Abstract
Digitization of histologic slides brings with it the promise of enhanced toxicologic pathology practice through the increased application of computational methods. However, the development of these advanced methods requires access to substrate image data, that is, whole slide images (WSIs). Deep learning methods, in particular, rely on extensive training data to develop robust algorithms. As a result, pharmaceutical companies interested in leveraging computational methods in their digital pathology workflows must first invest in data infrastructure to enable data access for both data scientists and pathologists. The process of building robust image data resources is challenging and includes considerations of generation, curation, and storage of WSI files, and WSI access including via linked metadata. This opinion piece describes the collective experience of building resources for WSI data in the Roche group. We elaborate on the challenges encountered and solutions developed with the goal of providing examples of how to build a data resource for digital pathology analytics in the pharmaceutical industry.
Keywords
This is an opinion article submitted to the Toxicologic Pathology Forum. It represents the views of the authors. It does not constitute an official position of the Society of Toxicologic Pathology, British Society of Toxicological Pathology, or European Society of Toxicologic Pathology, and the views expressed might not reflect the best practices recommended by these societies. This article should not be construed to represent the policies, positions, or opinions of their respective organizations, employers, or regulatory agencies.
Introduction
The nature and volume of histopathology data results that toxicologic pathologists can obtain from in vivo studies are expanding as computational methods are increasingly applied to digitized histopathology images.1-3 These recent advancements in computational approaches are reliant on access to high-quality whole slide images (WSIs) that meet the image-quality standard for their intended use, 4 pathologist review and interpretation, and often also require high-volume WSI data since state-of-the-art deep learning (DL) computational methods can require up to tens of thousands of image examples for model building.5,6
These promising applications, along with digital slide evaluation, have driven the emergence of the subfield of digital pathology in toxicologic pathology, which enables not only digital slide review, but also computationally-driven assessment of WSIs, and the potential for multimodal integration across additional data types (e.g., in vitro assays, and genomic, biochemical, and metabolic evaluations) as computational pathology methods mature. 7 In addition to creating opportunities for novel endpoints, the reproducibility of many computational methods enables more robust meta-analysis of findings across studies than was previously possible. Increased capabilities to identify, and ultimately predict, translational safety endpoints through computational analysis of WSIs and their associated data may result.
Organizations interested in building computational toxicologic pathology capabilities must first establish large-scale data management processes for the WSI files and their associated metadata (e.g., test material administered, species/strain, sample type, histopathology findings). The scope and complexity of building robust data resources for state-of-the-art digital pathology represents a significant challenge. Furthermore, any systems built today need to be flexible, scalable, and extensible given the fast pace of advancement in the field, with new platforms and approaches introduced routinely, and the near certainty that the needs of today will change tomorrow.
This opinion piece describes a collaborative approach to develop effective informatics resources for toxicologic pathology WSI data in research and early development across the Roche group, comprising Genentech (gRED) and Roche Pharma Research and Early Development (pRED). The discussion below describes the joint approach of gRED and pRED. In instances when different approaches/ways of working were used, they are described separately. Discussion of application of virtual microscopy using WSIs in Good Laboratory Practice (GLP) toxicologic pathology workflows is beyond the scope of this work and has recently been discussed in depth elsewhere.8-10
Metadata
Before discussing approaches and solutions to the technical challenges of obtaining and managing WSI data, the criticality of associating each WSI with identifying information (metadata, or “data about data”) following the FAIR (Findable, Accessible, Interoperable, and Reusable) data guiding principles must be highlighted. 11 At its core, a comprehensive and harmonized metadata model enables the four key concepts in data FAIRification, and is thus the foundation of a digital image data strategy.
The first step in building a FAIR WSI data repository is establishing a unique identifier for each WSI. A glass slide and its subsequent WSI can potentially be made findable through the use of unique barcodes, which is an approach used in our laboratories internally. However, glass slide barcoding is not universally performed by histology laboratories. To uniquely identify WSIs in the absence of barcodes, gRED and pRED have jointly specified the following metadata fields (Figure 1) required for us to uniquely identify each WSI file in our repository, and provide the links to other source systems following proposed data management best practices. 12 These metadata thus provide the key linkage between the WSI files and all relevant associated study data. Integrating the systems in this way is essential to manage the burden of metadata curation and minimize data redundancy across systems.
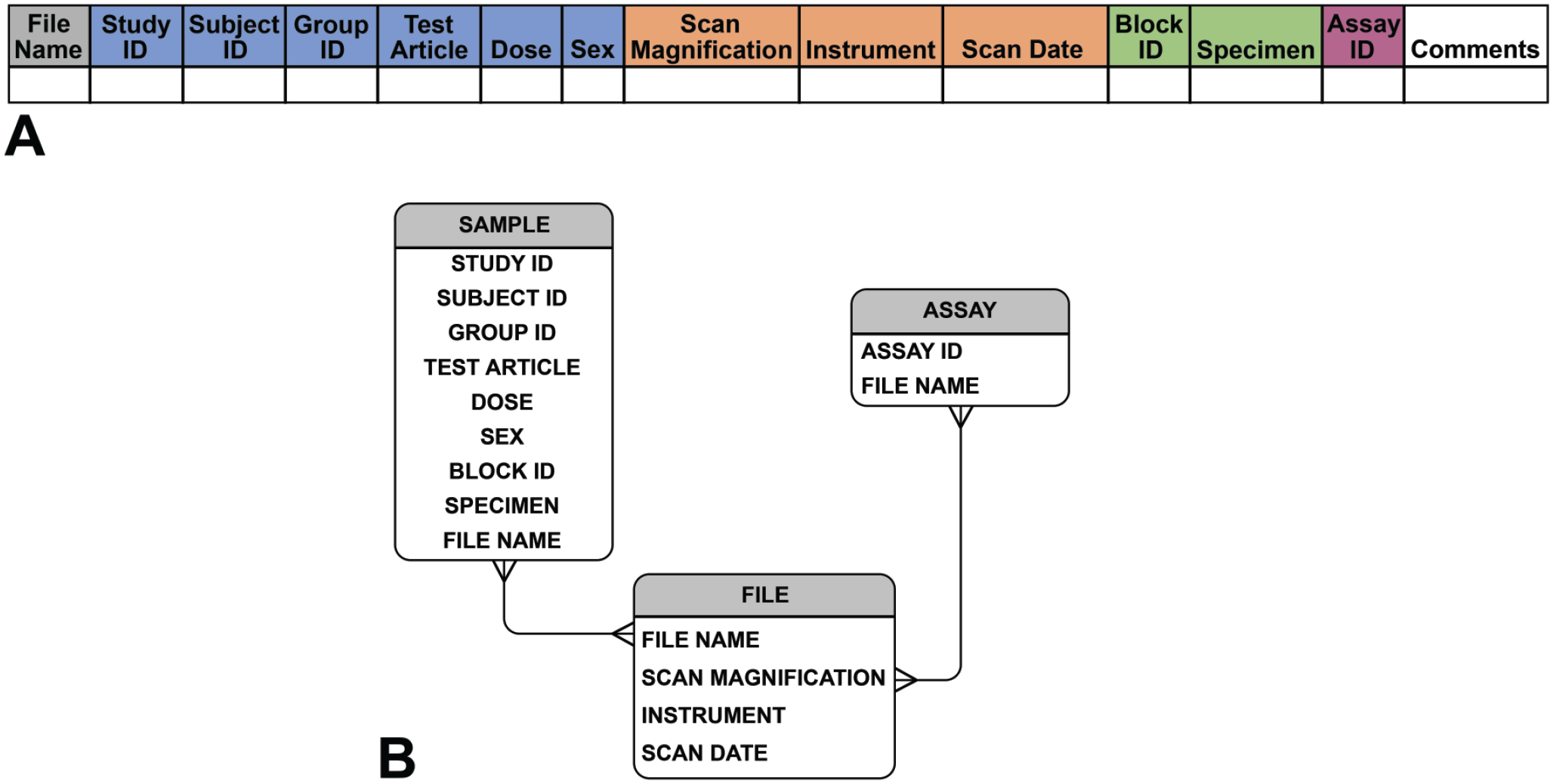
(A) Data collection sheet header shows our mandatory metadata example that ensures collected metadata are uniquely linked to each WSI file. Note that in the header example, study sample metadata fields are highlighted in blue, scan file fields in orange, slide fields in green, and a field for assay registration ID in purple if an assay registration system is employed. (B) Entity relationship diagram demonstrating example metadata fields sorted into the classes sample, assay, and file, and showing the relationship between these fields.
Although the established metadata are sufficient to link WSI files to data from other relevant systems in our context, for example, commercially available pathology Laboratory Information Management Systems (LIMSs), such as SLIMS (Agilent Technologies, Inc., USA) and Provantis LIS (Instem LSS Limited, UK), custom-built study data repositories, and digital image viewer systems, fully realizing efficient and complete data assembly has been one of our greatest challenges. Application programming interface (API) connectivity among our systems for direct interoperability has not always existed, and metadata interoperability has been inconsistent. In addition, there have been numerous instances in which, even when theoretically accessible, study data from one or even two systems have been incomplete, for example, in cases of missing histopathology findings (e.g., histopathology partially or completely skipped in early terminated studies), histopathology findings available only in legacy PDF file formats unreadable by our API, or incomplete data transfer agreements (DTAs) with contract research organizations (CROs) resulting in partial data transfers. In addition, we have found that data gaps can be introduced in instances when samples are processed through multiple labs, for example, when embedding and staining occur in different laboratories, if the laboratory management systems are not fully interoperable and/or data management best practices are not fully operationalized. Success of data assembly efforts requires full commitment at all levels to invest in developing and adapting systems and obtaining high-quality and complete study data to complement the WSIs.
Although not related to metadata, we have also confronted gaps in WSI data itself, such as missing images from scanned studies. This can occur, for example, in cases of absent or inadequate WSIs, particularly from archived slides with poor slide quality (e.g., air underneath the coverslip, excessively faded staining) resulting in substandard quality scans that cannot be used for artificial intelligence (AI).
Despite the many challenges, we have made significant progress in assembling an image data repository. Figure 1 shows the list of our mandatory metadata fields and captures a column header field from an example metadata spreadsheet. In rare instances, there may be a desire to generate more than one WSI from a particular slide, for example, when more than one focal plane must be scanned to capture tissue elements of interest. In these cases, an optional field is included to distinguish multiple scans of a single slide. Further examples of collection metadata fields are included in Online Appendix A. The foundation of robust metadata positions us not only to leverage WSI data within and between units, it also facilitates participation in large-scale digitized histopathology image repository development, such as the IMI Bigpicture project. 13
Controlled vocabularies and ontologies within each metadata field are critical to developing robust metadata. For example, discrepancies as simple as assigning the tissue liver the value of LV, LIV, Liver, liver, and so on complicate searching the tissue field in a database for all samples of liver. In toxicologic pathology, it is standard practice to reference the Clinical Data Interchange Standards Consortium (CDISC) Standard for Exchange of Nonclinical Data (SEND) standard controlled terminology 14 to curate vocabularies, with the recognized caveat that CDISC/SEND does not always include all the needed anatomical terms in all nonclinical species, and is extensible for this reason. We have found terminology discrepancies despite our reliance on CDISC/SEND, most notably in older legacy studies run both in-house and outsourced, and instances when needed terms are not within the lexicon. We have spent significant time aligning controlled vocabularies for metadata fields and registering required terms into our corporate terminology system. To generate a more comprehensive ontology, we referenced and harmonized published ontologies, and then registered and curated specimen ontology based on organism-specific code lists to avoid the need to fit all codes to those of a specific species. To retain consistency of anatomic reference terms across our organizations, we leverage an internal Roche Terminology System (RTS), which is harmonized across common codelists and is used across Roche sites and technology domains. At gRED, we take advantage of a semi-automated internally developed algorithm that maps and harmonizes terminology to this validated dictionary to speed the process of terminology mapping. The consistent terminology enables FAIR data practice and facilitates automation of data conformance checks and subsequent processing.
In addition, the need to record details of tissue preparation (histology) and labeling (e.g., specific histochemical or immunohistochemical [IHC] methods) was anticipated. To facilitate capturing this information at gRED, while streamlining metadata collection, an internally developed registration system was leveraged that allowed an assigned identifier to be obtained for each assay. Our metadata collection then simply references the correct assay identifier to link to all required details of slide generation, such as antibodies used in the case of IHC methods. To accommodate novel or unique assays, we built the assay metadata model to be flexible to collect this information. At pRED, the slide preparation, antibody information (e.g., target and lot number), and the staining protocol are captured in the pathology LIMS with unique identifiers. In the LIMS, all these modules are integrated within the vendor product to allow linking between staining information (name of stain/IHC target) and the tissue slide.
With the foundational element of established metadata requirements to uniquely identify each WSI file, associate it with the animal and study from which it was obtained, and record the procedure used to generate it, we were poised to build our WSI data repository for toxicologic pathology.
Associated Data
To take full advantage of computational methods to gain insights from digital histology slides, it is important to know as much as possible about the biological context and outcomes associated with the histologic samples. However, information about in-life experimental conditions and observations, and later tissue findings, are often stored in data systems that may be separate from each other and are often, if not generally, disconnected from the WSI data repository. Our organizations have been investigating solutions to these issues. Generally, developing mechanisms for direct communication between different repositories through APIs that can facilitate seamless integration and search across systems invisible to the user, while maintaining the database structure of each repository, is the preferred approach. Our teams are working to build infrastructure in which WSI files in our image data repository are discoverable based on searches using data fields from other relevant repositories without compromising the integrity of those database systems or creating data redundancies.
As a top priority in this effort across both gRED and pRED, associated data were first limited to microscopic findings taken from the respective gRED and pRED safety data warehouses, comprising in-house and CRO studies. We found it straightforward to associate findings with the appropriate tissue given the findings data structure in our data lake and data mart, which is based on the SEND data model. However, associating findings to a particular slide scan was sometimes problematic. For example, when a tissue was present on multiple slides, it was challenging to associate a finding to the slide it was identified on, since our findings database only specified the tissue a finding was identified in, and did not capture the slide information. In the future, we suggest that digital pathology will be best served if findings are captured by tissue and slide, so that, particular WSI files can be associated with specific findings. A SEND requirement that would propel development of data entry systems that capture findings by slide as well as by tissue could be helpful in addressing this issue.
As a second step to extend our data usage, we considered association of additional in vivo study data, such as hematology and clinical chemistry (“clinical pathology”) and organ weight data, with histologic slide scans. Associating these types of data has demanded interoperability between data management systems that has posed logistical challenges. In addition, harmonizing especially the clinical pathology data across all our studies has not been trivial. We have found there can be study design variability, for example, in tests and timepoints run for clinical pathology, which can cause data gaps across the studies. Assay methods (e.g., machines and reagents used) vary in our clinical pathology data over time and laboratories. Despite our attempts to leverage reference ranges to harmonize our data and include clinical pathology data without AI recognizable traces of characteristics such as study of origin, assay laboratory, or time-period, we have found this challenging and have ongoing efforts to address these issues.
Slide Scanning
Any analytics approach in digital pathology starts with access to high-quality WSIs. We committed to building our scanned-slide data resources at gRED and pRED, and launched projects to scan all slides from all of our contemporaneous and archived toxicologic pathology studies over the past 25 years. To achieve this, we fully resourced our internal slide scanning laboratories with equipment and staff, and further worked with our CROs to enable them to scan our study slides. Importantly, this effort required us to pay careful attention not only to procurement and installation of the appropriate equipment, but also to dedication of engaged and trained staff both in house and at CROs to lead these efforts, and to development of scanning-process guidelines to enable and unify our approaches.
We selected scanners that could produce high-quality WSIs, scan 40X WSIs with a high loading throughput, and provide good scanner servicing options. Details of scanner capabilities are beyond the scope of this opinion piece and have been reviewed previously. 15 The chosen equipment also provided software with the capacity to offer a quality control step to be integrated into the laboratory quality workflow. Between our organizations, we strategically placed scanners in diverse geographic locations, including North America, Europe, and Asia, and worked closely and collaboratively with all the facilities to develop effective scanning procedures. Our widespread scanner placement and cooperation between our units to leverage each other’s scanning capacity proved especially beneficial when the coronavirus disease 2019 (COVID-19) pandemic closed international borders to travel, and our pathology workflows (pathology peer reviews) were largely reliant on access to digital scans of study slides.
One of the evaluated considerations was whether to leverage the Digital Imaging and Communications in Medicine (DICOM) standards routinely used in picture archiving and communications systems (PACS) for other biomedical imaging modalities. DICOM enables FAIR data capture during image acquisition and standardization of file formats, and has been used in other applications, such as radiology, to standardize and FAIRify image data across the discipline.16,17 However, DICOM is not yet widely used in toxicologic digital pathology despite having been adopted by some newer scanner models, such as the Aperio GT 450 DX scanner (Leica Biosystems, Nussloch, Germany). At the time of our evaluation, DICOM WSI implementation was, and still is, lacking integrated upstream and downstream informatics solutions to fully benefit from its standardization and interoperability advantages. We chose to de-prioritize shifting to DICOM file formats in preference for ease of systems integration. Importantly, the file format produced by our scanners is readily readable into the multiple image viewers used in our organizations for applications as diverse as pathology digital slide review (e.g., peer review), pixel and region-based slide annotation using a variety of open-source and commercial platforms, and image analysis using both commercially available and internally developed approaches including advanced DL methods.
Integrating slide scanning into the study workflows has demanded planning to ensure study timelines are met. There are many approaches to this, for example, scanning the slides in batches to make them available to the evaluating pathologist step by step while scanning is ongoing, scanning after primary glass slide review, prioritizing scanning for open studies over archived ones, and so on. Staffing levels and scanner throughput have the greatest impact on timelines, and planning must be based on the study needs and these constraints. Implementing quality methods is critical under all circumstances to ensure consistency, and results in long-term efficiency as well.
Our teams found that detailed Standard Operating Procedure (SOP) and guidance documents were critical to enabling new external scanning labs to rapidly develop the expertise to scan our tissue slides and expediently and reliably deliver the metadata. It was also important to have dedicated technical teams available to develop the needed skill sets and establish collaborative relationships with our technical staff. Training time, with opportunities for knowledge sharing, was critical to the success of the new slide scanning teams. Technical teams shared documents and met in virtual discussions to clarify points of uncertainty and to refine processes to meet our needs and accommodate the environment and constraints at our partner organizations. The opportunity for dialog, supported by clear documentation of requirements and methods, enabled us to quickly align methods resulting in the generation of high-quality metadata in the required formats with a consistent quality level of WSIs. Online Appendix B is an example CRO slide scanning SOP and guidance document; it demonstrates the granularity of the specifications described. Upon receipt of the WSI files, any issues of image or metadata quality identified at our internal quality check (see the “Quality Check” section and Online Appendix A) are communicated back to the CRO teams to trigger correction/rescanning to ensure data quality, and to allow continuous process improvement.
We choose to perform our scanning at 40X magnification, the highest magnification routinely available on high-throughput commercial scanners. Despite the challenge 40X scans pose to storage capacity due to their large file-size (up to 4 gigabytes per slide on average with larger files possible), we made this choice because of the greater flexibility the high-resolution scans afforded to analytical methods in the long term; slide scans can always be down-sampled to lower magnification, whereas achieving an optical-40X scan from a lower resolution scan would require re-scanning.
Data Management
Transfer of data on the scale of numerous WSI files is daunting. Our teams were especially invested in developing effective data transfer procedures, both to receive digitized slides with corresponding metadata from CROs, and to share data between our teams to build analytics approaches using data-hungry DL methods. As a key component of effective data management, we have created new and integrated workflows for data transfer, either from external contracted organizations or bidirectional data sharing with internal collaborators.
For data transfer practices with external contracted organizations, gRED mainly leveraged the IBM Aspera FASP (Fast, Adaptive and Secure Protocol, IBM Aspera High-Speed Transfer Server v3.9.1), 18 infrastructure for high-speed big data handling. In addition, we have rolled out DTAs that specify the requirements for WSI files and metadata, data packaging instructions with naming conventions, and data transfer instructions for data providers. The DTAs serve as an approved agreement between the data provider and recipient that ensures data are FAIR compliant at the point of data acquisition. Furthermore, the data validation rules in the DTA serve as the baseline for both manually triggered and fully automated programmatic conformance checks of the metadata and the WSI file to ensure cross-validation of the datasets.
pRED uses Arvados, 19 an open-source platform, for managing, processing, and sharing large scientific and biomedical data (https://arvados.org/ discussed further below in V. Storage capacity), as the central place to store and share data. Internal collaborators from pRED and gRED can directly upload data to Arvados within Roche’s network. For CROs, we use Roche’s corporate Data Transfer Platform (DTP), developed internally for use only for Roche research and development data, which provides secure data upload from outside the network including a direct deposit into Arvados. From there, the Arvados API allows us to automatically locate relevant WSIs for sharing as well as to automatically synchronize the shared folder on Arvados with local copies at gRED and pRED. Files in Arvados can be copied inside the platform without creating physical copies of the data, such that, scans that are re-uploaded or copied within Arvados are physically only stored once. Thanks to this feature, the slides can be stored by study and again by collection for algorithm training without duplicating the physical data storage.
Storage Capacity
As alluded to above, the storage requirements for a data resource of WSIs pose significant challenges. A typical toxicologic pathology study can easily generate more than 1,000 glass slides and, with each slide scan at 2 to 4 GB in size at 40X resolution, 1 TB or more of storage capacity may be required. Demand for storage can increase by hundreds of terabytes annually if all of an organization’s studies are scanned and stored, as is desirable if a data resource suitable for developing DL analytics approaches is to be built. Scanning of tissue slides from archived studies to expand the data available for DL model building further increases the demand for storage.
The types of storage needed for the WSI data can be constrained by usage requirements. For example, high-capacity storage may have insufficient performance with respect to the volume of data that can be processed per unit time (throughput) and data processing speed (latency) to allow for efficient pathologist review of the WSIs or active model training on the data. Conversely, data that are not being actively used likely does not require high-performance storage. Meeting the need to store the vast amounts of WSI data in the data repository combined with the requirement to rapidly access data needed for particular projects necessitates a well thought out storage strategy that is cost-effective, avoids data duplication, and retains provenance (origin/history) of any WSI. It is important to state that whatever storage solution is in place, interoperability with other systems and software is crucial to connect the WSI file with metadata and other associated data. Furthermore, the storage solution(s) not only needs to allow easy access of WSIs for viewing, model training, and computational evaluation as mentioned above, but also should permit file transfer and sharing to enable collaboration.
Each of our organizations has taken a somewhat different strategy toward addressing the storage of our toxicologic pathology digital slide resources. pRED has implemented Arvados on premise to store WSI data. Apart from offering a distributed and secure storage, Arvados provides automatic deduplication of data to avoid reduplicative use of storage resources. From a FAIR data 11 point of view, Arvados enables pRED to tag data collections with a minimum set of metadata that makes data easier to find and track across multiple systems. For instance, for each collection of WSIs, the unique study identifier and the system in which this identifier is registered must be specified for every dataset. In this way, the WSIs for a study are easily findable on Arvados, and it is clear in which system additional information on the study can be found.
Cloud storage implementation for maximum performance, flexibility, and scalability is planned at gRED. However, to meet near and mid-term needs, the interim solution of leveraging on-premises infrastructure by combining a high-capacity staging area for data validation and processing, followed by a high-performance storage destination for integrated WSI viewing and advanced analytics is being employed for both in-house-generated and CRO-transferred datasets. This approach better prepares our infrastructure to be mirrored to a two-step cloud storage solution in the near future, in which we can decouple and accommodate different user groups’ specific needs within the data store at an optimized cost.
Storage options are rapidly evolving, and future best practices may be very different from today. Opportunities to transition from filesystems to object store that will (1) relax the constraint on searchability and scalability posed by folder hierarchy, simple structured metadata, and limited namespace and (2) enable smoother indexing and more cost-efficient storage of high-dimensional unstructured data by allowing higher granularity of metadata, are being investigated by the digital pathology community.20,21 Leveraging object storage strategies on cloud storage systems promises to enable “accessing data from anywhere, at any time” thanks to its simplified data management, geospatial-distributed storage, and robust data protection capability. Moreover, particularly relevant to the rapidly growing data volume needs of WSI data repositories, it is more cost-effective to scale object storage than filesystems. In addition, accessibility from multiple sites enabled by cloud storage allows data access from any geographic location, with the advantage that performance optimization is subject to local networks and can be locally optimized. In combination with next generation file formats, such as Zarr, that enable data accessibility in a more granular, non-monolithic fashion, this flexible strategy promises “cloud elasticity”; the ability to expand or decrease computational parameters and resources as needed to meet fluctuating demands for computing resources that would otherwise be constrained by conventional storage solutions. 22
Conformance Check
Effective data ingestion into our image management system from both internal and external sources requires assurance that data and metadata are correctly aligned, all files are accounted for, and data and metadata are error-free. This is challenging in the context of high-volume data and demands automated solutions both to complete conformance checks in a reasonable time and to avoid the introduction of human error.
gRED has approached this effort at multiple levels. First, upon receipt, a basic conformance check is performed for authenticity based on file properties (filename, size, checksum, and timestamp) as well as a metadata alignment with DTA requirements. From there, a layered cross-validation method is executed to ensure that information embedded in the WSI file matches up with the validated metadata, such as embedded technical metadata, label information, and downsample image patterns for specimen and assay type. For this purpose, an automated method was developed—Image Tale (patent pending) where a modular approach is adopted for: (1) technical metadata extraction and comparison; (2) label Optical Character Recognition (OCR)/decoder; (3) a DL-based bimodal assay color pattern classifier to automatically recognize H&E vs. non-H&E images; and (4) a DL-based specimen recognition to ensure data validation.
Similar conformance checks are done at pRED upon receipt of the scans. Once WSI files are imported, a spot-wise manual check (currently about 10% of WSI files based on experience) is performed to assess whether the metadata (associated and file-embedded) matches the WSI file.
Quality Check
The quality of WSIs has an important impact on the ability of pathologists to effectively examine the virtual slides and the success of computational scientists to develop algorithms or perform computational analyses on the slides. The first step in ensuring scan quality is addressing the quality of the physical slide that will be scanned; this has a significant impact on the eventual scan quality as mentioned in a recent review. 3 However, scanning can also introduce a variety of artifacts, 3 and quality checks of digital scans are additionally required.
Our organizations have employed a number of strategies to ensure that image quality meets a sufficient standard, 10 ranging from leveraging tools built into scanners, human evaluation of the WSIs, either by the pathologist or a technician, or by technical staff assisted by semi-automated detection methods for scanning artifacts.
As a first step (at both sites), the blurriness detection tool built into scanners is leveraged to quickly identify severely out-of-focus scans. The performance of this blurriness detection tool was internally evaluated with an extensive list of organs evaluated in nonclinical environments, and best practices were shared with the technical staff responsible for scanning. These recommendations included how to optimally establish focus areas and focus points during scanning processes, guidance on the interpretation of the QC blurriness tool, and potential strategies for re-scanning with improved quality. Specific tissues that are frequently misclassified as out of focus were highlighted in these protocols (e.g., homogeneous poorly cellular tissues, such as bone, lens, or fluid-filled structures). In a second step, additional QC is performed by the technical staff, including confirmation that the number of WSIs generated matches the total number of glass slides scanned, and a check for image quality and barcode accuracy is performed in a representative selection of WSIs, including confirming correspondence of the code in the slide label and the code read by the scanner. At pRED, correspondence with animal No., Block No., Organs, Study No,, and so on is also confirmed, whereas at gRED, since all our barcodes are generated in-house, this step has already been completed during bar code generation.
Recently, gRED has incorporated semi-automated computational methods into the workflow that speed up the assessment of out-of-focus regions, greatly increasing the capacity of our technical staff to QC slide scan quality. In the future, we hope to increasingly rely on fully automated image-quality assessment as methods for this process are refined and further developed.
At both our sites, we have found that performing QC at the scanning phase (when the slides are still within the scanner) dramatically decreases the need for subsequent time-consuming steps that require retrieval and re-scanning of slides. Based on our experience, for a batch of 360 slides and making use of the QC tool integrated in the scanner, approximately 8 to 12 hours are required for the QC check, although this time may broadly vary depending on several factors, including tissue types and size (rodents vs. large animals).
Searchability and Interoperability
The true utility of any data repository emerges when researchers and practitioners are able to quickly identify and retrieve data of interest for a wide range of applications. For example, a scientist may want to search for: WSIs from a specific study; certain organs (e.g., liver, kidneys) either within or across studies; WSIs associated with certain microscopic findings or other pathology data; and/or scans within a certain range of prediction values of an AI tool. To achieve maximal utility and flexibility, the repository should be searchable based on any information associated with the image either as metadata or as associated data in other information systems. We have found that the data we are interested in using to search our WSIs are most often housed in other data repositories. The best practice is to create interoperability between data storage systems that will allow association and efficient search of data among them without data duplication. Whenever possible, we aim to link databases through APIs that will allow us to, for example, search for WSIs using information contained within our in-life study data collection systems. Linking databases in this way requires careful assignment of identifiers (keys) within the database systems to ensure that the linkages between data are reliable and meaningful. As discussed earlier, it all comes back to careful metadata planning to enable this sort of interoperability.
Challenges
Common data management challenges that may not be specific to digital pathology have been considered extensively, 12 including (1) partial information availability for archived studies, such as missing data in studies performed prior to modern data management practices; (2) lack of a standardized vocabulary; (3) lack of the necessary details to distinguish similar datasets from different studies; (4) scarcity of streamlined end-to-end data management processes; and (5) dispersed data quality control. We have found that standardization of vocabularies has been particularly vexing at multiple levels of data alignment. Even resolving discrepancies in abbreviations used to identify particular organs has been a challenge, as mentioned previously. Orders of magnitude harder, some level of categorization of pathology diagnoses is also desirable to enable searchability. In toxicologic pathology, we are constrained by the SEND format and already utilize standardized nomenclatures for pathology findings. This use of curated glossaries to record microscopic findings helps unify terminology across time, practitioner, and instance, and makes finding classifications more useful for meta-analysis across studies. Links to CDISC nomenclature or other terminology service/data curation tools are extremely helpful in bringing terminologies into conformance for these activities. However, this requires additional work especially on old retrospective studies.
Conclusion
In summary, building the data resources to support digital toxicologic pathology in a pharmaceutical setting is a highly complex task encompassing everything from collecting and storing critical data about samples, slides and WSIs, to conceptually framing the relationships between the data across multiple databases. At our organizations, tackling these complex challenges has taken the dedicated efforts of large multidisciplinary teams, including data scientists, pathologists, database systems professionals, infrastructure architects, hardware specialists, and data cleaning specialists. In many instances, we have confronted challenges with retrofitting systems that were not built to work together to enable their interoperability, and analogously, shifting people to a new culture of unfamiliarly high levels of data stewardship. However, as this new resource of FAIR data and interoperable systems is built and can support the magnitude of data that digital pathology requires, the reward becomes the ability to leverage this information to build cutting-edge analytics approaches and advance pathology in our digital age.
Supplemental Material
sj-docx-1-tpx-10.1177_01926233221132747 – Supplemental material for Toxicologic Pathology Forum*: A Roadmap for Building State-of-the-Art Digital Image Data Resources for Toxicologic Pathology in the Pharmaceutical Industry
Supplemental material, sj-docx-1-tpx-10.1177_01926233221132747 for Toxicologic Pathology Forum*: A Roadmap for Building State-of-the-Art Digital Image Data Resources for Toxicologic Pathology in the Pharmaceutical Industry by Xing-Yue Ge, Juergen Funk, Tom Albrecht, Merima Birkhimer, Moritz Gilsdorf, Matthew Hayes, Fangyao Hu, Pierre Maliver, Mark McCreary, Trung Nguyen, Fernando Romero-Palomo, Shanon Seger, Reina N. Fuji, Vanessa Schumacher and Ruth Sullivan in Toxicologic Pathology
Footnotes
Acknowledgements
The authors would like to acknowledge the technical staff at the scanning labs for their assistance with data acquisition and annotations, our outsourcing teams, as well as the animal care and histology laboratory staff who contributed to the generation of the original histologic slides upon which this work is based.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors are employees of Roche/Genentech.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
ORCID iDs
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
