is available that was raised against the target antigen from the species of interest, care should be taken to ensure that the selected antibody raised against the target in another species also cross-reacts with the target antigen in the species of interest. Some antibodies will cross-react with the target protein from multiple species, while others might only react with the target antigen from one species. Cross-reactivity of antibodies for new animal species is checked by showing that the target antigen of interest exists in biospecimens (e.g., using orthogonal methods like ELISA or Western blots to detect protein antigens of interest for IHC) and localizes to expected cells and tissues. If there is no clear species cross-reactivity information available, trial-and-error testing must be performed to determine the level of cross-reactivity across species and to ensure both antibody and assay sensitivity and specificity in that species. 89 Due to the high degree of amino acid sequence identity between humans and NHPs, particularly within antigenic epitopes, antibodies developed against human proteins frequently exhibit cross-reactivity with the corresponding NHP orthologs.
Biological considerations in antibody selection
The physiological state of antigens within biospecimens influences the choice of primary antibody for a given target antigen. Depending on the scientific question, the reagent might need to be directed against a molecule in a particular state of activation, such as phosphorylated or dephosphorylated forms, rather than merely demonstrating protein localization. Similarly, some proteins occur in tissues in two or more splice variants, only one of which may carry the antigen of interest. In such cases, the primary antibody must be directed against the specific antigen (i.e., the phosphorylated epitope or splice variant) rather than against the native molecule. The dynamic nature of protein phosphorylation and splicing and the lability of such modifications in postmortem tissues represent challenges to verifying the IHC results. 99
While routine sampling and processing for nonclinical studies produces FFPE samples, some antigens are “delicate” and must be in their native state to be recognized by antibodies. In such situations, IHC assays are performed on flash-frozen biospecimens to avoid epitope distortion induced by cross-linking fixatives (e.g., aldehydes, acrolein) as well as the dehydration and thermal fluctuation that occur during histological processing. Flash-freezing requires advance planning to assemble appropriate materials (e.g., a tray of dry ice or a container of dry ice- or liquid nitrogen-cooled isopentane, aluminum foil or plastic film for wrapping frozen samples); a suitable freezer (–20°C or colder) for storing them until sectioning; and special cryomicrotomy equipment to prepare sections. Frozen tissue specimens for IHC may be immersed directly in the freezing solution or may first be covered with a suitable embedding substrate (e.g., OCT [Optimal Cutting Temperature] medium) prior to freezing. In some cases, conventionally immersion-fixed specimens may require cryoprotection (typically by overnight infusion at 4°C in a 30% sucrose solution in phosphate-buffered saline [PBS]) to minimize structural distortion from ice crystal formation during processing. In general, optimized IHC assays for frozen biospecimens will have different conditions relative to those for optimized IHC assays performed using FFPE samples.
Vendor considerations in antibody selection
If using antibodies from commercial sources, careful qualification of vendors is necessary to ensure that reagents are of suitable quality. Substantial variation can exist in the quality and reliability of antibodies (even for monoclonal antibodies of the same clone) from vendor to vendor and across antibodies as well as from lot to lot for a specific antibody from the same vendor. In general, when first developing an IHC assay, primary antibodies are often procured in multiple forms (polyclonal and monoclonal [from multiple clones]), and/or against multiple epitopes of the same target, from several different vendors to compare reagent performance, after which the best reagent is used in devising and optimizing the assay. Experience within one’s laboratory, critical review of well-published literature demonstrating the successful performance of a reagent in an assay, and word of mouth among immunohistologists and pathologists are important means for initially estimating antibody quality.
Selection of Control Materials
Quality positive and negative control materials are critical to demonstrate sensitivity, specificity, and reproducibility of the primary antibody for an IHC assay performed under a specific set of conditions. Use of controls is not as straightforward as it appears. More than one type of control is necessary in most instances, and a weight of evidence approach should be used in interpretation of results.58,61,125,133 Control materials should be selected carefully to ensure run-to-run consistency within a laboratory and inter-facility consistency for organizations with multiple histology sites. Appropriate positive and negative control specimens and reagents (Table 7) should be included in every assay optimization experiment and then also repeated regularly (ideally every time) during definitive staining runs due to the importance for interpretation. Some controls may function as tissue controls while others are considered assay controls.
Control biospecimens
Common control specimens for chromogenic IHC assays are positive (express antigen of interest) and negative (do not express antigen of interest) specimens. These controls may be a cell line, tissue, or identifiable tissue component (e.g., blood vessel, extracellular matrix). Where feasible, control specimens and test specimens will share comparable structures since tissue integrity does affect penetration of solutions used in fixation and processing. That said, cultured cells are often used as IHC control samples for actual tissue sections since the expression of antigens of interest can be confirmed by molecular engineering of the cells. When cells will be used as controls for IHC assays performed on test tissues, a common approach is to generate a cell pellet embedded in agar or another suitable matrix to produce a pseudo-tissue that can be fixed and processed in a fashion similar to actual tissues. Positive control cells that have been engineered to express various levels of the target antigen can be a powerful tool for establishing the sensitivity of the staining assay being optimized, and under certain conditions might be employed to help quantify antigen expression levels in target tissues.
A suitable
A suitable
A single tissue may suffice as both positive and negative tissue control for well-characterized antigens. For example, spleen serves as a double control for lymphoid populations expressing either CD3 (a T lymphocyte marker) or CD19 (a B lymphocyte marker) since white pulp domains positive for one marker are negative for the other.
Control tissue can be selected from within the study (study internal controls) or by using archived tissue (study external controls). In general, external controls tested during previous studies and/or used during assay validation should be incorporated in every staining run to ensure that the IHC assay is performing consistently between staining runs and over time. The addition of internal controls helps differentiate issues arising from preanalytical variables (such as fixation and embedding) from analytical components (the IHC reagents and conditions). 125 In other words, if the IHC assay performs as expected on the external control but not on the internal control, the assay itself is functioning appropriately even though the IHC results from the study tissues are different from those expected based on IHC staining patterns observed for historical tissues.
Antibody-related controls
Multiple control antibodies may be used for various steps in developing and optimizing chromogenic IHC assays and in verifying the consistent performance of such assays over time. The choice of antibody controls depends on the nature of the question, which typically involves demonstrations of antibody specificity. Control antibodies from various species can be acquired from many commercial vendors.
Positive control antibodies are employed for two purposes. One is to demonstrate the presence of the target antigen, which is a crucial step in validating that binding by a new primary antibody directed against the same antigen may be specific. In such cases, duplex IHC to show co-localization of both the positive control and test antibodies represents strong evidence that binding of the test antibody is specific if the two antibodies recognize different non-overlapping epitopes. The second purpose is to confirm specimen quality (if not already completed via separate tissue validation staining, as discussed above) by demonstrating that the collection and preservation methods successfully protected the integrity of one or several antigens (usually distinct from the target antigen of interest) within the sample. However, it is important to understand that the integrity of different proteins within the same tissue can vary.
Negative control antibodies (i.e., isotype controls) are used to demonstrate assay specificity as well as characterize nonspecific background staining. The negative control antibody is directed against an irrelevant epitope (commonly one not present in mammalian tissues) but otherwise should be as similar to the detection (primary) antibody as possible: sourced from the same host species, representing the same isotype and subisotype, modified with similar labeling (if applicable), etc.119,122 Similarly, the negative control antibody should be used at the same protein concentration(s) and applied under the same staining conditions (e.g., incubation time and temperature) for all control (positive and negative) and test tissues. While it is understood that negative control antibodies are unique entities from the detection antibodies, their use during assay optimization is still relevant for demonstrating that IHC steps to block Fc receptor and nonspecific blocking are adequate and that artifactual staining inherent to the IHC assay has been minimized or eliminated.
Omission of the primary antibody is another antibody-related control. This step evaluates staining inherent to the IHC assay or to the specimen to be stained. 27 While this approach has been used, it is not considered by some as robust or reproducible as the use of an isotype control in place of the primary antibody. 58 If desired, both approaches can be included on the same staining run.
Epitope blocking controls can be used to help illustrate antibody specificity, particularly in situations where the primary antibody has unexpected tissue binding. In these cases, purified target antigen (e.g., a nucleotide chain or peptide) is preincubated with either the detection and/or negative control antibody prior to incubation of the antibody with the biospecimen.14,61 Implementing this control might be complicated if binding of the detection antibody to the target epitope is dependent on antigen conformation; in these cases, it can be difficult to ensure that the purified blocking antigen has folded correctly so that adequate blocking is achieved. If the blocking antigen is large, experimental design of blocking controls may be difficult (in terms of antigen synthesis costs and the risk of protein precipitation if concentrations of the antibody/blocking antigen complex rise too high) due to the need to sustain a molar excess of blocking antigen to completely thwart primary antibody binding. Thus, such blocking experiments should be approached with care and interpreted conservatively since this weak control only demonstrates that the antibody is specific to the target molecule used in generating the antibody while binding in the tissue may represent either specific binding to the target molecule or nonspecific binding to structurally similar molecular targets.58,61,133
Optimization of Other Immunohistochemical Assay Steps
IHC assays involve many steps, each of which can and should be optimized in the context of the integrated assay. A detailed review of the key elements to optimize is beyond the scope of this “Points to Consider” publication but is discussed elsewhere.18,25,26,64,105,107,122
Other Considerations for Chromogenic Immunohistochemistry
Several additional parameters influence the development and optimization of IHC assays. These factors do not relate to particular assay steps but rather to establishing the specificity of the IHC assay as a whole.122,133
Sources of specimen variation
The quality and reproducibility of IHC data can be influenced by various factors that contribute to variance in IHC staining including the subjects (i.e., animal species, stock/strain/breed/geographic origin, size, age, sex, genotype, etc.); research environment; batch effects resulting from flawed research designs; or autolysis.43,88 These sources of variation are routinely minimized during prospective nonclinical studies by careful attention to the experimental design and conduct.
Lower limit of detection
During assay optimization, the lower limit of detection for the IHC assay must be established to ensure that the assay is sufficiently sensitive to detect the target antigen in the test tissues under the same or similar conditions of collection and processing. Determining the lower limit of detection is performed using control cells with varying but known levels of target expression (e.g., flow cytometry characterization). 125 Evaluating the limit of detection is sometimes repeated after initial optimization is completed to confirm that new antibody lots still afford suitable sensitivity.
Dynamic range
The dynamic range can be defined as the highest measurable output of the biomarker to the lowest measurable output. The dynamic range represents the ability to visualize variable degrees of color intensity, which equates approximately to differing amounts of antigen (where light staining means little antigen and intense staining denotes abundant antigen). A dynamic range can be determined using cells expressing varying levels of target (e.g., via flow cytometry characterization) in a manner similar to that used to determine the lower limit of detection. 118 Determining the dynamic range for an IHC assay may be difficult. Chromogenic IHC is generally considered to be nonlinear as the “amount of color” produced depends on the activity of enzymes that saturate easily, and because additional levels of signal amplification also involve nonlinear reactions. For this reason, chromogenic staining typically should be used more to answer “yes/no” questions (“Is an antigen present in the sample?”) rather than quantitative questions (“How much antigen is present in the sample?”) when it comes to interpreting staining intensity. 118 It is possible, however, to characterize an IHC assay in detail to find the narrow part of the dynamic range where the relationship between staining intensity and protein expression is linear and utilize that range for signal quantification. However, for meaningful signal quantification, the impact of preanalytical variables on the antigen of interest and the signal intensity/dynamic range need to be thoroughly explored as well.
Automated versus manual staining
The choice between automated vs. manual staining is often dictated by investigator preference, and each assay has its own pros and cons. Automated IHC staining minimizes or eliminates human variability both within and among staining runs as well as among technicians and across laboratories. However, many automated systems are “closed/semiclosed” systems that limit the user only to prepackaged proprietary buffers, antibodies, and other reagents from that vendor as well as limit the ability to vary or optimize individual steps in the staining protocol. Some vendors offer automated “open” systems that allow greater flexibility with respect to incorporating home-made or nonproprietary reagents and adjusting individual assay steps. Automated staining systems are capable of handling conventional 75 × 25 mm slides but might not be capable of handling slides of unusual size (e.g., 75 × 50 mm). In contrast, manual staining allows the greatest flexibility with respect to reagents that can be used (thus expanding the choice of antibodies and vendors), the order of reagent application, and the size of each staining run.
Preanalytical Variables
Many variables associated with specimen collection and processing can influence the outcome of IHC analysis (Table 8).26,40,121 Some major variables include biospecimen collection, sampling, fixation, trimming, processing, sectioning, and archiving (Figure 4). Optimizing as many of these attributes as possible is important as they affect the interpretation and reproducibility of IHC results.
Primary points to consider for preanalytical variables.
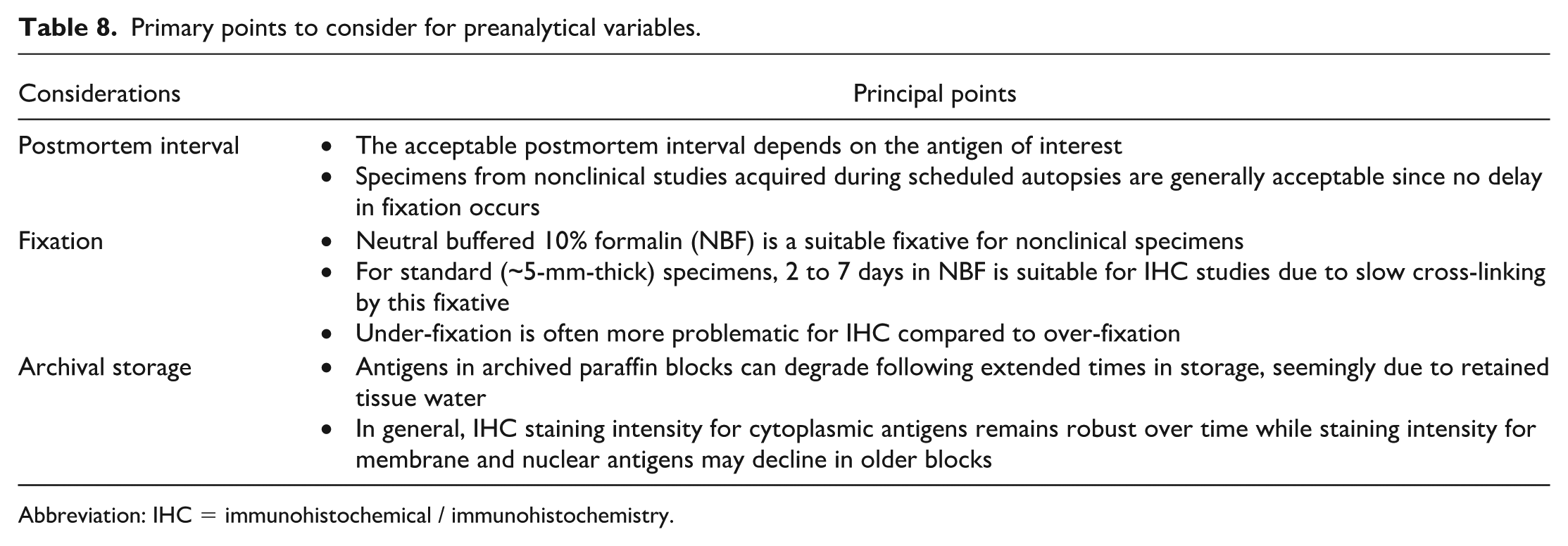
Abbreviation: IHC = immunohistochemical / immunohistochemistry.
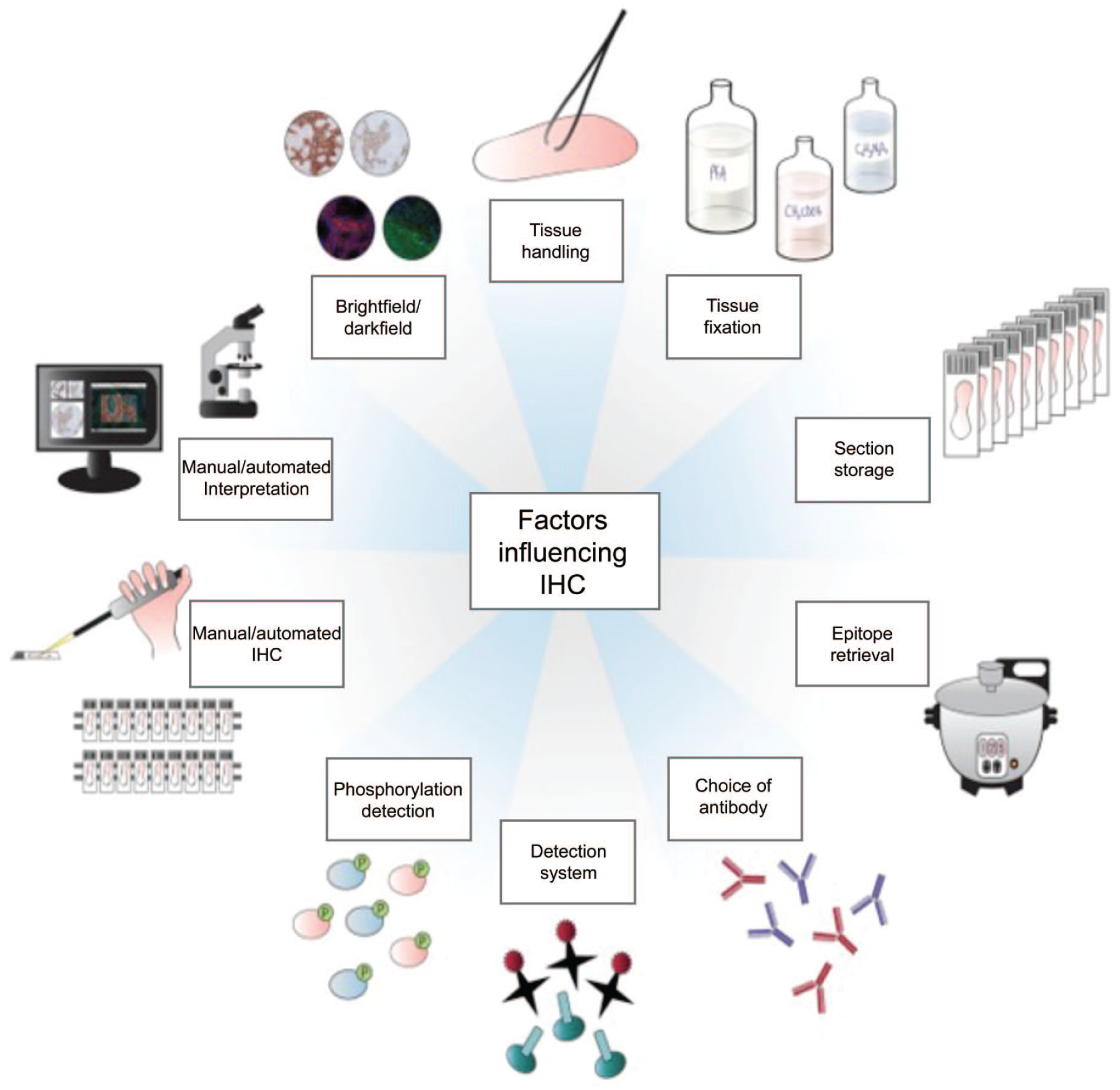
A schematic representation of various factors that may influence the standardization and reproducibility of the immunohistochemistry (IHC) process. Modified from O’Hurley. 99
Postmortem Interval
The delay between specimen removal and fixation, designated for autopsy samples as the postmortem interval or time to preservation, affects the outcome of chromogenic IHC assays. The acceptable cold ischemia time (CIT) differs depending on the antigen of interest and objective, so the time to preservation should be recorded as an estimate of specimen/antigen preservation. For example, in human samples, a time to preservation of 30 minutes can alter the protein levels relevant to certain markers of cancer progression while for others an interval of 1 hour for human biopsy specimens supports acceptable IHC staining. 29 This timing indicates that specimens from nonclinical studies that are acquired during a scheduled necropsy should be acceptable as no delay in fixation should occur in this scenario.
Fixation
Stabilization of cell and tissue constituents by appropriate fixation is the most important variable affecting the outcome of IHC. Poor fixation cannot be remedied later, so the effects of fixatives and the fixation process on antigens must be considered in advance. Key elements in this regard include the choice of fixative, duration of fixation, ratio of fixative volume to tissue mass, and fixation temperature.
In general, neutral buffered 10% formalin (NBF), which includes approximately 1% methanol as a stabilizing agent in most commercial formulations, is a suitable fixative for chromogenic IHC assays in terms of balancing superior morphological preservation and the ability to detect antigens.100,105,121 Fixation with 4% methanol-free formaldehyde (MFF, colloquially known as 4% paraformaldehyde [PFA] since it is constituted from PFA pellets or powder) is preferred by some researchers for optimizing IHC results, but MFF has several disadvantages (e.g., needs to be prepared right before use, polymerizes in solution over time, requires more time to penetrate) and often is equivalent to NBF in terms of tissue preservation.70,116 While NBF penetrates rapidly, actual tissue fixation is slow. During fixation with NBF, cross-linking occurs progressively over time and is impacted by variables such as specimen thickness, temperature, and pH.56,57,59 Tissues from nonclinical studies typically are fixed at room temperature (RT) since this approach offers efficient fixation. It is important to ensure that control and test samples are fixed under similar conditions.
When developing IHC assays, under-fixation is often more problematic than over-fixation. 50 In the authors’ experience, many epitopes lose little immunoreactivity with 2 weeks, sometimes more, in NBF. However, for some epitopes, leaving tissues in NBF for a very long time can lead to false negative IHC results due to epitope distortion caused by the progressive aldehyde cross-linking. 105 In contrast, short fixation times (<24 h) in NBF often result in aldehyde-mediated molecular cross-linking at the specimen periphery while the core undergoes coagulation fixation only by exposure to organic solvents (especially alcohol) during histological processing. 105 This variable fixation results in uneven specimen staining, which complicates the interpretation of the IHC results. Therefore, recommended fixation times in NBF are usually 24 to 48 hours, followed by embedding or short-term storage in alcohol.
Certain tissues require special fixatives (mixing aldehydes, alcohols, and acids) for optimal fixation, such as Bouin’s, Davidson’s, or modified Davidson’s solutions for eyes and testes. 77 Choosing the appropriate fixative for IHC in such thickly encapsulated tissues depends on the antigen of interest, 23 and IHC assays performed on specimens fixed in this fashion may need to be reoptimized since the fixative composition differs considerably from pure aldehyde-based variants like NBF and MFF. Similar considerations also apply to hard specimens (e.g., bone, teeth) that require decalcification prior to routine histological processing.20,85,111
Specimen Dimensions
As noted above, specimen thickness impacts the speed of fixation. A maximum thickness of 4 to 5 mm is ideal for most tissues as fixative penetration occurs quickly from both sides of planar samples. Fixation is further aided by immersion of specimens in an excess of fixative solution since more free aldehydes are available to react with molecules in the sample. An ideal ratio between the fixative volume and tissue mass is 20:1 while the minimal fixative volume to tissue mass ratio should be 10:1. 78
Reagent Turnover
Fixatives and reagents used in tissue processing have finite shelf-lives and must be replaced at intervals to ensure specimen quality. For example, aldehyde cross-linking (polymerization) in older fixative lots weakens the fixative strength by reducing the number of fixative molecules available to preserve molecules in biospecimens. Similarly, delayed rotation of alcohol baths in tissue processors leads to gradual accumulation of water, which in turn leads to inadequate dehydration of biospecimens during the embedding process. Water retention in stored FFPE blocks of tissue is associated with gradual antigen degradation. 134
Paraffin Selection
Different variants of paraffin wax used for specimen embedding melt at distinct temperatures. Conventional paraffin (melting temperature, ≥58°C) supports IHC assays for most applications. Paraffin variants that melt at low temperatures (≤54°C) limit the extent of heat-mediated molecular degradation in biospecimens during the embedding process.
Archival Storage Conditions
Antigens in archived paraffin blocks can degrade following extended times in storage (where the length of “extended” depends on the antigen). In general, IHC staining intensity for cytoplasmic antigens remains robust over time while staining intensity for membrane and nuclear antigens declines in older blocks. 51 Reduced staining may be ameliorated by sectioning deep into blocks and lengthening heat-induced epitope retrieval (HIER) pretreatment.
Environmental parameters during storage of paraffin blocks and slides affect the outcome of IHC.38,108 Storing blocks in an office-like environment (i.e., dry conditions protected from excess or insufficient humidity, elevated temperatures, and light) limits molecular degradation in specimens stored as uncut blocks. Precut unstained slides should be stored in enclosed cases away from dust and direct light. Storing slides at 4°C is common practice, and prolonged stability (6 months or more) has been shown for some targets/epitopes. However, in some instances, it can be detrimental as excess moisture can lead to hydrolysis of molecules exposed in precut unstained sections. Some laboratories practice paraffin coating of precut unstained slides before storage; however, this practice does not save time and has not proven to be beneficial in preserving antigen structure. Ideally, use of freshly cut sections will avoid potential variations in staining resulting from storage of precut sections. Alternatively, a constant but limited storage time (e.g., 2 or 4 weeks) can be tested and optimized for precut unstained sections under ideal conditions and then followed as a standard procedure in the laboratory. The storage conditions for precut sections may need to be optimized for each antigen/epitope.
Scoring Paradigms
Analysis of chromogenic IHC assays may be performed in several manners (Table 9), and these scoring approaches may be influenced by the tissue staining patterns (Figure 5). Visual scoring (also commonly referred to as manual scoring) yields qualitative or semiquantitative IHC data as values assigned by the pathologist. In the nonclinical sphere, visual IHC scoring helps investigate and characterize a target antigen. In the clinical setting, visual IHC scoring (e.g., for in vitro diagnostic [IVD] tests) supports treatment decisions, predicts patient prognosis, and determines clinical trial design, enrollment, and outcomes. A visual scoring system for IHC assays should be definable, reproducible, and produce meaningful results.1,87,88 Visual scoring strategies may also be devised ad hoc for certain studies. In recent years, theoretical and practical advances in computer and statistical science have allowed valid quantitative measurements of IHC staining. The choice among such options depends on both investigator preference and the study objective.
Primary points to consider for scoring paradigms.
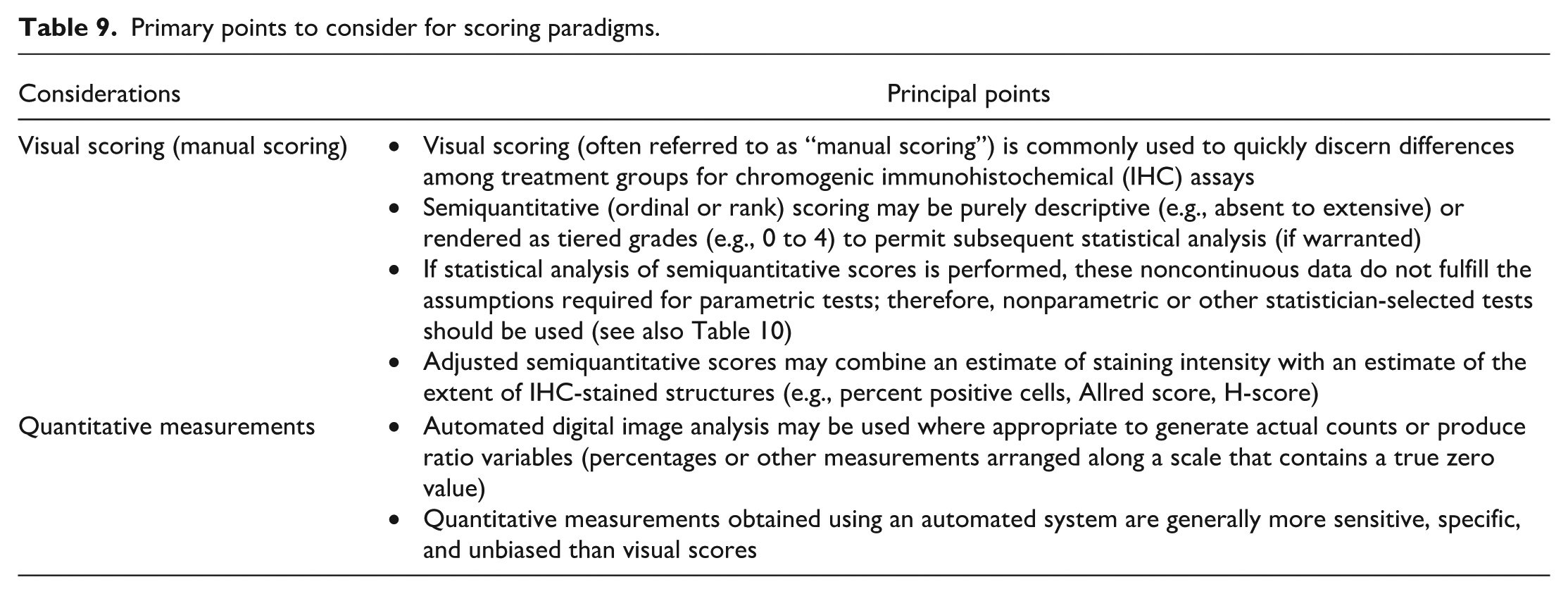
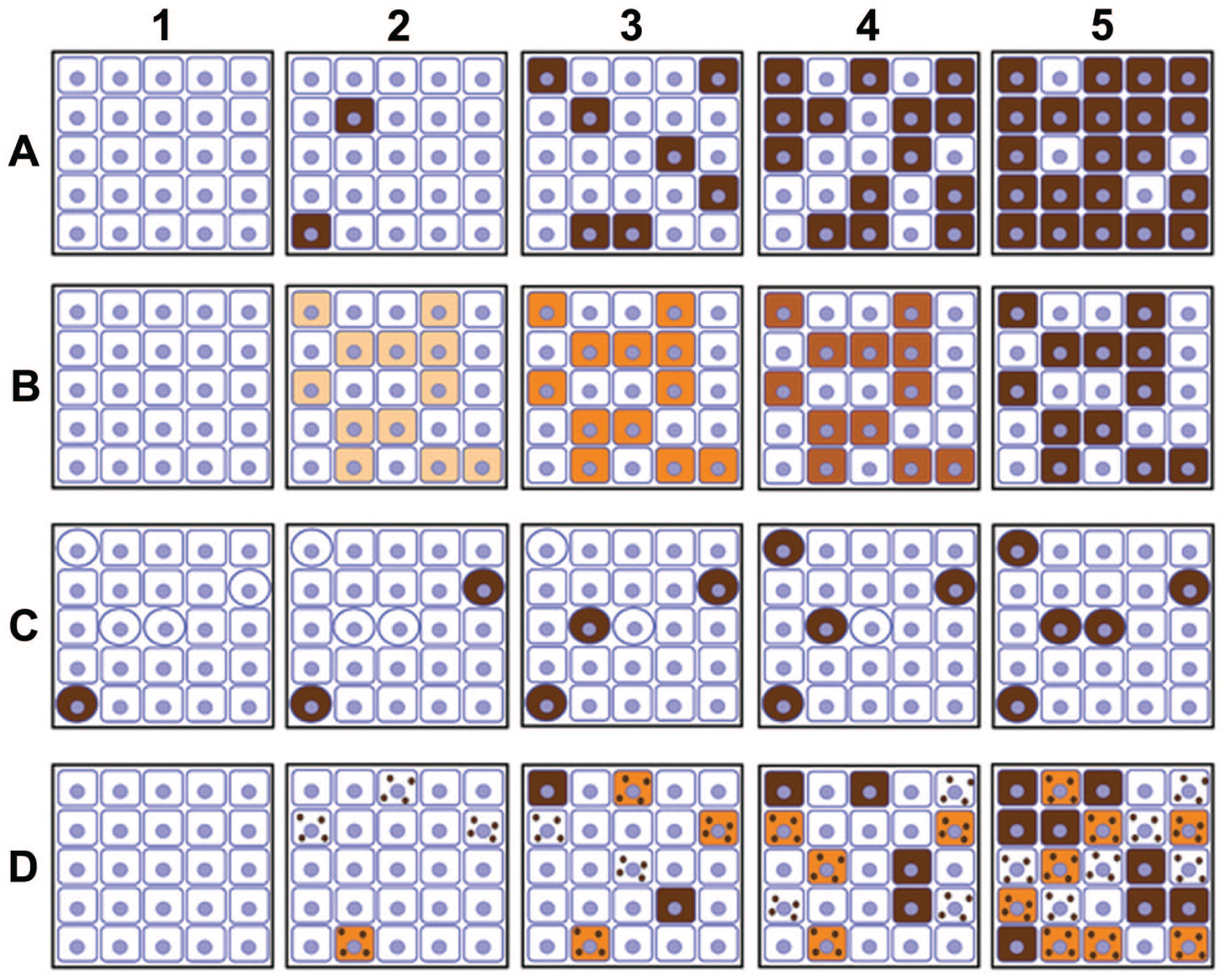
Types of cellular staining patterns (brown coloration) with differences in (A) frequency, (B) intensity, (C) cell type, or (D) mixed staining. Modified from Meyerhold. 88
It is important to recognize that toxicologic pathologists have a broad spectrum of functional roles in early biological discovery through final regulatory approval. Thus, regulatory guidelines and standards for pathologists can have different contexts in terms of allowable data usage and handling. One common example of this divergence is the use of GLP vs. non-GLP laboratory standards—each has distinct expectations in what is commonly needed and accepted. The uses and analyses of tissue scores can also necessitate different practices within the spectrum. For example, early discovery studies often utilize tissue scores (quantitative, semiquantitative) and their statistical analyses to help better understand the pathology data and guide the designs (including specimen collection and scoring approaches) of future studies. At the other end of the spectrum, regulatory policies typically define tissue scores as “descriptive” and “subjective,” so application of statistical analyses are not deemed as valid or useful. Therefore, when reading the subsequent scoring section of this article, the reader is encouraged to keep in mind that some applications (e.g., statistical analyses of tissue scores) may be of relevance for one segment of pathologists and not for another.
Common Scoring Approaches
Visual scoring is a common choice for analyzing chromogenic IHC staining during nonclinical studies and is performed for several reasons. First, visual scoring is typically used for general determination of differences among different treatment groups. In this regard, visual scoring based on carefully defined and biologically relevant criteria is an easy, efficient, and fast means for obtaining qualitative or semiquantitative answers to research questions. Second, visual scoring may serve as a tool for clinical diagnosis and prognosis. Visual scoring is typically the first-line choice for analysis of staining for chromogenic IHC assays for these two purposes. Finally, visual scoring may aid in identifying situations and target endpoints appropriate for quantitative assessment, such as high-throughput cell counts. In such situations, automated analytical systems to quantify IHC staining are often more efficient and faster than visual scoring. Importantly, IHC data should have biological relevance such that specific grades predict the presence and/or severity of findings that can be observed for the nonclinical model. 90
For any visually collected data (e.g., semiquantitative scores), these must be generated using clearly defined, reproducible, and objective scoring parameters. 31 Ideally, the scoring approach could be validated by more than one pathologist. Interobserver validation can act as quality control to ensure consistency (e.g., a grade of “1” is consistently applied by different pathologists).31,112
Nominal (binary) scoring
The most basic scoring paradigm for IHC staining categorizes entire tissue sections on a nominal (binary) scale as either positive or negative based on the present or absent expression of a particular biomarker. Binary scoring is usually used to produce a percentage case incidence for each group, and any staining observed that is above the background staining level classifies the sample as positive. An alluring feature of this approach is that it is less subjective than some other scoring methods since it is essentially a “Yes” or “No” assessment with less room for bias. Nominal data may be summarized in a contingency table for positive or negative cases relative to the total cases for each diagnostic classification with group analysis by chi-square or Fisher’s exact test.
Ordinal (rank) scoring
The rank (or “ordering”) scoring paradigm is a simple and quick approach in which each sample from a study is sorted from least to most substantially affected. As with severity grading for findings on H&E-stained sections, semiquantitative IHC staining determines the amount and/or intensity from least to greatest using several ranks (commonly called “grades”)—typically negative, weak (or minimal), strong (or moderate), and intense (marked or high). Ideally, ordinal scores should be based on only one parameter and should be framed by a set of discrete, easily discerned features; such well-defined criteria facilitate interobserver reproducibility.87,112 Some studies may try to combine multiple parameters together (e.g., markers for proliferation and cell death) to produce one score, but this approach should be avoided as it makes interobserver repeatability more challenging. Ordinal variables may be purely descriptive (e.g., absent to extensive) or rendered as tiered scores (e.g., 0 to 4) to permit subsequent analysis (if warranted). A masked (“blinded”) approach is common to minimize bias and “diagnostic drift” (i.e., a gradual change in scoring expression of a target antigen within a single study over time). 88 Masked evaluations can have context in their application.13,31,112 A masked evaluation of a study and its treatment groups can be beneficial when the toxicant has a known lesion spectrum and the pathologist has experience with the model; an example of this might include safety studies for the US Food and Drug Administration (FDA). In situations where a model (e.g., toxicant, dose, species, route of administration, etc.) and its lesion spectrum are not well-defined, pathologists are recommended to have knowledge of treatment groups at the beginning of the evaluation. Then, after initial assessment, pathologists can perform a targeted masked review of specific tissues or microscopic findings.
Percent positive scoring approaches
Another semiquantitative scoring approach is to estimate the percentage of positive staining for a particular cell population or tissue. This approach may be performed in various ways. An example of a simple 4-tiered system is as follows: score 0 means no staining, a score of 1 has less than 10% of cells with staining, a score of 2 has 10% to 50% of cells staining, and a score of 3 has greater than 50% of cells staining. 86 Alternatively, the “percent positive” is determined by counting the percentage of positive immunolabeled cells over the total cells in a selected area/region of interest. The denominator can either be the entire cell population present in the tissue section or a subcompartment (such as tumor cells within the tissue). 1
Percent positive scoring can be enhanced by integrating staining intensity to produce an immunoreactivity score (IRS). The IRS is calculated as the sum or product of the ordinal scores for “percent positive” tissue (available scores, 0 to 3) and “staining intensity” (available scores, 0 to 4). Multiplication is typically used, so the IRS can range from 0 to 12 based on frequency and intensity. 42
The H-score (histochemical score) is used widely in the clinical setting. Like an IRS, it combines an assessment of staining intensity with an estimation of the percentage of stained cells at each intensity. Staining intensity is scored as 0, 1, 2, or 3 (“bins”) corresponding to the presence of negative, weak, intermediate, and strong staining, respectively, and for each category the percentage of stained target cells is recorded. The final H-score is the sum of each intensity rating multiplied with its corresponding percentage, generating a score on a continuous scale of 0 to 300, where 300 = 100% of tumor cells staining strongly with a score of 3. 88 In the context of data/statistical analysis, it is important to understand that while the H-score appears to be on a continuous scale, its underlying assessment of staining intensity relies on scoring bins (i.e., not a continuous scale).
The Allred score (sometimes called the quick score), like the H-score, also combines the assessment of staining intensity and proportion of cells staining. The formula for the Allred score equals the intensity score
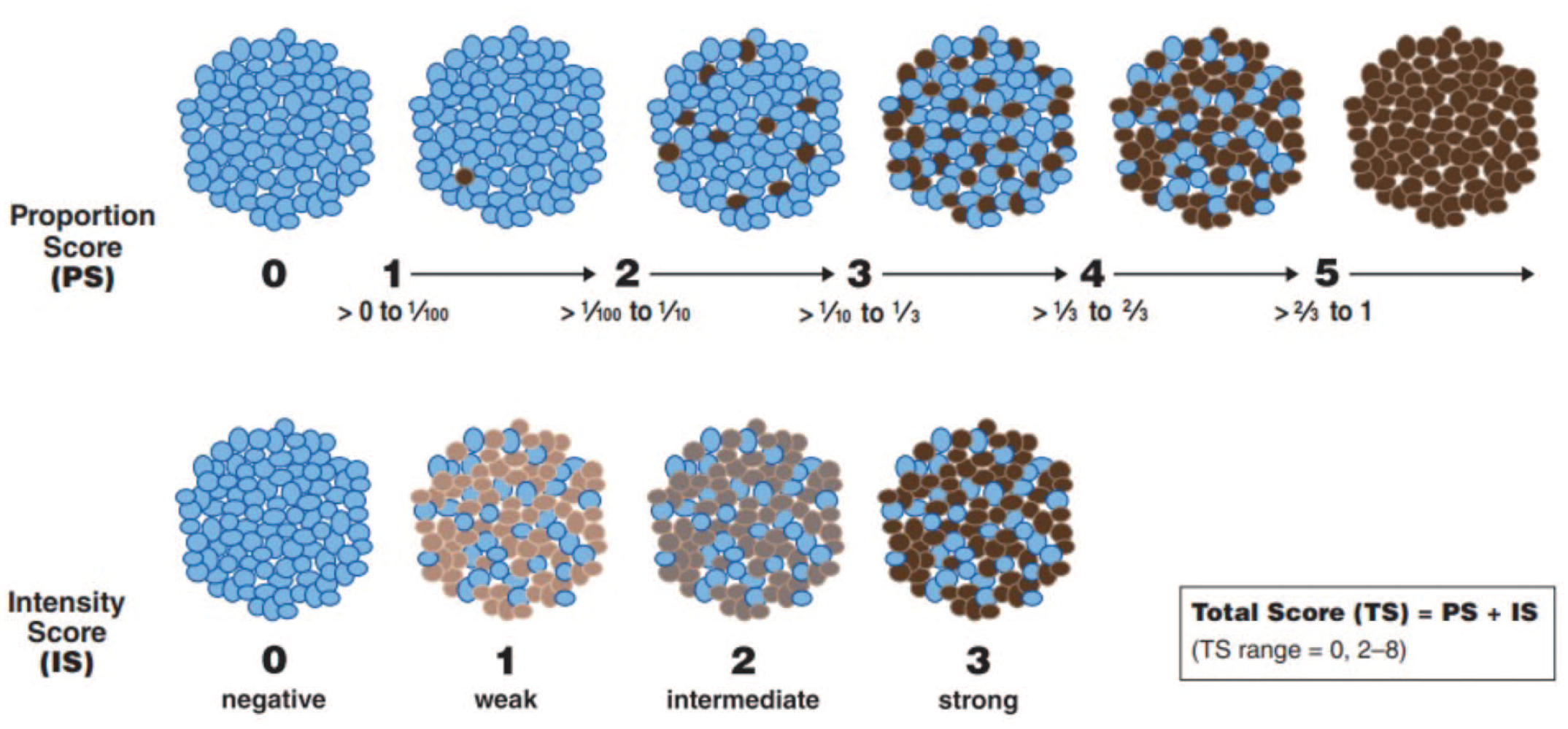
Allred scoring guideline. Based on the percentage of positive cells, the sample is assigned one of six possible proportion scores (PS; 0 to 5). Based on the intensity of most of the positive cancer cells, the sample is also assigned one of four possible intensity scores (IS; 0 to 3). The two scores are then added together for a total score (TS [i.e., the Allred score]) with 8 possible values. Note that a score of 1 is not a possible outcome. Modified from Chand. 22
Pathological assessment of tumor-infiltrating lymphocytes
Solid tumors often contain tumor-infiltrating lymphocytes (or TILs), which have emerged as a biomarker in predicting the efficacy and outcome of treatment. The TILs are divided into intratumoral and stromal populations that are usually quantified separately. Visual assessment requires extensive training by pathologists, and interobserver variability occurs. No single IHC stain highlights all lymphocytes with high sensitivity and specificity, so H&E remains the stain typically used in the routine clinical setting. Computational assessment is beneficial when IHC is involved in order to subtype TILs into cytotoxic, helper, regulatory, and other T cells. 4
Fit-for-purpose scoring
Investigator-defined approaches may be appropriate when researchers establish scoring systems based on the specific question they are asking. As noted above, such methods must be carefully designed and verified (i.e., definable, reproducible, and meaningful). The scoring system should match the range of IHC staining evident in the test tissues to be evaluated, which may entail fine-tuning the definition of each ordinal grade to fit the range of changes observed in the specimens. It has been suggested that such scoring systems should be comprised of four to five scoring levels to maximize detection and repeatability of the IHC assay. 42 Fewer scoring categories reduces the sensitivity of the system while increased categories lessens reproducibility since the distinctions between categories are less clear.
Quantitative analysis
While visual (especially semiquantitative) scoring often serves as a first-line analysis of IHC staining, quantitative measurements performed using an automated digital imaging and analysis platform may provide a more precise answer for some research questions. Automated digital analysis of IHC-stained sections may be used to generate actual counts or ratio variables, where the latter are discrete numbers (e.g., percentages or other measurements) arranged along a scale that contains a true zero value.2,91 Automated analysis may also be used to verify staining reproducibility.25,26 In general, validated quantitative measurements acquired using an automated system are similar in sensitivity and specificity to visual scores by pathologists. However, automated systems are more unbiased and can have a higher workflow compared to visual scoring. The emerging interest in artificial intelligence (AI) based approaches may reveal additional advantages and opportunities for use in tissue scoring in nonclinical settings.9,80,96 One approach to quantitative analysis of chromogenic IHC staining is to count structures (e.g., cell numbers or area dimensions) stained for an antigen of interest. For this purpose, the stain intensity is not relevant as long as the signal is specific.
A second approach is to quantify the amount of antigen present in a given specimen. As described above, extensive work during IHC assay development is required to understand not only the impact of preanalytical variables on the antigen of interest (e.g., time to preservation, time in fixative, impact of different fixatives, antigen stability in precut sections) as well as to hone assay conditions to produce staining intensity in the part of the dynamic range where there is a linear relationship between signal intensity and the amount of target antigen present. Once this has been established, within that range differences in staining intensity can be equated to differences in the quantity of the target antigen. This type of assay development usually goes beyond what is routinely done for research-use-only (RUO) IHC assay development. This approach is only valid when staining runs include multiple control specimens (commonly engineered cells), each of which expresses the target antigen at a known level, so that a broad range of paired expression levels and staining intensities may be used to generate a calibration scale. However, in our experience, quantitative data should be interpreted with care since data from a given IHC assay can rarely be extrapolated between studies (especially retrospective) or across species.
Automated IHC analysis may offer circumstantial advantages to visual scoring. Depending on the scoring paradigm, pathologist-derived visual scores can have good to excellent intra- and inter-observer reproducibility, but percentage estimation of stained areas often has only poor to good reproducibility. 109 For example, automated HER2 (human epidermal growth factor receptor 2) IHC measurements more closely replicate consensus visual scores by multiple expert pathologists and HER2 gene amplification data compared to individual visual scoring. 115 Furthermore, automated algorithms may be retained such that all subsequent images are analyzed using the same parameters. Automated methods are increasingly becoming standard practice in nonclinical settings, and it has been suggested that AI-aided platforms not only recognize different tissue structures but also can measure staining intensity accurately. 74 AI-aided analytical software may further improve predictive value by correctly (and rapidly) identifying cells of interest as well as differentiating among organelles, protein-specific labeling, and nuclear counterstaining while processing data in a high-throughput manner. 8
Considerations in Selecting a Scoring Paradigm
An obvious question to ask when choosing a scoring method is, “What is the assay’s intended use?” The more investigative/novel the question, the more leeway one has in devising the scoring system. Novel scoring systems may be disease- or model-specific and based on established methods, permitting an established (commonly accepted) scoring paradigm to be fine-tuned to satisfy a personalized medicine approach. An example of this latter case is a heterogeneity scoring approach (HetMap) designed to visualize an individual patient’s IHC (antigen) heterogeneity in conjunction with an HER2 H-Score. 102
Bias (i.e., a conscious or unconscious tendency to design or conduct experiments and/or to analyze, interpret, and/or communicate experimental results that favors a particular outcome) is to some degree nearly always present in experimental studies.21,68 Bias is independent of the sample size and statistical significance. Broadly, bias in research can result from visual traps and cognitive traps.1,75,110 Visual traps are a sort of optical illusion in which the perceived image differs from objective reality while cognitive traps represent tendencies to think in ways that lead to systematic errors or deviations from rational thinking. Evaluation of stained material from chromogenic IHC assays is subject to both categories of traps. 1
Sampling bias is the probability that structures of interest will be visible for evaluation and imaging after specimens have been sectioned and stained. Geometric bias (a subset of sampling bias) occurs when planar sampling of topologically complex specimens systematically misses structures of interest. In terms of chromogenic IHC assays, several questions may be asked when assessing whether sampling bias has occurred. Is tissue collection at necropsy random, based on defined anatomic landmarks, or limited to obvious lesions? Does scoring represent IHC staining over a whole section, one defined area in a single section, or an average over multiple smaller regions (e.g., five random microscopic fields) in a single section or across several sections? Ultimately, the goal of tissue sampling for chromogenic IHC is to best represent the true nature or quality of the IHC-staining pattern. Experiments with carefully devised designs and sampling procedures can help avoid sampling bias, which improves the IHC analysis and ultimate results. Automated whole-slide imaging and tissue image analysis can increase the sampled population of IHC-stained elements in a section, which minimizes the chance for systematic biases to affect outcomes. 1
Analysis and Interpretation
The quality of chromogenic IHC data is impacted by several analytical factors, which in turn influences data interpretation (Table 10). Ideally, a pathologist should be consulted at the study design phase for each prospective IHC assay to help plan for tissue collection (including potential internal study controls [discussed below]), fixation, and processing. Because IHC has technical limitations and the hypothesis being tested may require substantial assay development (which can be prolonged and resource-intense), pathology analyses other than or in addition to IHC assays can be discussed simultaneously. Inclusion of satellite samples for orthogonal data generation may be prudent at this stage for IHC assays that are not well-validated or with which the laboratory or pathologist has limited experience with the target antigen.
Primary points to consider for analysis and interpretation.
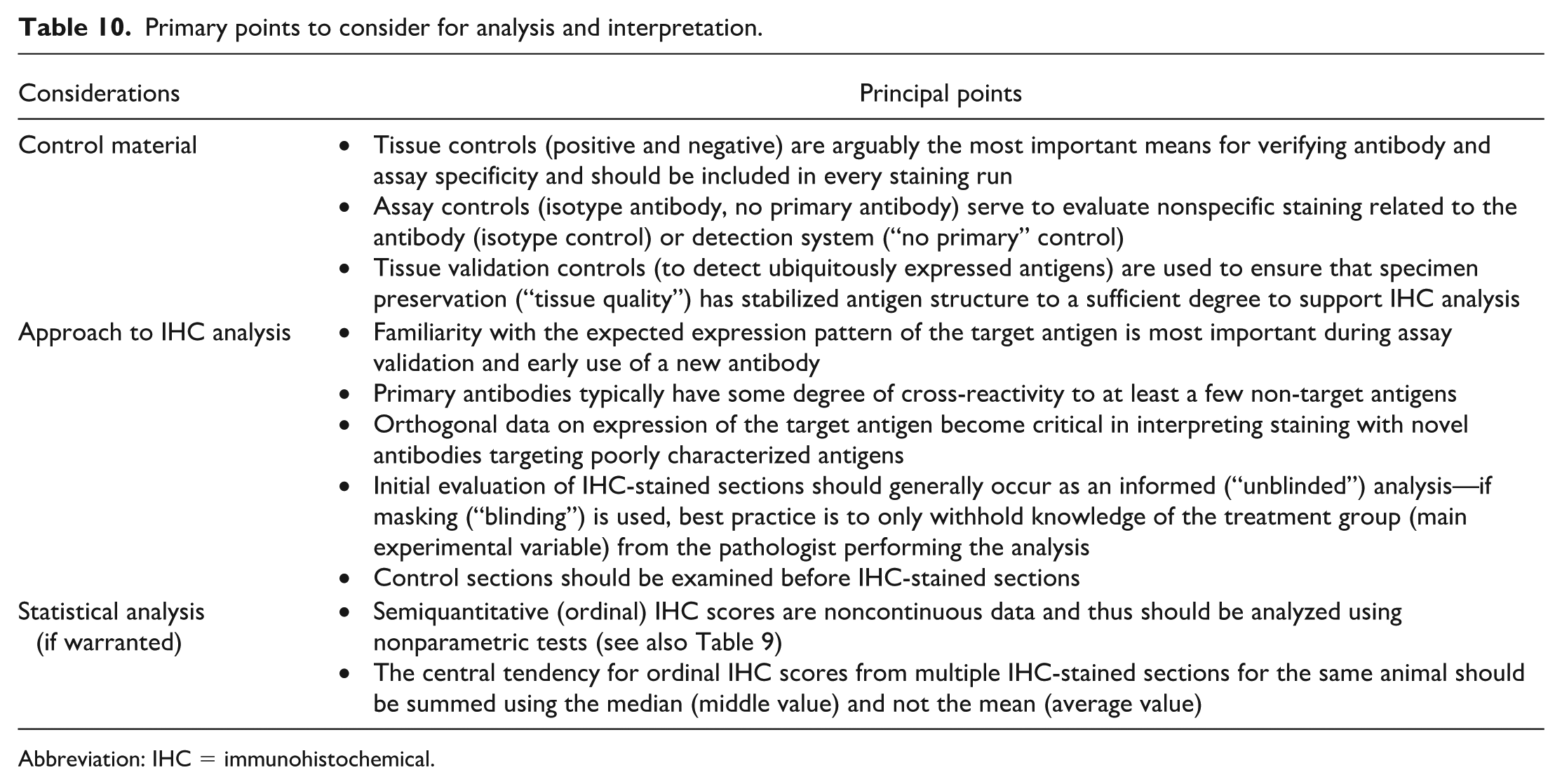

Abbreviation: IHC = immunohistochemical.
Control Materials
As discussed above in the section on “Staining Optimization,” incorporation of appropriate controls is critical for IHC interpretation (Table 11). The numbers and types of controls vary depending on the extent of prior characterization with the target antigen and the IHC assay. Controls include positive and negative tissue controls, assay controls, and IHC validation controls. Availability and review of appropriate controls are paramount to support accurate assessment of staining and data interpretation.
Principal control materials for chromogenic immunohistochemical assays.
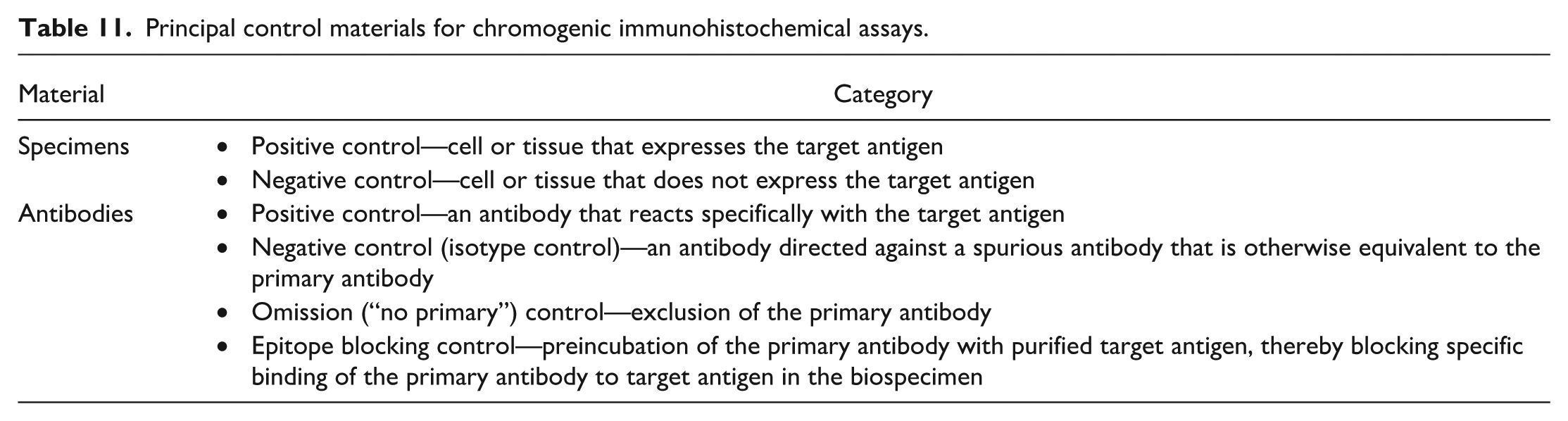

Approach to Chromogenic Immunohistochemical Analysis
Familiarizing oneself with the expected expression pattern of the target antigen detected by the primary antibody facilitates interpretation. This knowledge is more important during assay validation and early use of a new antibody than after its use is established by the laboratory and pathologist. Monoclonal primary antibodies bind to a specific epitope on the target antigen and tend to be more specific (i.e., produce less background staining), while polyclonal primary antibodies are more sensitive as the cocktail of antibodies is likely to bind multiple epitopes of a single molecule or recognize multiple related isoforms.105,107 Multiple bands of differing molecular weights on Western blots suggest that the primary antibody may not be specific to one target antigen unless the bands can be explained as differing isoforms or degradation products. 114 While often limited, the vendor product information sheet for the antibody may contain some of this information. The pathologist should familiarize themselves with available target expression data from prior in-house studies (if any), the published literature, and public databases (e.g., proteinatlas.org) as an aid to IHC interpretation.
Objective of the analysis
When progressing to test tissues, the pathologist should consider the objective of the study, expected tissue morphology, and ancillary factors when recording and interpreting IHC staining. For instance, if the objective is to semiquantify the extent of lymphocyte infiltration into a tumor using an IHC assay for CD3, cell morphology combined with specific staining can reasonably be used to score the extent of the infiltrate while excluding staining interpreted as not associated with lymphocytes. Thus, the pathologist can tolerate some degree of background or nonspecific staining while still meeting the objective of the study. This semiquantitative grading of IHC staining becomes far more problematic when the expressing cell type and distribution are not well known in advance, such as in a target tissue characterization to determine the expression of a given protein across a variety of tissues. Often the target antigen is a relatively novel protein, and little is known about its expression pattern in cells and tissues. Interpreting any specific staining as indicative of target antigen expression assumes that the IHC antibody binds only to its intended target, and this is unlikely to be true because antibodies typically have some degree of promiscuity.28,54 Therefore, one must discriminate “specific on-target” staining (CDR-mediated labeling of the intended IHC target]) from “specific non-target” staining (CDR-mediated labeling of a target other than the intended target [i.e., labeling of a cross-reactive protein]). Similarly, lack of staining may represent target expression below the lower limit of detection of the IHC assay, and that lower limit may be unknown. Orthogonal data on expression of the target antigen become critical in interpreting staining with novel antibodies targeting poorly characterized antigens. 114
Nature of the analysis
Initial evaluations of IHC-stained sections are generally done with knowledge of the study metadata (i.e., an informed or “unblinded” examination) consistent with suggested practices when evaluating nonclinical toxicity studies.13,31,58,87,97,103,112 This approach is particularly true for target tissue characterization where there is no group comparison but rather an evaluation of staining pattern to facilitate understanding of target expression; in the authors’ opinion, orthogonal data are far more important for IHC assay interpretation in this situation compared to performing a masked (i.e., “blinded”) review. For studies with group comparisons, an initial unblinded review to determine the basic antigen expression pattern and establish grading criteria may be followed (when warranted) by a blinded analysis to see if antigen expression is correlated with the group-specific treatments. If the animal model is well characterized, the IHC assay well-validated, and the pathologist experienced in evaluating IHC stains, the initial evaluation may use a blinded approach if the study was designed to test a focused hypothesis. When conducting a blinded evaluation of IHC-stained material, it can be completely blinded (“blind to all”), where each sample is individually evaluated independent of all other samples, or group-blinded (“blind to treatment”), where the pathologist knows that samples are in a common group but knows no details of the specific treatment for that group.13,87 The “blind to all” approach seems beneficial on the surface, but it significantly hinders the observer from discriminating subtle but known from subtle unexpected group-specific changes or identifying nonspecific background lesions. The advantage of group-blinding, which is the recommended approach for nonclinical studies in which a blinded evaluation is performed, 13 is that it more readily allows the pathologist to detect antigen expression patterns that vary among groups. A third option, the “post hoc blind to treatment” approach, is performed as a two-stage process: an initial informed analysis of all or selected specimens with full knowledge of the subject’s metadata (e.g., treatment, dose, and ancillary clinical and pathology data) so that scoring criteria may be established, followed by masked re-evaluation of all specimens for all treatment groups (including control groups) in random order to establish scores for each specimen.13,31,87,112 This third approach works well for most nonclinical studies and for evaluating new experimental models. However, if the number of specimens is small, the observer may collate discrete morphological features with the specific group assignment, thereby making masking ineffective.
Order of analysis
When evaluating an IHC assay, the control sections should be examined first. The staining on positive and negative control tissues should appear similar to previous studies or IHC runs including the stain distribution, labeled cell type(s), subcellular localization, and staining intensity. The negative control tissue should appear “clean” (no apparent specific staining) or have a low level of background staining. As previously stated, assay controls (if included) are typically clean. If the expected staining pattern is not present and/or the background is higher than expected in any of the controls, potential issues with the IHC assay should be explored and corrected prior to evaluating test tissues. A good guide to troubleshooting IHC staining may be found elsewhere. 106
Validation of the analysis
Erroneous interpretation of IHC results is thought to substantially contribute to the lack of reproducibility for IHC assays. 7 Therefore, after evaluating the IHC-stained sections, particularly during a study for target tissue characterization of a relatively novel IHC target, the pathologist should review the data in the context of other internal or publicly available data on expression of the target antigen. Do the IHC results align across studies? If not, is it possible that a portion of the staining for the current IHC assay was not specific for the intended target antigen? Generation of confirmatory orthogonal data should be considered prior to publishing or making a program-driving decision. The presence or absence of an antigen in a tissue should be based on the weight-of-evidence rather than simply the presence of positive IHC staining detected using one antibody.
Statistical considerations in immunohistochemical analysis
Several considerations can help guard against pitfalls and errors in statistical analysis of IHC data. Statistical tests each have specific assumptions that must be met to provide confidence in the statistical results and conclusions. The differing assumptions among tests for various data types are why scientists should not evaluate data using multiple statistical tests (“shopping for significance”) merely to find one that produces
Reporting of Immunohistochemical Assays and Data
As with scientific writing generally, descriptions of IHC in publications and study reports should follow the communication ABC’s: Accuracy, Brevity, and Clarity. The “Animal Research: Reporting In Vivo Experiments” (ARRIVE) guidelines provide generic requirements for designing high-quality experiments but provide little coverage for pathology endpoints. 71 More complete recommendations for describing pathology endpoints are available in the “Minimum Information for Publication of Experimental Pathology data” (MINPEPA) guidelines. 113 Comparable recommendations for collecting assay details have been recently released by experienced histologists. 24
For IHC, papers and reports should include sufficient detail that readers both achieve understanding and have a reasonable likelihood of being able to reproduce the work. 64 The same basic elements should be covered in both papers and reports: materials and methods, results, and discussion. This information (especially materials and methods) may be abbreviated in papers due to word limits applied by publishers, but details in reports should be comprehensive. Table 12 shows a list of key information to cover, identifying elements that are required for completeness while also including additional points that facilitate experimental replication. This information may be given in the main body of a paper or report, but inclusion as an appendix or supplemental file also is acceptable as long as the details are available for evaluation.
Primary points to consider for reporting of methodological details and data.
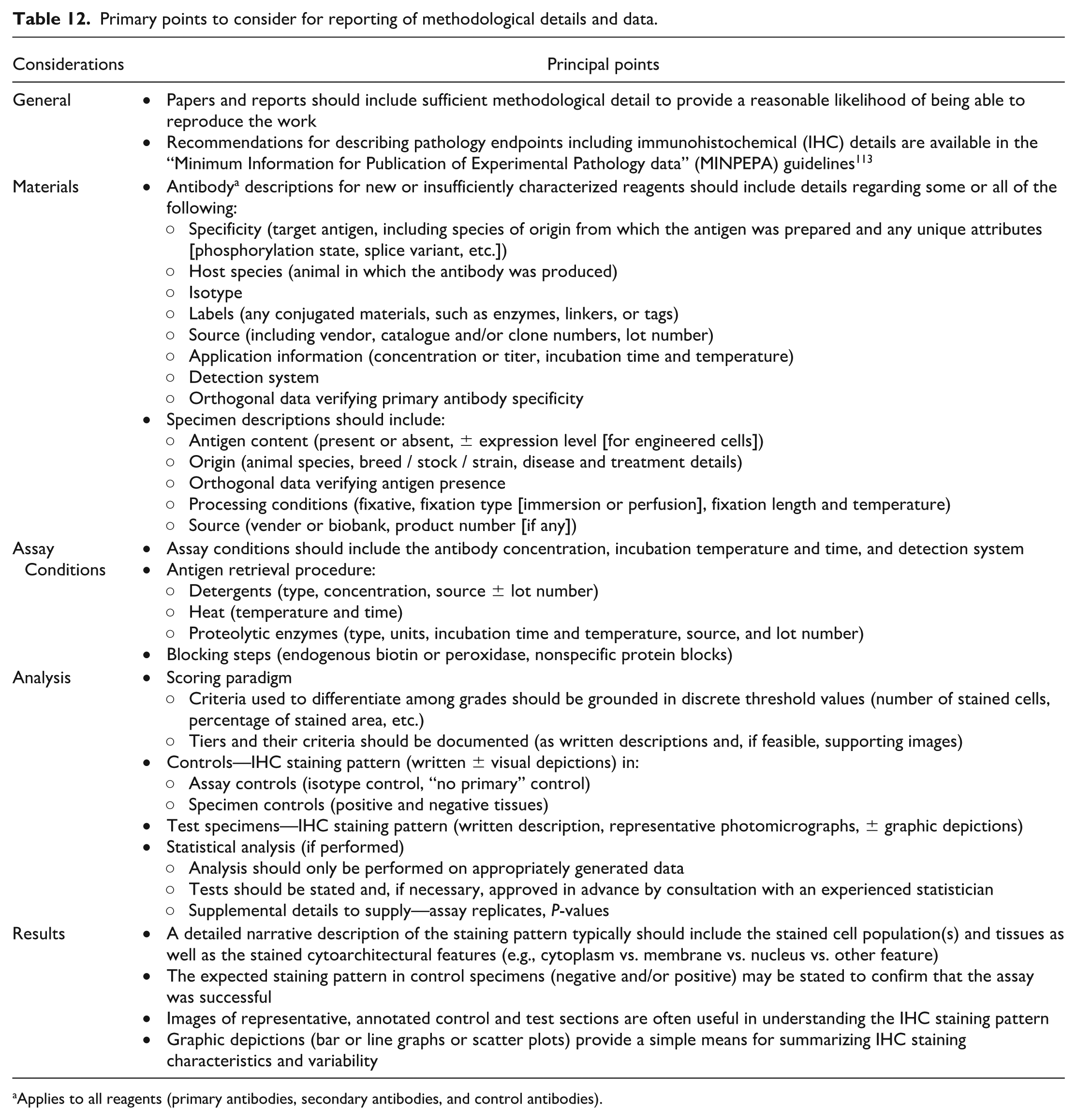
Applies to all reagents (primary antibodies, secondary antibodies, and control antibodies).
A detailed description of antibodies used in IHC studies should be given in the Materials and Methods section.25,26 This information may be presented as text, tables, or both in papers and reports. In general, information for antibodies should include the host species (in which the antibody was produced), product type, target antigen (protein and species ± any special attributes [e.g., phosphorylation state or splice variant] if known), isotype and structure, any conjugated materials, product number, and vendor. The application conditions for antibodies should include the antibody concentration, incubation length and temperature, and detection system. For example, a detailed text description for a primary antibody directed against the macrophage marker F4/80 might read: Rat polyclonal anti-mouse F4/80 IgG2b (Clone No. C1:A3-1; Bio-Rad, Hercules, CA) was applied at a concentration of 3.33 µg/L for 1 hr at room temperature (RT) followed by rabbit secondary anti-rat IgG antibody (1.66 µg/mL for 30 min at RT; Cat No. ab102248, Abcam, Boston, MA).
Fewer details often are sufficient for well-known antibodies with very reproducible staining patterns; a suitable description for the mouse macrophage marker F4/80 might be limited to “Anti-mouse F4/80 (Clone No. C1:A3-1, Abcam) was applied at a final concentration of 3.33 µg/mL for 60 min at RT.” In general, reporting antibody dilutions (e.g., “applied at a titer of 1:200”) in the absence of definitive concentrations is discouraged since dilutions will vary among personnel and autostainers. Similar parameters for any other IHC steps such as blocking, epitope retrieval, and autostainer protocols should be supplied in some detail, especially for multiplex assays where the extended assay length and many washing steps may impact the staining intensity. Reagent lot numbers typically should be included in reports.
The substrates (cells or tissues) to which the antigen is being applied should receive an equally thorough description in the Materials and Methods section. Key information should include the animal species, breed/stock/strain (as warranted), any treatment (including engineered expression of an antigen), the product number, the specimen source (vendor or biobank), and when possible the sample strategy. Details regarding specimen preservation (e.g., fixed or frozen, fixative, and especially fixation length and temperature) are essential in confirming that unexpected IHC results do not arise from suboptimal fixation or storage practices.
Scoring criteria are a key component of the Methods section. References (usually peer-reviewed publications) or institutional standard operating procedures (SOPs) may be sufficient for well-known, reproducible scoring schemes, but in many cases the scoring methodology is devised for a specific IHC study. Features used to discriminate among grades should be given in some detail so that other investigators including peer review pathologists can have a meaningful discussion regarding IHC findings. In general, discrete threshold values (number of stained cells, percentage of stained area, etc.) are preferred to anchor various grades in a tiered grading scheme. The scoring approach (informed [“unblinded”] vs. masked [“blinded”]), scoring process (visual vs. automated [including the software brand and version]), and any quality control steps (e.g., instrument analytics, peer review) should be stated.
Statistics should be noted in the Methods section (if warranted by the study design). For many IHC questions, statistics are not needed to attain a confident interpretation of the staining data; in such cases, statistics need not be performed. Statistics for IHC studies often are reserved for hypothesis testing to facilitate preparation of a peer-reviewed publication. The correct statistical test should be chosen to accord with the type of data supplied by the study. Nonparametric tests are employed for noncontinuous data with non-normal distribution (e.g., semiquantitative visual scoring, provided such data are collected in a consistent and unbiased fashion, adhering to clear, objective, and reproducible scoring criteria), while parametric tests are appropriate to continuous data (e.g., digital image analysis data sets) with confirmed normal distribution. Consultation with an experienced biostatistician prior to performing the IHC assay may help ensure that the study design is capable of addressing the study objective.
The Results section of papers and reports should provide a detailed narrative description of the staining pattern. Specific details to communicate include the stained cell population(s) and anatomic features as well as the stained cytoarchitectural constituents (e.g., cytoplasm vs. membrane vs. nucleus vs. other feature). The expected staining pattern in control specimens (negative and/or positive) should be stated for comparison as a means of affirming that the IHC assay was successful. A simple narrative description of the IHC findings is often suitable if the distribution (staining pattern) for a given antigen is well known and is comparable between the control and test sections. Presentation of visual data often is a substantial aid in achieving accuracy, brevity, and clarity in presenting IHC data if the staining pattern for an antigen is new, unexpected, or varies substantially between control and test specimens. Images of representative, annotated control and test sections are particularly useful in demonstrating stained structures. Graphic depictions such as bar or line graphs or scatter plots may demonstrate the degree to which IHC staining intensity varies across different variables. 26 Another layer of descriptive data is that of central tendencies for each group. 66 Mean (average value), median (value of the midpoint), and mode (most common value) provide insight regarding the organization and skewing of group data. In general, legends for figures depicting IHC-stained material may be limited to the assay (e.g., anti-Ki67 IHC) and details describing key elements and any structural details visible in the figure that inform the interpretations made from the IHC data.
Regulatory Considerations
Although IHC endpoints are essential elements in GLP-compliant tissue cross-reactivity studies, 79 other IHC-based assessments in GLP-compliant nonclinical toxicity studies are typically defined as non-GLP endpoints in the study protocol (Table 13). The rationale for electing to perform non-routine IHC assays under GLP exceptions is based on the difficulty in performing GLP validations of complex IHC assays to address one-time scientific questions. This approach of including highly specialized non-routine IHC assays as non-GLP endpoints is generally acceptable to health authorities if the rest of the study is GLP-compliant, the assay is of high quality and transparently disclosed as being non-GLP, and documentation describing the assay materials and methods as well as the data and their interpretation is detailed in the study record. This approach to address the inclusion of complex nonroutine assays in GLP studies is described in the ICH S12 guidance on biodistribution studies. 63
Primary points to consider for regulatory documentation.
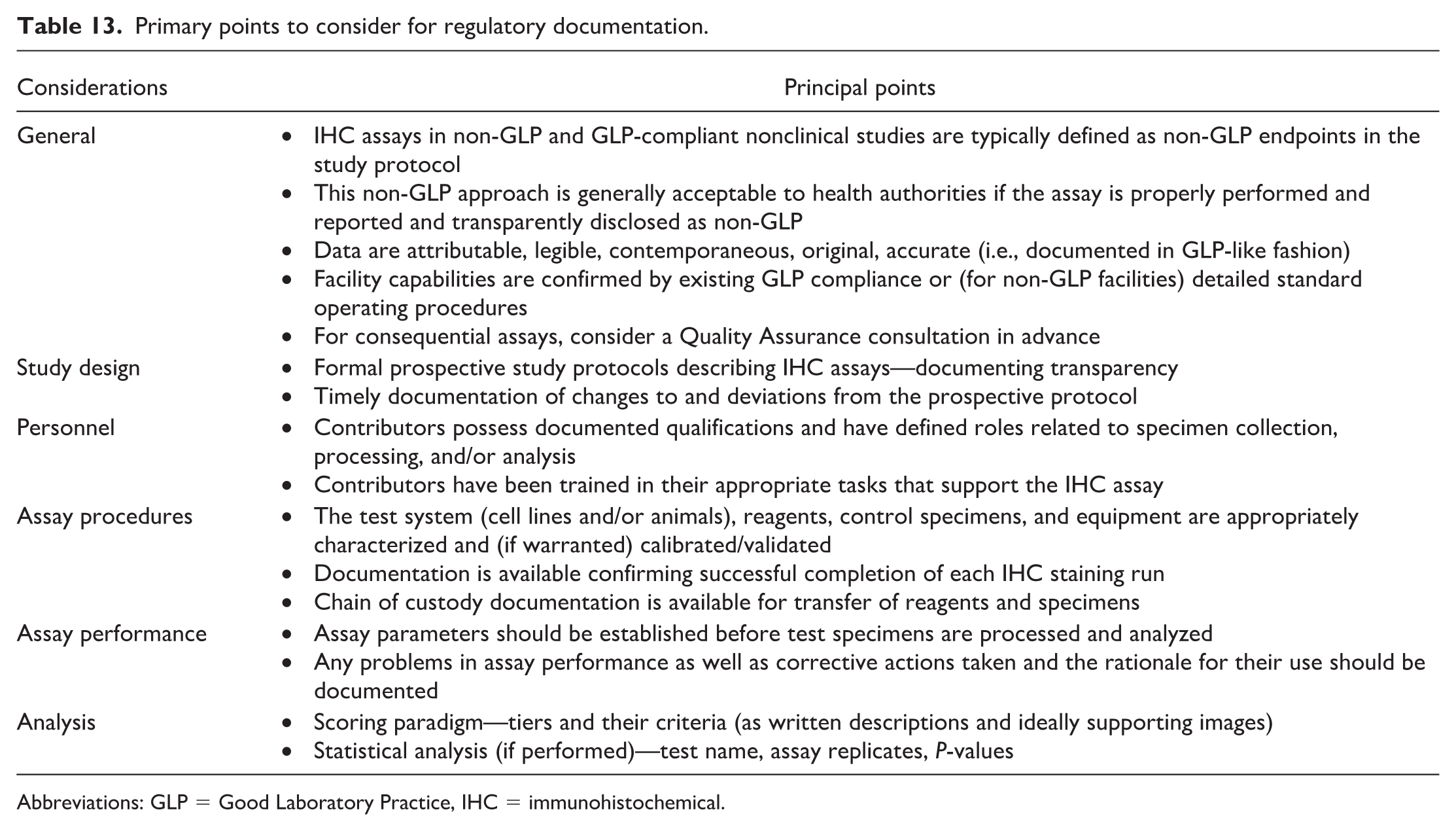

Abbreviations: GLP = Good Laboratory Practice, IHC = immunohistochemical.
When considering which quality attributes to include in a nonroutine IHC-based assay intended for inclusion in a GLP-compliant study, general GLP principles should apply as no guidance has been offered by regulatory agencies (FDA, PMDA [Pharmaceuticals and Medical Devices Agency]) or international consortia (ICH [International Council for Harmonisation of Technical Requirements for Pharmaceuticals for Human Use], OECD [Organisation for Economic Co-operation and Development]) specific to the conduct of ICH-based laboratory-developed tests based on RUO reagents and systems. Moreover, the quality compliance effort undertaken during the IHC assay and included in the study report should reflect the importance of the IHC endpoint in the study interpretation and the public health implications of the outcome. Study endpoints conducted in the spirit of GLP (i.e., intended to align with GLP principles despite the lack of full GLP documentation) should include adequately supported methods (as described in preceding sections), documentation, and record retention, as summarized in Table 13.
In general, investigators conducting a non-GLP IHC assay should endeavor to demonstrate the scientific validity of the procedure by making clear to an outside observer that all IHC study-related activities were prospectively planned (e.g., outlined in a protocol amendment) and adequately controlled, thereby ensuring that the resulting data are accurate and credible. Parameters critical to the assay performance (antibody concentrations, incubation times and temperatures, control materials, analysis cut points, masking procedures for microscopic evaluation, statistical analyses, etc.) should be established before the work is undertaken on test specimens, which may involve the conduct of pilot experiments. When problems are encountered, any corrective actions taken and the rationale for their use should be well-documented. For IHC assays that generate semiquantitative or quantitative data, the methods employed for data acquisition and statistical analysis are critical to the quality of the interpretation. In these scenarios, it is often prudent to involve a professional statistician in planning the experiment. For more consequential studies, details of the statistical analysis may be provided in a separate subreport.
If the laboratory that developed the IHC assay is inexperienced at generating data for use in regulatory submissions, investigators should consider technology transfer to an experienced GLP-compliant test facility for performance of the definitive IHC assay. Some aspects of GLP work in chromogenic IHC are not immediately obvious and often are overlooked by academic and start-up industrial laboratories. Furthermore, setting up and maintaining an adequately GLP-compliant laboratory environment is expensive with obligations that can last for many years.
Conclusion
Chromogenic IHC plays a pivotal role as a molecular localization tool in the realm of biomedical research and nonclinical drug development. The amalgamation of pertinent literature with the collective expertise of the seasoned Working Group members resulted in a comprehensive exploration of factors influencing data quality, with particular emphasis on sample selection, general assay considerations, and nuanced aspects of data generation and interpretation. The resulting “Points to Consider” article aims to provide specific and practical topics for thought and application tailored for pathologists, histologists, and allied scientists using light microscopic chromogenic IHC methodology within the context of nonclinical drug development. In understanding the fluidity of this topic with respect to technological advances (e.g., AI), the authors predict this article will lead to improved robustness and reproducibility of chromogenic IHC studies, facilitating informed decision-making in the nonclinical development of biomedical products.
Footnotes
Authors’ Notes
Chromogenic IHC assays label molecules by using antibodies to place a colored agent at sites of antigen localization in cells and tissues. IHC labeling is often referred to as “IHC staining” in scientific parlance even though IHC “labeling” is technically distinct from “staining” (where histochemical dyes [“stains”] directly bind cells and tissues while IHC labels develop at sites where antibody-bound enzymes convert noncolored chromogens to deposited colored products).
The 10 IHC Working Group members have had pathology careers involving chromogenic IHC assay development and/or usage ranging in length from 17 to 33 years in such practice settings as academia, biopharmaceutical firms, contract research organizations, government research laboratories, and private consulting.
Author Contributions
Authors contributed to conceptualization: FA, BB, MB, BSB, TF, KJ, KM, DKM, SP, MCR; writing—original draft: FA, BB, MB, BSB, TF, KJ, KM, DKM, SP, MCR; visualization: FA, MCR; writing—review and editing: FA, BB, MB, BSB, TF, KJ, KM, DKM, SP, MCR; project administration: FA; and supervision: FA, BB, KJ.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article. The author BB is a senior advisor to the Editor of
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.