Abstract
Pathology, a fundamental discipline that bridges basic scientific discovery to the clinic, is integral to successful drug development. Intrinsically multimodal and multidimensional, anatomic pathology continues to be empowered by advancements in molecular and digital technologies enabling the spatial tissue detection of biomolecules such as genes, transcripts, and proteins. Over the past two decades, breakthroughs in spatial molecular biology technologies and advancements in automation and digitization of laboratory processes have enabled the implementation of higher throughput assays and the generation of extensive molecular data sets from tissue sections in biopharmaceutical research and development research units. It is our goal to provide readers with some rationale, advice, and ideas to help establish a modern molecular pathology laboratory to meet the emerging needs of biopharmaceutical research. This manuscript provides (1) a high-level overview of the current state and future vision for excellence in research pathology practice and (2) shared perspectives on how to optimally leverage the expertise of discovery, toxicologic, and translational pathologists to provide effective spatial, molecular, and digital pathology data to support modern drug discovery. It captures insights from the experiences, challenges, and solutions from pathology laboratories of various biopharmaceutical organizations, including their approaches to troubleshooting and adopting new technologies.
Keywords
Introduction
Pathology laboratories in research and development (R&D) units of biopharmaceutical companies and Contract Research Organizations (CROs) operate with workflows, technologies, regulations, and collaborations tailored to their business portfolio and functional areas. These labs are well-established and routinely support drug discovery and development with therapeutic target identification and validation, efficacy and safety testing, translational biomarker development, molecular mechanisms, and exploring novel investigative techniques. Through meticulous laboratory design and strategic operations, with state-of-the-art instrumentation, skilled personnel, scientific rigor, quality, and regulatory compliance, these laboratories have shaped modern health care practices by improving efficacy and safety of therapeutics.93,148,165,187,199
Modern drug R&D entails emergent novel therapeutic modalities such as cell, nucleic acid, and gene therapies among others. 30 Inherent to new modalities are research questions that demand new ways of answering them, including questions related to target expression, pharmacokinetics and pharmacodynamics (PDs), efficacy, and toxicity risk prediction. In this context, the Modern Pathology Laboratory (MPL)135,164 is uniquely poised to leverage multidimensional, high-plex tissue-based (spatial) molecular technologies4,5,15,26,45,55,64,71,97,126,133,137,140,151,160,167,178,184,190 combined with whole slide imaging, and digital and computational analytics1-3,6,10,20,37,56,70,74,77,84,89,90,100,112,113,131,143,144,153,157,166,203,211 to deliver biology insights. Accordingly, biopharmaceutical R&D pathologists are required to advance drug development by adapting to, and adopting, such novel approaches while ensuring rigor and excellence in data generation.
This article provides an overview of the traditional R&D pathology laboratory activities and specifics of the MPL concept and shares considerations for the implementation of the next generation of molecular approaches and for the integration of multimodal data visualization and downstream analytics platforms and pipelines. As an example, the current state of spatial molecular biology (ie, spatial “-omics”) allows for the simultaneous detection and visualization of high-plex transcriptomic and proteomic signatures on a single slide.5,45,26,71,126,137,140,151 Evolution in this field brings new challenges to the traditional pathology laboratory and to pathologists. First, prior to investing in any novel expensive technologies, the pathologist must become familiar with the specifics and performance of an extensive repertoire of high-plex spatial platforms. Second, relevant lab personnel expertise needs to be matrixed into multidisciplinary teams that can implement and integrate fit-for-purpose analytical pipelines to generate multidimensional data. Last, a digital repository infrastructure must be built and maintained for accessing, visualizing images, image co-registration across modalities, and interrogating their pertinent data sets with full sample traceability.
The goal of this article is to raise awareness of how the biopharmaceutical industry evolution has shaped the scientific demands for pathology labs and to inspire the current and future generation of lab leaders to rise to the occasion and reimagine the MPL to future-proof the discipline of pathology and augment its impact in the industry setting. In addition, we hope these perspectives will be helpful in the training of pathology residents and other pathology scientists entering the biopharmaceutical sector or established practitioners seeking to update their knowledge base.
Evolution of the Research and Development Traditional Pathology Lab into the Modern Pathology Lab
Over the last decade, the field of pathology has been propelled forward by significant advancements in laboratory technology and analytical tools pertaining to multiplex detection of targets in tissue sections and to digital and computational analyses.1,10,37,55,70,74,77,84,90,131,153,166,167,178 The depth of information derived from each sample and the throughput of samples analyzed have been significantly scaled up by these advancements.5,71,77,190 Below, we highlight the changes that helped traditional R&D pathology laboratories transition into molecular and digital anatomic pathology labs in the industry setting, referred to herein, as the “MPL.” Figure 1 provides an overview of the sample workflow and pathology-oriented activities of the MPL.
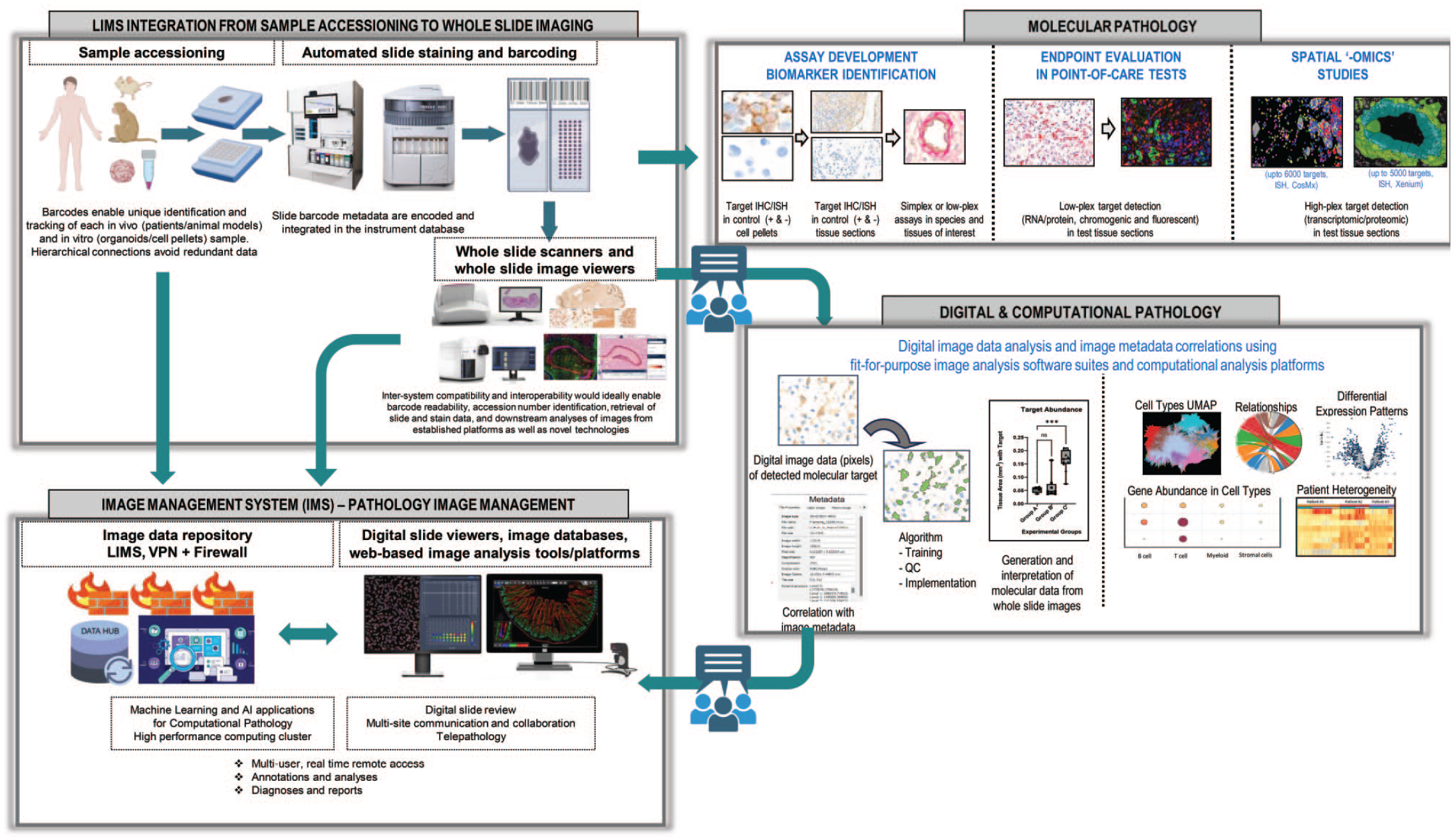
Overview of workflows and team expertise of pathology-oriented activities of the MPL. This figure was created with BioRender.com.
The Histological Specimen Continues to Remain the Foundation From Which the Modern Pathology Laboratory Derives Data
Histology laboratories, which are specialized laboratories that receive, process, and stain specimens on glass slides, remain at the core of MPL. The MPL remains equipped with the typical standard histology laboratory instrumentation, albeit with ever higher level of automation, and staffed with histotechnologists tasked with processing and sectioning tissue samples, developing, and performing tissue assays. Despite the rapid evolution of noninvasive techniques such as circulating tumor DNA (ctDNA) analysis, high-resolution radiological imaging, and real-time live cell imaging, tissue-based assays remain the benchmark for confirming the results obtained from these emerging technologies. The routine histochemical stain of hematoxylin and eosin (H&E) remains the standard procedure that enables histomorphologic evaluation of tissues, usually performed on formalin-fixed paraffin-embedded (FFPE) tissues, cell pellets, or organoids.21,83,145,162,163 Organoid models, primary cell cultures, and cell lines can partially recapitulate the morphologic, molecular, and mechanistic features of various organs in physiologic or pathologic states. Histologic characterization of these new models necessitates collaboration between research and pathology teams. Current progress in artificial intelligence (AI) has led to the availability of several “virtual” stains that are purported to eventually replace actual histochemical or immunohistochemical stains.12,101,147 However, such advanced computational techniques still require quality tissue sections.
Tissue Biobanking is a Vital Resource in the Modern Pathology Laboratory as It Enables Access to Quality-Controlled, Well-Annotated Normal and Disease-Specific Tissues
Surgical biopsy and autopsy samples from human patients and necropsy samples from animal models are the primary source materials for tissue biobanks.19,23,25,122,124,145,174,201 Banked animal and human tissues are typically sourced from external providers (CROs and commercial tissue vendors) who use standard protocols and regulatory-compliant procedures for biospecimen collection, handling, proper fixation, and processing. Quality control (QC) of tissue specimens done by the pathologist is critical to confirm the presence of morphologic and pathologic features of the disease of interest, ensuring biospecimen relevance.18,22,52,53,66,67,85,88,91,99,125,134,149,159,175,186
For commercially sourced tissue samples intended for histopathology, immunostaining (IHC), in situ hybridization (ISH), and spatial assays, pre-procurement QC is often performed on digital whole slide images from H&E-stained tissue sections made available by the vendor. A post-procurement QC is then performed based on the slides made in the recipient laboratory including assessing sample performance in IHC and ISH. Typically, this comprises IHC for at least one common nuclear antigen (eg, Ki67) and one common cytoplasmic/membranous antigen (eg, CD31 in vascular tissues, glial fibrillary acidic protein [GFAP] or synaptophysin in nervous tissues, or pan-cytokeratin in epithelial tissues), and ISH for PPIB (peptidyl-prolyl cis-trans isomerase B) mRNA or equivalent gene. These steps confirm general protein and mRNA integrity and quality, which are contingent on proper prior specimen handling (including cold ischemia time during harvest and time in fixatives such as aldehydes and solvents such as ethanol). Routine QC steps help to maintain a high-quality biobank and to confidently run in-house exploratory/pilot studies relevant to a particular disease.19,23,122,149,174,186,201
Demographic and biologically relevant information related to the tissues obtained from de-identified human patients or healthy donors should also be included in the documentation provided to the pathologist and the research team. Furthermore, all ethical and regulatory aspects related to human tissue procurement (eg, documented consent and protocol language, institutional review board approval, local, state, and national ethical guidance) must be fully adhered to. All archived biospecimens should be carefully inventoried and stored in optimal conditions. Current practices for maintaining a disease-focused and tissue-focused biobank also include the customization of tissue microarrays (TMAs), which can be built from a diverse panel of qualified donor blocks selected in collaboration with pathologists. The TMAs add value by increasing throughput and sparing resources with the trade-off of focused anatomical coverage of the cores. There are multiple options for tissue storage and archival ranging from the traditional cabinets that house labeled tissue blocks and slides according to a variety of systems (project number, tissue type, accession number), to more advanced archiving systems using bar coding and that are directly integrated with the laboratory information management system (LIMS) or laboratory information system (LIS).104,142,173 We recommend the use of the latter to streamline retrieval and automate tissue inventory and track sample accession and lifecycle.
Quality Programs Are Fundamental to Maintaining High Standards of Performance and Accuracy in the Modern Pathology Laboratory
Quality Assurance (QA) in this setting encompasses proficiency testing, internal audits, external quality assessment schemes, and regular training for laboratory staff. Quality control encompasses monitoring sample quality and assay performance, validating test methods, maintaining data integrity, and troubleshooting technical issues. De-identified LIMS data should be the frame of reference for the usage of patient tissues in any laboratory that runs assays on clinical trial samples or samples from Good Laboratory Practice (GLP) toxicology studies. In the case of the former, auditing and adherence to informed consent forms are required, and in the latter, sample custody should be tracked and documented. Although biopharmaceutical industry laboratories can operate outside of GLP regulations or other accreditation requirements, most medical molecular pathology labs are subjected to stringent guidelines and requirements prescribed by accrediting bodies such as the College of American Pathologists (CAP) and the International Organization for Standardization (ISO), and regulatory bodies such as the Clinical Laboratory Improvement Amendments (CLIA), and the US Food and Drug Administration (FDA).35,38,87,102,183 A few industry and contract research laboratories also fall under the scope of FDA GLP regulations. 182 Compliance with these regulations is essential for obtaining accreditation, ensuring patient safety, and maintaining public trust. Accredited labs must adhere to standards for personnel qualifications, test validation, documentation, and proficiency testing. This greatly influences the type of equipment, assays, and assessment provided.
Developing Single or Combined Assays for the Detection of Proteins, mRNA, and/or DNA in a Tissue Section is the Mainstay of the Modern Pathology Laboratory
These spatial assays are directed toward a variety of research questions, including but not limited to characterizing physiological pathways in normal and diseased conditions, verification of new molecular targets, investigating the underlying mechanisms of disease, efficacy and/or toxicity of novel therapeutics at the subcellular, cellular, and tissue levels, interrogating reporter expression in biodistribution studies of non-viral (eg, lipid nanoparticles) or viral vectors (eg, Adeno-associated virus [AAVs]), and novel tissue biomarkers.64,97,133,151,184,201 In that respect, MPLs develop highly specialized protocols that are reagent-specific (ie, chromogenic vs immunofluorescence [IF]; simplex vs low-plex vs high-plex) and require optimization and validation using in-house or commercially available antibodies and mRNA or DNA probes, and digital pathology support. Most laboratories run assays on automated immunostainers (eg, Bond [Leica], Ventana [Roche], Dako [Agilent], etc) to ensure reproducibility, reduce intra-assay and inter-assay variability, and enable upscaling of assay output.41,130,132,150,156
Of note, molecular pathology assays are not simply off-the-shelf “stains” to be used strictly according to instructions from the manufacturer. An assay requires testing a range of conditions against an array of qualified samples to understand and demonstrate the ability of the reagents and procedures to perform as intended relative to specificity, sensitivity, and linearity. The development of an assay must also be done in the context of its use. Therefore, it is highly recommended that assays are developed and optimized by a cross-disciplinary team comprised of biologists, histotechnologists, histomolecular scientists, imaging scientists, and pathologists. These experts are not only familiar with molecular biology and histological procedures but are experienced in sample handling, processing, and staining (chromogenic IHC, IF, ISH, proximity-based, etc) and the scientific principles that underlie each assay. This team will collaborate to define the research question to be addressed by the molecular pathology assay to ensure parameters are fit-for-purpose and that pitfalls for each tissue type or disease are addressed.
Robust Assay Development in the Modern Pathology Laboratory Demands a Defined Workflow and Experimental Plan
Molecular confirmation of protein biomarkers in tissues is usually performed via indirect methods of protein detection, such as chromogenic immunohistochemistry using 3,3’-diaminobenzidine (DAB) as a chromogen and orthogonal assays (enzyme-linked immunosorbent assay [ELISA], Western blot, mass spectrometry [MS]).36,111,120,121,179,206 The IHC workflow tests a primary antibody’s ability to specifically detect the intended target with sufficient sensitivity. In brief, an initial chromogenic testing in frozen and/or FFPE cell pellets generated from positive and negative cell lines with known expression of the specific target, synthetic peptide arrays, and/or other suitable control matrix is recommended prior to tissue testing. For the development and implementation of assays intended for relative quantitative endpoints (such as relative scoring of protein expression levels), the inclusion of cell lines, each with different protein expression levels, is also indicated. Typically, such protein expression gradients are determined orthogonally by MS, quantitative western blotting, ELISA, and so on. In instances where the positive control cell line used for the initial assay validation is an overexpressing cell line (eg, a transgenic cell line expressing the protein at the highest gradient), bridging the IHC/IF expression patterns to those in tissues with endogenous (and typically lower) level of expression is also recommended to ensure broader interpretability.36,111,120,121,141,179,206 For IHC and other antibody-based biomarker assays, it is imperative to ascertain fit-for-purpose specificity, sensitivity, and, if applicable, dynamic range of the test system. Proper documentation of the final assay protocol should meet good laboratory principles for data integrity and reconstruction, including the instrument platform used, as such assays are often not easily transferable to another platform.
More complex spatial assessments (like identification of cell subtypes, cell targeting of viral vectors, tumor microenvironment, cell-to-cell interactions, neighborhood analyses) are increasingly being used to support investigative and mechanistic de-risking studies. Chromogenic assays can be transferred to single fluorescent IHC (IF-IHC/ISH) or multiplex chromogenic or IF-IHC/ISH assays with appropriate sequence optimization of chromogens and fluorophores. The automation of bench processes as well as added capabilities for low-plex IHC or ISH stains, or combined IHC-ISH kits, have resulted in a significantly higher throughput compared to traditional anatomic pathology labs. The role of the pathologist at this stage is to collaborate with the lab scientists to ensure all QC steps are met and that the assay development considers the intended application (ie, what are the minimum performance specifics that the assay must meet) to address the specific project questions. 130
The Modern Pathology Laboratory Has an Indispensable Role in Validating Biomarkers
This helps understand PDs, pharmacokinetics, and efficacy relationships in pre-clinical and clinical studies, to support a recommended phase 2 dose (RP2D) in clinical trials, assist in prognostication and stratification, inform reverse translation efforts and combination regimens, as well as help identify and manage safety hazards associated with novel therapeutics.13,27,33,58,62,71,85,146,152
Although important attention is given to peripheral biomarkers, assays in tissue biopsies continue to play an important role in trial management, data collection, and dose regimen selection. The most used tissue-based PD biomarker assays are designed to identify a specific drug target or cell(s) of interest in a tissue sample and correlate their distribution with efficacy. Classic examples of tissue-based biomarkers used in patient stratification include the multiple PD-L1 IHC assays approved by the FDA as companion diagnostics for immuno-oncology therapies (ie, the 22C3 anti-PD1 IHC for pembrolizumab), and the anti-Her2 IHC assay for Her2 low breast cancer for treatment with the ADC Enhertu (fam-trastuzumab deruxtecan-nxki). Tissue PD biomarkers may also include assays to investigate chromosomal translocations or gene truncations, fusions, and amplification. Proximity assays are used for investigative purposes in the development of bispecific antibodies. In the blossoming field of targeted protein degradation, quantifying the target protein in the tissue of interest after treatment and over time is an essential part of drug discovery and best done with a combination of IHC and proteomics. Understanding target engagement and PD using tissue-based assays can be a fundamental part of determining the RP2D for clinical trials.
Pathologists play an important role in biomarker discovery and in biomarker assay development and validation.33,58,71,85,152 For instance, it is common for research groups to cast a broad net to investigate multiple exploratory biomarkers in immune-oncology research. These assays are normally designed in the multiplex format to reduce the amount of tissue needed and to locate cells that express two or more biomarkers (ie, T regulatory cells, exhausted T cells, specific subsets of myeloid suppressive cells). These complex assays are then analyzed retrospectively to correlate with treatment response. Input from a well-trained pathologist in assay design and interpretation is vital to ensure a high-quality assay and guide bench scientists in the establishment of the final protocol. Pathologists can also propose novel biomarkers based on morphologic features of a tumor (ie, looking for KRAS mutations in mucinous solid tumors, or MYC and BCL2 amplification in anaplastic B cell lymphomas).
Similarly, the MPL plays a critical role in diligent molecular and histologic characterization of animal models, which are quintessential for modern drug discovery research.14,16 Genetic background, method of transgenesis, as well as husbandry conditions can greatly influence histologic and molecular phenotype of such disease models and should be duly considered during all stages of molecular pathology assay optimization and interpretation.40,75,81,95,123 The use of in vitro complex models such as organoid models (derived from embryonic stem cells, induced pluripotent stem cells, or patient tissues)73,80,196 and organ-on-a-chip 86 are increasingly being used in the MPL as alternative translational experimental models to animal testing. Advances in organ-on-a-chip microfluidic culture devices can recreate organ-level structures and recapitulate mechanistic features of human organs in physiologic or pathologic states and evaluate patient drug responses. 86 Standard laboratory workflow used for tissue-based histology can be easily adapted to support the validation of organoids and other New Approach Methodologies (NAMs).176,181,185,198
Digitization of Slides Into Whole Slide Images at Scale Emerged in the Past Decade and Has Allowed for the Incorporation of Digital Pathology Workflows to Molecular Pathology Labs in Industry and Academia
With the digital revolution, an integral team member of the pathology lab of the future emerged, the digital image data analyst.2,20,56,69,89,90,100,143,144,203,211 Spatial data scientists are specialized in extracting data from whole slide image (WSI) using commercially available or public software tools integrating the digitalized slides with AI and machine learning workflows for automated qualitative and quantitative assessments, ultimately streamlining workflows is a current industry focus (eg, automatization of tissue visualization, and segmentation of areas and cell types selected for analyses). These capabilities have enhanced collaboration and real-time exchange of data, increased the amount of information that can be derived from a slide, reduced variability among pathologists in interpretation, and allowed image analysis scientists to support anatomic pathology with quantitative assessment of morphological and molecular phenotypes together with the expert judgment of the pathologist. Lab staff training in this setting has benefited from beta testing various state-of-the-art digital imaging analytical software and from interacting with image analysts. Currently, most MPLs are equipped with one or more scanners that fulfill their specific workflow needs. Supplementary Table 1 lists many of the commercially available whole slide scanners and their specific properties.
Anatomic Pathology Laboratories Have Efficiently Implemented Low-Plex Assays Coupled With Whole Slide Image-Digital Pathology as part of Their Routine Workflows, Making It an Accessible Reality for the Modern Pathology Laboratory to Implement High-Plex Assays for Detecting Dozens of Combined Protein or Hundreds of Genes on the Same Tissue Section (ie, Spatial Biology and Spatial “-Omics”)
Academia has been, to date, the largest customer of many of the available spatial biology platforms.5,15,26,45,64,97,126,133,137,140,151,184,190 However, the usage of these platforms in translational biopharmaceutical research and clinical trials in industry settings is increasing as preliminary studies yield promising results and the technologies and analytical pipelines become more accessible and are further validated. Continued advancements in scalability, standardization, and clinical utility are expected to drive growth across all stages of R&D.140,184,190
This technological explosion shaped the field of spatial biology and the MPL with more than dozens of sophisticated, unique, and costly high-plex platforms for protein, mRNA, or both.32,39,44,63,69,72,79,98,109,115,117,128,129,139,154,172,180,189,194,200,202,204,205 As exciting as it is to witness this revolution in spatial molecular pathology tools, testing these technologies is very disruptive, expensive, and time-consuming. It is beyond the scope of this article to review all the available technologies32,39,44,63,69,72,79,98,109,115,117,128,129,139,154,172,180,189,194,200,202,204,205 in detail, but we have dedicated a section to provide an overview of this still-emerging field and discuss some of its caveats. The challenges with adapting toward these high-plex assays are not only to decide which platform meets your research needs best but also how to build downstream analytical platform tools that help deconvolute such complex data sets and analyze it efficiently. Section “Considerations Toward Building a Spatial Biology Multimodal Pipeline in the Modern Pathology Laboratory” provides a broad overview of the properties, advantages, and disadvantages of some that are currently under evaluation in our laboratories.
Additional Research Technologies That Might not Be an Integral Part of the Modern Pathology Laboratory, Yet
Occasionally, research questions may require a more specialized approach, instrumentation, and user expertise, often with a higher cost and lower throughput. Examples have been tabulated in Supplementary Table 2 and include transmission electron microscopy (TEM), cryo-electron microscopy (Cryo-EM), confocal laser scanning microscopy, laser-capture microscopy, expansion microscopy, light sheet fluorescence microscopy, RAMAN microscopy/spectroscopy, single-cell and multiomics approaches, MS Imaging, MALDI Multiplex IHC, and atomic force microscopy (AFM).5,17,26,31,34,42,45,54,59,64,65,68,82,92,97,105,106,118,126,133,137,140,151,158,169,170,184,191-193,197,208,209 Some of these modalities, which are currently seen as extremely specialized, are on course to becoming more routine in multimodal discovery applications. In scenarios that call for multimodal integration of histopathology and light microscopy with higher resolution advanced imaging modalities, the pathologist is encouraged to seek collaboration with internal or external subject matter experts, some of whom might be located within specific CROs or at academic centers.
Considerations Toward Building a Reliable Data Repository Ecosystem
It is essential that LIMSs of the MPL support full digitalization of their different activities as they evolve in complexity.103,177 The MPL, in collaboration with the information technology (IT) team, needs to delineate how the various image format files will be identified, stored, and removed, if necessary, and make sure results are delivered on time to the relevant stakeholders. An interactive and efficient system will avoid dissociation which often results in duplication of large numbers of images destined for use by different groups, which adds to the already steep cost of maintaining and storing petabytes of images, and auditing images from clinical trials. For example, images from slides used for assay development will often be numerous and no longer useful once the report has been generated, so they can be archived in cheaper storage conditions (ie, Glacier). Images from clinical trials, on the contrary, need to be retained as raw data, may be audited, and must be deleted in the instance a patient withdraws informed consent.
The Modern Pathology Laboratory Image Viewer Infrastructure Should Support the Various Laboratory Activities and Interdisciplinary Teams at a Global Scale
Ideally, this setting will enable automation of image registrations from different modalities and an interactive viewer for combined visualization of features and of data sets across the multiple modalities (Figure 1). Because WSIs are often very large files, viewing and analyzing images can be a challenge for groups that are geographically distant. This is an important problem for many global companies that have interdisciplinary teams spread around the world and that need to work with large histological images. Remote access to high-quality digital slides (or WSI) allows pathologists quantification of simple molecular pathology endpoints or analyzes complex endpoints such as multiplex immunofluorescent IHC images in which a single cell may express two or more overlapping markers that are impossible to deconvolute with the human eye. Despite these digital pathology workflows, some challenges remain around integrating digital processes with the remainder of the lab workflows, such as requesting assays and reporting results, and with implementing scalable and secure IT infrastructures.
The Modern Pathology Laboratory Will Need Centralized Access to Both the Pathology Images and Artificial Intelligence Tools to Enable Curation and Archival of Data Sets
With the rapid proliferation of deep learning and generative AI models, incredibly powerful image classification models are being built through artificial neural networks (ANNs) that rely on extensive image databases such as ImageNet, ResNET, UNet, and many others.7,47,61,136,155,161,168,171,195 Evolution of supervised learning AI tools that can integrate spatial molecular expression patterns within H&E-stained sections could successfully be used to predict gene expression from tissue sections where only a high-resolution H&E image is available. This approach has proven useful for the identification of gene expression variations in breast cancer Xenium data sets and for subsequently generating virtual spatial transcriptomics (ST) data from histological images using the iSTAR (Inferring Super-resolution Tissue ARchitecture) model. 205 In that respect, the role of the pathologist will be key in supervised annotation of multimodal stained tissues that can be used for training and building algorithms, and “closing the loop” of digital and computational pathology in clinical research. 157
The Modern Pathology Laboratory That Carries Out Spatial-“Omics” Is Required to Be Adaptable to Scale-up, and Yet Provide Laboratory Information Management System and Sample and Technology Interfaces
Ideally, the MPL will be equipped with a multi-omics spatial cloud-based software solution that is compatible with data sets generated by different technologies and that allows for combined visualization of high-resolution images and downstream data analyses. In that context, multimodal platforms will encompass automated landmark registration, allowing for the overlay of different modalities and comparing data generated by them. Ultimately, multimodal spatial “-omics’ studies need integration of all data sets. Visualization of various samples at the same time with synchronization for each technology facilitates sample evaluation for the desirable endpoints. Analytical tools include cell segmentation, colocalization of targets, neighborhood analyses, cell-to-cell intercations, dimensionality reduction, clustering, and spatial visualization. Figure 1 provides a snapshot of various analyses that these platforms can provide.8,43,48-51,94,107,108,114,119,138,210
Considerations Toward Building a Spatial Biology Multimodal Pipeline in the Modern Pathology Laboratory
The field of spatial biology evolved through the synergy of molecular biology and histopathology, and advanced with the breakthrough of spatial “-omics” technologies, which allowed the detection of large numbers of RNA and/or protein targets in a tissue section.5,15,26,45,64,97,126,133,137,140,151,184,190 These spatially resolved platforms offer high-plex mini-bulk approaches that can interrogate the whole transcriptome in multicellular areas of 2 to 55 µm in diameter (ie, GeoMx, Visium, Visium HD) and up to 6K genes at a single-cell and subcellular resolution in 2D images. In this rapidly evolving field, current technologies under development are addressing DNA detection and 3D spatial resolution.24,29,46,110,207
Broadly, these spatial “-omic” profiling methods can be divided into (1) targeted or multiplexed nucleic acid probe-based (Xenium, CosMx) or antibody (Akoya PhenoCycler, previously CODEX, Comet from Lunaphore, Image Mass Cytometry like Fluidigm/Standard Bio Tools CyTOF or multiplexed ion beam imaging (MIBI) from IonPath)-based microscopy imaging with single-cell resolution and (2) transcriptome-wide or next-generation sequencing (NGS)-based approaches (Visium, GeoMx, STOmics Stereo-seq) with multicellular or nanoscale single cell resolution.32,39,72,79,98,109
Spatial Proteomics
Investigating and understanding the loclization and dynamics of proteins in tissues is the core focus of spatial proteomics, which is crucial for advancing cell and tissue biology. Concurrent detection of dozens to hundreds of protein signals in their tissue location through antibody targeting is enabled by imaging-based spatial proteomic methods. Tissue imaging-based spatial proteomic technologies use classical IHC/IF methods in combination with various antibody tagging approaches, including fluorescent dyes, oligonucleotide barcodes visualized by fluorescence microscopy, or metal tags detected by MS. The fluorescent-based spatial proteomic workflow typically consists of cyclic staining, imaging, and dye inactivation, with 2 to 3 markers being imaged during each cycle and potentially visualizing up to 100 proteins. By comparison, MS-based spatial proteomics methods require only one cycle of staining and enable imaging for up to 40 markers. Independent of the modality, parameters such as antibody specificity and sensitivity marker panel selection, and tissue quality are key to ensuring the accuracy of the spatial data generated. In this context, integration of spatial proteomic workflows in the MPL requires upstream antibody verification, relevant tissue preparation and annotations, and multiplex panel design, before execution of the spatial assay.44,63,69,79,109,115,117,129,172,180
Examples of imaging-based spatial proteomics technologies that rely on antibody binding to protein targets include Cell DIVE (Cytiva, Leica), MACSima (Miltenyi), MIBIscope (IonPath), MultiOmyx (Clarient, GE Healthcare), PhenoCycler (formerly CODEX, Akoya), PhenoImager (Akoya), and COMET (Lunaphore, Bio-Techne). Cell DIVE exemplifies a technology in which antibodies to the targets of interest are bound and stripped in a cyclical/sequential manner. In CODEXm antibodies to all the targets are bound together and fluorescence imaging is performed in a sequential or cyclical manner. The Lunaphore COMET performs a series of stains, image acquisition, and elution with an integrated analytical pipeline. Its proprietary gentle hyperplex cyclic IF allows for the serial detection of proteins and mRNA transcripts upstream or downstream single-cell ST testing on the same slide. The MIBI is a technology in which antibodies to all the targets are bound together, and all the signals are imaged simultaneously by MS-based imaging.26,45,137
Spatial Transcriptomics
Like single-cell RNA (scRNA)-sequencing technologies, current ST methods have been instrumental in validating biological findings, elucidating gene expression patterns, and discovering cell subtypes and functional states, ultimately impacting the discovery of new diseased targets. Although most spatial “-omics” technologies work in FFPE and in fresh-frozen tissue sections, they vary in their spatial resolution, area of tissue coverage to be analyzed, scale and throughput (number of slides and profiling speed including benchwork preparation time), multiplexing capacity (breadth of molecular entities profiled simultaneously), species that they covered, and the cost of running one slide. 98 Without aiming to endorse any specific technology over another, Table 1 provides a high-level overview of the final product for some of today’s most established technologies in the field of spatial “-omics.”
List of commercially available in situ spatial “-omics” technologies to be performed on FFPE tissue sections.

Abbreviation: WT, whole transcriptomics.
Deciding on which technology to invest in poses a current challenge for several reasons: there is not a single platform that fits all research questions and laboratory processes; new possibilities for multiplexing at a higher resolution and better gene coverage continue to emerge; and these technologies remain cost prohibitive (instruments may run $250K-$500K and test runs between $2k-6K/slide depending on the technology and laboratory). Analysis benchmarking comparisons on various in situ spatial molecular profiling platforms conclude that, overall, most currently available platforms concord with orthogonal scRNA-seq data sets in performing spatially resolved cell types despite experiencing some rate differences in cell segmentation errors, false discovery rates, and various degrees of subclustering.26,72,194
Adopting and Integrating Spatial-“Omics” Technologies and Analytics in the Modern Pathology Laboratory Pipeline
Investing in high-plex spatial “-omics” approaches can be difficult to justify based on the scope of work of some MPL, as their laboratory implementation is time-consuming and their cost and throughput are high and low, respectively. Also, consideration of the species of interest is advised since most available ST platforms are currently best suited for research in human, non-human primate (NHP), and mouse specimens with very limited options for other species like canine or the rat. Data workflow informs target abundance in tissues of interest and differential cell expression, thus accelerating target discovery and ultimately impacting the drug discovery pipeline. Similarly, for exercises like cellular atlas building, these approaches might be hugely beneficial. Instead, for hypothesis testing, laboratories may adopt targeted technologies with customization of a panel with dozens of proteins and/or a few hundred genes that can be built to fit the project needs. Nonetheless, the MPL may prioritize combining sequential low-plex spatial assays in serial sections as a valuable and best approach. For example, detecting three transcripts and one protein in one slide, six proteins in the serial section, and another three proteins in the next serial section, all by purchasing off-the-shelf kits and using a standard automated stainer, may result in a faster turnaround, higher throughput, and lower cost.
In terms of throughput and headcount, although molecular scientists can run both wet lab and on-instrument processes, most of these techniques take 3 to 5 days to be completed, and only for 2 to 4 slides at a time. Thus, depending on the laboratory workload and workflow, spatial “-omics” studies with a large number of samples may require 1 to 2 dedicated persons to routinely run these technologies for several weeks until the full study is completed. The role of the pathologist is to confirm the approach that is needed for the questions at hand and then design a single modal or multimodal approach that provides the best data set outcome using the minimal tissue from the block. Figure 2 provides a schematic overview of the workflow that entails performing multimodal spatial “-omics” studies in the MPL. In that context, running high-plex spatial “-omics” profiling studies requires justification and strategic thinking in their design, since generating these data once could be a one-time-only experiment since data are saved permanently and can be interrogated to answer future questions. As part of the study design phase, the pathologist will primarily: (1) contribute to the multiomics spatial study design, (2) design and evaluate the custom gene panel together with the bioinformatics/computational team gene panel customization, (3) evaluate a priori the sample integrity and the mRNA/protein quality, (4) annotate the areas of interest from each sample that need to be cut, (5) decide on the number and order of serial sectioning to accommodate different technologies, (6) supervise the quality of the tissue sections and decide the final “omics” multisample lay-out slide with the histotechnologist, and (7) coordinate with the molecular biologists the timeline for the different runs. Finally, the pathologist can review the QC data of the run with the molecular scientists and perform a preliminary evaluation of the raw data by using the imager and visualization software provided by the vendor. To an extent, available software allows for visualization of multimodal data, ie, Xenium explorer allows for uploading Xenium transcript data sets and co-register the Xenium image with its H&E-stained and its multiplex IHC images run on the same section or in serial sections.
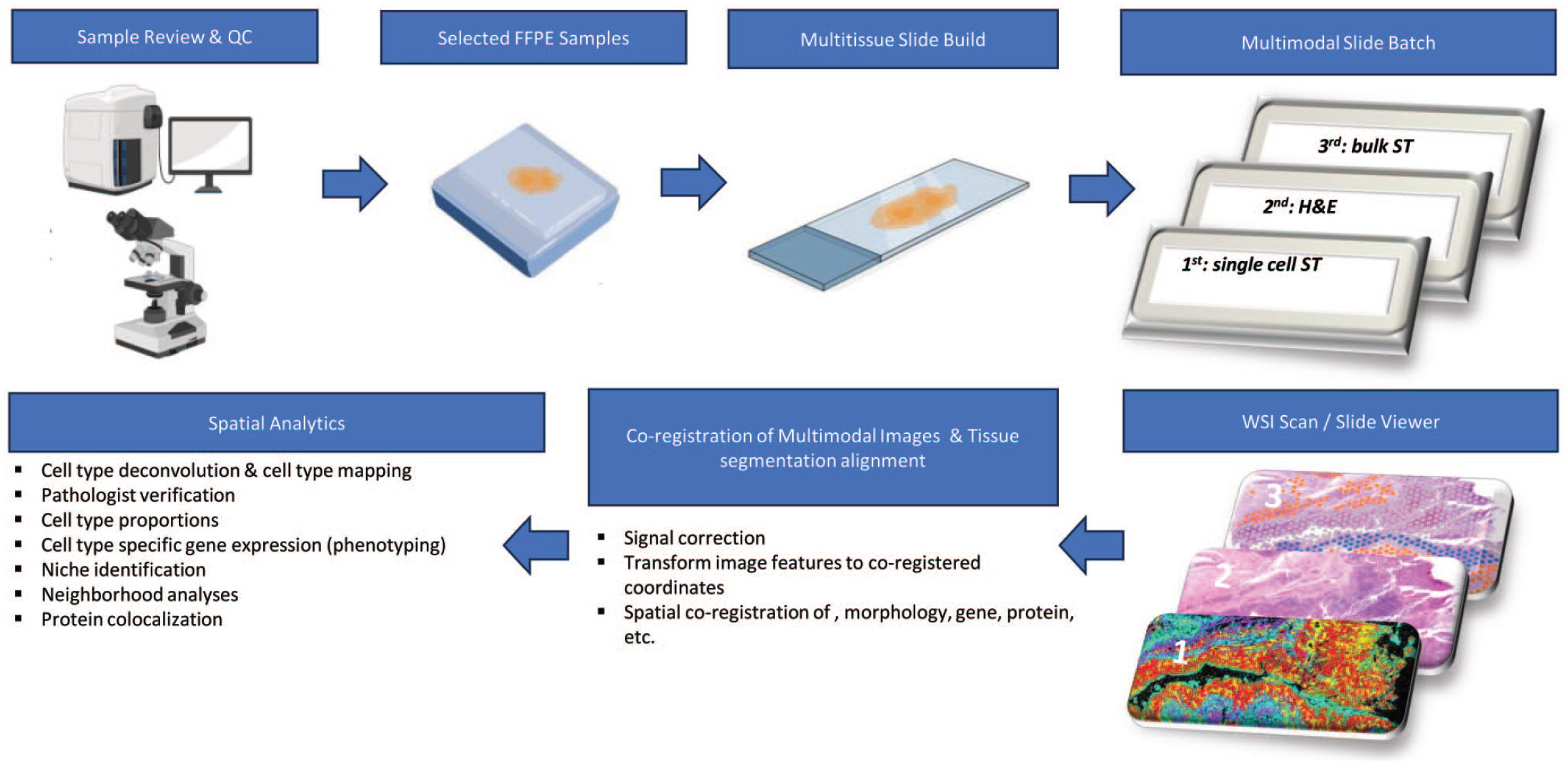
Workflow for performing multimodal spatial “-omics” studies.
Another important consideration is that analyzing ST data, particularly in elucidating cell-type compositions within tissue environments, necessitates advanced computational deconvolution techniques and building a fit-for-purpose downstream visualization system and analytics pipeline. At this stage of the process, a team of image data scientists, bioinformaticians, and computational biologists are the drivers of the downstream analytics in collaboration with the pathologists.
Spatial Transcriptomics Downstream Analytic Pipeline in a Nutshell
The advantages of ST over scRNA-seq methods include the interrogation of cell-to-cell proximity and interaction with high confidence, groups of spatially co-varying genes that can identify niches and gene signatures associated with pathological features, among others. Platforms for data preprocessing, and visualization tools that offer interactive exploration of ST data sets, are becoming integral to the workflow. These tools facilitate the standard analysis and enable researchers to customize their approaches to suit specific experimental needs, paving the way for innovative studies in tissue biology, disease pathology, and developmental biology. Recent advances in ST technologies have also introduced sophisticated machine learning models such as graph convolutional networks (GCNs) that leverage the spatial context more effectively. These models integrate multiple types of data, including histological images and detailed spatial annotations, to enrich the transcriptomic data analysis. Such integrations allow for a more nuanced understanding of the spatial and functional organization of cells within tissues, bridging the gap between molecular data and biological function. Through these comprehensive and integrated approaches, ST is set to revolutionize our understanding of biological systems at an unprecedented spatial resolution.
The ST data analytical framework will vary depending on the platform used and is evolving rapidly. It provides multiple options for graphs and spatial feature visualization, interactive visualization with exploratory data browsing, and visual inspection of results and analyses. Providing varied formats aids the exploratory visualization of and sharing of the data and analyses with collaborators. References for available ST computational tools are provided.51,49,98,108,119,137,138 Summarized below are some of the key steps needed for ST analyses. The analytic process begins with the acquisition of raw data represented by gene-by-cell count matrices accompanied by spatial coordinates. This step includes QC measures such as filtering genes and cells based on specific thresholds, normalization to adjust for technical variations, and possibly transformation of count data to align with downstream analysis requirements. Following preprocessing, deconvolution is pivotal as it discerns the mixture of cell types contributing to gene expression. Several spatially resolved cell-type deconvolution techniques have been developed, including SPOTlight, spatialDWLS, RCTD, SpatialDecon, CARD, and SPADE just to name a few. 114 Cell-type identification and spatial pattern analysis involve the use of dimensionality reduction techniques to simplify complex data into more manageable forms, followed by clustering algorithms to group similar expression profiles or categories. These steps are critical for distinguishing different cell types or cell states within the tissue, which are essential for understanding tissue architecture and cellular interactions. The next phase focuses on the analysis and visualization of these spatial data to draw biological conclusions. This includes mapping cell types back to their original spatial coordinates, exploring cell-cell interactions, and identifying spatially variable genes. Advanced computational tools and algorithms are employed to handle these tasks, enhancing the resolution and depth of the analysis. One important point is that cell-type annotations are based on publicly available sources that require to be confirmed by internal computational biologists and correctly annotated for mapping cell types in the ST experimental samples. This process will need to be generated for each of the organs and disease of interest and is best when initiated prior to running an ST experiment and might be required at the time of contacting the vendor for purchasing of custom gene panels. 44
A final but important consideration when applying ST techniques to clinical trial samples is the correct management of patient samples in collaboration with clinical operations teams in order to maintain appropriate validation reports filed in the electronic trial master file (eTMF), ensure that assays performed are included in the informed consent forms, and that results are reported in the correct format for the trial databases and remain blinded when necessary. All patient privacy regulations should be strictly adhered to, and samples must be destroyed when consent is withdrawn.
Concluding Remarks and Considerations for Training the Anatomic Pathologist
Current Landscape
Drug discovery has shifted to new therapeutic modalities and toward personalized and precision medicine, requiring advanced techniques beyond traditional methods. This shift requires adopting advanced research techniques and interdisciplinary collaboration, especially in the MPL, which integrates molecular and computational biology, imaging, data sciences, and AI. The MPL can help to simultaneously visualize the spatial distribution of morphological features, drug targets and therapeutics using simple tissue-based assays or deploy complex spatial assays to map each molecule of the human body, in health and disease. Significant investment in infrastructure and expertise is needed to establish an MPL, but the potential for valuable, long-term data is high. Scientists must carefully consider resource allocation for these advancements. Modern pathology data are complex, multidimensional, and large, requiring interdisciplinary collaboration and robust IT support for effective metadata extraction, to ensure sufficient storage, ease of data mining, and data quality.
What is not Changing?
Despite advancements in data generation and analyses, quality knowledge in pathology relies on rigorous scientific methods and clear project strategies. Laboratory scientists and pathologists are essential in maintaining these practices throughout the research process. Although new technologies are important, traditional methods remain crucial for addressing most pathology questions. Novel research modalities are expected to enhance, not replace, traditional techniques.
Looking Ahead
The MPL is a bespoke research unit with multidisciplinary scientists and advanced technology, designed to handle increasing data complexity in drug discovery. Scientists and pathologists of the MPL will need new skills and support to work with large-scale data sets. Such skills and collaborative interdisciplinary opportunities include:
Integrative biological insight: the ability to synthesize information from molecular data, clinical and histopathological observations, offering a holistic view of disease mechanisms and tissue organization.
Mastery in multidimensional data analysis: proficiency in interpreting complex data sets, including spatial, genetic, and phenotypic information, to draw comprehensive biological insights.
Computational and bioinformatics skills: competency in bioinformatics and computational biology to manage, analyze, and visualize large-scale ST data sets.
Advanced technical expertise: a deep understanding of spatially resolved transcriptomics and proteomic high-throughput technologies, coupled with the ability to innovate and adapt these tools for novel applications.
Collaborative and leadership abilities: strong leadership skills to spearhead multidisciplinary projects, advanced communication skills to integrate and convey complex biological data succinctly and clearly, combined with the ability to collaborate effectively with researchers from diverse fields.
Training for veterinary pathologists in molecular pathology is mostly informal and self-driven. Research-oriented doctoral programs offer advanced training, but most learning happens through self-directed methods like webinars, lectures at pathology society meetings, and presentations by companies. Special interest groups within professional societies (eg, ACVP, ASIP, BSTP, ESTP, STP, AMP, ASGCT, DPA, and ESDIP) also provide valuable resources for ongoing education. Discussions and presentations featured in molecular pathology special interest groups within the Society of Toxicologic Pathology (STP MP SIG) or the European Society of Toxicologic Pathology (ESTP Omics subcommittee under Pathology 2.0) are proving to be informative resources for informal continuing education in molecular pathology.
Amidst modest progress, there remains an ever-growing and overwhelmingly unmet need for structured training and formal certification in molecular pathology for trainees and practitioners of veterinary pathology. This requires ongoing collaboration among academia, government, and industry to develop formal programs. For pathologists, following primary anatomic or clinical pathology residency training, interested trainees or new diplomates could pursue tailored PhD programs and/or fellowships at designated academic and industry centers with training in contemporary molecular, digital, and computational pathology approaches. Frameworks could be drawn from medical molecular genetic pathology and pathology informatics training programs.9,11,28,57,60,76,78,96,116,127,188 An important distinction to be made is that diagnostic medical/veterinary pathology needs to follow an individual case-based approach, whereas training for modern discovery and safety pathology practice in academia and industry would follow an experimental, cohort-based comparative approach and that distinction needs to be woven into the training framework. A new generation of scientists and pathologists will be called upon to realize the vision of the MPL, and it is imperative that our professional societies and residency and graduate training programs prioritize expanding and updating their curricula to empower this next wave.
Supplemental Material
sj-docx-1-tpx-10.1177_01926233241303898 – Supplemental material for Trends and Challenges of the Modern Pathology Laboratory for Biopharmaceutical Research Excellence
Supplemental material, sj-docx-1-tpx-10.1177_01926233241303898 for Trends and Challenges of the Modern Pathology Laboratory for Biopharmaceutical Research Excellence by Sílvia Sisó, Anoop Murthy Kavirayani, Suzana Couto, Birgit Stierstorfer, Sunish Mohanan, Caroline Morel, Mathiew Marella, Dinesh S. Bangari, Elizabeth Clark, Annette Schwartz and Vinicius Carreira in Toxicologic Pathology
Footnotes
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: SS and AS are the employees of AbbVie and may hold shares and/or stock options in the company. CM and DSB are the employees of Sanofi and may hold shares and/or stock options in the company.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
