Abstract
Mass Spectrometry Imaging (MSI) is a powerful tool to understand molecular pathophysiology and therapeutic and toxicity mechanisms, as well as for patient stratification and precision medicine. MSI, a label-free technique offering detailed spatial information on a large number of molecules in different tissues, encompasses various techniques including Matrix-Assisted Laser Desorption Ionization (MALDI), Desorption Electrospray Ionization (DESI), and Secondary Ion Mass Spectrometry (SIMS) that can be applied in diagnostic and toxicologic pathology. Given the utmost importance of high-quality samples, pathologists play a pivotal role in providing comprehensive pathobiology and histopathology knowledge, as well as information on tissue sampling, orientation, morphology, endogenous biomarkers, and pathogenesis, which are crucial for the correct interpretation of targeted experiments. This article introduces MSI and its fundamentals, and reports on case examples, determining the best suited technology to address research questions. High-level principles and characteristics of the most used modalities for spatial metabolomics, lipidomics and proteomics, sensitivity and specific requirements for sample procurement and preparation are discussed. MSI applications for projects focused on drug metabolism, nonclinical safety assessment, and pharmacokinetics/pharmacodynamics and various diagnostic pathology cases from nonclinical and clinical settings are showcased.
This “Points to Consider” article is a product of a European Society of Toxicologic Pathology (ESTP) Pathology 2.0 Working Group. It has been reviewed and approved by the Committee of Regulatory and Scientific Standards (CRSS) and Executive Committee of the ESTP and endorsed by the Executive Committees of the British Society of Toxicological Pathology (BSTP) and Society of Toxicologic Pathology (STP), but it does not represent a formal best practice recommendation of the Societies; rather, it is intended to provide key “points to consider” for the toxicologic pathology community. The opinions expressed in this document are those of the authors and do not reflect views or policies of the employing institutions. Readers of Toxicologic Pathology are encouraged to send their thoughts on these articles or ideas for new topics to the editor.
Introduction
Drug metabolism and pharmacokinetics (DMPK), including Absorption, Distribution, Metabolism, and Excretion (ADME), are essential in optimizing drug efficacy and safety during drug development. ADME studies guide regulatory approval and the selection of candidates for late-stage trials by predicting pharmacokinetics and potential toxicity. Inadequate or excessive accumulation in tissues can affect the therapeutic effect and lead to toxicity, underscoring the importance of precise distribution assessments for risk evaluation.
Traditionally, knowledge of tissue localization has relied on techniques like whole-body autoradiography and homogenate liquid chromatography-mass spectrometry (LC-MS) analysis. 37 However, in recent years, mass spectrometry imaging (MSI) has emerged as a powerful tool for in situ chemical detection and tissue distribution. 8 This technique enables researchers to visualize the spatial distribution of compounds and their metabolites within biological tissues, providing detailed maps that can reveal how drugs are distributed, metabolized, and disposed of at the cellular level, and the potential link with efficacy or toxicity/ histopathology. In addition, MSI can detect biomarkers in diseased tissues and/or compound/metabolite-related histological findings in toxicity studies. Furthermore, MSI can simultaneously localize and quantify thousands of pharmacodynamic markers in the tissue microenvironment simultaneously with the drug. This can help identify new therapeutic targets, assess surrogate markers of drug efficacy, and predict the potential benefits of candidate compounds. Therefore, investigating tissue-based biomarkers is a crucial step in preclinical and clinical drug development.
The three most important and impactful MSI techniques are Matrix-Assisted Laser Desorption Ionization (MALDI), Desorption ElectroSpray Ionization (DESI), and Secondary Ion Mass Spectrometry (SIMS), all suitable for application in diagnostics and toxicologic pathology.35,45
These methods enable the label-free in situ detection of both endogenous and exogenous molecules across a broad spectrum of molecular weight and polarity range. The sensitivity of MSI techniques enables the reliable detection of molecules within the physiological concentration range and the resolution is steadily approaching the cellular dimensions.
In targeted MSI, the experiment focuses on detecting and quantifying specific molecules that are preselected based on prior knowledge: these molecules could be known biomarkers, metabolites, drugs, or proteins that are relevant to the biological or clinical question being studied. In untargeted MSI, the goal is to perform a comprehensive analysis without prior knowledge of which specific molecules to focus on: the experiment aims to detect a wide range of molecular species present in the sample, allowing for the discovery of unknown or unexpected molecules. In this type of experiments, the spectra that can be covered are limited.
MSI requires not only experts in the field but also other scientists, in particular pathologists. Pathologists have been and are important contributors due to their deep knowledge in tissue morphology and their involvement in tissue sampling and processing, as well as their ability to draw correlations between molecular features and microscopic changes observed in the tissue. This article includes high-level discussions about the principal characteristics of the most commonly used modalities for spatial metabolomics, lipidomics and proteomics and also includes how data generated by MSI can be implemented in the pathologist’s workflow.
Some of the most impactful demonstrations of MSI technologies within discovery-based biomarker research, precision medicine efforts, as well as applications within drug metabolism and pharmacokinetics/pharmacodynamics and toxicologic pathology are highlighted.
Since MSI is an evolving field, this review provides the current situation without claiming completeness.
Factors concerning the use of mass spectrometry imaging to address biologic questions in diagnostic and toxicologic pathology for label-free detection of molecules are outlined in Figure 1.
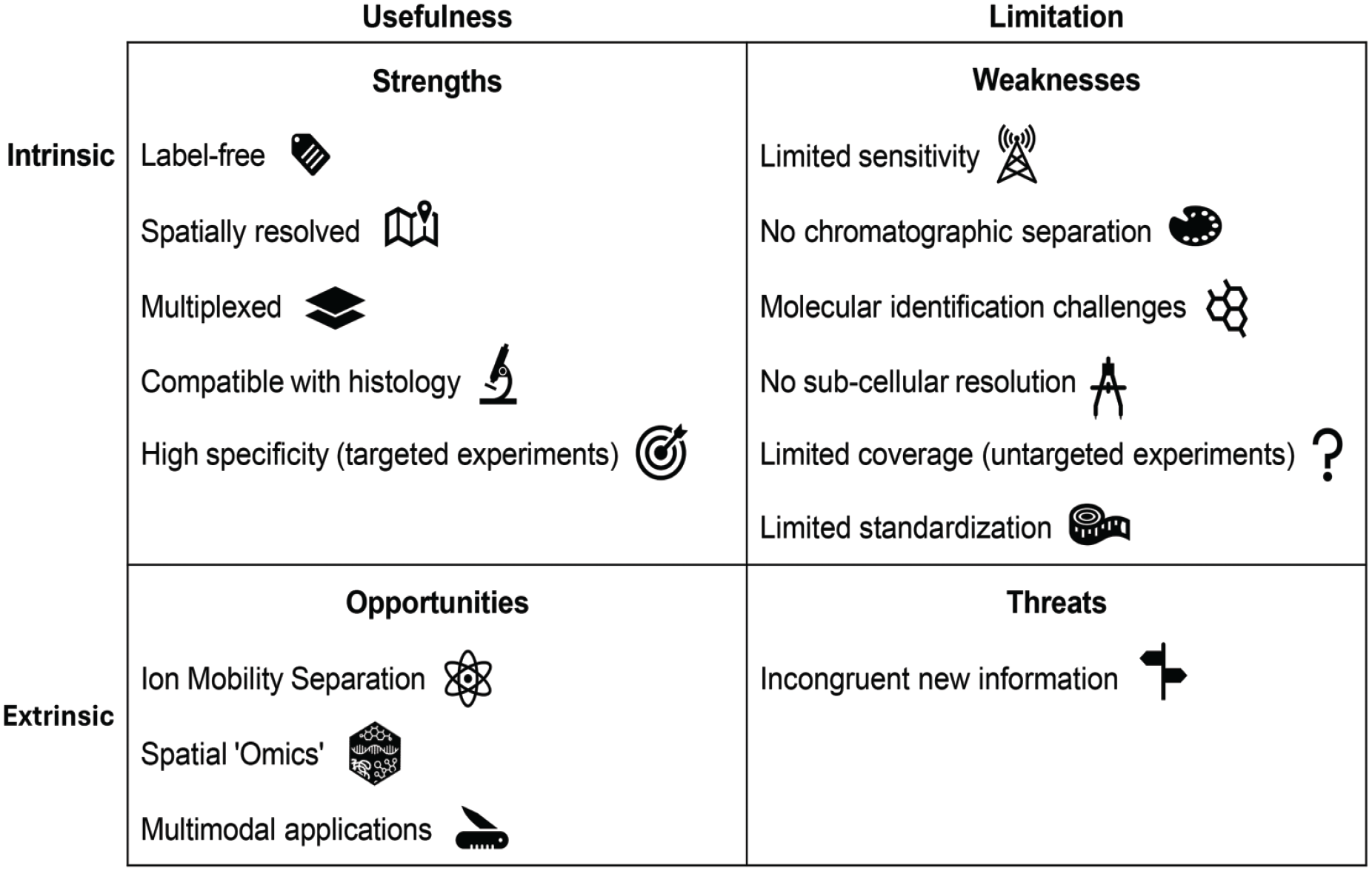
Factors concerning the use of mass spectrometry imaging to address biologic questions in diagnostic and toxicologic pathology for label-free detection of molecules. The strengths of mass spectrometry imaging (MSI) include: label-free detection (ability to detect molecules without external labels, tags, or dyes, commonly used in other imaging techniques like immunohistochemistry), spatial resolution (ability to simultaneously detect molecules while preserving their locations within the tissue sample), multiplexing (ability to simultaneously detect multiple molecules in a single experiment), compatibility with standard and molecular pathology techniques (such as immunohistochemistry) to characterize spatial resolution and high specificity (ability to detect molecules with a defined molecular weight, particularly in targeted analyses of preselected molecules based on prior knowledge). The weaknesses of MSI include: limited sensitivity (limited ability to detect low-abundance molecules within a sample), lack of chromatographic separation (inability to effectively separate complex mixtures of molecules, as overlapping spectral information from multiple analytes may occur within the same pixel of the tissue being analyzed), molecular identification challenges (difficulties in characterizing molecules due to technical limitations of direct mass spectrometry), no subcellular resolution (inability to visualize structures at the subcellular level, such as organelles within a cell), limited coverage in untargeted experiments (the spectrum that can be analyzed is limited in case of comprehensive analysis without focus on specific molecules) and limited standardization (lack of consistency across laboratories, leading to variability in results and making comparisons challenging). The fast-growing new techniques (advances in ion mobility separations, Spatial Omics, multimodal applications) presents many opportunities for new applications of MSI. However, a major threat remains the challenge of interpreting incongruent new information, where the data may not align with existing knowledge or expectations.
Instrumentation and Technical Information
MS, the core of MSI, is an analytical technique used to measure the mass-to-charge ratio (m/z) of ions, referring to the ratio of the mass (m) of an ion to its electrical charge (z): this ratio allows mass spectrometers to separate ions based on their mass and charge, enabling the identification of different molecular species.
As mentioned above, MALDI, DESI and SIMS are the major techniques used nowadays in MSI. All can generate detailed molecular information concerning morphologic changes in tissues, with the added value of spatial resolution. For more detailed information, including technical advancements and applications, the reader can refer to dedicated technical reviews, such as those by Maciel et al. 35 and Schulz et al. 45
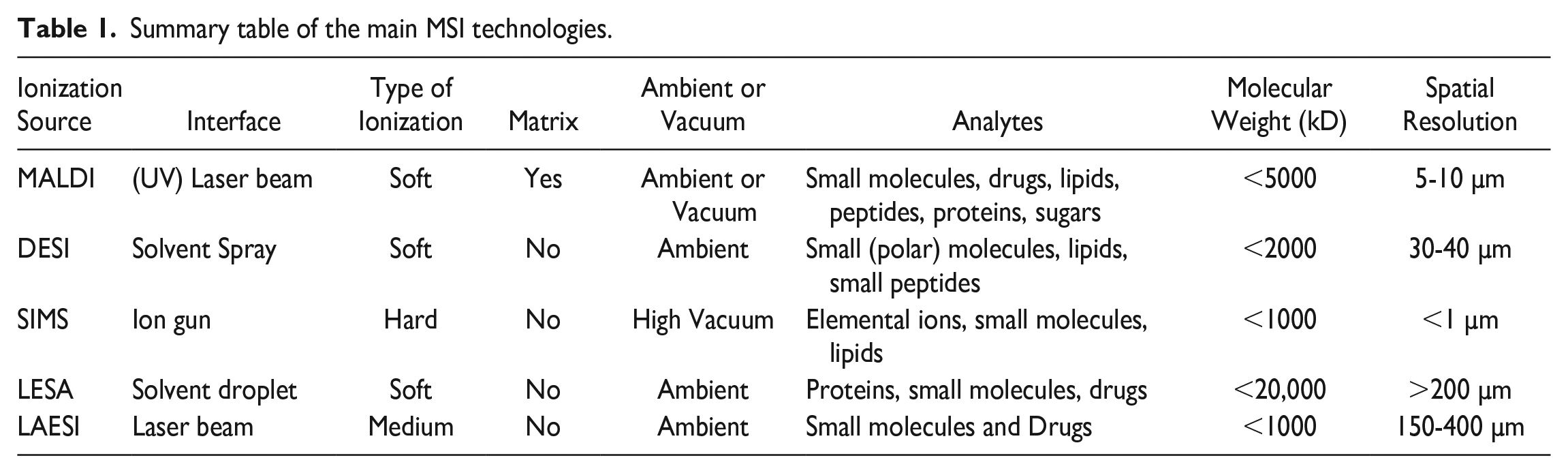
The main ionization techniques are discussed across key parameters (spatial resolution, sensitivity, sample preparation requirements, and typical applications) and summarized in Table 1, according to the authors’ experience.
Summary table of the main MSI technologies.
MALDI
MALDI is the most common MSI technique used not only in pathology, but also in pharmacology, microbiology, plant, food, forensic and materials sciences. It can detect molecules in frozen or formalin-fixed and paraffin-embedded (FFPE) tissue sections, but sensitivity is optimal in fresh frozen tissue. This method currently offers a maximal spatial resolution of 5 to 10 μm, although these limits are being challenged by adapting hardware to reach 1-µm resolution level. As a trade-off, the high spatial resolution comes with lower sensitivity, which is already apparent at 5 to 10 μm but is especially true for adapted hardware reaching 1-µm resolution level.
The basic principle of MALDI is exposing thin (10-20 µm) tissue slices, typically coated with an ultraviolet (UV)-absorbing matrix, to a UV-laser beam, leading to formation of charged species such as protonated or deprotonated molecules and some labile fragments, which are subsequently sampled into the mass spectrometer due to their charge. Although the MALDI technique can effectively ionize small molecules, lipids, peptides, and some proteins, the yield of this ionization process is limited (<20%). This is why postionization laser ionization, using a secondary laser (named MALDI-2) is being used in several laboratories, to enhance the signal intensities of a variety of molecules.
To be noted, MALDI-MSI in negative ion mode (that is the ionization and detection of negatively charged ions, ionized from the analyte) is much preferred for the analysis of compounds that ionize more efficiently as anions, such as acidic compounds, several phospholipids, and other molecules that can easily gain an electron or lose a proton.
DESI
DESI is the second most popular technique for MSI. It consists in principle of an electrospray plume coming from a regular electrospray ionization (ESI) head, which is directed onto the uncoated tissue slice, where it interacts with the tissue, resulting in the desorption of molecules and ions from the surface. These ions are then sampled by the MS through a collection capillary positioned just above the tissue surface. The biggest advantages are the operation of DESI in atmospheric conditions, the wide range of solvents which can to be used to enhances selectivity of this method, and the absence of a matrix. Hence, desorption is directly from the tissue, resulting mostly in an improved sensitivity when compared with MALDI. The downside of this technique is that the spatial resolution is limited to approx. 40 µm.
SIMS
The choice between SIMS and MALDI depends on the specific requirements of the analysis, such as the desired spatial resolution, the type of molecules being studied, the nature of the sample, and the desired speed of analysis. SIMS is particularly effective in the detection of small molecules, inorganic compounds, and elements, as it offers superior spatial resolution (sub-micron level), it is, however, highly surface-sensitive (useful for analyzing the outer layers of a sample) and it can provide detailed elemental and isotopic composition. In addition, it does not require a matrix for ionization, avoiding potential matrix-related artifacts. SIMS involves the bombardment of a sample with a high-energy primary ion beam, which interacts with the top layers of atoms and molecules at the surface and hence, causes the emission of ions, termed secondary ions (SI) -for distinguishing from the primary ions, and neutral particles from the surface of the sample. Traditionally SIMS was and still is, employed as a surface analysis technique and is limited to the analysis of elements like carbon, sulfur, nitrogen, oxygen, and fluorine, among others. However, modern advancements have enabled the visualization of lipids and small molecules in tissue and even single cells. The technique’s ease of use is not comparable with MALDI and or DESI. Acquisition times for only a 5 × 5 μm image easily takes up to 15 to 20 hours.
Others (LESA, LA-ICP, . . .)
Liquid Extraction Surface Analysis (LESA) is an ambient mass spectrometry technique that allows direct examination of analytes from solid substrates. This method is especially effective for analyzing intact proteins. 23 Single chemical elements are a suboptimal target for MALDI, which could be complemented with laser ablation inductively coupled plasma (LA-ICP) imaging for the detection of endogenous elements (including gadolinium, platinum, lithium, sodium, potassium, magnesium, calcium, iron, cobalt, nickel, copper, zinc, and silver). These techniques, albeit unique in their possibilities, are very uncommonly used in the diagnostic and experimental pathology fields.
Among the many available technical possibilities using MALDI apparatus, two recent applications, viz. imaging mass cytometry (IMC) 20 and Immunohistochemistry (IHC)-MALDI 50 deserve to be mentioned. Both detect (membrane) proteins using an antibody tag of “rare earth”-metals or synthetic peptides, respectively. After antibody incubation the tag is released either by feeding the laser ablated material into an LA-ICP MS, in the case of IMC, or by UV radiation, as in IHC-MALDI. Either the metal ions or the synthetic peptides are detected by an LA-ICP MS or by MALDI-MS. Both of these techniques substantially widen the scope of applications for MSI, as they can identify biological therapeutics (including antibody-drug conjugates, antibody fragments, bispecific antibodies, and monoclonal antibodies), with spatial resolution reaching subcellular level. 4 For both MALDI and IMC, sample preparation involves several steps: a cryostat for frozen sections or a microtome for paraffin sections and employing a histological automatic processor and stainer. The readout is based on absolute quantification (see paragraph on quantification in the following section about data management).
Experimental Considerations
MSI has been extensively reviewed by textbooks 11 and comprehensive journal articles,35,45 therefore here follow those points to be considered when approaching this technology from the pathologist’s perspective, without aiming to cover the whole spectrum of variables and conditions. The MSI workflow is outlined in Figure 2.
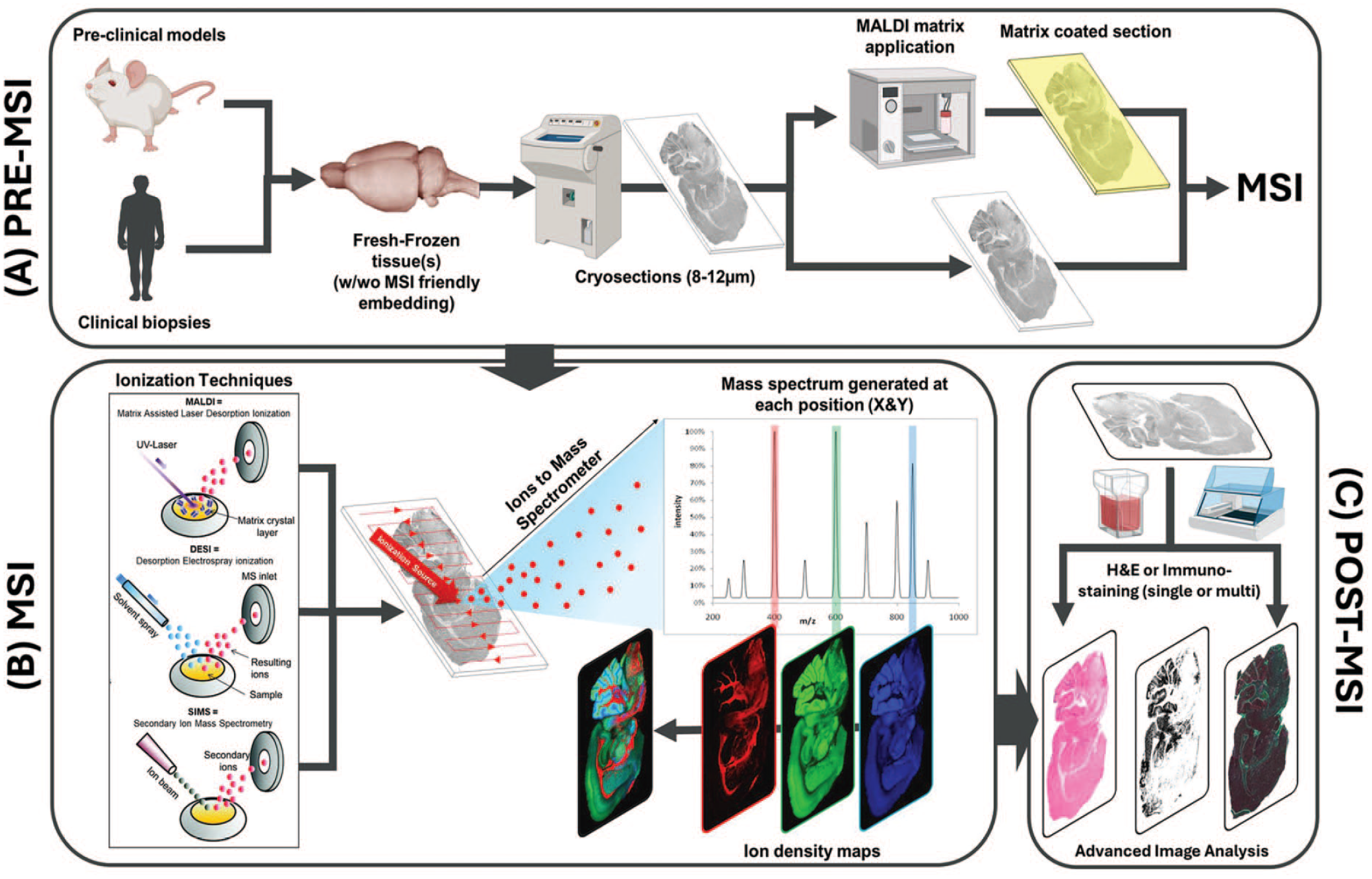
MS imaging workflow. (A) Sample processing Pre-MSI, (B) Principle of MSI. Graphical representation of the main ionization techniques for MSI and principle of scanning the tissue, (C) Post-MSI, integration of other imaging technologies after MSI experiment. Modified from figure adapted with permission from Porta Siegel et al. 41 and Vaysse et al. 46 , Icons created with BioRender.com.
Sample preparation
Tissue samples destined for MSI can be of different types according to availability. Ideal conditions as well as suboptimal but still proficient conditions are discussed in the following paragraphs. 13
MSI frozen tissue sampling techniques
In general, the tissue is placed in a cassette covered with aluminum-foil and placed on dry ice (or in tin-foil floating on liquid nitrogen) to freeze it. Tissue is stored at −80°C. Solid tissues (such as brain, liver, or kidney) can generally be cryo-sectioned without any embedding.
For delicate tissues, embedding may be required to preserve the morphology during freezing process. Embedding media compatible with MSI: low melting agarose gelatin and carboxymethylcellulose (CMC). Optimal cutting temperature (OCT) medium, which is commonly used for cryostat sectioning, should be avoided, due to ion suppression. 22
MSI FFPE tissue sampling techniques
In toxicologic pathology studies, tissue specimens are usually FFPE blocks for routine histopathological diagnosis and long-term storage. While the formaldehyde-induced crosslinking between biomolecules helps to maintain the tissue structure, it may impede biochemical or molecular analysis. Furthermore, as crosslinking continues in paraffin blocks, storage time of FFPE samples has a significant impact on MALDI-MSI analyses. 14 Although standard MALDI-MSI protocols can be used for the successful detection of certain small molecules and their metabolites in FFPE tissues, 34 the analysis of large biomolecules often requires special pretreatment and matrix materials.
Heat-induced antigen retrieval (HIAR) in citric acid or Tris-EDTA buffer, widely used for IHC on FFPE tissue sections, is known to enhance the efficiency of protein and solvent-resistant lipid detection by MALDI-MSI.18,24,25 Although HIAR partially reverses methylene bridges formed by formaldehyde, the mechanism of retrieval is not yet fully understood, and chemical modification of biomolecules may occur during the process. A recent study by Lee et al. 29 demonstrated that hydroxylamine hydrochloride can effectively remove formaldehyde-induced peptidyl crosslinks in acidic TRIS buffer, enabling detection of pancreatic islet hormones even in more than 5-year-old FFPE human tissue samples.
In situ enzymatic digestion of tissue by micro-spotting or spraying of enzymes is necessary to produce proteolytic fragments in FFPE samples, especially for large molecule detection. Not only trypsin but also other enzymes such as chymotrypsin, elastase, pepsin, and Peptide—N-Glycosidase F (PNGase F) can be used. The enzyme type and concentration, digestion buffer and digestion time should be optimized based on the tissue type, target analytes and sample storage time.9,28
Among the variety of matrix compounds developed for MALDI-MSI, α-cyano-4-hydroxycinnamic acid is widely used for peptides and protein analysis in fresh frozen tissue sections. However, use of this matrix in FFPE sections results in lower signal intensity or peak resolution compared with fresh frozen tissue samples. Lemaire et al. 30 discovered that these issues can be overcome by neutralizing aldehydes using 2,4-dinitrophenylhydrazine mixed with α-cyano-4-hydroxycinnamic acid in equal proportion.
Data Management
MSI generates complex data sets that require robust computational approaches for meaningful analysis and interpretation. Interpreting MSI data involves multiple steps from normalization and analyte identification to advanced supervised and unsupervised computational approaches. Different tools are provided by the platform-associated software, even though customized computational approaches are often applied, beyond the purpose of this article. Anyhow, here follows a general guidance on computational approaches for data interpretation related to MSI.
Data Normalization
MSI data can be influenced by experimental variations, including matrix effects, ion suppression, or instrument instability. To correct for these, signal intensity normalization is due: when applicable, known internal standards helps normalize the data more precisely by compensating any matrix effects and technical variability; in addition, normalizing based on the spatial characteristics of the sample can adjust the intensity in case of nonuniform ionization or differences in tissue thickness; finally, Total Ion Current (TIC) normalization is adopted to scale the intensity of detected ions by the total signal in each pixel to reduce variability across the data set. Data normalization, including considerations on calibration and validation strategies, remains a critical step that should be tailored for different experimental settings. 48
Analyte Annotation
After raw data acquisition, identifying the molecular species is a critical step. MSI software, such as MSIReader, SCiLS Lab, or MALDIquant, provide interfaces for annotating mass spectra with putative molecular identities. Then, the so called “molecular annotation” can be achieved by using mass spectral libraries and reference databases such as METLIN, Lipid Maps, or HMDB, aiming to match detected values with known compounds. If no exact match is available in libraries, molecular formulas can be predicted based on mass measurements. Ultimately, for increasing the confidence, tandem mass spectrometry (MS/MS) can be used to confirm the structures of molecules, based on fragmentation patterns.
Quantification
Quantification approaches can be relative or absolute.
Relative quantification compares signal intensities and ion abundance of particular analytes across different regions or conditions in the tissue sample, assuming that the ionization is consistent.
Absolute quantification adopts internal standards with known concentrations: matching internal standards to analyte-specific regions, the absolute concentration of molecules can be estimated, even though this approach is challenging time-consuming, and challenging due to ion suppression and matrix effects.
Data Analysis
Unsupervised analysis is often used for exploratory identification of patterns without prior knowledge of the biological question: principal component analysis (PCA) highlights trends, such as regions of interest or clusters of related spectra, visualizing also the variance in large data sets; then, hierarchical clustering algorithms are used to group similar spectra or spatial regions based on molecular profiles: this can reveal biochemical heterogeneity in tissue samples, such as tumor subregions with distinct metabolic signatures. Statistical software, such as R or Python, with libraries like scikit-learn, NumPy, and Matplotlib, can help in this context.
Supervised analysis is instead useful when there is a hypothesis or known biologic question as a guidance: partial least squares discriminant analysis (PLS-DA) is often employed to discriminate between predefined groups, such as normal versus diseased tissue, by correlating the spectral data with prelabeled pathology annotations; otherwise, machine and deep learning models, including convolutional neural networks (CNNs), have been used to correlate MSI with histopathological data in automatism.
Pathology correlation and multimodal integration
MSI can be integrated with histopathology data as any other molecular methods given spatial resolution, by aligning images generated by different techniques, possibly by machine learning, too, as this computer aid can help also in this phase of the workflow.
Data interpretation and visualization
Visualization techniques such as ion intensity maps (heatmaps) help in interpreting spatial distribution of specific analytes, as well as in sharing and divulgating the results to a diverse audience, such as the senior management: these heatmaps are color-coded overlays that represent the abundance of selected ions across tissue sections. In addition, some advanced MSI software reconstruct 3D molecular images by stacking 2D ion maps from serial tissue sections, similar to tomographic techniques for medical diagnostic (eg, using DICOM images).
Applications
To showcase the extensive range of MSI applications, various diagnostic pathology cases from nonclinical and clinical settings (eg, renal amyloid deposits, lipids, N-glycans, and tryptic peptides), as well as toxicologic pathology cases in drug development (eg, drugs, metabolites, crystal deposits, biomarkers for phospholipidosis, phosphorothioate (PS)-containing antisense oligonucleotides, needle-shaped material in intestines, liver morphology biomarkers, long-acting injectable formulations, ovarian corpora lutea, antibody-drug-conjugate (ADC) lung toxicity, and ocular toxicity assessment are presented below). These cases are not meant to represent an exhaustive list of all possible applications, but to be considered as examples that include important discussion points, critical for MSI.
The focus on MALDI in use cases within MSI, compared with techniques like DESI or SIMS is mainly due to the fact that MALDI has been more extensively refined over time, being one of the first to gain widespread adoption in biomedical research. Its maturity has led to a larger body of use cases here reported compared with DESI, SIMS or others. In addition, MALDI is suitable for a wide range of applications, excelling at analyzing large biomolecules as well as small molecules. Its versatility allows to visualize the molecular distributions in tissue samples. Not less important, compared with DESI and SIMS, MALDI is generally more straightforward in terms of sample preparation and data acquisition, contributing to its popularity for real-world applications. However, DESI requires complex setups for precise solvent delivery, while SIMS can cause fragmentation of biomolecules due to its higher energy ionization method, limiting its use in biological applications. In addition, MALDI is particularly well-suited for detecting larger biomolecules, like peptides, proteins, and lipids, which are often considered to be interesting biomarkers. Again, the commercial availability of MALDI instruments and software, along with continuous improvements from vendors, has also driven its prevalence in MSI studies, because the instrumentations for MALDI are highly optimized for user-friendly operation and routine analysis. Given that MALDI has a longer history and a broader user base, the authors prioritized this technique, in the presented use cases. However, DESI or SIMS are not less important, but MALDI is currently the predominant tool in biological MSI applications.
Cases including animals were based on experiments conducted in accordance with institutional and governmental guiding principles in their use. In detail, all nonclinical specimens were obtained from studies approved by the local animal ethical committee and conducted in facilities accredited by national institutions adhering to international guidelines (eg, AAALAC). Cases including human samples were approved by appropriate institutional review boards and complied with all relevant local governmental regulations. In detail, human specimens were procured as part of biobank initiatives. Patients gave written informed consent for biobanking and research use of surgical specimens.
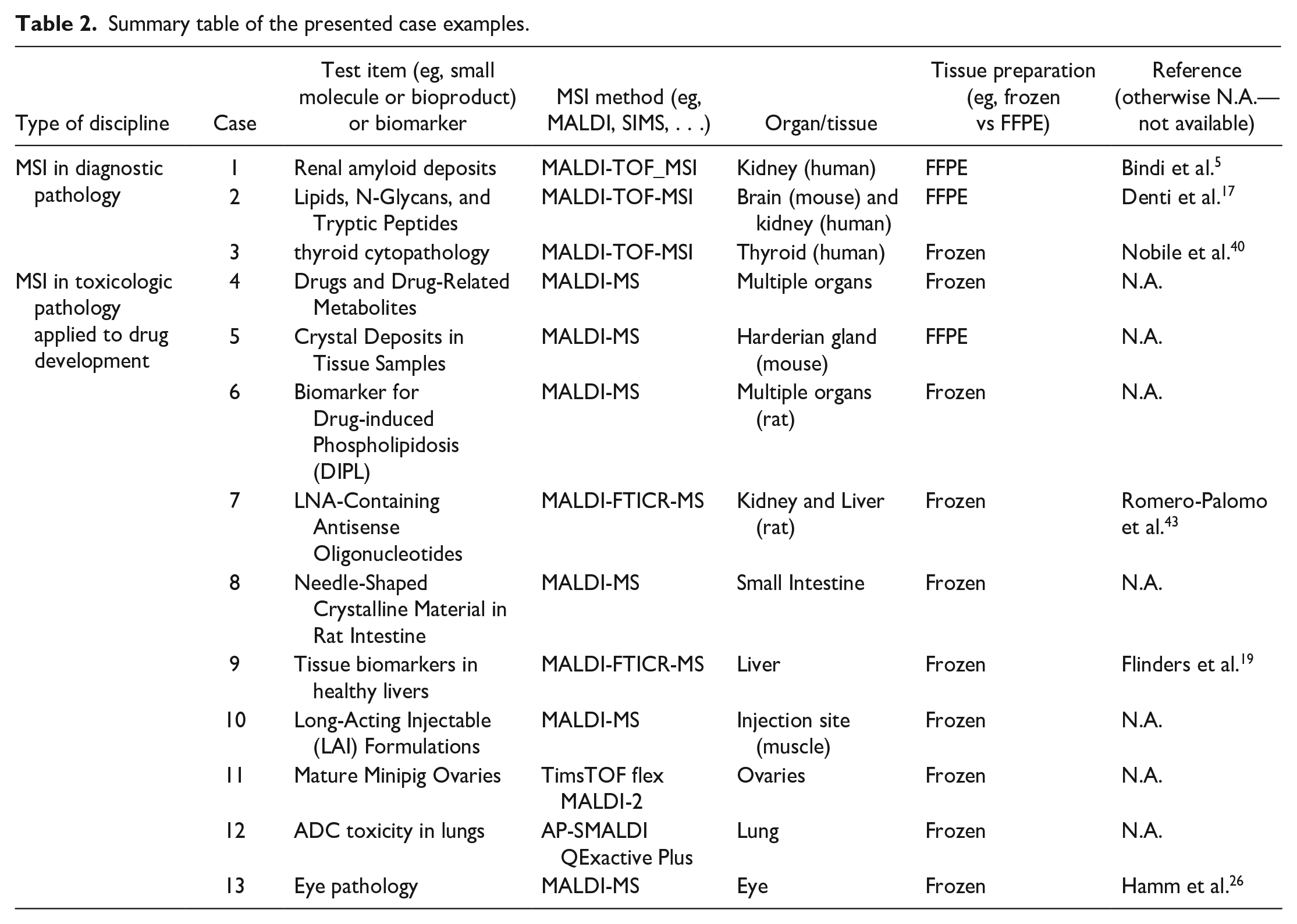
The presented case examples are summarized in Table 2.
Summary table of the presented case examples.
Case Examples of MALDI-MSI Applications in Diagnostic Pathology
As we transition toward the era of personalized medicine, there is an increasing demand for molecular tools that can assist pathologists, facilitating a more precise disease classification. In this context, MALDI-MSI could be included in the digital pathology workflow due to its spatially resolved characteristics and high throughput capacity, offering a supplementary molecular perspective of pathological tissue. Over the past few decades, the volume of research dedicated to identifying new diagnostic, prognostic, and predictive molecular biomarkers using MSI has grown significantly. When combined, particularly with computational pathology methods, these efforts demonstrate considerable potential in advancing the field. In the next section, we discuss various promising application areas and briefly touch upon methodological advancements that may provide pathologists with a more holistic overview of diseased tissue.
Case 1. MSI-Based Proteomics for the in Situ Detection and Typing of Renal Amyloid Deposits
Within the field of renal pathology, the early detection and accurate definition of renal amyloidosis, one of the “monoclonal gammopathy of renal significance” (MGRS), can impact the final treatment of patients. In challenging cases, amyloid precursor identification (typing) may be compromised by the inherent limitations of immunofluorescence (IF) technique. Under such circumstances, MALDI-MSI may represent a valid alternative. In fact, Bindi et al. 5 highlighted that this technology can detect amyloid deposits in FFPE renal biopsies, even within tubulointerstitial regions, based upon the colocalization of tryptic fragments derived from amyloid fingerprint proteins (apolipoproteinE [ApoE], apolipoprotein A1 [ApoA1], apolipoprotein A4, [ApoA4], serum amyloid P (SAP), and vitronectin [VTNC]). When guided by the predominant expression of these fingerprint proteins, the regions highlighted matched well with the annotations provided by the renal pathologist on a routine Congo Red-stained section. Moreover, by extracting the protein profiles specific to the renal amyloid deposits using MALDI-MSI, it was possible to distinguish renal biopsies of Amyloid Light chain (AL)λ-amyloidosis (n = 3), ALκ-amyloidosis (n = 3), and Amyloid A (AA)-amyloidosis (n = 3) based on the relative intensity of the tryptic peptides deriving from these precursor proteins. Considering both the presence, or absence, of amyloid fingerprint proteins and amyloid precursor proteins, it was possible to correctly assign two cases of light-chain deposition disease as nonamyloid, while concurrently identifying the presence of the correct lambda light chain, as confirmed by the M-spikes observed in IF.
More recently, the same authors also explored the potential of using MALDI-MSI with Bouin-fixed paraffin-embedded (BFPE) renal biopsies, which still represents the primary tissue archiving strategy in the vast majority of specialized nephropathology centers worldwide. BFPE maintains the morphological intricacies of renal structures but preserves antigenicity only poorly. In this work, the authors underlined the ability to leverage the colocalization of amyloid fingerprint proteins to detect amyloid deposits also in BFPE renal biopsies, showing good agreement with the annotations provided by the renal pathologist on the Congo Red stained tissue and with comparable reliability to an FFPE counterpart. Collectively, these results indicate that renal pathology archives containing numerous BFPE biopsies from cases of renal amyloidosis can be utilized to further improve the capability to detect and type renal amyloidosis using MALDI-MSI and extend its utility to rarer subsets of precursor proteins with amyloidogenic potential.
Spatial in situ molecular characterization of various amyloid deposits using MSI is a significant advance over previous techniques, enabling precise biochemical diagnosis and better downstream therapeutic management. Collectively, these works highlight the advantage that can be offered by using MALDI-MSI to perform the spatial detection and characterization of renal amyloid deposits, especially in instances where deposits are in their early stage and may be overlooked using traditional histology techniques. Moreover, this is also relevant for alternative MS-based approaches that employ laser capture microdissection coupled with mass spectrometry (LCM-MS), given that a greater amount of material is required to achieve sufficient sensitivity and it still relies on the guidance of Congo Red staining. These advantages, along with the possibility of performing direct histo-molecular correlation, suggest that this technique could enable more precise diagnoses and better downstream therapeutic management.
Key Insights
MALDI-MSI represents a feasible tool for the spatially resolved detection and subtyping of renal amyloid deposits, even in instances with minimal material available and at the earlier stages of formation. This has the potential to offer distinct advantages with respect to traditional histology and MS-based techniques.
Case 2. Spatial Multiomics of Lipids, N-Glycans, and Tryptic Peptides on a Single Section of FFPE Tissue
Recent trends indicate a movement toward integrating spatial omics data sets that span various molecular levels, including the lipidome, N-glycome, and proteome. Integrating these multiomics MSI data sets may enhance the ability of molecular classifiers to discriminate and provide a more comprehensive, untargeted mapping of the molecular complexity in diseased tissue. Of note, Denti et al. 17 have demonstrated the feasibility of conducting spatial multiomics analyses of these three molecular classes on a single FFPE tissue section within the heterogeneous context of clear cell renal cell carcinoma (ccRCC). This highlights that each molecular level provided complementary information and characterized different histological regions. Moreover, when the spatial omics data was integrated, the different histopathological regions of the ccRCC tissue could be better discriminated, especially concerning the imaging data set of any single omics class. This further reinforces previous evidence showing that the use of classifiers spanning multiple omics levels has potential in enhancing the correlation of molecular features with clinical outcomes and refining patient stratification. It is also noteworthy that such a sequential workflow is particularly valuable when clinical tissue samples are limited, such as in rare diseases or cases that require multiple histological and immunohistochemical evaluations for routine diagnosis.
In addition, new diagnostic avenues such as multiplexed imaging of cell-specific protein expression markers have advanced. Given that single-plex imaging in conventional IHC assays is limited and IHC is frequently required in routine pathological workflows, multiplexed imaging could markedly improve diagnostic efficacy and throughput. MALDI MS-based high-plex immunohistochemistry (MALDI-HiPLEX-IHC) is capable of mapping tens, if not hundreds, of protein expression markers with the aid of photo cleavable mass tags. It may promise to provide significant diagnostic potential in the precision medicine and digital pathology era, particularly in the context of tumor diagnostics. To the best of the authors’ knowledge, high spatial resolution MALDI-MSI, MS/MS data acquisition and lipid, N-glycan and peptide annotation and identification are reported on different tissue sections, 39 while the current case describes the experience on the same tissue section.
Key insights
The possibility to ingrate spatial multiomics data, obtained from one single FFPE tissue section, can help to generate a more complete molecular overview and better discriminate cells of diverse pathological states. Moreover, the recent development of the MALDI-HiPLEX-IHC workflow has opened new avenues within the realm of clinical diagnostics and has the potential to serve as a high-throughput alternative for performing routine clinical IHC assays.
Case 3. Toward Implementation of MSI-Based Proteomics for the Classification of Cytological Thyroid Specimens
In addition to tissue-based pathology, MALDI-MSI may also be applied to cytological samples. This technology has been adapted for the diagnostic evaluation of indeterminate thyroid nodules whose malignant nature is often difficult to ascertain and inadequate diagnostic evaluation results in inefficient patient management. 40 With this scope in mind, a more robust sample preparation workflow, ensuring sample stability over longer periods of time, was initially developed. This method also facilitates sharing samples from multiple centers and can be more easily collected and processed by individuals without expertise in MSI.
The same authors proceeded to construct a least absolute shrinkage and selection operator (LASSO) based statistical model that was trained using cytological samples obtained from frequently encountered thyroid lesions, including benign (hyperplastic) and malignant cases (papillary thyroid carcinoma), as well as those with an inflammatory background (Hashimoto’s thyroiditis). Of note, this classifier was designed to consider the entire cytological sample through a pixel-by-pixel approach in MALDI-MSI data, minimizing the necessity for manual annotations from the pathologist. Despite including thyroid samples with inadequate cellularity in the validation set (n = 170), the approach demonstrated a specificity of 82.9% and a sensitivity of 43.1%. Sensitivity increased to 76.5% when focusing on fine needle aspiration biopsies with adequate cellularity.
Recently, Capitoli et al. 7 proposed a novel workflow, which describes how spatial proteomics, using MALDI-MSI, could be introduced into clinical practice for cases of indeterminate diagnosis. They suggested a three-level diagnostic classification for indeterminate nodules based on the percentage of malignant and benign pixels in the sample. A low number of malignant pixels (<7%) indicates a benign sample requiring ultrasound follow-up at 12 months, while a high number of malignant pixels (>16.7%) suggests a malignant sample necessitating thyroidectomy. For nodules with a gray-zone percentage of malignant pixels (7.0%-16.7%), strict ultrasound follow-up and potential repeat biopsy are recommended.
Key insights
MALDI-MSI can also be applied within cytological contexts, in particular, for the classification of thyroid nodules. Recent works have demonstrated a specificity and sensitivity of 82.9% and 76.5%, respectively, when applying a LASSO statistical model to proteomics MALDI-MSI data and work is ongoing to provide more clear classification guidelines when seeking to define nodules based upon the percentage of malignant and benign pixels in the specimen.
Case Examples of MALDI-MSI Applications in Toxicologic Pathology Applied to Drug Development
Case 4. MALDI MS Imaging of Drugs and Drug-Related Metabolites: A Workflow Tutorial
MSI can support nonclinical drug safety (ie, toxicological) studies in pharmaceutical research and development. It provides evidence on tissue exposure, preliminary information on drug metabolism, and, most importantly, can associate local concentrations with morphology, especially in the case of pathological findings. Here, a robust workflow that was developed to integrate MSI as one of many work packages in a standard toxicological study is described. The workflow is commonly applied to early studies (eg, 1- or 2-week repeat dose) in rats and mice but can easily be adapted to other nonclinical species. It is easier to integrate nonstandard investigations like MSI into early (non-GLP) studies due to fewer regulatory and reporting requirements. Integrating the sampling requirements for a growing number of analyses, be it mandatory (eg, histopathology, toxicokinetics, and clinical chemistry) or optional tests (eg, MSI, genomics, metabolomics, and proteomics), requires careful planning, especially when animal numbers are kept low, in line with animal welfare guidelines.
It is recommended to sample tissue from at least three animals of each dose group, including the mock-dosed control group. As the main metabolizing organs, liver and kidney s always should always be collected. Other tissue samples can be selected according to project specific needs and may include target organs or tissue considered at risk for off-target effects. Samples should be frozen in liquid nitrogen-cooled isopentane and stored at −80°C until further use. The time of sacrifice in relation to the pharmacokinetic profile of the investigated compound is an important consideration. Therefore, it is recommended to collect samples close to the compounds tmax (time of highest plasma exposure), as tissue concentrations would be highest. However, sampling at later time points, even after a recovery period, may be sensible when investigating tissue accumulation. When individual animals are removed from the study (eg, due to acute toxicity), the time of sacrifice cannot be precisely coordinated and can become a serious confounder in the analysis. If such samples cannot be excluded from the analysis, it is helpful to correlate drug concentrations in plasma (derived from toxicokinetic investigation of the same study) with MSI signal intensities to evaluate the effect of different sampling time points.
During sectioning, care must be taken to obtain similar sectioning planes per organ for each animal investigated, as it facilitates the comparison of distribution images later. Different tissue types are analyzed separately, but for each tissue it is recommended to create a complex layout of tissue sections to enable controls and minimize the number of slides required.
The individual elements are (Figure 3) as follows:
MALDI matrix with reference compounds: freshly made matrix solution is spiked with reference compounds that cover the desired mass range (eg, Chloroquine, Reserpine, and Rosuvastatin for small molecule analysis) at low µM concentration (ie, 1-10 µmol/L). The spiked matrix is used in all experiments and provides stable reference for intensity normalization and assessment of mass accuracy.
On-tissue dilution series to determine limit of detection (LoD): 8 to 10 concentration steps (ranging from 0.1 to 100 µmol/L) of the formulation solution used in the study are spotted on a liver section from a control animal. This section is sprayed with matrix and analyzed by MSI, which provides information on the LoD and linearity of signal intensity for the target compound. The linear dynamic range is usually limited, covering 30- to 50-fold changes in concentration. Spotting on glass or steel surfaces usually leads to a significant overestimation of the LoD. Instead, liver tissue provides a realistic biological background and is homogenous enough to limit spot-to-spot variation. Sensitivity between compounds may vary significantly: 20-fold differences in LoD are not unusual. This setup can also be used to optimize sample preparation, including choice of MALDI matrix, solvent and spraying conditions. It should be noted that although a spotted dilution series provides reasonable information on LoD, it does not perfectly mimic compound extraction from tissue, and therefore absolute quantitation of signal intensities by this approach is not recommended. Using formulation solution instead of pure reference compound, drug-related signals that derive from contamination, chemical degradation or in-source fragmentation of the target compound can be detected. Otherwise, these could erroneously be assigned as drug metabolites if detected in dosed animal tissue.
Control tissue sprayed with target compound: spray tissue sections from a control animal with MALDI matrix, adding ~10 µmol/L of the target compound. The concentration should be adjusted to achieve approximately the same signal intensity as the tissue of the highest dose group. This sample serves as a positive control for the detection of the target compound, and more importantly, it can assess tissue-specific ion suppression, which is of particular concern in heterogenous tissues, such as brain or skin.
Sections from all investigated animals and dose groups: if possible, mount a single section from each animal (across all dose groups) on a single slide. This allows easy comparison of signal intensity within and across groups and limits the influence of technical variability. Consecutive sections are collected on additional slides to allow for replicate measurements (typical n = 2-3), histological staining, and additional measurements (changes in mass range, instrument polarity, choice of matrix, on-tissue MS/MS). For larger tissue sections, samples have to be distributed to multiple slides, but care should be taken that a dilution series, positive and negative control sections are included on each slide to assess slide-to-slide variability.
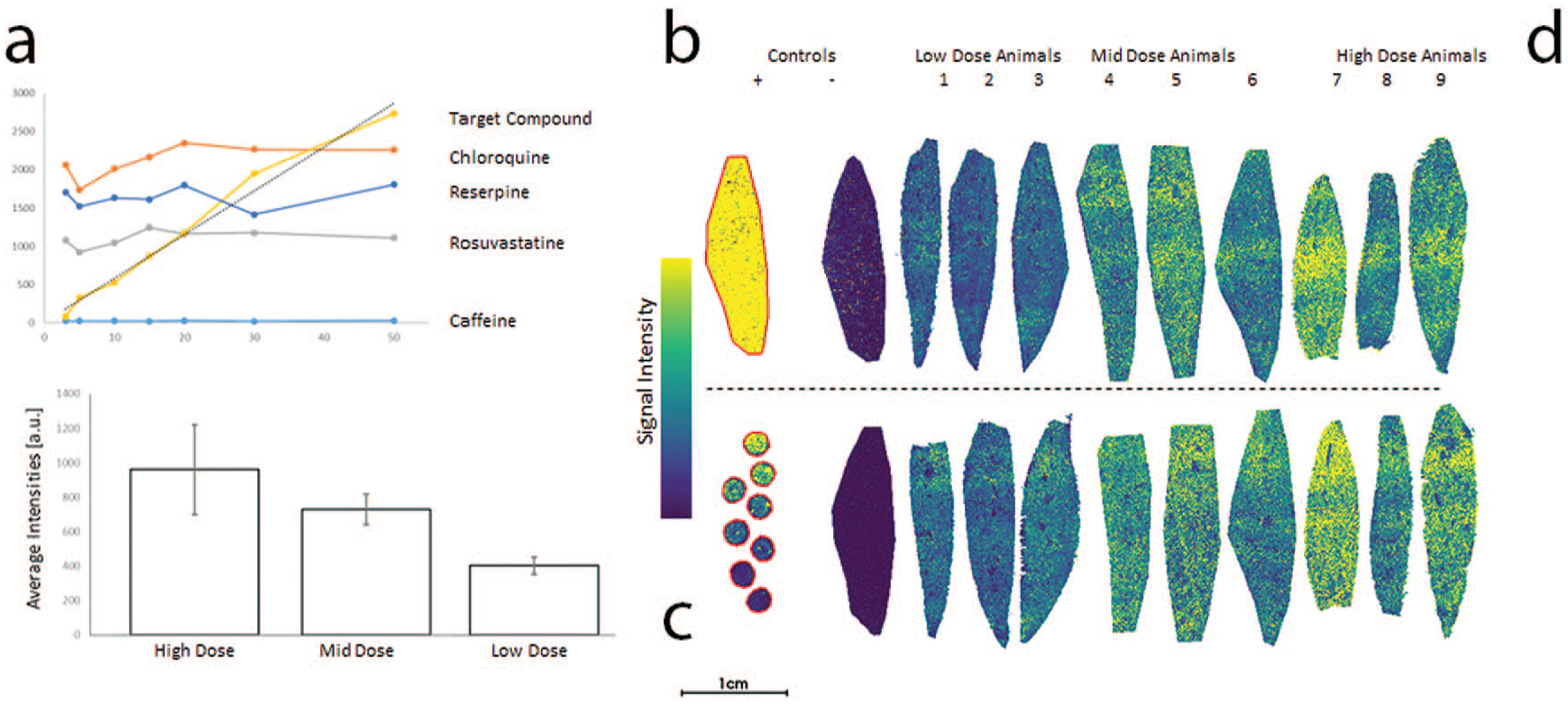
Proposed slide layout for MSI of drugs and drug metabolites. The samples shown are from rat liver, collected after repeat dosing in four different dose group (including control). Distribution of the parent compound is shown. Individual elements as described in the text: (A) Intensity of reference compounds spiked into the MALDI matrix at fixed concentration. (B) Control animal tissue section sprayed with formulation used in the study as positive control. (C) Dilution series of the target compound spiked on top of control liver tissue. The resulting signal intensities are shown in (A). (D) Tissue sections from animals of all dose groups mounted on a single glass slide. Consecutive sections on a second slide as technical replicate (lower row of tissue samples). Average intensities (per group) for the parent compound are shown in the bottom left corner, a dose-dependent increase in signal intensity is clearly visible. (Case 4)
Due to the (semi-) targeted nature of this approach, data analysis is straightforward. Initially, a list of target m/z-values should be generated, with a tolerance for mass accuracy (low ppm range). Each resulting interval can be used to extract signal intensities from the MSI data. Potential bio-transformations of the target compound, generated by prediction tools (eg, Mass-MetaSite, Molecular Discoveries) form the basis of the target list, typically include 100 to 300 putative metabolites. This can be expanded by other entries, such as known metabolite m/z (eg, from in vitro studies), reference compounds, and endogenous compounds of interest that can be monitored alongside drug-derived targets. The target list can then be filtered according to several criteria to extract relevant signals, which are then reported. Based on the author’s experience, the following suggests a putative metabolite:
Average intensity 3-fold or more above noise (ie, signal intensity in the control tissue) as drug-derived signals should be entirely absent in control tissue. This requirement is essential and helps to remove false-positive results due to isobaric endogenous compounds.
Average intensity at least 1% of target compound signal, as very low abundant metabolites are usually not relevant for drug safety considerations.
Average intensity at least 3-fold higher than in the spotted formulation solution. This helps to rule out nonbiologically formed drug-related signals (ie, contaminants, degradants, or MS-based artifacts).
Most small molecules undergo only a limited number of relevant bio-transformations, and some metabolites are present only in specific tissues. Therefore, the number of putative metabolites that pass the selection criteria is typically 10 or fewer. It is of course possible to analyze the MS data generated by such an approach in an untargeted fashion, elucidating all signals that change in comparison to the control. Such changes typically include a number of endogenous molecules that are altered as an effect of the treatment and therefore may serve as toxicological or pharmacodynamic markers.
Key insights
This workflow addresses several key requirements for targeted MSI measurements in a comprehensive way: limit of detection and dynamic range for the target compound, potential ion suppression effects of specific tissue areas, interference of endogenous background signals and slide-to-slide variability. The semi-targeted approach using a well-defined list of targets (such as putative drug metabolites) utilizes the highly multiplexed capabilities of MSI and allows for quick and straightforward data analysis.
Case 5. Analyzing Crystal Deposits in FFPE-Tissue Samples
Colocalizing drugs and drug-related material with morphological changes observed in tissue sections is a key application for MSI. In standard toxicological studies, histopathological assessment is almost always conducted on FFPE sections, and even if cryopreserved material is collected during the study, the FFPE material assessed by pathologists is often from different regions of the organ or from different animals of the same group. Since morphological changes can be highly localized within a tissue or restricted to specific animals within a group, MSI analysis of FFPE-tissue sections might be required. Analysis of FFPE tissue by MSI has been demonstrated before for peptides and endogenous small molecules, but represents a particular challenge for the analysis of small molecule drugs, which are often selected for good solubility and therefore are subject to wash-out and delocalization during dehydration and fixation process. However, when analyzing drugs and other xenobiotics, reference material for the compound of interest is usually available, so sample preparation and mass spectrometry methods can be optimized easily.
In the example presented here, MSI was used to investigate crystalline deposits in the Harderian gland of mice after 13 weeks of treatment with a drug candidate. Precipitation and crystal formation may indicate high local concentrations of drug-related material combined with unfavorable physicochemical environment and can be a safety concern. As MSI investigations were not originally considered when planning the study, no dedicated material had been collected and the investigation was limited to FFPE samples. High local concentrations of drug-related material incorporated into the crystalline deposits help to offset the limitations imposed on MSI performance by FFPE material. Although good sensitivity was achieved for the parent compound during method development, only very low signal intensity was detected in the Harderian glands of animals dosed at 300 mg/kg (Figure 4). The investigated compound is mainly metabolized by the formation of a carboxylic acid metabolite (Δm = 14 Da) and the signal for this metabolite was significantly higher than that of the parent compound (Figure 4, Table 3). Comparing the intensity of two different compounds can be error prone due to potential differences in ionization efficiency. As the carboxylic acid metabolite was synthetically available, an additional control experiment was conducted to determine differences in ionization. Parent and metabolite were sprayed on Harderian gland sections from control animals at identical concentrations (10 µg/mL) (Figure 5) and the differences in signal intensity and molecular weight were used to calculate a correction factor that was applied to all intensities measured in dosed animals. In addition, prior to data analysis, sections were stained with H&E and annotated by a pathologist to separate crystalline deposits, unaffected tissue and pigments which were then evaluated separately (Figure 6, Table 4). In the crystal deposit, the corrected metabolite was 22.9-fold higher than that of the parent compound, while the ratio was ~1 (and signal intensities close to limit of detection) in the remaining tissue. In this way, it could be clearly demonstrated that the crystalline deposit contained significantly higher amounts of the drug metabolite and only limited amounts of the parent compound. In this particular project, the result helped in de-risking the finding, as the carboxylic acid metabolite had much higher exposure multiples than the parent, making precipitation at physiological doses very unlikely (unpublished results). This application is an example of how spatially resolved metabolite date combined with histopathology data and other project information (like metabolism data) can support pharmaceutical development.
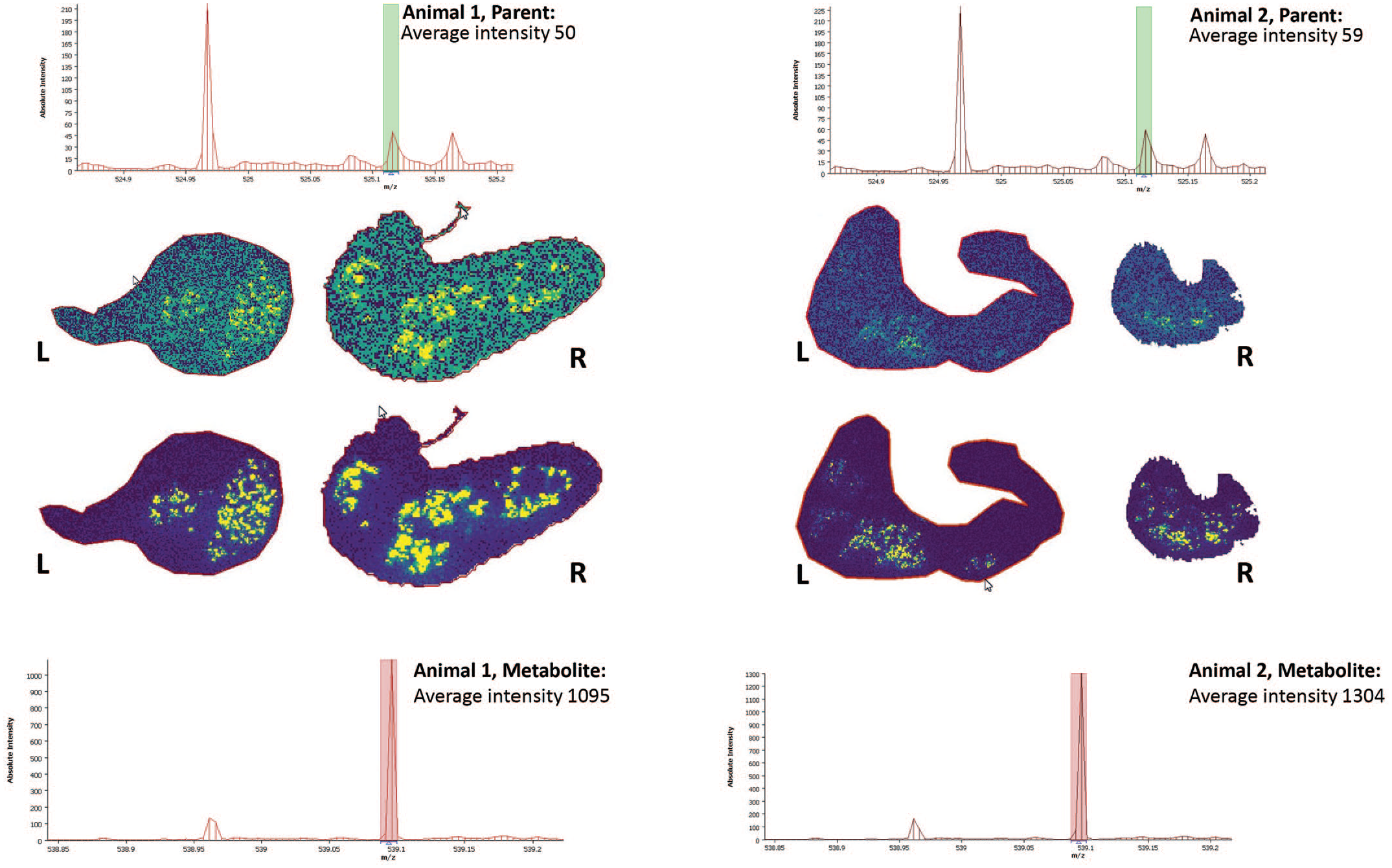
FFPE-tissue sections of mouse harderian glands after 13-week repeat dosing (300 mg/kg; once daily) of a drug candidate. The distribution of the parent drug (top half) and a drug metabolite (bottom half) were analyzed by MSI. The left and right gland of two animals from the same dose group were analyzed. The average signal intensities (per animal) is shown in the spectra. Additional signals in the spectra shown do not match any of the drug-related compounds investigated and most likely represent endogenous background that was not further evaluated. (Case 5)
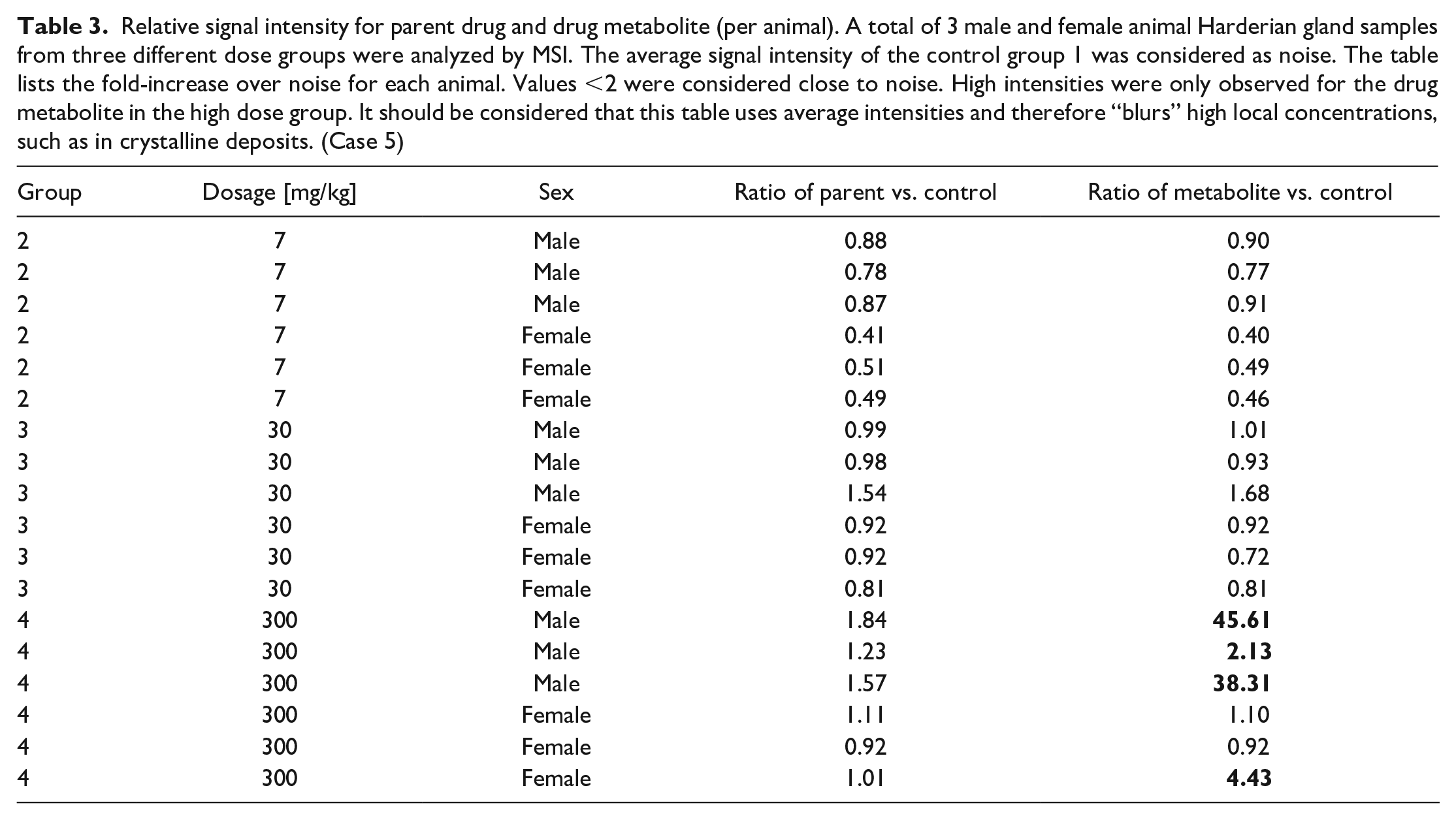
Relative signal intensity for parent drug and drug metabolite (per animal). A total of 3 male and female animal Harderian gland samples from three different dose groups were analyzed by MSI. The average signal intensity of the control group 1 was considered as noise. The table lists the fold-increase over noise for each animal. Values <2 were considered close to noise. High intensities were only observed for the drug metabolite in the high dose group. It should be considered that this table uses average intensities and therefore “blurs” high local concentrations, such as in crystalline deposits. (Case 5)
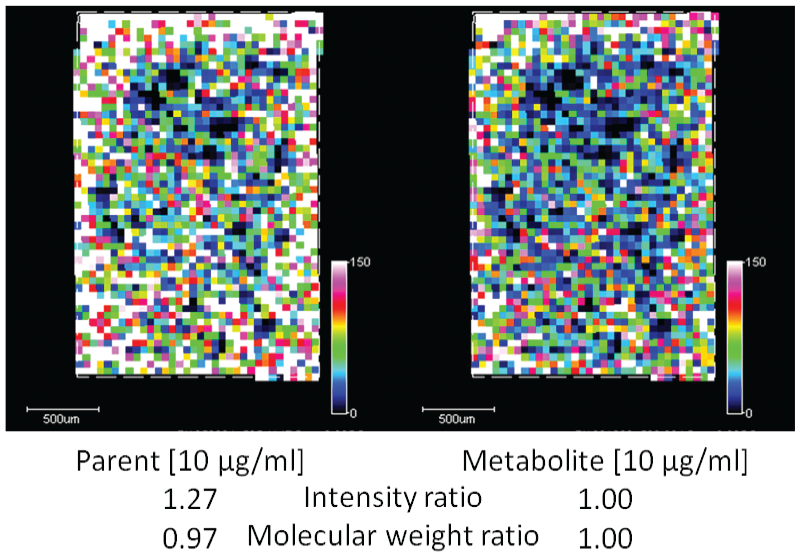
Intensity correction when comparing two different molecules in MSI data. Identical concentrations (10 µg/mL) of reference compound for parent drug (A) and drug metabolite (B) were sprayed on top of a tissue section from a control animal. Due to differences in ionization efficiency, the average signal intensity for the parent drug was 1.27-fold higher. Moreover, given that the parent drug had a lower molecular weight with respect to that of the drug metabolite (a factor of 0.97), which therefore equates to a greater number of molecules per µg, the metabolite intensity factor was adjusted accordingly to take this into consideration, resulting in an overall correction factor of 1.24. This correlation factor was then used in order to more appropriately compare the intensity of the parent drug with that of the drug metabolite. (Case 5)
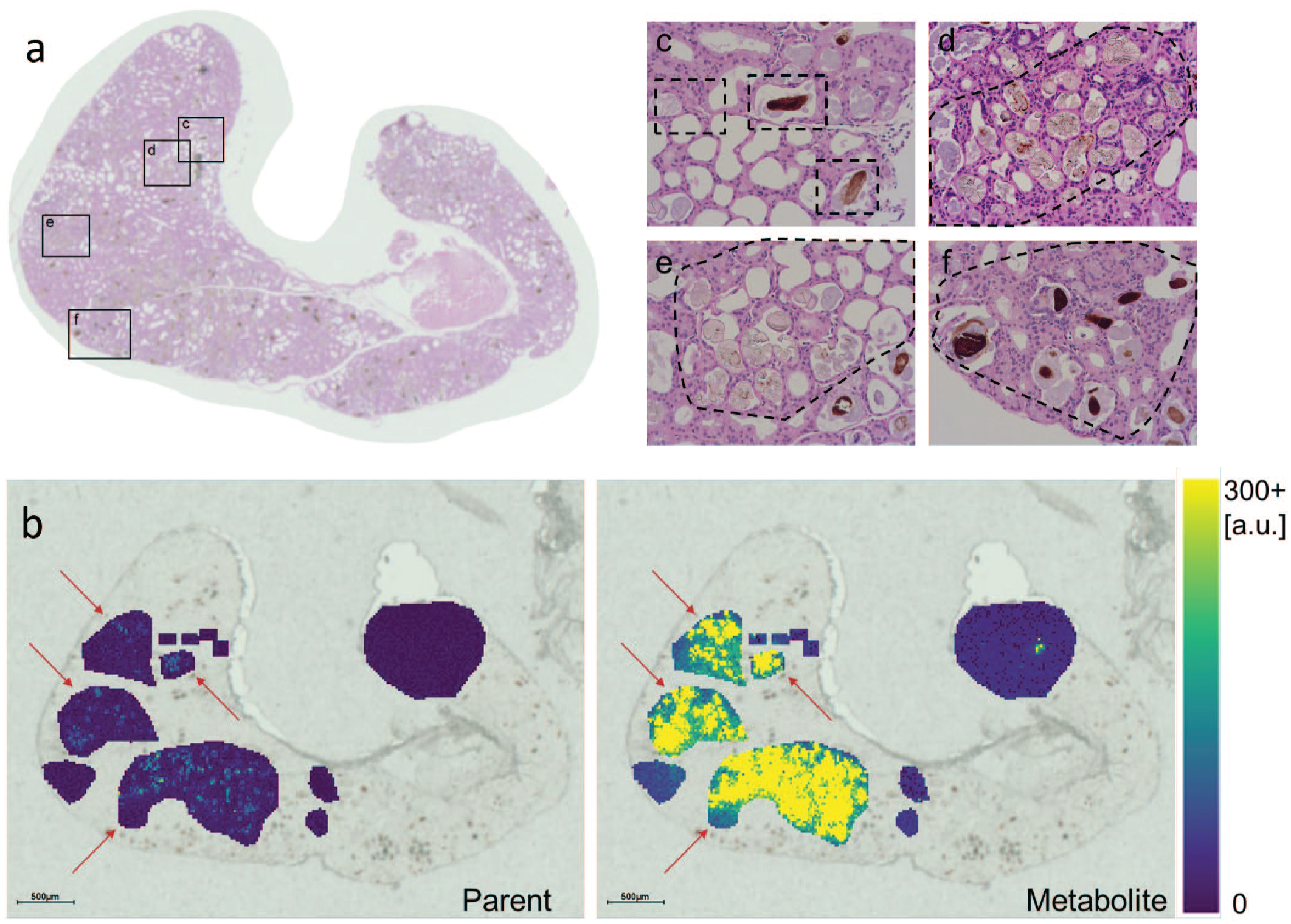
Histology-guided MSI acquisition. (A) With the aid of a pathologist, specific sample areas were identified on a H&E-stained Harderian gland section from the high dose group (300 mg/kg). (B) MSI analysis was performed on a consecutive section and the different sample areas analyzed independently for parent drug and drug metabolite signal intensities. Red arrows point to measurement areas with crystalline deposits. The areas considered were normal gland tissue (C), pigmented areas (C and F), and areas with multiple crystalline deposits (D and E). (Case 5)
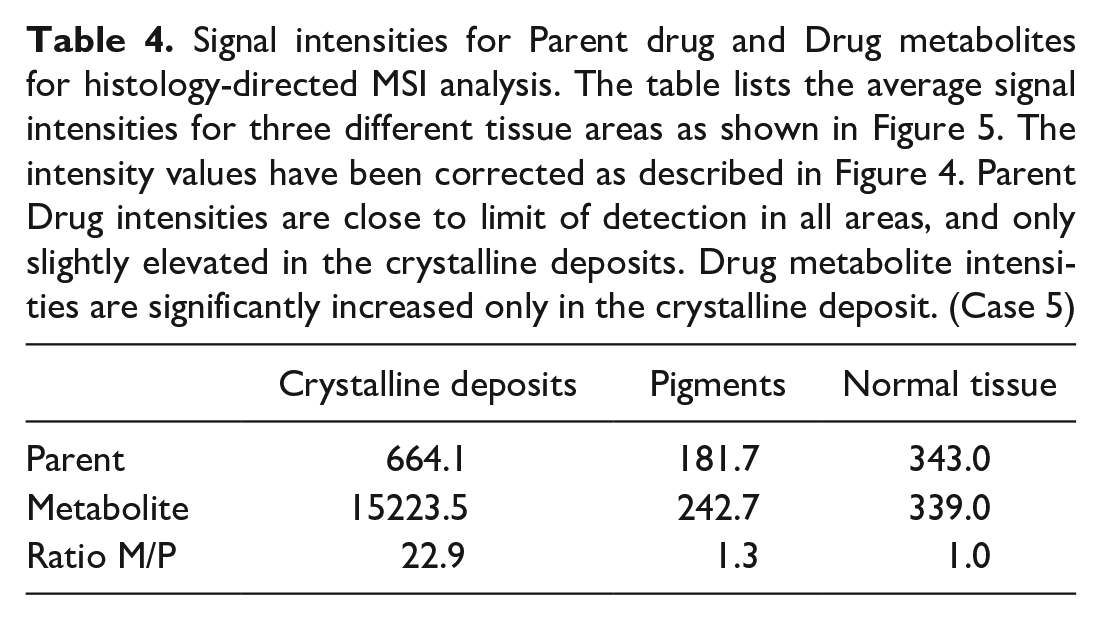
Signal intensities for Parent drug and Drug metabolites for histology-directed MSI analysis. The table lists the average signal intensities for three different tissue areas as shown in Figure 5. The intensity values have been corrected as described in Figure 4. Parent Drug intensities are close to limit of detection in all areas, and only slightly elevated in the crystalline deposits. Drug metabolite intensities are significantly increased only in the crystalline deposit. (Case 5)
Key insights
FFPE tissue are the most common specimen in histopathological investigations, but the sample processing involved can be detrimental to MSI results, especially when investigating small and/or soluble molecules. The use of reference compounds can alleviate these limitation as they allow specific optimization of experimental conditions. Reference compounds enable the comparison between different molecules by determining their respective ionization efficiencies. In addition, this case example shows the importance of interpreting the results of MSI in the context of additional results (such as light microscopy or pharmacokinetics data).
Case 6. Biomarker Elucidation for Drug-Induced Phospholipidosis
Drug-induced phospholipidosis (DIPL, also referred as drug-induced PLD or PL) is a condition induced by drug treatment, primarily by cationic amphiphilic compounds.3,42 The underlying mechanism is not fully understood, but the main effect observed is a massive accumulation of phospholipids in lysosomes. DIPL can affect almost any organ including lung, liver, kidney, but is itself not considered a serious adverse finding in nonclinical drug safety studies, especially as it is often reversible after cessation of dosing. However, since DIPL can be associated with more serious conditions such as degenerative or inflammatory changes, it can represent a challenge for the development of a drug candidate, especially if long-term or chronic treatment is envisioned. The gold standard to identify DIPL is histopathology and electron microscopy. Affected cells have a finely vacuolated and/or “foamy” appearance microscopically, which correlates ultrastructurally with cytoplasmic lamellar bodies characterized by concentric, myelin-like, structures.
To identify an early onset marker for DIPL, a mechanistic study with four discontinued drug candidates that had induced DIPL in previous animal studies was performed. Rats were dosed for 14 consecutive days at two dose levels per compound. The higher dose was selected to induce clear phospholipidosis, whereas the lower dose was selected to be below threshold for DIPL. Tissues (kidney, liver, and lung) were cryopreserved for analysis by MSI: 10-µm-thick sections were mounted on indium tin oxide (ITO) slides, coated with 9-AA matrix using a TM-Sprayer (HTX Technologies) and analyzed on solariX 7 Tesla FTICR-MS in negative ion mode at 50-µm pixel size. We found numerous molecular changes in all high dose animals when compared with the placebo-dosed control group. Due to the complexity of spatial distribution, intensity variations and large number of samples, we adopted a severity grading (similar to those used in histopathology) to describe the changes for each m/z-value, dose group, and tissue type (Figure 7, Table 5). Because of the large number of changes observed, it was decided to focus on those m/z-values exhibiting the most pronounced changes (ie, “++” in Table 5) for further analysis.
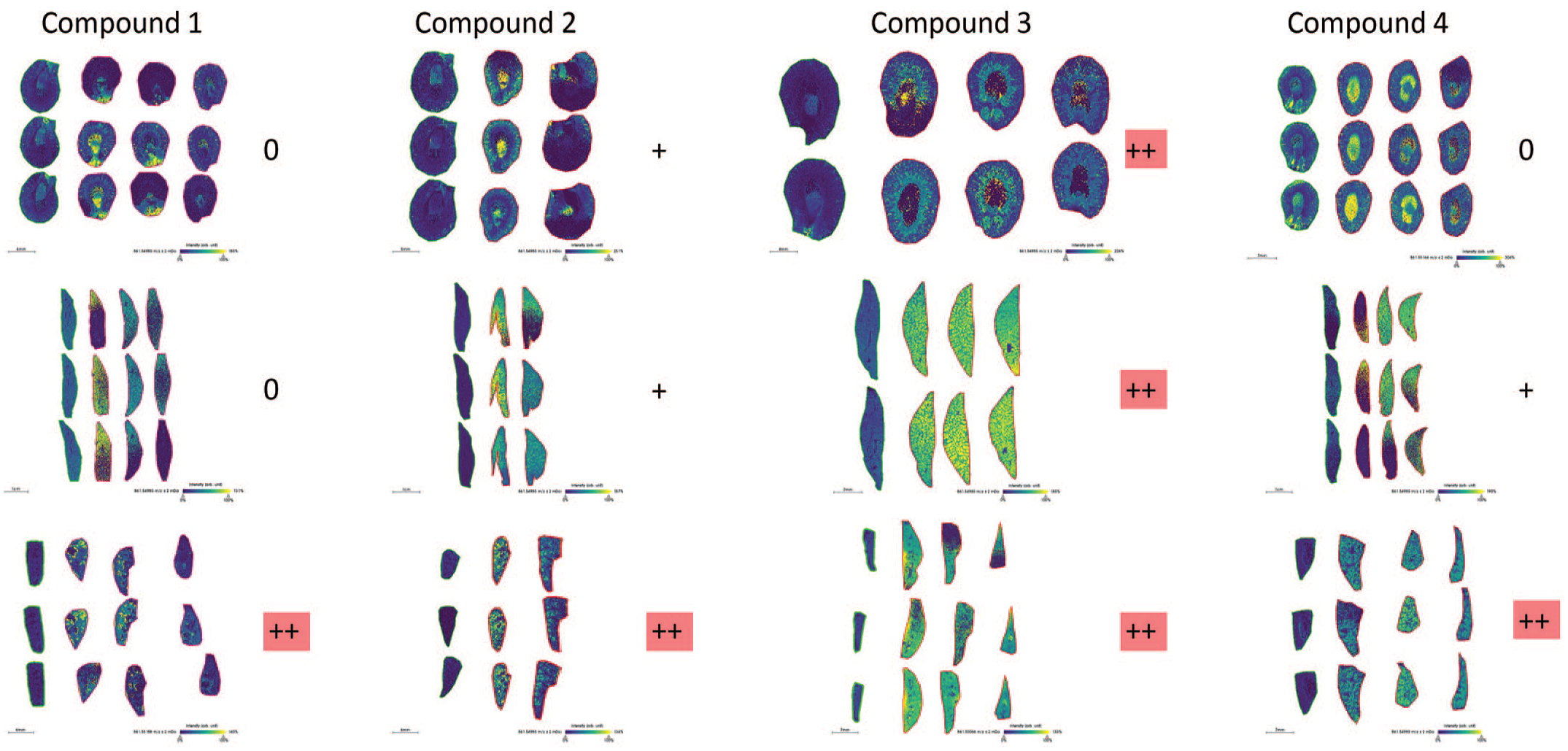
Changes of m/z 861.5 in tissue after treatment with drug-induced phospholipidosis (DIPL)-triggering compounds. m/z 861.5 is selected as a representative example of how molecular changes were evaluated and assigned a severity rating of “–” (decrease in comparison control); “0” (no or no consistent change); “+” (mild but consistent change); and “++” (strong and consistent change). Both changes in intensity but also distribution were considered. Each column shows tissue samples from one of the four compounds investigated, each row shows a different tissue type (kidney, liver, and lung from top to bottom). Within each panel, each row is one of 2 to 3 technical replicates, and each column represents an individual animal in the treatment group. The leftmost column of sections in each panel is from control animals. (Case 6)
Severity grading, reflecting complexity of spatial distribution and intensity variations, describing changes for each m/z-value, dose group, and tissue type (m/z-values exhibiting the most pronounced changes are labeled by “++”; note that some samples showed such pronounced changes more frequently than others). (Case 6)
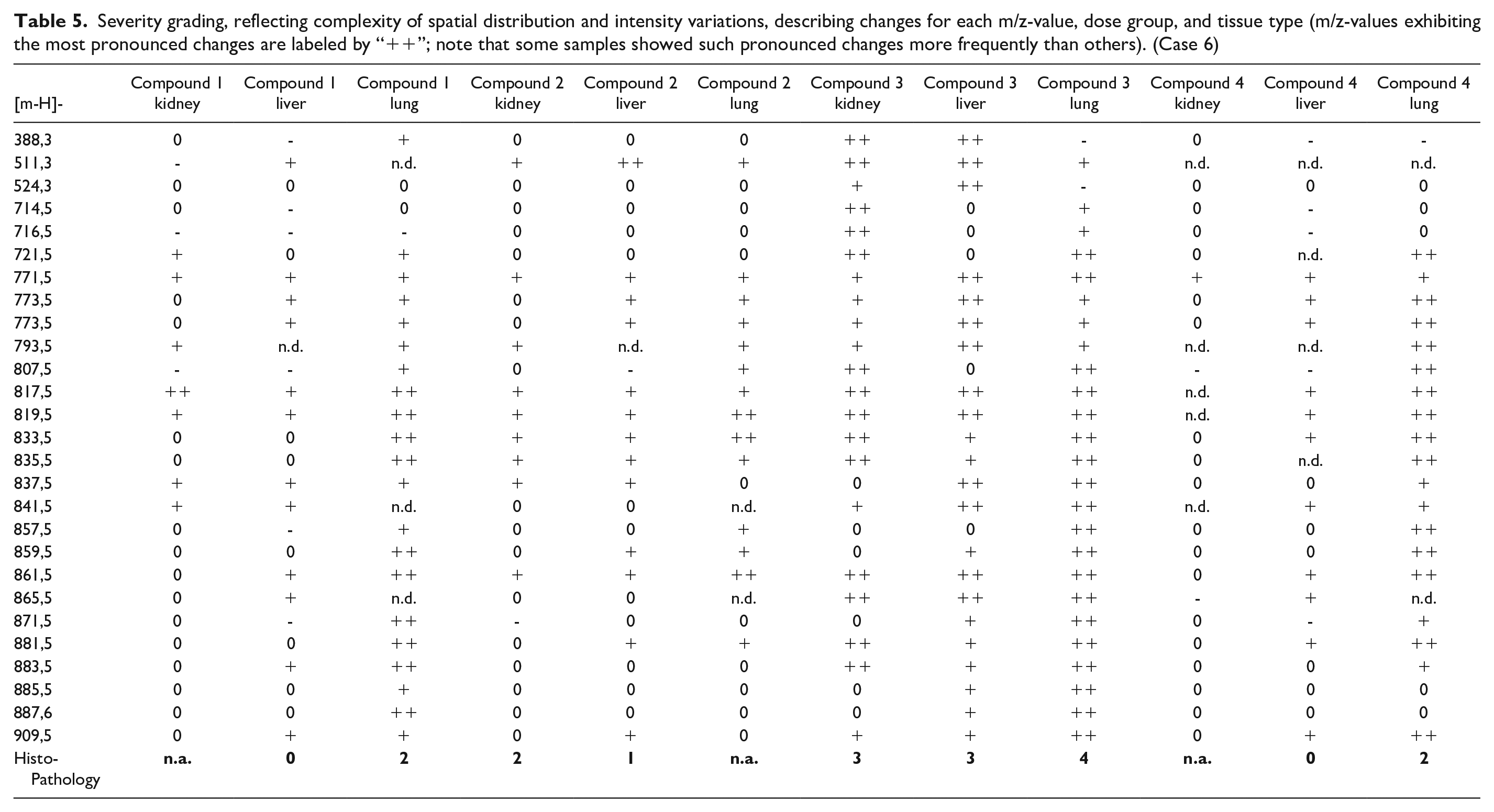
The changed m/z-values were not fully consistent across the four compounds and three tissue types analyzed, but some samples clearly showed a higher frequency of pronounced changes than others. To further evaluate if these m/z-values qualify as DIPL markers, each sample was assigned an independently determined histopathological severity grade, represented by the median severity of all animals investigated (on a scale from 0 [no change] to 4 [severe changes consistent with phospholipidosis]). In general, a severe histopathology grade was associated with a larger number of pronounced molecular changes. Interestingly, samples in the low dose group also showed clear but less pronounced changes for several of the m/z-values, but there were no, or only nonconclusive morphological changes reported in these tissues. These illustrate that molecular changes can be detected by MSI before any morphological effects of DIPL manifest in the animal, and therefore changes in these molecules may serve as early warning signs of a compound’s potential to cause phospholipidosis.
To elucidate the molecular identity of the changes, exact mass matching (±3 ppm) to the Lipid Maps (lipidmaps.org) and Human Metabolome (hmdb.ca) data bases was used. Not surprisingly, many of the annotations were for phospholipids. To increase confidence in the annotations, a combination of on-tissue MS/MS experiments and commercially available reference compounds was used. However, most of the prominently changed compounds could not be unambiguously identified by either of these methods, as they were assigned to isomeric pairs of phosphatidylglycerols (PGs) and bis(monoacylglycero)phosphates (BMPs). BMPs have been previously associated with DIPL, so it is likely that the majority of changes can be attributed to BMPs. To confirm this, a chromatographic method using a hydrophilic interaction liquid chromatography (HILIC) column and commercial standards of isomeric BMPs and PGs was developed. This method was able to clearly resolve PGs and BMPs by retention time, and showed that the mass spectrometric sensitivity for both lipids was similar, allowing semi-quantitative measurements. Subsequently, the remaining tissue from the high dose groups was homogenized and the lipid extracts analyzed with the established HILIC method, showing that for all PG/BMP isomers, most of the changes were indeed due to BMP increase.
In addition, the altered BMPs in plasma samples collected during the study (data not shown) could be detected. Plasma represents an amalgamation of changes from multiple organs; the level of changes was therefore less pronounced. However, the analysis of plasma samples allows easy observation of changes over time and ultimately may allow the transfer of the proposed markers to clinical samples. This study is an example of how the combination of MSI with other analytical methods (LC-MS/MS in this case) can help to develop its full potential. Direct analysis of tissue samples in combination with histopathology can reveal molecular changes that are less pronounced in extracts or body fluids, but once markers are known they can be measured more precisely and efficiently with more routine analytical methods.
Key insights
While this study highlights the strength of nontargeted, discovery style MSI investigations, it also demonstrates some of challenges of such studies. Unambiguous identification of proposed markers, especially in the case of isomeric compounds is still challenging, and may require additional methodology, such as LC-MS/MS. Time course studies are difficult to set up, as most tissue sampling can only be conducted during necropsy. In addition, while the image information provided by MSI can be a powerful asset, the automated analysis across many samples remains very challenging as the biological variations from section to section are often substantial.
Case 7. Imaging of PS-Containing Antisense Oligonucleotides in Rat Kidney and Liver With MALDI-MS
MSI is well known for its ability to detect small drug molecules and their metabolites as well as low molecular weight cellular metabolites. However, this technology can also be extended to larger molecular weight entities eg, therapeutic antisense oligonucleotides (ASOs) by using diagnostic fragments. An example of such an application is described by Romero-Palomo et al., 43 using the PS backbone (m/z 94.9362 Da) as a diagnostic fragment. They analyzed locked nucleic acid (LNA)-containing PS-linked ASOs on frozen sections of rat kidney and liver by MSI and found signals of ASO-drug material in the renal cortex for all 5 compounds with different signal intensities (Figure 8), which was in good agreement with parallel applied quantitative methods like LC-MS/MS using tissue homogenates. The PS modification is widely used in ASO drugs to increase both stability against nuclease degradation and binding to protein. A detailed methodological description can be found in Romero-Palomo et al. 43 It is noteworthy that this MSI method was performed in an untargeted way. Thus, it is generally applicable independently of the ASOs nucleotide sequences, chemical modifications, potential conjugations (eg, GalNAc), metabolism and/or degradation to shorter oligonucleotides(-mers), as long as the imaged molecular analyte contains a part of the phosphodiester backbone with the diagnostic PS modification.
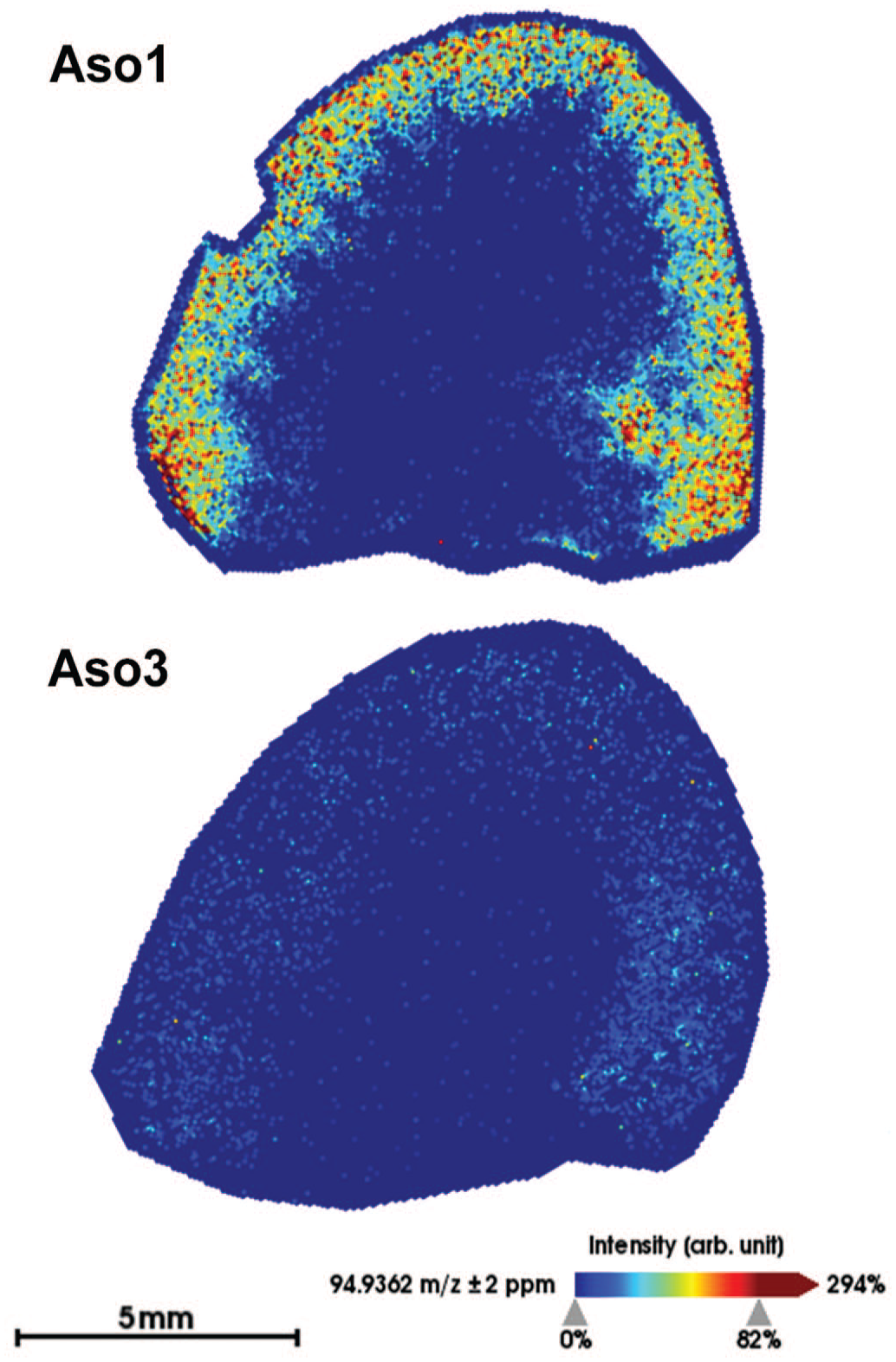
MALDI MS molecular imaging in rat kidney tissue showing the spatial distribution of m/z 94.9362, a fragment specific to phosphorothioate-linked oligonucleotides, administered to the animals. (Modified from Romero-Palomo et al. 43 ). ASO: antisense oligonucleotide. (Case 7)
Key insights
Despite their proven therapeutic value, some ASOs are associated with toxic effects that restrict their use. In this study, the authors aimed to characterize mechanisms of toxicity of tool compounds with different safety profiles, and tracing their accumulation and metabolism was a key component of these investigations. There is no gold-standard technique for quantifying ASOs in biological matrices, and several aspects should be considered when selecting the appropriate techniques including spatial tissue resolution, sensitivity, or differentiation between parent and metabolized compounds. MALDI MSI contributed in this study to determine that the degree of accumulation of ASOs in tissues is not necessarily associated with their safety profile.
Case 8. MALDI-MSI of Needle-Shaped Crystalline Material in Rat Intestine
During an exploratory 14-day oral (gavage) toxicity study in Wistar rats, needle-shaped crystalline material from unknown origin, was observed in the jejunum of all high dosed males. This needle-shaped material was mainly located in the lamina propria often at the tip of the villi and was associated with macrophage aggregates, including multinucleated giant cells, engulfing the crystalline material (Figure 9). MALDI-MSI analysis of these crystalline materials confirmed the presence of the parent compound, as well as its sodium and potassium adducts (Figure 10). Overlaying the hematoxylin and eosin (HE), optical image and MS data clearly showed a high degree of colocalization of these signals, confirming a direct link between the dosed compound and the observed needle-shaped crystalline material. Recently, intestinal lamina propria depositions from unknown origin associated with histiocytosis, were reported in association with metal deposits.36,44
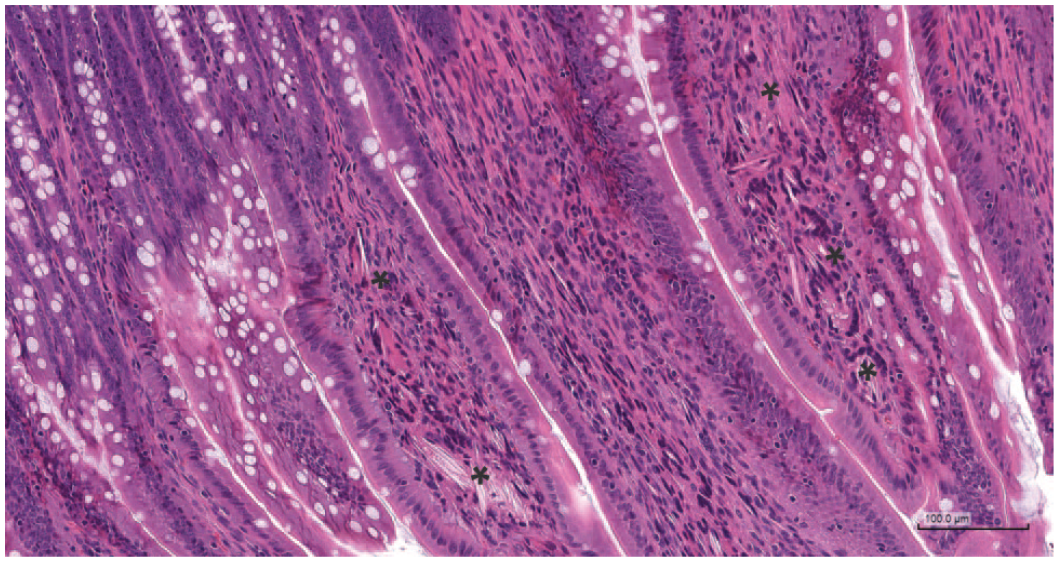
HE stain of rat jejunum, showing needle-shaped crystalline material (*) in the lamina propria, associated with multinucleated giant cells. (Case 8)
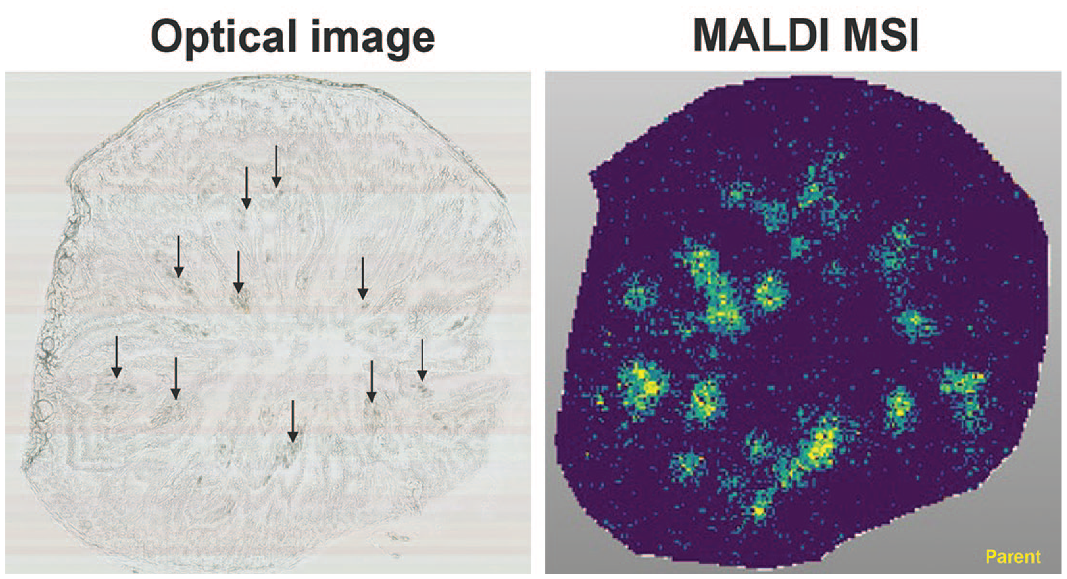
MALDI-MS imaging of rat jejunum. Showing (I) crystalline aggregates in apical villus tips indicated by black arrows (optical image), (II) the compound (parent; protonated form and potassium adduct), colocalized with the observed crystals. (Case 8)
Key insights
MALDI-MSI demonstrated a direct link between the (spray dried) compound and the needle-shaped crystalline material indicating intestinal absorption and precipitation of compound in the lamina propria.
Case 9. Multimodal Imaging of Healthy Livers to Identify Tissue Markers and Their Inclusion in Livers From Primary Sclerosing Cholangitis (PSC) Patients
Bile acids control liver homeostasis, liver disease and are often associated with liver toxicity. To allow in-depth investigation of functions of bile salts (bonding bile acids to potassium or sodium), which are important for signaling as well as detergent actions in healthy and diseased liver, the spatial distribution of bile salt species within the liver needed to be studied. Therefore, the aim of this study was to determine hepatic bile salt distribution and identify specific lipid markers that define the structural elements of the liver. To this end, MALDI-MSI was used to monitor the spatial distribution of bile salts and lipids in liver sections of rat, dog, and human liver samples with unaffected and cholestatic parenchyma. MALDI-MSI in negative ion mode (that is based on ionizing and imaging the distribution of negatively charged ions) showed the local presence of a variety of bile salts, predominantly taurine-conjugates, as localized patches of varying sizes (representing the bile ducts) throughout the liver tissue (Figure 11, panel IV). Specific molecular markers were identified for the connective tissue (phosphatidic acids, eg, [PA (18:0_18:1)-H]-), the liver parenchyma (phosphatidylinositols, eg, [PI (18:0_20:4)-H]-), and the bile ducts (hydroxylated-sulfatides, eg, [ST-OH (18:1_24:0)-H]-). (Figure 11, panel V, VI, and VII, respectively). One of these sulfatides (at m/ z 906.6339) was uniquely localized in a thin lining on the inside of the bile duct, colocalized with cytokeratins, and encased luminal bile salts (Figure 12). A similar distribution of the aforementioned sulfatide was observed, albeit in constricted ductular structures, in the liver of a patient with a mild clinical phenotype of primary sclerosing cholangitis (PSC). In contrast, sulfatides were virtually absent in the liver of patients with more severe PSC (Figure 13, panel X) showing a severe clinical phenotype, with (atypical) cholanoids (eg, the bile alcohol 5-cyprinolsulfate) abundant in the extra-ductular space and glyco(cheno)deoxycholic acid-3-sulfate localized to fibrotic connective tissue. The latter two molecular species were able to discriminate between healthy liver tissue (n = 3) and tissue from PSC patients with a severe clinical phenotype (n = 3). In conclusion, the distinct structural elements of the mammalian liver are characterized by specific classes of lipids. Based on these results, (hydroxylated-)sulfatides could be specific molecular markers of the bile duct. 19
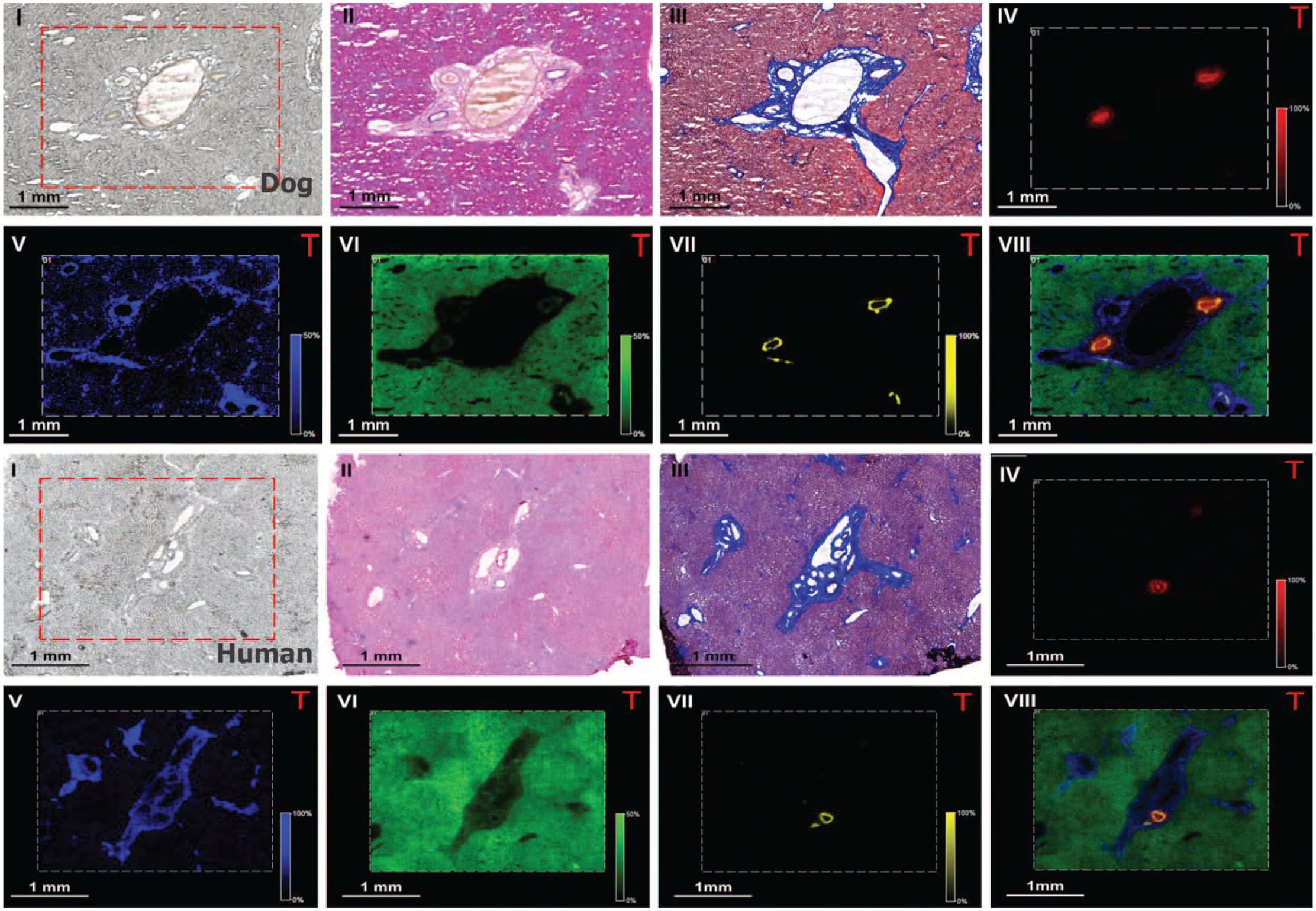
Multimodal imaging of healthy dog/human liver tissue at 15 µm spatial resolution (dog: first and second row; human: third and fourth row). (I) Optical image, (II) H&E, (III) Masson’s trichrome. MALDI-FTICR-MS images showing the distribution of selected molecular species in the (IV) bile duct lumen ([TCA-H]- at m/z 514.28); (V) connective tissue ([PA (18:0_18:1)-H]- at m/z 701.51); (VI) parenchyma ([PI (18:0_20:4)-H]- at m/z 885.55); (VII) bile duct ([ST-OH (18:1_24:0)-H]- at m/z 906.63) and (VIII) overlay of the selected species. Figure modified from Flinders et al. 19 (Case 9)
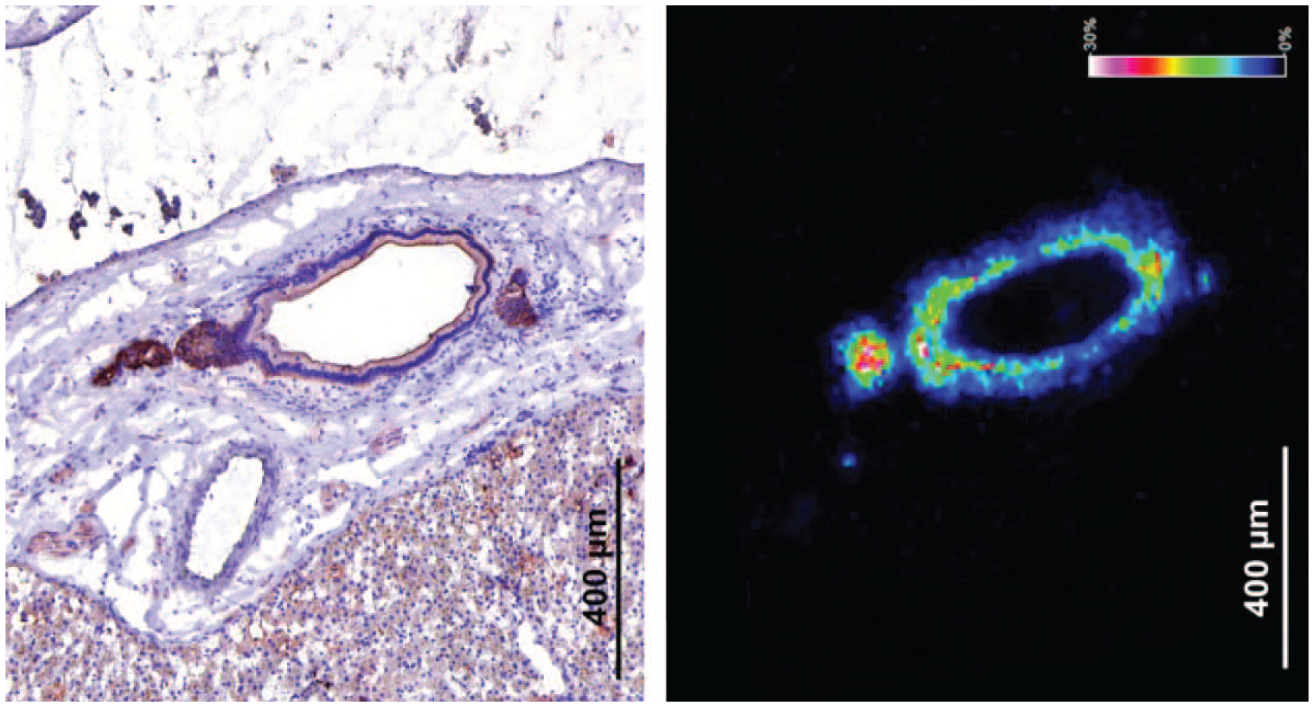
The sulfatide [ST-OH (18:1_24:0)-H]- was found to be uniquely localized in the differentiated bile duct epithelium, colocalized with cytokeratins, and encased luminal bile salts. Figure modified from Flinders et al. 19 (Case 9)
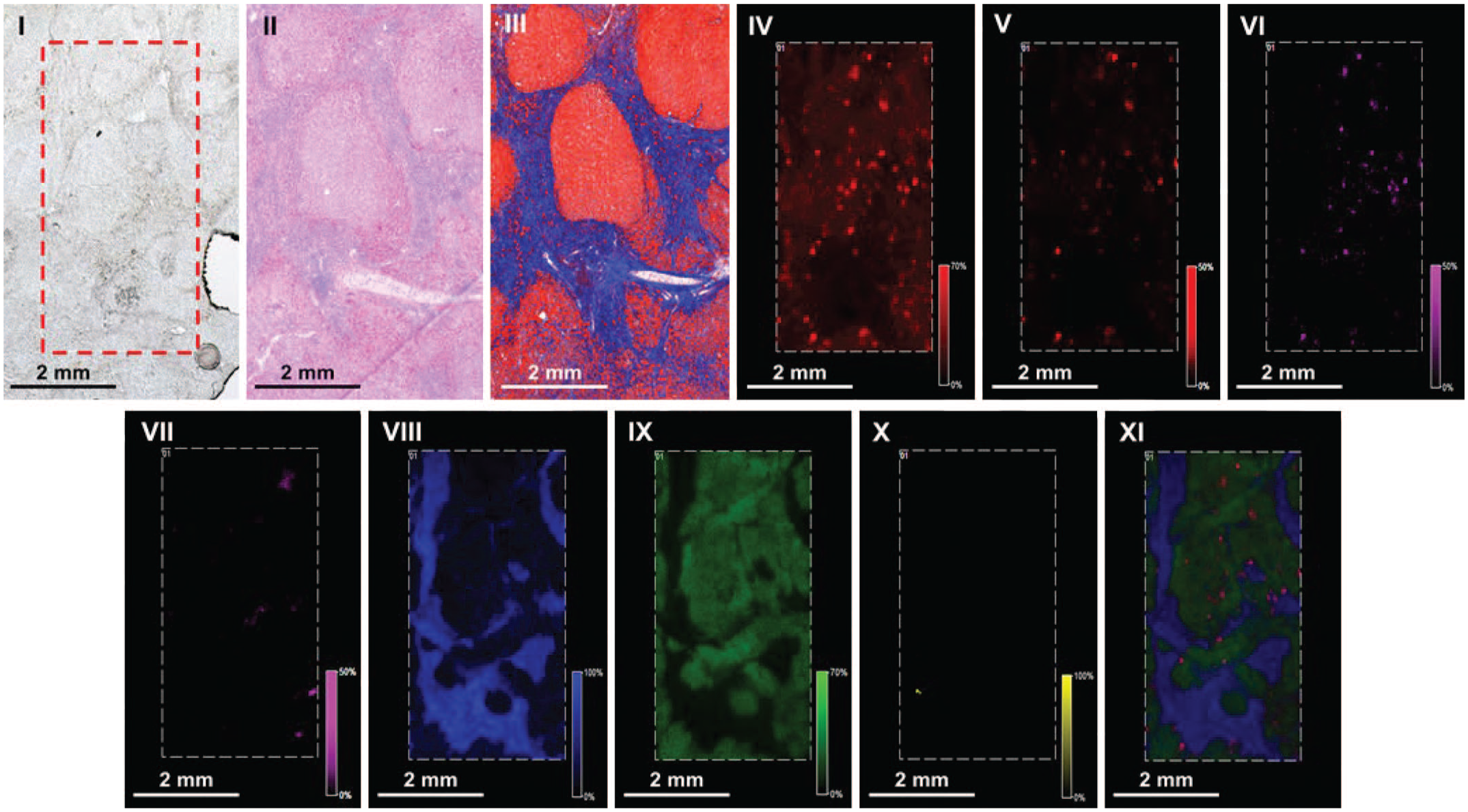
Advanced primary sclerosing cholangitis (PSC) patient had liver test abnormalities indicating cholestasis and liver injury. Histology: Nodular hepatocellular regeneration, diminished number/absence of interlobular bile ducts, portal to portal bridging fibrosis, bile ductular proliferation in portal areas and fibrous septa with mononuclear infiltrates. (I) Optical image; (II) HE; (III) Masson’s trichrome. MALDI-FTICR-MS: distribution of selected molecular species in the (IV) [TCA-H]- at m/z 514.2855; (V) sulfated bile acid ([M-H]-) at m/z 531.3004; (VI) bilirubin diglucuronide ([M-H]-) at m/z 935.3218; (VII) heme ([M-H]-) at m/z 615.1704; (VIII) [PA (18:0_18:1)-H]- at m/z 701.5128; (IX) [PI (18:0_20:4)-H]- at m/z 885.5504 and (X) [ST-OH (18:1_24:0)-H]- at m/z 906.6353. (XI) Overlay. Figure modified from Flinders et al. 19 (Case 9)
Key insights
MALDI-MSI allowed us to identify general healthy liver tissue markers and a specific hydroxylated-sulfatide (m/ z 906.6339) as biomarker for differentiated bile ducts.
Case 10. MALDI-MSI of Long-Acting Injectable Formulations
After a single intramuscular (IM) injection of 200 µL/kg body weight of an aqueous prodrug nano-/microcrystal suspension to male Wistar rats, the active drug plasma concentrations displayed biphasic flip-flop pharmacokinetics. It was hypothesized that this resulted from a transient absorption that was driven by the high intrinsic dissolution rate of individual prodrug nano-/microcrystals, followed by a slower release/absorption modulated by the physical behavior of the formulation depot (eg, reduced effective dissolution surface area due to particle agglomeration promoted by effusion of the vehicle) on one hand and physiological variables (eg, formation of macrophage rim around the long-acting injectable (LAI) depot, neovascularization and fibrosis) on the other. 15 The active drug absorption rate during the second, slower but predominating absorption process, appeared to be inversely related to the rate and extent of the IM inflammatory reaction, especially to the macrophage recruitment and infiltration of the Prodrug-LAI depot. To combine the molecular information (prodrug—active) with the localization in the injection site, MALDI-MSI was used. Histopathology of the IM injection site, one week after administration, showed the presence of delivery material surrounded by a rim of granulomatous inflammation. MSI demonstrated that the prodrug was restricted to the formulation depot, correlating with the delivery material at histopathology, while the active drug was mainly concentrated in areas of granulomatous inflammation (ie, infiltration of phagocytic macrophages) and in particular surrounding the LAI depot (Figure 14). The colocalization of the active drug with the macrophages, suggests a possible phagocytosis driven enhancement on the conversion of the prodrug into the active moiety by infiltrating macrophages. 15
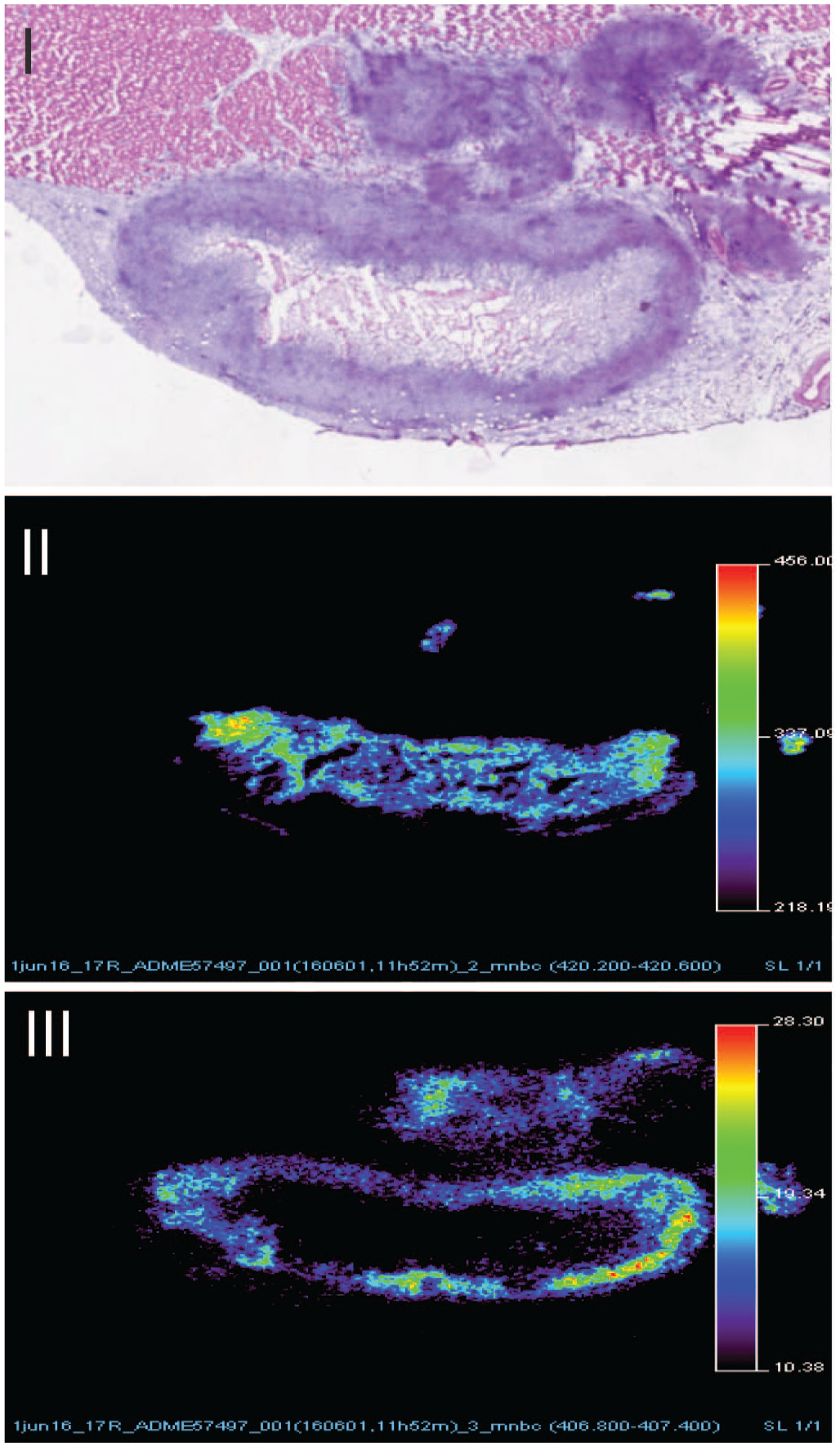
MALDI-MS imaging of IM injection site (rat). Showing (I) HE image of central microcrystalline depot surrounded by an inflammatory rim, (II) the prodrug in the center, and (III) the active drug in the inflammatory rim. (Case 10)
Key insights
MALDI-MSI demonstrated that the histopathologically observed central amorphous material in the injection site after one week, corresponded to the delivery material (prodrug) and that conversion to the active drug mainly occurred in the surrounding granulomatous rim, indicating an important role for the macrophages in the conversion of the prodrug to the active drug.
Case 11. MALDI-MSI on Mature Minipig Ovaries
In a pilot study, we wanted to evaluate the performance of the system for the analysis of several “difficult” analytes (eg, caffeine, prednisolone and estradiol). Here we focused on the detection of different steroid hormones, like estrogen, androgen, progesterone and their precursor, cholesterol, by using TimsTOF flex MALDI-2 (Bruker). Traditional, LC-MS analysis of these compounds is already troublesome due to several factors like, eg, their low ionization efficiency, their lipophilic character, their large dynamic range, their broad polarity range and the large amount of regio- and stereo-isomers. Hence, derivatization strategies are often employed to overcome some of these issues. Nonetheless, such analysis are time-consuming and require dedicated workflows to obtain robust and reproducible data. The spatial information is mostly lost. Hence, completing the analysis to also include this spatial info is desirable. Here, ovaries (one FFPE and one frozen) from several mature minipigs throughout different estrus cycle stages were collected. On-tissue derivatization of the steroid hormones using Girard’s T reagent prior to MALDI matrix spraying to increase their sensitivity was performed. Several steroids like cholesterol and progesterone showed a strong signal (Figure 15), other intermediates, like 17a-hydroxyprogesterone and estradiol, were also observed, indicating active steroidogenesis. In addition, a panel of lipid markers (eg, phosphoethanolamines, phosphoserines and phosphoinositols) were monitored to visualize some anatomic regions like the ovarian medulla containing large blood vessels, the cortical interstitium, developing follicles (primordial-, primary-, secondary- and tertiary follicles) and formation of corpus luteum. The latter is also a site of intense angiogenesis reflected by the peripheral heme signal in large corpora lutea, enabling the hormone-producing cells to obtain oxygen, nutrients and hormone precursors necessary for synthesis and release of progesterone. These results demonstrate the power of post-ionization with a TimsTOF flex MALDI-2, to visualize and localize a variety of steroid hormones and cholesterol, one of their most important pre-cursors, in maturing follicles and developing corpora lutea.
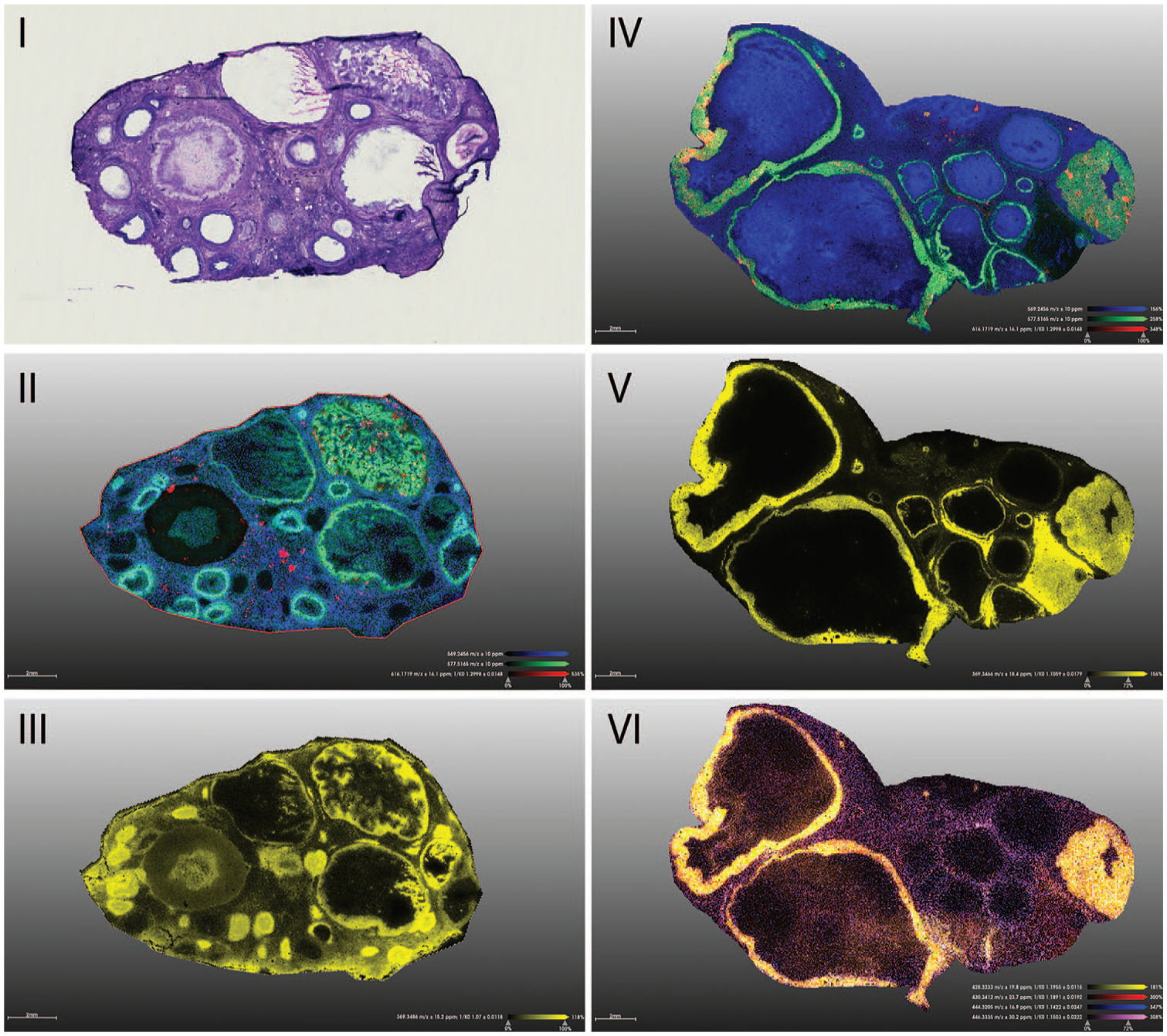
MALDI-MS imaging of 2 mature minipig ovaries. Showing (I) HE stain, (II and IV) lipid tissue markers (green) and heme (red), (III and V) cholesterol (yellow) and (VI) progesterone (yellow). (Case 11)
Key insights
TimsTOF flex MALDI-2 analysis on minipig ovaries confirmed the ability to detect and localize certain steroid hormones and their precursor cholesterol in this proof-of-concept study.
Case 12. Untargeted Biomarker Analysis for Lung Toxicity of Intravenous ADC in Cynomolgus Monkeys by MALDI-MSI
The aim of this study was to evaluate ADC induced lung toxicity, when administered to cynomolgus monkeys as fractionated doses. For this purpose, MALDI-MSI was applied as a complementary technology to the microscopic evaluation identifying potential drug-related metabolites and/or biomarkers, present in lung tissues, and colocalized with microscopically confirmed lesions.
Four cynomolgus monkeys, one male and one female per group, received three administrations of vehicle control or ADC at 0.62 mg/kg, on days 1, 4, and 7 (30-minute intravenous infusion followed by a saline flush). The animals were euthanized 4 weeks after the last administration. At necropsy, there were no macroscopic findings in the lungs, however the treated male presented a 2-fold increase in the absolute and relative lung weights, correlating microscopically with the marked interstitial edema associated with slight mixed cell inflammatory cell infiltration. The female presented similar changes, albeit at a minimal severity. Only male groups were measured. Lung samples from control animals and from treated animals with marked microscopic findings and a 2-fold increase in lung weight were evaluated by MALDI-MSI.
Tissue sample preparation and MALDI matrix application were performed according to protocol published by Hinsenkamp et al. 27 The tissue slices were measured on a Q Exactive Plus mass spectrometer, equipped with an atmospheric MALDI source (AP-SMALDI, Thermo Scientific, Transmit). Ion images were acquired in the positive mode at a lateral resolution of 50 × 50 µm and a pixel size of 40 µm. The mass range was restricted to 300 to 1000 Da, and the data were calibrated using a 2,5-dihydroxybenzoic acid (DHB) matrix cluster as internal lock mass (m/z 444.09252). Data evaluation was conducted by in-house developed software applications, and the biomarker candidates were assigned as described previously 31 (Figure 16).
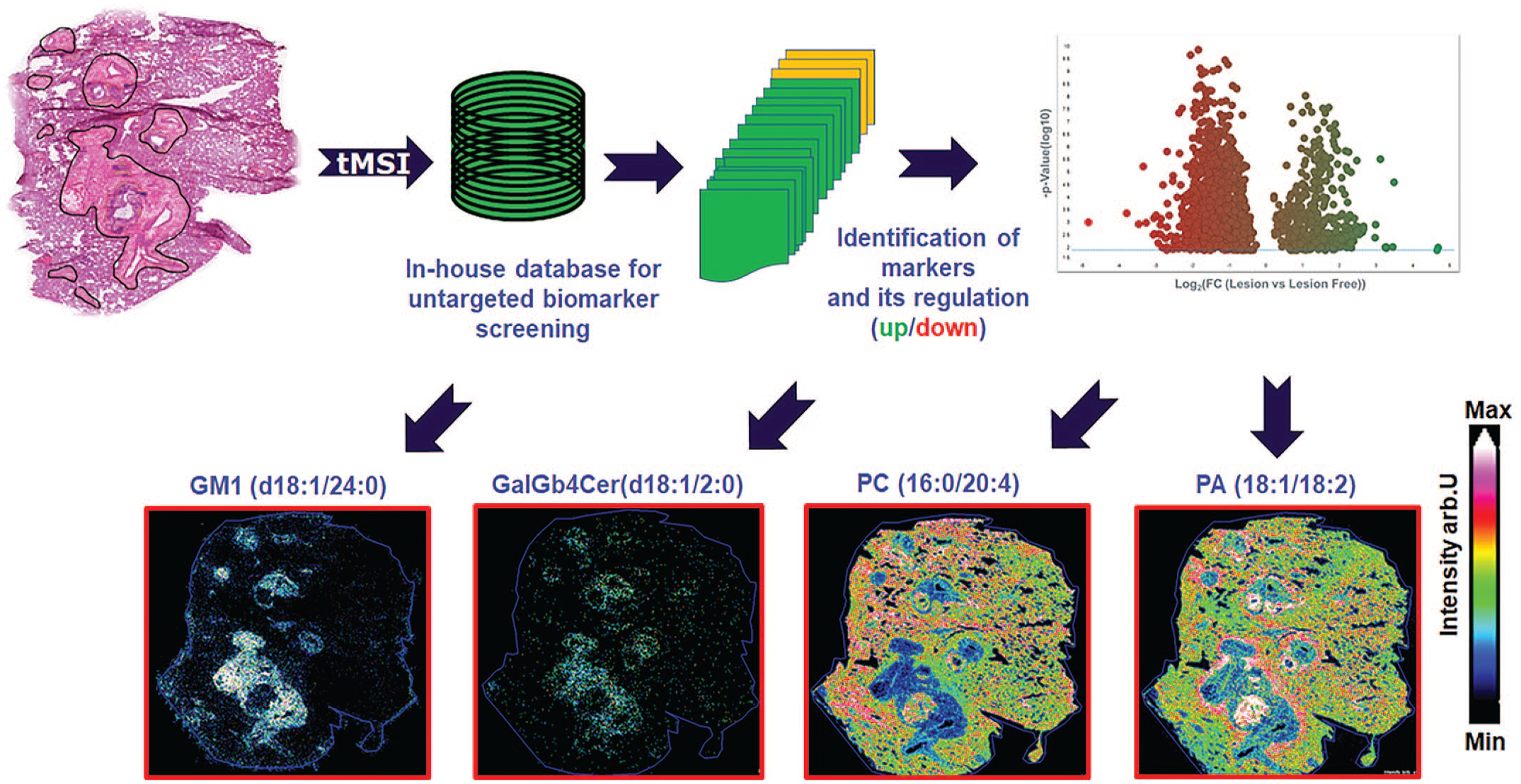
General MALDI-MSI strategy to identify potential biomarkers which colocalized with microscopically confirmed lesions. tMSI: tissue mass spectrometry imaging; GM1: Ganglioside GM1; GalGb4Cer: Globoside GalGb4Cer; PA: phosphatic acid; PC: phosphatidylcholine. (Case 12)
In total, more than 3000 biomarker candidates were assigned. An unsupervised biomarker analysis was performed to evaluate major differences between control and test article-treated and between lesion and lesion free environment.
Major effects indicate an increase in sphingolipids, reflecting a good correlation with the area of alteration such as Ganglioside GM1 (d18:1/24:0) and Globoside GalGb4Cer (d18:1/2:0). These sphingolipid candidates are described to be highly abundant in neutrophils 51 and showed strong correlation with phosphatidylinositol-3,4,5-trisphosphate, a signaling molecule involved in neutrophil activation. 33 These molecular observations are in agreement with histopathological observations of infiltrating immune cells in lung tissue lesions in the animal after ADC treatment.
Besides sphingolipids, glycerophospholipids (GPL) represent an important lipid class, with diverse functions. GPLs are major components of the cellular bilayer membrane and play an important role during cell regeneration.21,52 The distribution for glycerophospholipids such as phosphatic acid (PA) (eg, 18:1/18:2) and phosphatidylcholine (PC) (eg, 16:0/20:4) was reflected in an increase in test article-treated animal compared with the control and a strong decrease within the lesions, suggesting a disturbed tissue regeneration process.
Key insights
Standardization and rigorous quality control standards have enabled the establishment of a robust Ultrahigh-Mass Resolution (UHR)-MALDI imaging-MS workflow to evaluate not only the distribution of pharmaceutical drugs and their putative metabolites but also to provide spatial driven untargeted biomarker information based on a specific manual curated internal MALDI-MS biomarker database. This makes it possible to monitor and label thousands of endogenous metabolites in a single measurement, to assign them to activated or disrupted metabolic pathways, and finally, to improve our understanding of the mechanism of action of the cytotoxic effect of a drug. 31 This technology was successfully applied to correlate ADC lung toxicity findings in cynomolgus monkeys with endogenous biomarker distributions and regulation, adding an additional level to histopathology by demonstrating accumulation/depletion of metabolite species specific to the lesion areas.
Case 13. MSI for Ocular Toxicity Assessment
MSI has recently attracted significant attention in ophthalmic drug discovery, particularly for preclinical research.16,38 It enables rapid and effective assessment of ocular treatments in terms of efficacy, safety, and toxicity. This technology can simultaneously examine a drug candidate’s biodistribution 32 and its associated metabolites, along with potential changes in efficacy or toxicity biomarkers such as peptides, lipids 47 and metabolites. In addition, MSI’s high spatial resolution offers precise localization of compounds within the eye’s anterior and posterior segments, which consist of several histologically different structures of only a few micrometers in size. Consequently, MSI has emerged as a preferred method for evaluating new targets and understanding the mechanisms of drug action in ocular diseases, including age-related macular degeneration (AMD), 2 uveal melanoma, 10 glaucoma, 6 and diabetic-related retinopathy or macular edema (DME).
Combining traditional staining and IHC with advanced MSI provides potent tools for examining the distribution of various compounds, such as amphiphilic eye drop excipients that are known for their harmful effects. 6 This combination proves particularly valuable in pharmacological and toxicological preclinical studies. 12 With the high spatial resolution offered by MSI, researchers can track drugs, their metabolites, and potential biomarkers to evaluate the efficacy and/or toxicity of treatments. MSI can deliver dependable safety and toxicity data regarding ocular drug distribution or biomarker modulation within small histological regions of the eye. This capability ensures precise and detailed insights, enhancing the overall assessment of ocular treatments.
Such an application has been recently described by Hamm et al. 26 MSI was combined with classical histopathological examination (hematoxylin and eosin [HE]) and ultrastructural change assessment (electron microscopy) to study a highly selective inhibitor of MERTK, a receptor tyrosine kinase (RTKs), playing an essential role in homeostasis of the retina via efferocytosis. Retinal degeneration has been linked to loss-of-function of MERTK. The retina layers consist of multiple compartments with unique and specialized properties. Consequently, MSI was used to identify if the compound was distributed in those retina layers to assess if they could be impacted by MERTK inhibition. Figure 17 shows the multiscale analysis of a treated rat eye section from whole tissue (top panel) to small regions of the retina (bottom panel). This example demonstrates the advantage of MSI to avoid misinterpretation of the “real” amount of compound in the eye as shown in Figure 17F when comparing whole eye versus posterior region. This is particularly true when measuring drug exposure by bulk LC-MS. There, a high lateral resolution is needed to differentiate drug exposure in retina layers. Those layers can be defined thanks to lipid biomarkers, as illustrated by Anderson et al. 1 The pyridinium bisretinoid, N-retinylidene-N-retinylethanolamine or A2E (Figure 17D and G) is commonly used to define the retinal pigment epithelium (RPE) but also to evaluate some local functional disorders. In this study, the compound was found to be mainly located within the RPE, which was in line with lesions identified at the interface between RPE and the photoreceptor layers by histopathology and EM. In addition, those types of data are crucial to evaluate the Blood-Retinal Barrier (BRB) crossing of drugs reaching the eye to assess their potential toxicity. It can also be used to trigger additional diagnostic tests such as an electroretinogram (ERG) measuring retinal disorders.
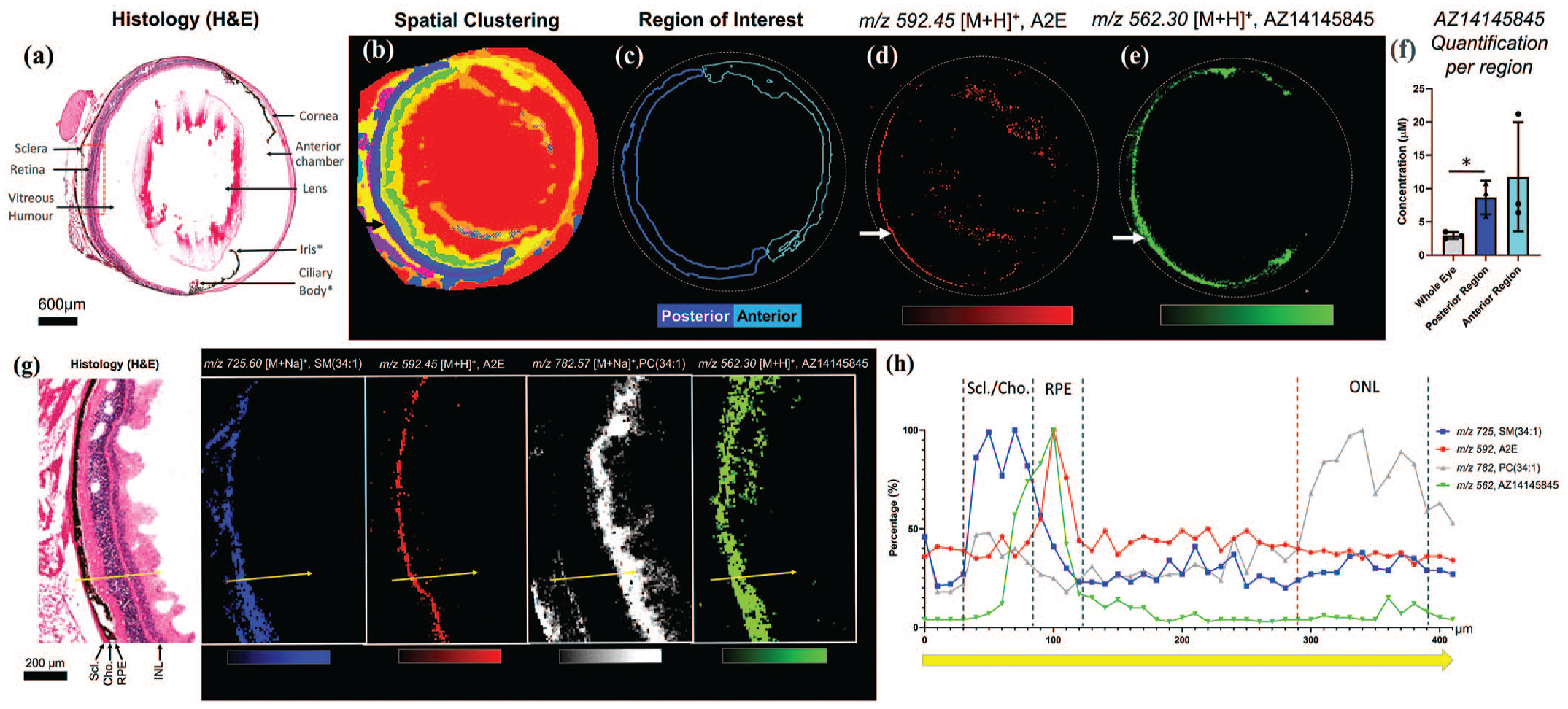
MALDI Mass Spectrometry Imaging analysis of whole eye sections and retina layers from a mouse treated with AZ14145845 (MERTK inhibitor). (A) Histological staining (HE) of adjacent eye sections with corresponding histological features. (B) Spatial segmentation based on MSI data at 20-µm spatial resolution. (C) Region of interest describing the posterior and the anterior part of the eye used for the quantification of AZ14145845 and based on spatial clustering. (D) N-retinylidene-N-retinylethanolamine (A2E) molecular image. The white arrow shows the individual retinal layers. (E) AZ14145845 molecular image from MSI. (H) Quantification of AZ14145845 in the whole eye, posterior and anterior segments in µM. (G) Histology of the retinal layers with corresponding 10-µm spatial resolution molecular images of Sphingomyelin, SM(34:1), A2E, Phosphatidylcholine PC (34:1) and AZ14145845. (H) Line scan data from the region shown by the yellow arrow in (G) and selected tissue of interest labels. Adapted from Hamm et al. 26 (Case 13)
Key insights
The use of MSI in drug-induced ocular toxicity provides detailed spatial and molecular information, within ocular tissues. While MALDI and LA-ICP can be destructive (inducing extensive tissue drying, affecting morphology), DESI MSI is a nondestructive technique (preserving the tissue integrity for further histopathology examination): this is particularly important in ocular studies where sample availability might be limited. This positive view is supported by numerous studies demonstrating the technique’s efficacy in providing valuable insights into ocular pharmacology and toxicology. Finally, the eye is peculiarly situated, and it is compartmentalized in aqueous humor and vitreous body, separated by a barrier represented by a lens and iris with ciliary body: these compartments escape the standard body vascularization, implying a peculiar biodistribution within the same organ. To summarize, the utilization of MSI in ocular pharmacokinetic studies as well as to investigate toxicity mechanisms nowadays is challenging the position LC-MS as method of choice thanks to an improved spatial resolution, a better sensitivity of detection of wider class of molecules and a more reliable instrumentation together with optimized data analysis solutions.
Discussion
Applications and Advantages of MSI
MSI has several versatile applications due to its species-agnostic nature, enabling the translation of molecular biomarker assessments from preclinical species to humans. This technology is especially advantageous for analyzing complex and heterogeneous tissues and organs, such as brain, skin, kidney, liver, or tumors. Unlike traditional bulk MS methods, MSI provides better lateral resolution, spatial distribution and avoids issues with metabolite or drug degradation product detection, as it does not require labeling. This makes it superior to quantitative whole-body autoradiography and qualitative microautoradiography.
Moreover, MSI can be integrated with other imaging modalities, including spatial proteomics (such as multiplex immunofluorescence, IHC, and IMC) and spatial transcriptomics (such as Visium and GeoMx). The ease of feasibility tests, especially when reference compounds are available and target concentrations can be estimated, allows MSI to be employed whenever image information and molecular distribution can provide valuable insights. In certain cases, it serves as a rapid method to analyze solid samples using MS.
Selecting the Appropriate MSI Technology
The choice of MSI technology depends on the specific requirements and available equipment. Both MALDI and DESI MSI are similar in capabilities, with the selection often based on the availability of the technology. The type of MS analyzer coupled to the ion source—such as high-resolution analyzers, linear time-of-flight (TOF), or triple quadrupole (QQQ)—also influences the choice. SIMS differs significantly in instrumentation and use cases. It is more suited for smaller molecules and offers higher resolution, but is more destructive and requires higher vacuum conditions, making it less user-friendly. Other technologies, LESA or LA-ICP MS, are less commonly used.
Feasibility and Detection Capabilities of MSI
MSI technology allows the detection of various molecules, including endogenous biomarkers, parent compounds and their metabolites. The main challenge in MSI concerns the detection and identification of unknow metabolites (eg, in drug discovery) or biomarkers (eg, in early or clinical-stage development of compounds), as mentioned in the introduction about targeted and untargeted investigations.
Several factors affect the successful application of MSI in pathology, but the authors would like to emphasize that tissue concentrations of around 10 µg per gram are needed, and samples are typically prepared on ITO-coated glass slides for MSI and Superfrost Plus glass slides for special stains; in addition, the feasible resolution varies with the technology used: approximately 10 µm for MALDI, 30 µm for DESI, and less than 1 µm for SIMS. 49 Frozen tissue sampling is preferred to maintain molecular integrity and localization.
Time Required for MSI Analysis
Instruments for MSI typically sample sequentially, pixel by pixel, with scanning rates ranging from 1 to 30 pixels per second depending on the instrument and settings. The actual acquisition time is determined by the area and pixel density of the sample. For example, a rat kidney section at medium resolution requires approximately 1 to 3 hours for measurement. Overall, processing a single slide, including the entire workflow, can usually be completed within a day. For untargeted studies, multiple replicates are needed, increasing the time required for measurements and data handling/analysis.
Costs Associated With MSI
The consumable costs for standard MSI experiments are relatively low, ranging from 10 to 20 euros per slide. However, costs can increase with on-tissue chemistry requirements. The instrumentation itself is expensive, costing over 1 million euros, with maintenance costs exceeding 50,000 euros annually. Despite these high costs, small-scale or feasibility experiments are relatively easy to run once the instrumentation is in place. Data storage requirements are also significant, often requiring several terabytes per year. MSI instrumentation, particularly axial MALDI-TOF-MS instruments, are more cost-effective and tailored for smaller scale workflows, (eg, Bruker neofleX bench top instrument, or BioTyper and Vitek MS benchtop instruments—normally used for bacterial identification), allowing high-quality untargeted and targeted spatial omics on a budget.
Importance of MSI Expertise
Successful implementation of MSI goes beyond the access to advanced instruments and relying on pathologists’ insights. MS scientists with specialized technical skill sets in mass spectrometry, data interpretation, and tissue preparation are needed to ensure the proper application of MSI. The interdisciplinary nature of the collaboration between pathologist and MS scientists should be considered as an opportunity to broaden the spectrum of knowledge, aiming to address a scientific question, rather than a limitation. This necessary expertise includes the aware selection of the different MSI methods (eg, between MALDI, DESI, or SIMS), the ability to tune instrumentation to meet specific requirements, the management of complex data sets that require computational methods for accurate molecular imaging and quantitative analysis. Meaningful MSI readouts are complex and cannot be easily obtained without trained personnel who understand sample preparation, ionization techniques, and spatial data analysis. Without the proper expertise, there is a risk of generating data that could be misinterpreted, impacting negatively the outcomes of the project.
MSI is a highly specialized technique that is not routinely used across all mass spectrometry-equipped laboratories. While many laboratories may possess general mass spectrometers, MSI requires additional dedicated instrumentation and custom workflows. As a result, MSI remains a technique reserved for specialized research institutions or highly focused studies, limiting its availability to those with the necessary infrastructure and expertise. Small and medium-sized institutions can consider outsourcing MSI as a service, albeit this might be expensive, especially when high-resolution imaging or large-scale studies are required: the cost includes not only the technical expertise needed to operate the instruments but also the advanced data processing and interpretation services; this expense can represent an economical barrier for research projects with limited funding, therefore it is essential to carefully weigh the cost-benefit ratio of MSI readouts when planning a study. their studies.
Role of the Pathologist in MSI
Pathologists play a crucial role in MSI by initially identifying microscopic lesions and suggesting MSI analysis as second tier to further characterize the lesion or mechanism of action to support a project. Their expertise in systems biology, tissue morphology, sampling, and orientation is essential for the successful outcome and interpretation of MSI results, in particular when dealing with frozen tissues as opposed to FFPE tissues that are commonly available from toxicity studies. They can also review HE stained frozen consecutive sections prior to MSI and are critical in selecting the appropriate regions or samples for analysis by MSI. Pathologists play multiple roles in MSI, beyond just the technical aspects: the understanding of disease mechanisms and how various toxins affect biological tissues is essential for interpreting MSI data, as they can correlate the molecular findings with pathological changes observed in tissues; in addition, pathologists can translate findings from animal models to human diseases, bridging preclinical and clinical studies. Finally, the integration of MSI data, not only with anatomic and clinical pathology but also with molecular biology, is provided by pathologists.
Value of MSI for Pathologists
MSI provides a label-free technique that combines molecular information with spatial localization within tissues. Unlike traditional technologies such as whole-body autoradiography, which require radiolabeling and do not differentiate between parent compounds, metabolites, and biomarkers, MSI allows the simultaneous detection and identification of a large number of molecules, including endogenous molecules, like tissue biomarkers and compound breakdown products. Images can be generated for every molecule within the measured mass range and ionization mode and can be correlated with tissue morphology assessed by HE, IHC, or special stains and support the interpretation of findings. MSI provides complementary molecular information, that may not be highlighted by histological or IHC staining, as well as not be sufficiently characterized by other multiplexing techniques that only provide partial information.
Research Questions Addressed by MSI
MSI is a powerful tool that can address a variety of critical research questions in biomedical and pharmaceutical sciences. One of its primary applications is the ability to link microscopic lesions to specific compounds or their metabolites, providing insights into disease mechanisms and drug effects at the molecular level. In addition, MSI can determine the chemical composition of crystalline materials within tissues.
MSI is also instrumental in analyzing the distribution of compounds within tissues, such as tracking the localization of inhaled substances within the lungs or the dispersion of drugs following intra-tumoral administration. This feature is vital for assessing the efficacy and safety of therapeutic interventions. Furthermore, MSI facilitates the study of prodrugs by revealing where, when, and how these compounds convert into their active forms, particularly in the case of subcutaneous or intramuscular long-acting injectables. 11
The application of MSI across different classes of compound, from small molecules to large biologics, is linked to some technical considerations that are well covered by the scientific literature and are out of this article’s purpose.
Comparative analysis of healthy versus diseased tissues using MSI allows researchers to identify potential biomarkers, which can lead to the development of diagnostic tools and targeted therapies. This comparative approach helps in understanding disease pathology and the molecular changes associated with various conditions, thereby contributing to advancements in personalized medicine.
MSI can also guide experimental design, often requiring separate studies or prior knowledge of the compound and a preference for frozen tissues over FFPE samples. FFPE samples, although increasingly accepted, offer suboptimal results and significantly lower sensitivity compared with frozen samples: while for some analytes both type of tissue fixation are comparable, for others characterized by chemical instability formalin might bias the quality of the results; in addition, while traditional mass spectrometry has a proven track of success when using FFPE-tissue samples, the evidence in MSI application is less consolidated.
Future Trends in MSI
MSI is rapidly evolving, with several emerging trends potentially revolutionizing its applications in pathology and in general in biomedical research. The current review is destined to come to an age; in the meantime, it is interesting to draft the expected future perspectives and their impact on pathology.
Combining MSI with single-cell omics, such as single-cell RNA sequencing (scRNA-seq) and proteomics, is an emerging field of MSI applications that might help understanding the heterogeneity of cell types within tissues. Such single-cell technologies have already enhanced gene expression and proteome analysis. When applied to MSI the localization of molecules within the tissue architecture could be further characterized. For example, this integration could allow pathologists to correlate single-cell gene expression profiles with spatial molecular patterns, or it could identify molecular alterations in specific cells within a diseased tissue, enhancing our understanding of disease progression and treatment responses on a cellular level. Nevertheless, the technical challenge of aligning single-cell data with MSI spatial resolution and the need for further powerful computing remain a major challenge.
As mentioned in the data management paragraph above, machine and deep learning as well as artificial intelligence (AI) are applicable to automate MSI data analysis, especially for high-throughput pathology applications: these advancements could speed up diagnostic processes, complementing the traditional histopathology readouts, and at the same time improving reproducibility while reducing human error. Nevertheless, to achieve a point when machines could help, for the training phase large and high-quality annotated data sets are needed to train models effectively, as for any other application; in addition, such models should be validated across different laboratories and instruments.
Recent advancements in SIMS are pushing the spatial resolution of MSI below the micrometer scale, enabling the detailed study of molecular distributions within individual cells or subcellular structures: despite requiring highly specialized equipment and extensive sample preparation, the increased spatial resolution of MSI will allow pathologists to study subcellular molecular alterations, for example, in pathologic conditions or in response to drugs administration.
The increasing interest in multiomics including MSI could provide a more comprehensive molecular picture of pathological tissues, improving the ability to identify useful biomarkers. Integrating multiomics data though, represent a significant computational challenge, but scientific computing solutions are evolving even faster than MSI technologies, and this challenge might be only hypothetical. For example, MSI is already compatible with traditional techniques such as IHC: combining MSI with IHC enables the simultaneous localization of molecular information and specific proteins of interest within tissue samples, allowing pathologists to correlate molecular information with protein expression and morphological features, even though the co-registration and data integration between different techniques remains technically complex, and very hard to be standardized.
Another emerging area of MSI research is its application in studying the microbiome, and especially the spatial distribution of microbial communities within tissues. MSI can detect metabolites and biomolecules secreted by bacteria or fungi, providing insights into host-microbe interactions, but a very high specificity and sensitivity in complex tissue environments is difficult to achieve, and further refinement of MSI protocols and data analysis methods are still needed.
As last applications on the horizon, researchers are exploring the use of MSI in real-time during surgeries, especially in oncology, and MSI is raising an increasing interest in personalized medicine by enabling the spatial mapping of biomarkers associated with drug response or disease prognosis, but these goals are not expected in the near future.
Footnotes
Acknowledgements
The authors acknowledge Fabienne Battilana for the medical writing and editing support; Andreas Brink and Bernd Steinhuber for their valuable contribution to the case about an antisense oligonucleotides; Constanze Holz, Chelsea J. Nikula, Jens Riedel, Franck Chanut, and Thomas Bartels for the material concerning the case about an antibody-drug conjugate; Farid Jahouh, Kathleen Allaerts, and Raffaella Capobianco for the data and material on the minipig ovaries and rat intestines; Bryn Flinders for the experimental support in the multimodal imaging of healthy livers in relation to primary sclerosing cholangitis; and Ronald de Vries for his contribution to the long-acting injectable formulations.
Author Contributions
Authors contributed to conception, introduction, discussion, and conclusion (AS, BL, BS, CC, CT, EV, FRP, GH, MB, MVH, RV, SWY, TL); application case examples (AS, MB, FRP, MVH, RV, CC, GH); and reference management (FRP). All authors reviewed and approved the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
