Abstract
The 2019 manuscript by the Special Interest Group on Digital Pathology and Image Analysis of the Society of Toxicologic pathology suggested that a synergism between artificial intelligence (AI) and machine learning (ML) technologies and digital toxicologic pathology would improve the daily workflow and future impact of toxicologic pathologists globally. Now 2 years later, the authors of this review consider whether, in their opinion, there is any evidence that supports that thesis. Specifically, we consider the opportunities and challenges for applying ML (the study of computer algorithms that are able to learn from example data and extrapolate the learned information to unseen data) algorithms in toxicologic pathology and how regulatory bodies are navigating this rapidly evolving field. Although we see similarities with the “Last Mile” metaphor, the weight of evidence suggests that toxicologic pathologists should approach ML with an equal dose of skepticism and enthusiasm. There are increasing opportunities for impact in our field that leave the authors cautiously excited and optimistic. Toxicologic pathologists have the opportunity to critically evaluate ML applications with a “call-to-arms” mentality. Why should we be late adopters? There is ample evidence to encourage engagement, growth, and leadership in this field.
Keywords
The “Last Mile” metaphor was first used by the early land-based telecommunications industry 1 to describe the difficulty of connecting end-user homes and businesses to the main telecommunication network. One of the main barriers was the cost of installing and maintaining this infrastructure, because it could only be amortized over 1 subscriber, compared to many customers in the main trunks of the network. Other challenges of “Last Mile” delivery included ensuring transparency with the customer; following guidelines from local, state, and federal regulatory agencies; increasing efficiency of the workflow; and improving infrastructure supporting the network. Interestingly, these 4 challenges overlap with common ones we currently face as toxicologic pathologists when confronted with the notion of applying machine learning (ML) 2 in the digital histopathology space. In this review, we highlight both published medical pathology examples and our personal experiences to date with ML applications in toxicologic pathology, which indicate that we are starting to overcome some of the “Last Mile” challenges. As most of us work in a highly regulated environment and are at least cautious about implementing new approaches or technology because of the perceived burden of qualification or Good Laboratory Practice (GLP) validation, we also share personal thoughts on early qualification and validation efforts and the challenges therein.
In our opinion, and as pointed to in the 2019 Special Interest Group manuscript, 3 a continued rapid growth in each of the 3 key ingredients of ML: (1) massive computer power, (2) big data and (3) inherent knowledge, has fueled accelerated use of artificial intelligence (AI) and ML in almost all areas of science. More and larger partnerships between AI scientists and medical specialties are being established, and the inference (causal and counterfactual) and probability (prediction) output of machines are married with the high-level decision-making and reasoning of humans at an increasing frequency.
Yet we need to be cautious of the hype surrounding all things AI. In its June 2020 edition, the Technology Quarterly of the journal
Based on the authors’ experiences of developing and applying ML solutions to digital histopathology data, as was the case for ML-based image analysis and stereology solutions, we feel that general adoption of ML is achievable and the potential return on investment favorable. However, ML is not a panacea and, like all scientific methods, should be applied when its use adds clear scientific or operational value. We currently see several application areas and opportunities for ML growth in digital toxicologic pathology: (1) abnormality detection, (2) decision support, (3) tissue lesion screening, (4) diagnostic scoring simplification, (5) counting automation, and (6) object quantification. Examples of these are shown in Figure 1 and described in the following paragraphs.
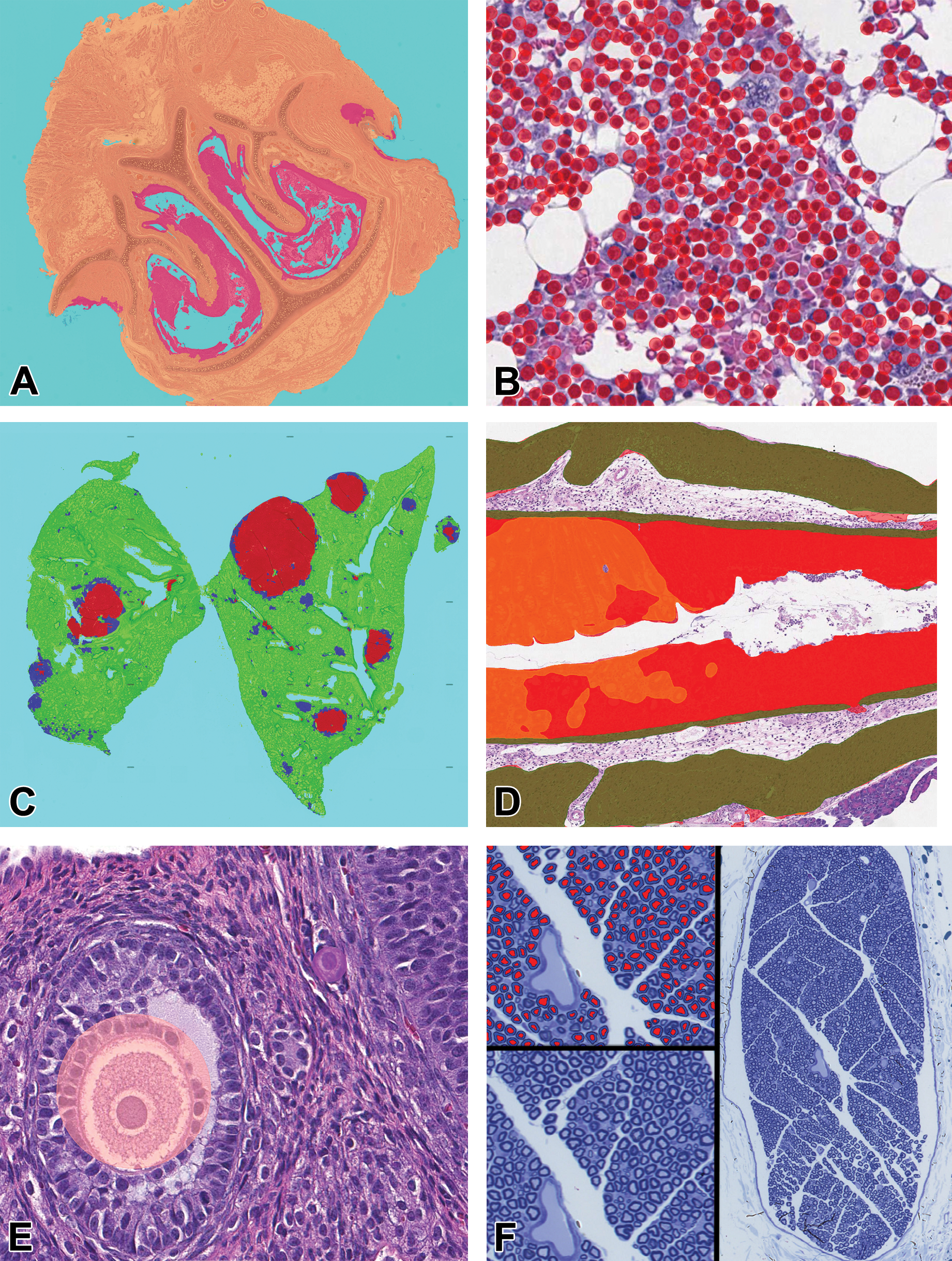
(A-F) Potential machine learning applications in toxicologic pathology. A, Abnormality detection—Colored masks highlight abnormal regions (red) of the rat anterior nasal cavity (Courtesy of Sam Neal). B, Decision support—Bone marrow masks mark hematopoietic cells in cynomolgus monkey bone marrow, allowing for enumeration in support of qualitative scoring. 5 C, Lesion screening—Machine learning (ML) masks identify proliferative lesions in TgRasH2 mice (blue = hyperplasia; red = adenoma). 6 D, Scoring simplification—ML segments mucosa (red and orange) and muscle/serosa (green) and measures mucosal inflammation area (red) and unaltered mucosa (orange) in murine efficacy model. 7 E, Counting automation—An ML-based object counter quantifying different types of ovarian follicles (pink = antral; purple = primary) (Courtesy of Lauren Prince). F, Object quantification—ML-based segmentation of myelin for calculations of axonal-G ratio analysis (Courtesy of Michael Staup).
Abnormality detection. Toxicologic pathologists evaluate as many as 40 tissues per animal in a toxicology study, and a majority of these tissues are normal or display common spontaneous lesions. Using ML algorithms could help pathologists efficiently identify potential target organs and the dose response within a study. The ML algorithms have been shown to accurately and quickly detect morphologic changes in the layers of the retina, 8 for example.
Decision support. Toxicologic pathologists determine the diagnostic terminology, thresholds, and scoring paradigm for every histological finding noted in a study. Maintaining consistency within and across studies can be difficult. The ML algorithms have the potential to assist a pathologist in diagnostic criteria, thresholds, and scoring criteria. Examples include spermatogenic staging, 9 evaluating the number and phenotype of cells in bone marrow histology 5 (Figure 1B), and quantification of immunohistochemical markers. 10 The ML applications could also support the technical staff creating the digital whole-slide images (WSI). As is the case for glass slides, the quality control of WSI is an important best practice but is time-consuming and manual. The ML-based methods are being developed to automate digital slide quality control review and could increase the efficiency of this workflow dramatically. 11,12
Lesion screening. ML algorithms can support the targeted evaluation of specific diagnostic end points such as hepatocellular hypertrophy, 13 TgRasH2 model proliferative lesions 6 (Figure 1C), and rat cardiomyopathy. 14 These types of ML algorithms could benefit both early discovery (ie, lead optimization) and screening toxicology efforts and later stage evaluations, the latter when a specific finding is expected. For screening and lead optimization studies, scientists are iterating on a series of potential compounds where data speed and consistency are paramount. Microscopic end points are usually rate limiting in these studies and accompany other safety and efficacy biomarkers. The ML algorithms that are tuned to a specific finding have the potential to facilitate the pathologist’s evaluation of these studies. As test agents progress in development, a pattern of microscopic findings often emerges, and ML algorithms could be developed that screen for these expected changes and provide a pathologist a rapid picture of the presence, dose response, and progression of the expected microscopic change.
Scoring simplification. Efficacy models scored by pathologists often have complicated scoring systems that are based on combining several qualitative or semiquantitative scores into a summary score. The ML algorithms could be used to both simplify and improve quantification of the scoring in these types of models and were used for colitis in a mouse model 7 (Figure 1D). Scoring simplification also has possible applications in general toxicology studies. Certain tissue responses can be complicated (eg, cardiomyopathy, nephropathy, nasal cavity, testicular and injection site changes in multiple species), requiring a pathologist to record and score several diagnoses. 14 Designing ML algorithms that provide support to the pathologist to simplify these evaluations could be beneficial.
Counting automation. Toxicologic pathologists will often engage in manual, counting activities as part of either discovery or toxicology studies. Example where ML algorithms have been used to automate these processes based on our own experience as well as published reports include counting mitoses, 15 ovarian corpora lutea or follicles, 16 apoptotic figures, AAV capsids, and specific inflammatory cell infiltrates (numbers of specific types of leukocytes). Specifically for using ML algorithms to identify and count ovarian follicles (Figure 1E) and mast cells and leukocytes (unpublished data), we have found both reproducibility and speed to be improved considerably using ML.
Object measurements and quantification. As was the case for counting, certain discovery and toxicology study designs require quantitative measurements that are difficult and/or time-consuming to complete manually by the pathologist. The ML algorithms that can standardize and automate these quantitative changes would be beneficial to the pathology workflow. Examples include those described in this digital pathology special issue such as mammary gland epithelial proliferation, 10 hepatocellular hypertrophy, 13 and, in our own experience, an automated method for measuring nerve fiber axonal-G ratio (Figure 1F).
The emergence of the Clinical Laboratory Improvement Amendments qualified and validated AI applications in medical imaging and clinical diagnostic pathology has been motivated by the need to improve timelines and efficiency in clinical care. 17,18 Although our previous experience as well as the abovementioned examples suggest potential advantages for ML in driving efficiency and reproducibility in the digital toxicologic pathology workflow, we need to be careful and deliberate when considering ML. The ML needs to address well-defined, focused end points such as identification and characterization of specific changes in animal efficacy and safety models. 6 In our experience, the ML algorithms that appear most helpful in the workflow for a toxicological pathologic study evaluation are those that provide an automatic discrimination of normal and abnormal tissues (“abnormality detection”). Nonclinical safety studies present the pathologist with complex, multidomain data that drive the pathologist to make considerations on the safety of the investigated compound. Understanding and systematizing this complex series of mental queries and using it as a basis for training a computer to complete specific tasks, in an iterative manner, may bring us closer to developing ML tools for abnormality detection. This could significantly accelerate the pathologist’s review of normal and abnormal tissues and increase the transparency of the process based on the algorithm’s output.
The development of ML algorithm requires considerable effort and quality control and qualification procedures. Once the intended use for the algorithm has been clearly defined, the appropriate training data must be collected and collated in order to build the algorithm. Each tissue from the different species will require a well-curated and heterogenous collection of WSI. Pathologists are best suited to guide the data set selection in that their review of the WSI has much greater depth than key term searches of databases. Separate data sets should be used to teach, test, qualify, and validate an algorithm. The European Innovative Medicines Initiative “Big Picture” project 19 is poised to be a novel source of large data sets and may be a resource for the validation of algorithms, allowing their use in the nonclinical environment when tied to appropriate regulatory frameworks.
Although we believe the ML examples described earlier have great potential to impact our field, there are some inherent challenges in applying ML to digital histopathology. Three in particular include (1) classification across magnifications: For many classification tasks, the context needed by the computer to accurately identify a specific class is limited by training at a single magnification 6 ; (2) grading uncertainty: The classes of some microscopic changes (eg, lung hyperplasia vs adenoma) differ much more subtly than other classes, causing annotation and classification uncertainty 6 ; and (3) rare event measurements in toxicologic pathology; some classes may be rare, such that obtaining adequate amounts of training data for such classes is challenging. 6
To address the first challenge (classification across magnifications): Multimagnification analysis methods that resemble how pathologists analyze histologic slides using microscopes have already been developed and implemented in breast cancer diagnostics 20 and are being developed in the nonclinical space. 21 This multimaginfication approach combines the patch-level information with the information gathered from the context of larger fields of view at lower magnification and have shown improved performance in comparison to single-magnification classifiers. 21 In the TgRasH2 mouse model work, stomach papillomas and hyperplasia were not well differentiated at the ×10 magnification, and when a lower training magnification was employed (×2.5), the additional context resulted in high performance (F1 scores). 6
For the second challenge (grading uncertainty), ML algorithms could help standardize our individual grading and thresholds across a variety of studies or even across pathology groups and laboratories. The guidance an ML algorithm provides on grading or thresholds is based on probability, and although the algorithm may indicate a strong likelihood for a certain class (a specific grade or diagnosis), this prediction can still be highly uncertain, mislead the pathologist, and result in false-positive or false-negative predictions. As pathologists, we will not see the numbers behind the prediction, and this lack of transparency is often referred to as the “black box” nature of deep neural networks. Although beyond the scope of this minireview, techniques are starting to be adopted in computational pathology that will help the pathologist understand the source and amount of uncertainty related to a class prediction. 22
The last challenge (rare events) is especially relevant for a toxicologic pathologist. Is it possible to design a computer algorithm that could assist a pathologist in identifying a rare novel or subtle lesion? The so-called “out-of-distribution-detection” methodology is being developed that produces an ML model that has an internal distribution of normal and background lesions (ie, a “reference range”), both which account for most of what is observed by a toxicologic pathologist. Using this ML-based model built-in tissue-reference range, the algorithm could find the changes that have a low likelihood of belonging to the distribution (ie, “outliers”); hence, they are presented to the pathologist as potential important changes. Several recent publications have described the application of these methods to WSI, for example, allowing a breast cancer metastasis detection system to flag lymphomas, even though it was not originally trained to recognize them. 23 We think this is a field that will grow in importance as we are able to classify common disease patterns more reliably, but keep struggling with rare entities due to limited data sets.
Another area that is impacting the “Last Mile” of toxicologic pathology digitalization is the rapid evolution of both supervised and unsupervised ML deep learning (DL) techniques (part of a broader family of ML methods based on artificial neural networks with representation learning, which can be supervised, semisupervised, or unsupervised) in digital tissue image analysis. 24 Due to the nature of toxicologic pathology, we believe that DL applications optimized for sensitivity are critical so as to not miss any of the affected samples. However, the specificity of these tools cannot be neglected, as identification of too many “false alarms” decreases the toxicologic pathologist’s work efficiency and potentially defeats the purpose of using the tool itself. In the unsupervised DL approaches, data points with similar features are grouped (classified) together without any supervision (examples) provided by the experts. The danger associated with this approach in ML is that classes identified may not correspond to classes containing relevant pathological or biological information. There is also a risk of identifying classes based on uninterpretable artifacts. This is why interpretation and later labeling of the resulting data classes by pathologists and life scientists is crucial. A discipline which addresses this problem is explainable AI (XAI). Explainable AI produces a summary to the pathologist of the AI-based criteria for classification without sacrificing AI performance. 25,26 In addition to predicting the class for the pathologist, the computer provides the list of attributes (eg, color, size) that support the prediction. The XAI approaches are important when the decisions produced by AI models are applied to disciplines such as precision medicine or in identification of adverse health effects in the drug development and safety assessment process. Recent European regulations require this type of AI transparency be included in the global data protection regulations. 27
Since the publication of the 2019 Special Interest Group manuscript, 3 we still perceive that qualification and GLP validation continues as a barrier in ML implementation for digital toxicologic pathology even though ML approaches supporting image analysis and stereology have been a part of our workflow for several years. In January 2020, the Food and Drug Administration published a white paper entitled “Artificial intelligence and machine learning in software as a medical device.” 28 Although focused on clinical applications, the white paper has links to several important guidances, a regulatory framework discussion paper, and public workshop minutes that demonstrate the strong interest our regulatory partners have in this area. We need not fear the qualification and validation process. There is a systematic approach that we can take as an industry that insures these technologies are implemented responsibly and effectively. The IMI BigPicture 19 repository may be a greater resource of images for qualification and validation, which could significantly accelerate these processes.
In conclusion, although the “Last Mile” challenge of ML application in digital toxicologic pathology is real and should not be underestimated, the authors believe that ML algorithms have several applications and can be implemented safely in our regulated environment. Although perceptions of low/dubious return on investment still linger and limit some from taking a first step toward digital toxicologic pathology, more and more affordable tools are becoming available, and we feel that the potential benefits justify the initial investments. Good science should always demand a level of skepticism regarding large claims and rapid changes. Although we should move with appropriate caution, it is our opinion that we should not hesitate. Building ever stronger relationships within the computational pathology community and with the health authorities will help usher in an exciting new dawn for quantitative, minable, and predictive histopathology data for toxicologic pathology.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed no financial support for the research, authorship, and/or publication of this article
