Abstract
Toxicologic pathology is transitioning from analog to digital methods. This transition seems inevitable due to a host of ongoing social and medical technological forces. Of these, artificial intelligence (AI) and in particular machine learning (ML) are globally disruptive, rapidly growing sectors of technology whose impact on the long-established field of histopathology is quickly being realized. The development of increasing numbers of algorithms, peering ever deeper into the histopathological space, has demonstrated to the scientific community that AI pathology platforms are now poised to truly impact the future of precision and personalized medicine. However, as with all great technological advances, there are implementation and adoption challenges. This review aims to define common and relevant AI and ML terminology, describe data generation and interpretation, outline current and potential future business cases, discuss validation and regulatory hurdles, and most importantly, propose how overcoming the challenges of this burgeoning technology may shape toxicologic pathology for years to come, enabling pathologists to contribute even more effectively to answering scientific questions and solving global health issues.
*This article is a product of a Special Interest Group of the Society of Toxicologic Pathology (STP). The views expressed in this article are those of the authors and do not necessarily represent the policies, positions, or opinions of the STP.
Keywords
Introduction
Artificial intelligence (AI) has begun to augment and improve growth, productivity, and profit across both medical 1 and nonmedical industries. 2 A 2018 Harvard business review opined that “AI will change radiology, but won’t replace radiologists.” 3 In 2019, Wu et al showed that a hybrid model, averaging probability of malignancy predicted by a radiologist and a neural network combined, was more accurate than either of the 2 separately. 4 Litjens et al reviewed how medical imaging of the brain, eyes, and chest has been transformed by recent AI developments. 5 The notion that AI and machine learning (ML) might augment the quality and efficiency of our work as toxicologic pathologists is exciting but balanced by the fear that our day-to-day contributions might be replaced by the output of machines. Historically, progress in AI and ML were limited by the cost of sufficient computational power combined with an inability to efficiently digitize histopathology data. However, digital toxicologic pathology might be at an inflection point: Progress in computational pathology is rapidly evolving and the advent of multiple convolutional neural networks (CNNs), spurred by global image classification competitions (eg, ImageNet and CAMELYON challenges 6 -9 ), has created an environment where we are rapidly moving up the “slope of enlightenment.” So, we are at a point where medical imaging fields, both human and veterinary, are embracing these technologies and producing impressive, human equivalent, or even “better-than-human” results.
Artificial intelligence is consistently superior to humans at completing repetitive, detailed tasks rapidly and accurately and algorithm performance has been shown to be comparable with that of an expert pathologist interpreting whole-slide images (WSIs) without time constraints. 10 However, it is important to consider the high level of cognition the pathologist provides versus the computational advantages of AI. Although the importance of cognition may become less important in the future, as was described in the above example for the radiologist, the complementary impact or even synergies between medical, veterinary, and toxicologic pathologists and AI are potentially very powerful. For example, medical pathologists working in partnership with AI were able to identify metastatic breast cancer cells in lymph nodes more accurately (99.5%) than either AI-unassisted pathologists (96%) or AI alone (92%). 11 Toxicologic pathologists, in particular, should appreciate how study design and workflow may radically and advantageously change upon the advent of effective AI-human partnerships. The authors’ assess that, based on the integration of AI into other medical fields and current trends, AI will not replace all human tasks but rather will augment human speed and accuracy. Consequently, there is a pressing need to consider the best management of AI in highly regulated environments. 3
Definitions
Before considering the topic further, we will define some basic AI terms. Random forests or random decision forests were some of the first AI-type algorithms developed. They are an important form of ensemble classification, a data mining approach that utilizes a number of classifiers that work together to identify the class label for unlabeled instances. 12 We will not expand on these more established terms here however but will define below the more relevant terms of AI, ML, artificial neural networks (ANNs), and deep learning (DL) and their relationships with each other (Figure 1).
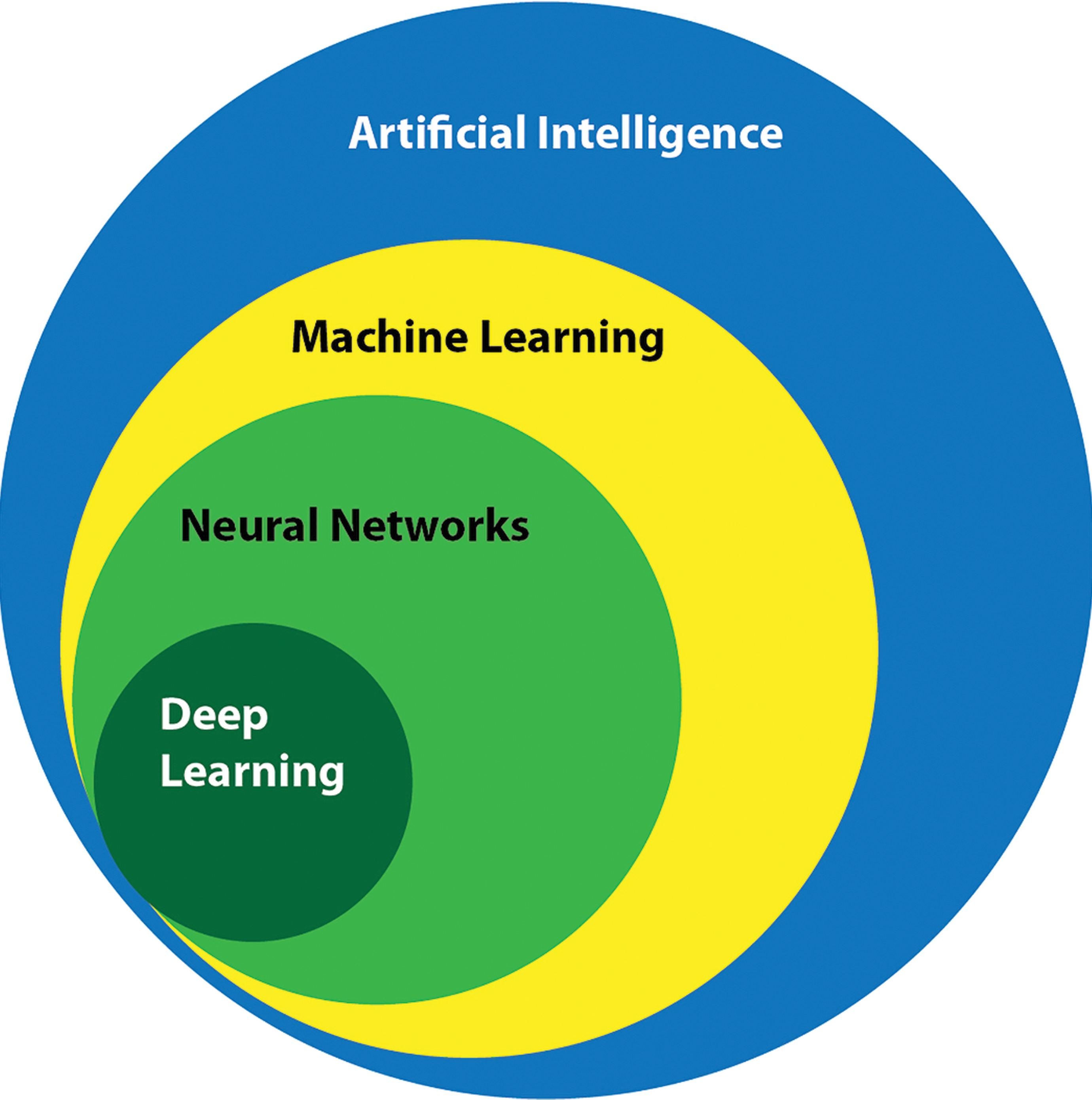
Overview of the relationships between disciplines within the field of artificial intelligence discussed in this article.
Artificial Intelligence
In the 1940s and 1950s, the development of digital computers led to the creation of programs that performed elementary reasoning tasks. These first computer programs could, for example, play chess, 13 and today, computers can play and outperform humans at the vastly more complex board game GO. 14 The term “artificial intelligence” was initially introduced by John McCarthy at a 1956 Dartmouth conference on AI, defined then as “the science and engineering of making intelligent machines.” 15 Artificial intelligence represents a broad category that encompasses all computer-based algorithms advancing decision-making processes. It operationalizes a computer system or a group of integrated systems to perform one or more tasks that historically would require human intelligence. 16 In the realm of histopathology, these tasks include, but are not limited to, interpretation of image data to define and label structures, tissues, and abnormal changes within these images.
Machine Learning
Machine learning is a discipline within AI that gives computers the ability to self-learn without explicit programming and to improve performance by continuously learning from experience. 16,17 More simply, ML refers to the ability of computers to analyze data and identify novel patterns without additional human programming. This type of computational self-learning may improve algorithm performance as measured by advances in innovation, efficiency, or quality.
By blending computer science and statistics to solve problems not previously solvable, ML applications are rapidly expanding their impact in medicine, and new ML health-care applications often top lists for medical funding. Machine learning approaches are gaining hold in image-based diagnosis, disease prognosis, and risk assessment. 18 There are 3 ML learning categories: supervised, unsupervised, and reinforcement (Figure 2). 16,19 -22
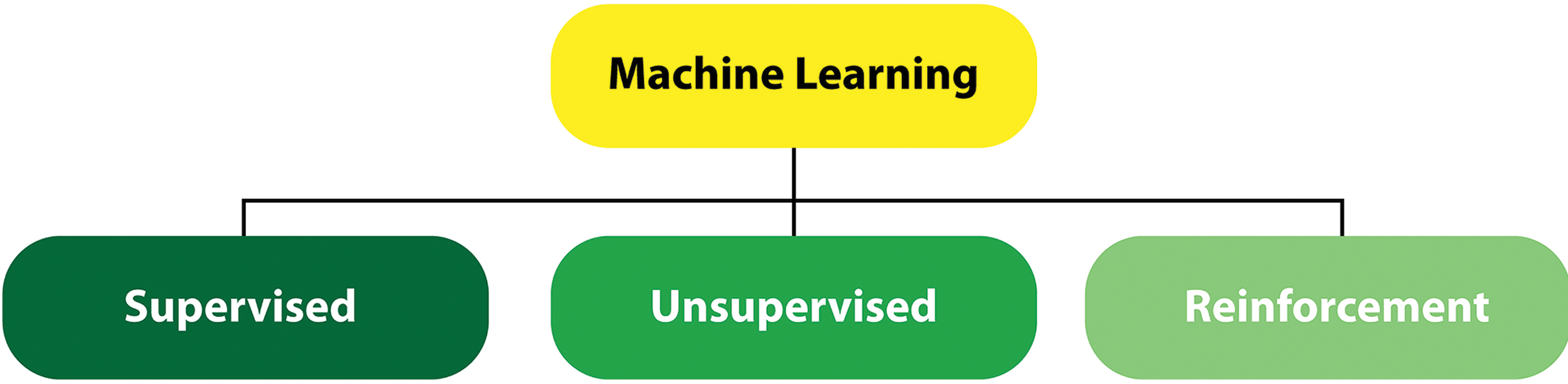
Machine learning encompasses 3 separate categories of learning.
In supervised learning, the computer is provided with the features related to the target (input) to train the algorithm. The label (ie, name, output, etc) of the target is also supplied. Three data sets (training, validation, and test data) are used in supervised learning. The computer is trained with a training data set that provides representative features as input. For example, to use supervised learning to identify a liver, a set of images of normal livers with normal major representative subfeatures such as hepatic lobules, hepatic cords, sinusoids, portal regions, and central veins are presented to the computer and the label (ie, liver) is provided. The algorithm is then trained to identify the representative subfeatures of the liver and the performance of the algorithm is tested using a validation data set. Validation data are distinct from the training set and contain labeled data; in this example, a collection of liver samples is used to fine-tune and choose the best algorithm. This will enable a more accurate prediction of an unlabeled tissue when presented to the algorithm. Finally, the algorithm is challenged with an independent test data set, for example, from different studies. In this example, the test set will consist of unlabeled normal livers. By identifying learned representative subfeatures in the test data, the algorithm should be able to predict the label “liver.” Thus, supervised learning is task driven. 23 The accuracy of properly labeling unknown targets often increases with the size of the training data set. 16
Unsupervised ML is used in instances where it is difficult or impossible to define characteristic features of biological tissues. In this approach, the computer is presented with a collection of different tissues from various organs and the computer identifies patterns in the tissues and clusters those with similar patterns together. The overall goal of unsupervised ML is to identify structures in the input data without prior user definition of the output, making it a data-driven and hypothesis generation approach. 16,21 The patterns identified in unsupervised learning not only provide answers but also raise questions that may not have been conceived by the investigators. While the innovation here is exciting, the patterns require evaluation by either a human (ie, a pathologist) or a supervised learning task. 17 A common application of unsupervised learning is to explore complex interrelationships between genetics, biochemistry, histology, and disease states. 17 There are many other subtypes of supervised and unsupervised ML 21 ; however, these are beyond the scope of this review.
The third broad category of ML is reinforcement learning, wherein the computer learns from its mistakes. In this approach, the computer is supplied with an unlabeled input and forced to predict the output. If the prediction is incorrect, the computer is provided with the correct label. The computer then uses these data for future predictions. 22 Reinforcement learning is thus another paradigm of learning—it is about learning an optimal policy. Examples of outcomes from reinforcement learning include self-driving cars and game playing (eg, recent headlines regarding alpha-GO, etc).
Artificial Neural Networks
Artificial neural networks are computer systems designed to model the neural networks and learning ability of the human brain. 24 They contain a number of processing elements (neurons) that are highly interconnected in a unidirectional pattern. 25 On their own, these neurons can perform only simple tasks, but as a network, they can execute more complex tasks. 26 Artificial neural networks receive outside information via an input layer, and information is propagated by neurons arranged in “hidden layers.” The transformation occurs through the weights and biases assigned to one layer which influence if and how information is moved to the next layer. The output layer contains the calculated output. The training of the network relies on the user to define the weights and biases that determine the strength of connections between neurons. 24,25 This structure allows a computer to teach itself how to learn by reviewing a large data set. 27 These data sets can be raw and unstructured (ie, lacking known or defined features). 16 Artificial neural networks are a topic of intense research within the field of biomedicine, as they learn by experience and do not require detailed input to fulfill a task. 25,26
Deep Learning
Deep learning is a class of ML that learns through neural networks in supervised and/or unsupervised patterns with multiple levels of representations that correspond to different levels of abstraction. As such, DL cannot be considered a single technology but rather is the application of multilayered ANNs or deep neural networks (DNNs) to a wide range of problems, from spatial recognition to image analysis. Notably, a DNN is an ANN with numerous layers between the input and output layers 28 -33 (Figure 3). The DNN finds the correct mathematical manipulation to turn the input into an output, whether it is a linear or nonlinear relationship. In recent years, DL techniques have become the state of the art in computer vision. A special neural subtype, the CNN, has become the de facto standard in image recognition and is approaching human performance in a number of tasks. 30 These systems function by learning relevant factors directly from huge image training sets, often containing millions of images. This contrasts with more traditional pattern recognition techniques, which rely on manually crafted quantitative feature extraction.
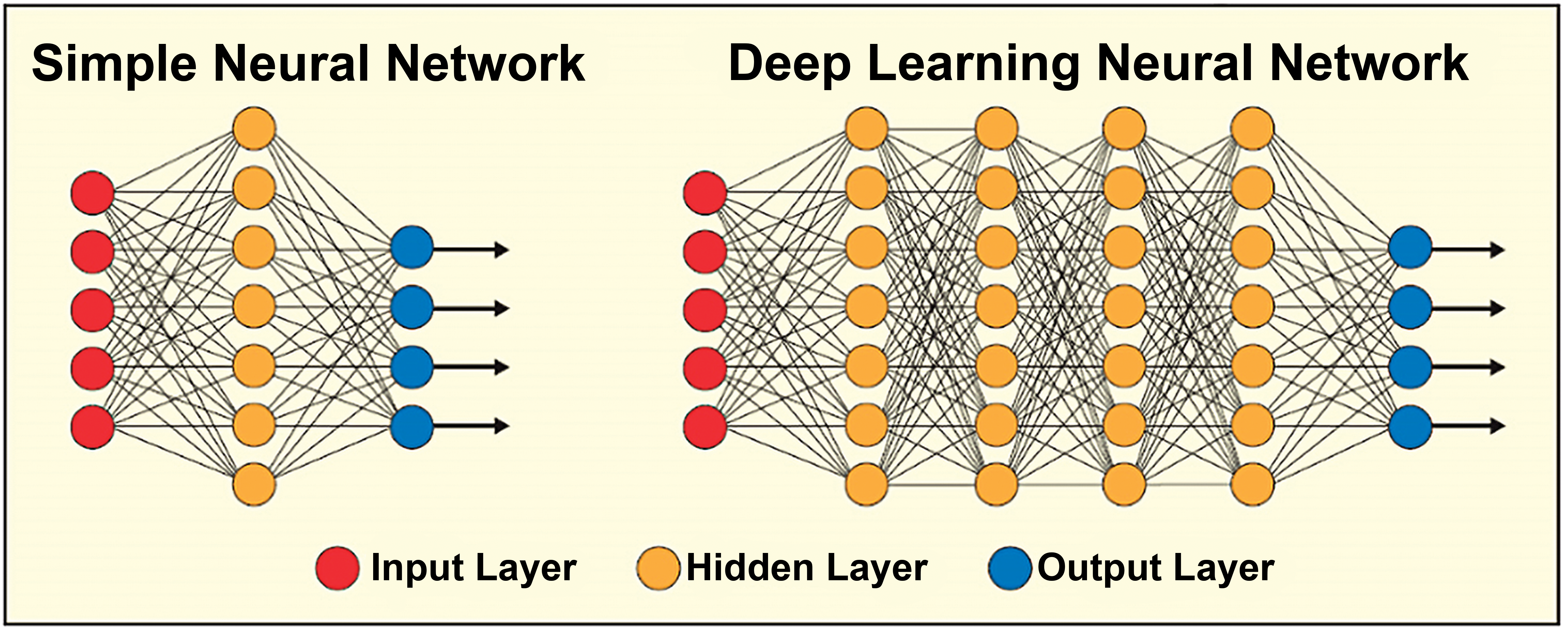
A simple neural network has up to 2 layers hidden between the input and output layers; systems with more than 2 layers are called deep learning neural networks, which can be used to model complex, nonlinear relationships. Reprinted with permission from the association for computing machinery. 34
Specifically, DL iteratively improves upon learned representations of the underlying data with the goal of maximally attaining class separability. 35 -41 Separability is the capacity of a system to use classifiers (eg, approaches, methods, etc) to categorize input data in ML, using distinguishing patterns, features, and so on, in appropriate feature space. 35,37 In DL, the term separable comes from image processing, where spatially separable convolutions are sometimes used to save on computational resources. A spatial convolution is separable when the 2D convolution filter can be expressed as an outer product of 2 vectors. This lets us compute some 2D convolutions more cheaply. In the case of DNNs, the spatial filter is not necessarily separable, but the channel dimension is separable from the spatial dimensions. 41 The modern concept of separability includes, at minimum, the Separability Index (a computational index used to classify the degree of discrimination between objects from different classes), the Hypothesis Margin (the largest distance the sample points can travel without altering the label of the new instance), and Linear Separability (the ability to separate 2 sets of points with a single line), including Support Vector Machines (supervised learning models for classification and regression with associated learning algorithms that maximize the hypothesis margin). 35 -37,39,40 The initial data/data sets are classified/categorized from distinguishing features of data objects in the data set, and in DL, the system learns pattern features to continue classifying and categorizing input data/data sets. This is possible if the main pillar of induction systems, based on the dictum, “similar objects tend to cluster together,” is true. The determination of the separability of the data/data sets is conducted and completed iteratively, based on the criteria, classifiers, approach, and so on, of the DL system used.
Every DL network relies on the assumption of random initialization, and for each iteration, data are propagated through the neural network to complete its respective output. The output is compared to the desired output (eg, determining whether a particular pixel belongs to the parameter image of interest), and an error is computed per parameter, so that there are no preexisting assumptions about a particular task or data set. This is done in the form of encoded domain-specific insights or proportions, which guide the creation of the learned representation. 28,31,33,42 The DL approach involves deriving an appropriate feature space only from the data itself. This is a critical attribute of DL methods, as learning from training examples establishes a path that allows the application of the learned model to other independent test sets. Once the DL network has been trained with an adequately powered training set, it usually generalizes well to novel situations, without requiring additional manual engineering features. Lastly, DL algorithms also have the potential to unify multiple tasks in digital pathology and tissue image analysis, as they are generally as good as, if not superior to, other methodologies for detecting mitosis, classifying tissue, and analyzing immunohistochemical staining. 28
Model Interpretation
A common reservation about DL is that neural network-based models are difficult to interpret and create inscrutable “black boxes,” relative to models based on decision trees or logistic regression. 43 While this may be somewhat true for some DNN types, CNNs, which are the most relevant DNN type for the visual domain and pathology, are extremely useful for interpretation and visualization, as they represent visual concepts.
Before briefly describing several commonly used approaches, we need to consider why it is important to interpret these models. Although one might think it is sufficient to evaluate predictive performance based on a validation data set using metrics such as accuracy, sensitivity, positive predictive value, and/or receiver operating characteristic curves, 44,45 these metrics may be misleading because a model might accurately perform its task, but it might be using features with no intrinsic relationship to the task. That is, the features are predictive only because of undetected sources of bias in the data sets. This caveat was demonstrated in striking examples by Nguyen et al who constructed artificial images that resemble random noise to humans, but that were predicted, with high confidence, to be recognizable objects by a DNN. 46 Model interpretation is also important because scientists focused on ML are asked for insight into the “black box” by their collaborators—and presumably, will increasingly be asked for the same insight by regulatory agencies. For these reasons, it is important not only to evaluate the accuracy of a trained model but also to try to define what the model has learned.
Many approaches for interpreting and visualizing representations learned by CNNs have been proposed. The most popular approaches fall into 4 categories (see Chollet for details
47
): Analysis of intermediate activations. The outputs of intermediate layers in a CNN can lead to an intuitive understanding of how the successive CNN layers transform their inputs. Visualization of CNN filters. The filters indicate the visual patterns to which a CNN is receptive. More “upstream” filters typically correspond to lower level concepts such as edge and corner detectors, while more “downstream” filters often correspond to higher level, “semantic” concepts. Class activation mapping. Methods such as Grad-CAM (Gradient-weighted Class Activation Mapping
48
) can be used to identify which parts within a given image sample were most relevant for the class prediction of a CNN and can be used, for example, to assess whether a network relies on counterintuitive visual clues. Mapping the embedding learned by a neural network: t-Distributed Stochastic Neighbor Embedding (t-SNE), Uniform Manifold Approximation and Projection for Dimension Reduction (UMAP). A fourth approach for investigating what a network has learned and how it “sees” different samples relative to each other consists in analyzing the “embedding” layer, which is usually understood as the penultimate layer of a network (before the final class prediction layer). If inputs are images, they would be mapped into “embedding space” by the network, where each image is now represented by a vector. For example, in the commonly used VGG16 network architecture, the embedding vector has dimension 4096. This dimensionality is far lower than the dimensionality of typical input image sizes, which might include 224 (rows) × 224 (columns) × 3 (colors, ie red, green, and blue) = 150,528, but is still far too high to be visualized directly. In recent years, a number of nonlinear dimensionality reduction techniques, highly suited for the task of projecting the embedding space into lower dimensional spaces, have been proposed. These include t-SNE
49
and UMAP.
50
From a user’s perspective, UMAPs can be envisioned as nonlinear versions of principal component analysis (PCA). Like PCA, UMAPs do not require substantial parameter adjustments to create useful results. In addition, the results of the dimensionality reduction (typically, but not necessarily to dimension 2) can be interpreted similarly to the first 2 principal components of a PCA plot; samples in the plot can be colored by their true class, with a good classifier revealing distinct “clusters” (ie, point clouds) corresponding to individual classes. A single class can also be divided into several clusters, and classes that are similar to each other may be grouped into a single point cloud. Interesting features apparent from the t-SNE or UMAP plots can be investigated further by visually exploring the actual images corresponding to their dimensionality-reduced network embeddings.
Figure 4 shows an example of the t-SNE visualization technique. 51 Each point in this t-SNE plot corresponds to a small image patch, with patches taken from rat tissues of different types (eg, liver, kidney, lung, etc). Patches are colored according to their true tissue type. The projection is obtained by taking the 4096-dimensional neural network embedding of each patch, and using t-SNE to project the embedding vector into the 2-dimensional plane.
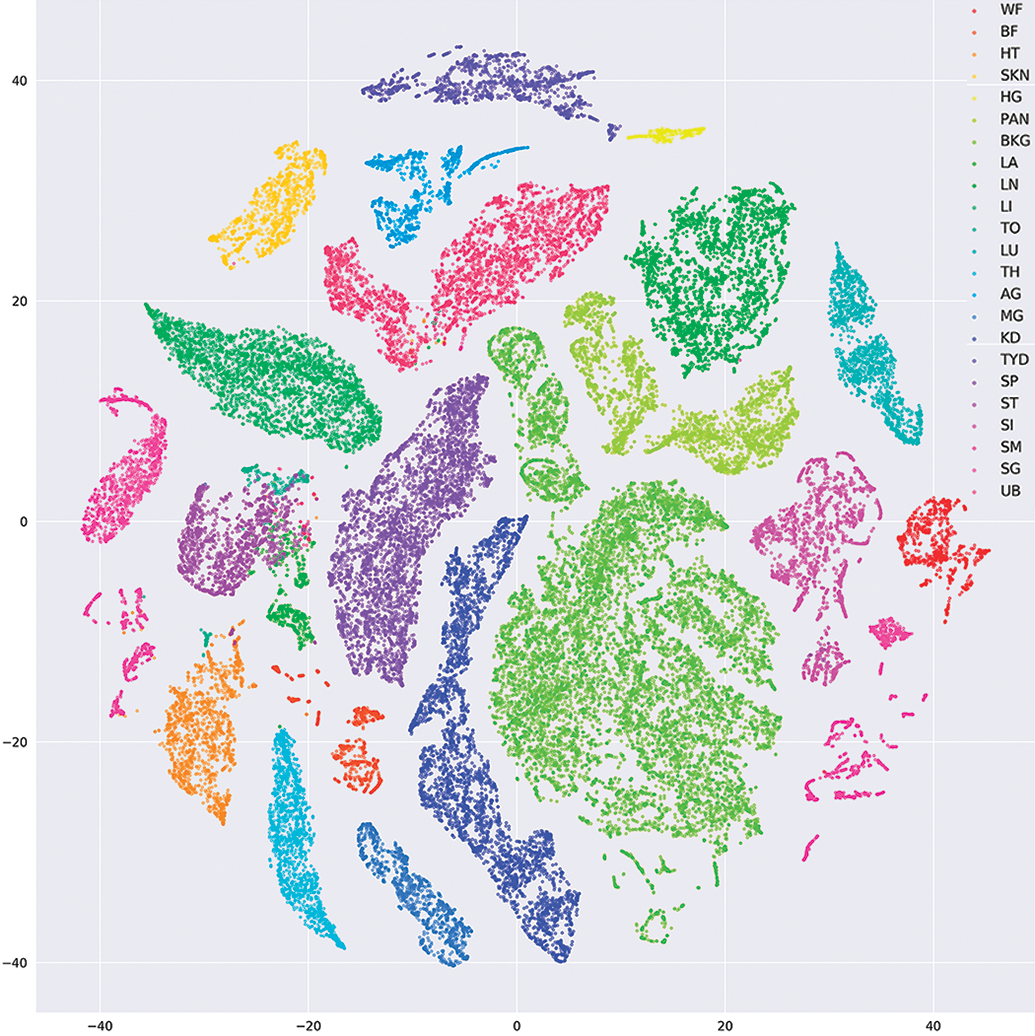
t-Distributed stochastic neighbor embedding visualization of the embedding space learned by a neural network trained to recognize rat tissues of different types. each point in this plot corresponds to a small image patch that is color-coded according its true tissue type. Each cluster and color represents a different tissue. Most points are within distinct, non-overlapping clusters. WF indicates white fat; BF, brown fat; HT, heart; SKN, skin; HG, harderian gland; PAN, pancreas; BKG, background; LA, large intestine; LN, lymph node; LI, liver; TO, tongue; LU, lung; TH, thymus; AG, adrenal gland; MG, mammary gland; KD, kidney; TYD, thyroid gland; SP, spleen; ST, stomach; SI, small intestine; SM, skeletal muscle; SG, salivary gland; UB, urinary bladder. Figure provided courtesy of Sing. 51
There are times when a trade-off between predictive performance (eg, accuracy) and interpretability of models may exist. However, contrary to the widely held notion that neural networks are simply “black boxes,” CNNs, arguably the most relevant models for the visual domain, are amenable to interpretation and visualization, as shown through the 4 approaches discussed in this section. The interpretation of learned representations is an active field of research, making it likely that novel methods will be developed in the next few years.
Medical and Biopharmaceutical Applications
Improvements in whole slide imaging (WSI), cloud-based computing, image analysis technology, and digital data archiving capabilities have enabled some form of digitalization of the pathology workflow in most pathology laboratories. Concomitant with this development, there has been exponential growth in the use of AI and ML approaches to interrogate pathology slides beyond the automated image analysis for generating quantitative pathology data. There are those that herald AI as the next great revolution in pathology. 52 In recent years, with its application to drug discovery, toxicology, and related fields, ML has led to what some describe as a “DL revolution” in drug discovery and development. Machine learning has already been used to predict novel biomolecular drug targets; off-target and the on-target toxic effects of drugs; and innovative, environmental chemicals, food, and household products. 28 -31
In this section, we provide an overview of several notable applications of AI and ML in the biopharmaceutical and applied research fields of oncology, diagnostic pathology, and toxicologic pathology. We also briefly describe the emergence of digital pathology companies and nonprofit/academic organizations that promote digital pathology applications.
Clinical Oncology and Cancer Research
The rapidly expanding scientific literature that documents applying ML algorithms to WSI of cancer tissues highlights the exciting potential of utilizing AI for evaluating histopathology images. Readers are encouraged to refer to individual citations for specific algorithms/models and descriptions of the training/test images used in their development.
In this era of precision medicine where accurate, objective, and reproducible subclassification and grading of cancer are imperative, ML algorithms have shown immense potential. For example, a DL classifier was developed for automated detection of estrogen receptor-positive breast cancer by examining >7000 fields from 174 WSIs to measure the ratio of the number of tubule nuclei to the overall number of nuclei (tubule formation indicator) and test the ratio’s correlation with the oncotype diagnosis (ODX) category. 53 The results suggested that a combination of the tubule formation indicator and other automated features (eg, nuclear architecture) might enable the AI algorithm to correctly predict the ODX category. Similarly, a Combined model with Active Feature Extraction, consisting of 2 logistic regression algorithms, was developed to distinguish benign usual ductal hyperplasia from malignant ductal carcinoma in situ lesions. 54 In this model, active features such as nuclear perimeter and eccentricity were used to make the diagnosis. Likewise, several papers have addressed efficient segmentation of nuclei to extract novel features for the classification and grading of cancer. 55 -59 Other studies applied DL tools to objectively and reproducibly grade prostate cancer, especially those cases with heterogeneous Gleason scores. 60 -63 Machine learning algorithms have also shown potential for extracting novel hidden features and helping establish clinicopathological relationships that pathologists and researchers might miss. 28,64 A new ML approach termed C-Path (Computational Pathologist) combined with DL neural networks was used to analyze breast cancer tissues, from a large cohort of patients, to identify morphological features in cancer stroma that independently predicted patient survival. 65,66
Several other studies show that ML algorithms performed either on par with or slightly better than pathologists in detecting cancer. 30,67 -69 Moreover, these algorithms were not only accurate in detecting and classifying a cancer subtype, but could also predict a cancer-specific genetic mutation based on pathology images. 70 Such ML algorithms have the potential to increase diagnostic sensitivity while reducing sample turnaround time, variability between clinical interpretations, the pathologists’ workload, and the overall cost per diagnosis. In contrast to these “improvements” in workflow, Montalto and Edwards wisely point out that histologic classification of poorly differentiated non-small cell lung cancer by microscopic review typically requires an immunohistochemistry panel to facilitate accurate classification, so that diagnoses are typically rendered within 1- to 2 days, such that the added value of an ML algorithm for improving the speed of initial diagnosis is unclear. 71
Large advancements came in 2017 and 2018 with studies showing that ML tools could be used to predict clinical outcomes and mutated genes by combining histopathologic images with genomic profiles. 72 -74 Mobadersany et al (2018) demonstrated that survival CNNs, created by integrating information from both hematoxylin and eosin (H&E)–stained images and genomic biomarkers (eg, isocitrate dehydrogenase mutations), surpassed the prognostic accuracy of human experts who used the current clinical standard for classifying diffuse gliomas. In another study, a statistical model was built to predict a genetic mutation in the gene encoding speckle-type POZ protein through DL on H&E images of prostate cancer 74 ; and in another study, estrogen receptor status, RNA-based molecular subtype (basal-like vs nonbasal-like), and risk of recurrence score could be predicted with approximately 75% to 80% accuracy based on DL analysis of H&E breast cancer images. 72 Finally, Coudray et al (2018) demonstrated that a DL neural network model (Inception v3) not only diagnosed lung adenocarcinoma and lung squamous cell carcinoma with 97% accuracy but also predicted the presence of mutations in specific genes when the neural network model was trained on images from patients with lung adenocarcinoma coupled with their mutational profiles. These studies imply that using ML tools to assess H&E images and perform correlative analyses that integrate other complex genomic/transcriptomic and epidemiologic data may help clinicians identify patients who could benefit from further molecular testing.
Machine learning algorithms have also been used to quantitate immunostained slides,
75,76
detect mitotic figures in H&E-stained images
77
with
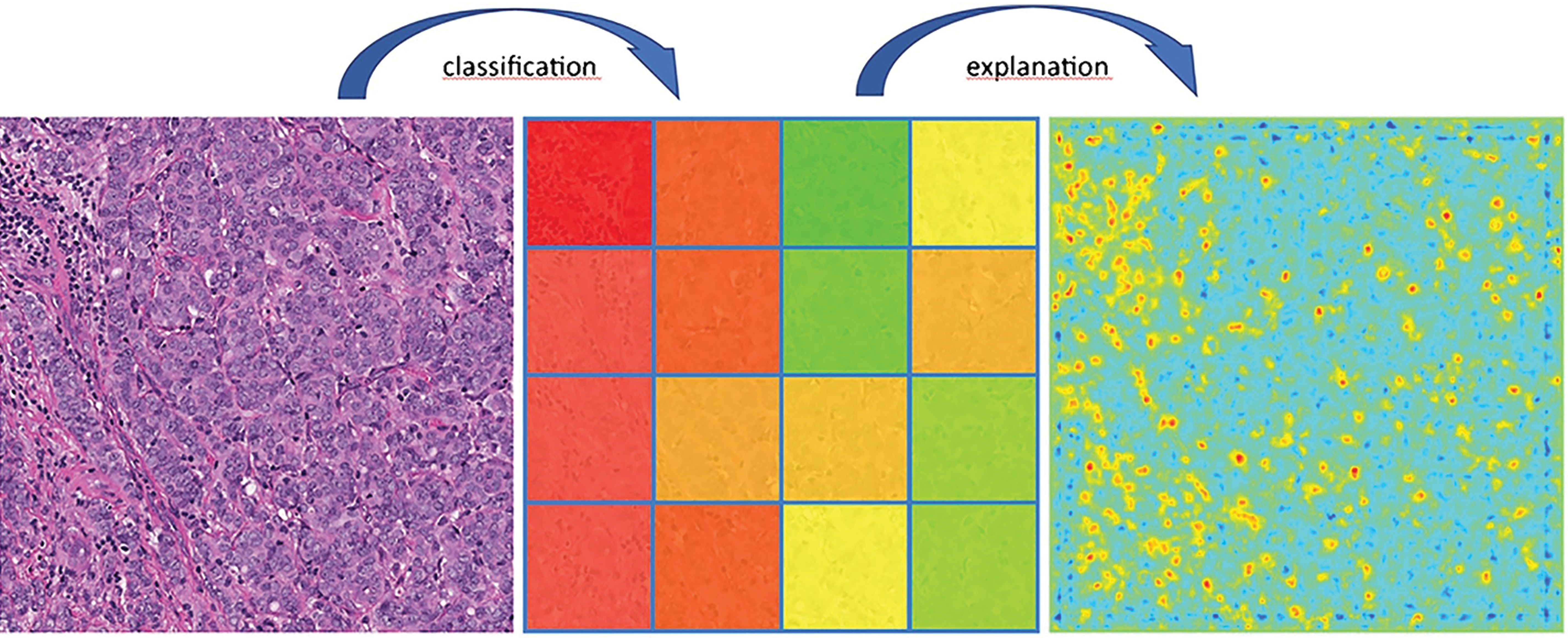
Machine learning–based tumor infiltrating lymphocyte evaluation. Left: Original hematoxylin and eosin breast cancer image. Center: Machine learning–based classification/scoring of regions with respect to their lymphocyte content (green: no lymphocytes, red: high content of lymphocytes). Right: Heatmap explaining classifier decisions with pixel-wise resolution demonstrates how lymphocytes are identified and localized based on layer-wise relevance propagation. Lymphocyte counts can be derived from the heatmap or directly during classification (blue to green: no evidence of lymphocytes; yellow to red: increasing evidence of lymphocytes). Reprinted with permission from the academic press. 80
Anant Madabhushi and colleagues have pioneered new methods to measure tumor-infiltrating lymphocytes and proposed an Image-based Risk Score, which computes the probability of disease aggressiveness using features mined from medical images for a variety of cancers. 82 This and related work created the field of Pathomics, which, akin to Radiomics, is “the comprehensive quantification of tumor phenotypes by applying a large number of quantitative image features.” 83 Recently, these 2 fields have been merged into the new area of RaPtomics. 84
The Cancer Drug Response profile scan is a novel DL model that predicts anticancer drug responsiveness. 85 It was developed using a large-scale drug screening assay and applied to evaluate 1487 approved drugs on 787 human cancer cell lines. It identified new potential cancer indications for 14 oncology and 23 nononcology therapies. Deep learning was also applied in early drug discovery to streamline the identification of cell line phenotypes and identify potential candidate molecules faster and more accurately than conventional image analysis. 85
Diagnostics (Human and Veterinary)
Accurate and sensitive interpretation of H&E slides has remained the foundation of pathological analysis and diagnostic medicine for over a century. 32 Most pathologists are trained to follow algorithmic decision trees to stratify patients based on tumor type and aggressiveness. 32 Despite this training, data interpretation is not always consistent between pathologists. Novel ML technologies have the potential to reduce human interobserver disagreement over the interpretation of H&E slides. Algorithms based on ANNs have yielded diagnostic interpretations for H&E slides of human skin cancer biopsies, for example, with accuracy similar to those of board-certified professionals. 86 Litjens et al conducted an extensive survey of applying DL methods to histopathology images and transmission electron micrographs. A few examples from human and veterinary medicine are summarized below. 5,87
There are many diseases of the blood and cardiovascular systems and several algorithms have been developed to aid in detection of abnormalities in these systems. Durant et al implemented a deep CNN-based approach for classification of erythrocytes based on morphology. 88 Erythrocytes were manually classified into 1 of 10 classes using a custom-developed web-based application. Using deep CNNs consisting of >150 layers and with dense shortcut connections, erythrocyte morphology profiles were measured with a high degree of accuracy (∼90%). Further optimization of the practical performance of these CNNs in a clinical environment is the next step toward their development as a diagnostic tool. Deep learning has also been used to detect malaria-infected red blood cells, which traditionally has required highly trained microscopists. Artificial intelligence aided the development of the Easyscan Go scanner (Motic, Hong Kong, China), which automates malaria detection in blood smears, allowing timely patient diagnosis and treatment, especially in countries with limited resources (https://www.motic.com). Another CNN was developed to detect clinical heart failure from H&E-stained WSIs with high sensitivity and specificity. 89 Although endomyocardial biopsies (EMBs) are the current gold standard for evaluating myocardial disease, manual evaluation of EMBs by pathologists is fraught with high interobserver variability. A CNN developed by Nirschl et al used 104 training samples and 105 distinct samples for independent testing. They reported 99% sensitivity and 94% specificity for their classifier on the test set for identifying patients with heart failure/severe cardiac pathology. Importantly, the CNN outperformed 2 expert pathologists by nearly 20%, suggesting that DL algorithms can be used to predict the cardiac outcome.
Identifying disease states affecting organs, like lung and kidney, and structures within such organs may also benefit from new AI approaches. Heinemann et al, describe a technique to score lung fibrosis and inflammation in a mouse model system using 2 different deep CNNs, manually trained using 14,000 and 3500 labeled tiles, respectively. 90 They achieved an accuracy of 79.5% for scoring fibrosis and 80.0% for scoring inflammation which was similar to the pathologist’s results. Sheehan and Korstanje developed an ML-based high-throughput method for automatic glomeruli detection in mouse kidney and subsequently for quantitative analysis of glomerular features. 91 This method, developed using free open-source software, required minimal human interaction between steps and created a data acquisition tool independent of user bias. This method can be applied to different animal species following training on relevant data sets. The application of DL to discover unidentified histopathologic features is one of the approaches of the Kidney Precision Medicine Project, 92 and these findings will be combined with extensive data provided by the analysis of proteomics, transcriptomics, genomics, lipidomics, metabolomics, and clinical parameters to better characterize the lesions, adapt the treatments, and identify new drug targets and biomarkers. 93
Beyond the obvious diagnostic applications, ANNs have been used to extract different morphologic features, not readily detected by the human eye and, to reliably differentiate between neoplastic and non-neoplastic lesions. 56,94,95 Olsen et al showed that a CNN algorithm could differentially diagnose nodular basal cell carcinoma, dermal nevi, and seborrheic keratoses with more than 99% accuracy. 96
In the veterinary diagnostic pathology setting, Hattel et al used a 2-stage supervised image algorithmic framework to correctly identify normal tissue and specific tissue lesions 97 with a classification accuracy ranging from 84% to 95% for acute suppurative pulmonary inflammation, normal kidney, chronic kidney inflammation, normal spleen, acute suppurative splenitis, normal liver, acute suppurative hepatitis, and acute hepatic necrosis. More recently, Awaysheh et al applied ML to improve automated classification of structured histopathology reports for feline small intestinal disease. 98 The authors tested the machine classification accuracy of unstructured (free-text) microscopic findings with those represented in the World Small Animal Veterinary Association (WSAVA) Gastrointestinal International Standardization Group format with a diagnosis of inflammatory bowel disease and alimentary lymphoma. Free-text duodenal biopsy pathology reports from 60 cats, 20 each from 3 groups: inflammatory bowel disease, alimentary lymphoma, and normal were reviewed. Biopsy samples from these cases were then scored following the WSAVA guidelines to create a set of structured reports. Diagnosis classification accuracy for the 3 algorithms was compared using the structured and unstructured reports. Using naive Bayes and neural networks, unstructured information-based models achieved higher diagnostic accuracy (90% and 88% respectively) compared to the structured information-based models (74% and 72%, respectively). Overall, the study suggests that discriminating diagnostic information was lost when the current WSAVA microscopic guideline features were used, but that the addition of free-text features (eg number of plasma cells) via ML helped identify candidate microscopic features that could be used in structured histopathology reports.
Toxicologic Pathology—A Preclinical Advantage
In the last several decades, pathology evaluation for nonclinical safety studies has become a well-standardized process with established guidelines and/or recommendations, including organ sampling and trimming schemes (https://reni.item.fraunhofer.de/reni/trimming), histopathology evaluation, 99 peer review, 100 and diagnostic nomenclature. 18,101 -112 However, it is labor intensive and time-consuming for a pathologist to microscopically review 100s to 1000s of slides, per study especially considering that most tissues/organs lack test article-related changes. In addition, normal background lesions need to be recognized as such, many of which are sex and age-dependent and further influenced by species and strain. As highlighted in a 2013 review, because of their comparative medicine training, toxicologic pathologists will play a key role in multidisciplinary teams across the medical industry. 113 Artificial intelligence and ML may provide the means to sort normal from abnormal samples thus allowing toxicologic pathologists to spend more time microscopically evaluating altered phenotypes instead of normal tissues, thus reducing the weeks to months necessary for toxicologic pathology evaluation and/or the peer review processes that can substantially affect drug development timelines. Also, AI and ML may assist in decreasing inter- and intralaboratory variability associated with the semiquantitative grading systems used in the industry. It is feasible that AI and ML technology might reduce or eliminate the estimated 10% error/discordance rate in the characterization of microscopic lesions by toxicologic pathologists. 114 Implementation of these novel tools is now possible due to the availability of WSI, advances in computer hardware and software, and the decreased cost of handling and storing large data sets. In this regard, several private companies are developing DL algorithms to screen digital slides for lesions in tissues. 115,116 The objective of these DL algorithms is to tag the slides as “normal” or “abnormal,” based on training the algorithm on sets of normal tissues. Although these algorithms need further validation, especially on large tissue sets and preclinical samples, in the future, they could be used to automate the classification of specific lesions commonly observed in nonclinical toxicity studies. This will benefit the toxicologic pathology community by reducing the time devoted to microscopic slide review while increasing the time available to define mechanisms of toxicity and/or participate in other drug development activities. Recent DL models have automated classification of WSI of different, normal, non-diseased rat tissues with 94.7% to 98.3% accuracy, thus providing a potential foundation for future models of toxicologic pathology. 51
The benefits of DL algorithms in investigative toxicologic pathology have already started showing great promise. For instance, as opposed to conventional image analysis algorithms, CNNs can precisely detect and segment glomeruli within digitized kidney sections. 117 Investigators have also used DL methods to quantitate mesangial matrix deposition per glomeruli in a diabetic nephropathy mouse model. Another potential application of DL methods is the quantitation of renal interstitial fibrosis in Masson trichrome-stained sections and its correlation to clinical pathology measures. 118 In 2018, Penttinen et al developed a deep CNN algorithm for counting tyrosine hydroxylase immunoreactive neurons in the substantia nigra of mice. The algorithm performed comparably to human observers and the results correlated with previously published estimates, based on unbiased stereology. 119 Taken together, the potential applications of AI and ML methods in toxicologic pathology are extensive.
The notion of robust DL algorithms being adapted to investigate the mechanisms of drug-related toxicities by integrating large data sets from histopathology images, clinical pathology and gene expression changes, and clinical data would be transformative. Integrated analysis of such large-scale data sets is challenging and is only beginning to be explored. As automated image analysis of WSIs becomes more common in nonclinical toxicology studies, AI and ML will follow suit. In 2018, an automated image analysis method was used to quantify bone marrow cellularity in the sternal bone marrow of 576 Sprague-Dawley rats from 17 nonclinical studies. This automated method was faster and more sensitive than manually scoring bone marrow depletion. 120,121
Another area where ML tools could make a big impact in toxicologic pathology is with Content-based image retrieval (CBIR 122 ) systems. CBIR retrieves WSIs, visually similar to regions of interest in a query image, from a database. CBIR will open new possibilities to the toxicologic pathology field. Lesions can be compared across studies, species, and databases for educational, diagnostic, or research purposes. One can easily imagine it could also help to better understand the progression of a complex lesion over time of drug exposure (likelihood of progression, premonitory changes, morphological evolution, primary insult, and so on). In this regard, DL-based CBIR already shows encouraging results for mining histopathology images. 123,124
In an extensive review of the opportunities and obstacles for DL in biology and medicine, Ching et al, (2018) concluded that “DL has yet to revolutionize biomedicine or definitively resolve any of the most pressing challenges in the field,” but that “promising advances have been made on the prior state of the art” and they “foresee DL enabling changes at both bench and bedside with the potential to transform several areas of biology and medicine.” We believe this is also the current state for digital toxicologic pathology and agree that recent progress indicates that DL methods will provide valuable means for accelerating or aiding human investigation. In addition, they raised very important issues that also remain to be addressed in toxicologic pathology, especially regarding interpreting these models to make testable hypotheses about the system under study, and the challenges presented by the existence of only a limited amount of well-labeled data for training. 31 And further to this, Tizhoosh and Pantanowitz in a 2018 review, outline 10 challenges and 8 opportunities that currently face AI and digital pathology. Concluding that “The danger of overselling AI is still omnipresent. Nonetheless, there is clear potential for breakthrough with AI in medical imaging, particularly with digital pathology, if we suitably manage the strengths and pitfalls.” 125
New Industries
How does this new industry currently impact the average toxicologic pathologist? At this stage, it is highly variable and depends on many factors including access to the technology. Although a few companies are already offering solutions, AI applied to pathology is a new and dynamic industry. Most of today’s companies are involved in streamlining the process for the primary diagnosis of human cancers, specifically, breast cancer, 126 breast and prostate cancers, 126 and oral and skin cancers, 127 by providing support and guidance to the pathologist.
Machine learning techniques are essential components of medical imaging research. The advancements in Big Data 128 and DL have played an important role in the development of ANNs and are supported by hardware breakthroughs, such as graphic processing units (GPUs), to provide greater calculation power. Traditionally, GPUs have been used for gaming and design, as the computing part of a graphics card. Following the creation of Compute Unified Device Architecture, a parallel computing platform and application programming interface by NVIDIA, researchers can now more easily use GPUs for parallel programming and nongraphical purposes, which is particularly well-adapted to the training of ANNs running multiple, simultaneous operations. 129,130 This led to the creation of general purpose graphic processing units (GPGPUs), which have a broader range of uses including computational chemistry and analysis of high-content imaging systems. 131 The exponential advances in chip processing capabilities in recent years have played a large part in the advancement of ANNs. 129 Moreover, these computational resources are becoming more accessible with the rise of cloud computing which provides greater access to high capacity GPGPUs, bypassing the need for a supercomputer to analyze large, complex data sets. 132 This rapid advancement in computing power coupled with current image analysis informatics, is leading to massive growth in digital pathology-related industries.
Growing business sectors related to digital pathology can be divided into 3 major categories: (1) research and development; (2) venues and means for reporting and sharing information; and (3) education and training. The first class includes small and moderate-sized computational digital imaging companies, which often collaborate with pharmaceutical/biotechnology and/or big computer companies; pharmaceutical companies, which often collaborate with computer companies, hospitals and/or academic research organizations; and governments that collaborate with and/or support research centers. The second class includes professional journals, organization, associations and societies devoted to the subject; new firms supporting global professional conferences and meetings; online information firms, which report on and analyze current industry news and trends; and individuals and companies who report, write blogs, or create podcasts and webinars. The third class includes new academic and medical education and training programs (often supported by large pharmaceutical/biotechnology companies or government) and other forms of education provided through webinars, online training, so on It is important to consider key areas emerging within these sectors. Two of note for the toxicologic pathologist are computational pathology and pharmaceutical and AI company collaborations:
Computational Pathology
Computational Pathology is a direct offshoot of Biomedical Informatics. In 2010, Thomas Fuchs, the “father of computational pathology” defined this novel field as “comprising aspects of ML, computer vision, clinical statistics and general pathology.” 133 This has been further developed by, for example, the American Medical Informatics Association, in July 2014 at a meeting, attended by pathologists, pathology department-based informaticists, and pathology department chairs from major medical institutions in the United States. 134 Computational pathology is focused on computational methods that incorporate clinical and anatomic pathology and molecular pathology and genomic pathology data sets to improve health care. Specifically, “Computational pathology is an approach to diagnosis that incorporates multiple sources of data (eg, clinical electronic medical records, laboratory data, including ‘-omics’ and both radiology and pathology imaging); extracts biologically and clinically relevant information from these data; uses mathematical models at the molecular, individual, and population levels to generate diagnostic inferences and predictions; and presents this clinically actionable knowledge to customers through dynamic and integrated reports and interfaces, enabling physicians, patients, laboratory personnel, and other healthcare system stockholders to make the best possible medical decisions.” 134 A computational approach, using adaptive feedback to train algorithms with visual patterns and molecular biomarkers simultaneously, can be used to enhance prognosis prediction accuracy in cancer. 73 In 2019, the Digital Pathology Association defined computational pathology as the “omics” or “big-data” approach to pathology and proposed best practices and recommendations for regulatory guidance. 135 And also in 2019, Fuchs and colleagues showed that a clinical-grade computational pathology approach with multiple cancer types (prostate cancer, basal cell carcinoma and breast cancer metastases to axillary lymph nodes), using weakly supervised DL on WSIs has the ability to train accurate classification models at unprecedented scale. Reporting that its clinical application would allow pathologists to exclude 65% to 75% of slides while retaining 100% sensitivity. 136
Pharmaceutical and AI Company Collaborations
Although key players in digital pathology-focused AI have not yet emerged, private companies already offer off-the-shelf DL algorithms 137 -139 that are being explored by pharmaceutical companies and research institutes as aids for drug discovery and preclinical testing. In addition, by collaborating with companies that are developing other AI technologies, Pharma is seeking to capitalize on the insights of some of the largest players in the AI world to improve not only drug discovery but also preclinical and clinical outcomes. Such collaborations include those of Pfizer 140 and Novartis 141 with IBM; Novartis with PathAI, 142 Genentech (Roche) with GNS Healthcare 143 ; Sanofi 144 and GlaxoSmithKline 145 with Exscientia; and Johnson & Johnson with BenevolentAI. 146
Future Impact of AI on Digital Toxicologic Pathology
Given these many waves of change, toxicologic pathologists will be expected to be at least very familiar with, if not fluent in AI and ML technologies. Pathologists must understand how these technologies will impact the processes and tools they commonly use today. In turn, this understanding will shape how these processes, including image analysis, 3D reconstruction, multispectral imaging, and augmented reality (AR) evolve. In addition, how student pathologists and toxicologic pathologists will be educated and how the regulatory environment will change to accommodate these technologies must be considered. Here, we briefly reflect on how these areas are being impacted by AI and ML.
Image Analysis and 3D Reconstruction
Digital pathology enables the possibility of 3D histology; however, a key computational hurdle in creating accurate image reconstructions lies in precisely registering sequences of 2D images. It is likely that AI will inform and improve future image analysis. For example, feature-based algorithms could be implemented to register serial sections, thus decreasing the need for manual input. 147 There have been numerous image-registration algorithms employed for 3D histology 148 including one used by an imaging system that scans each section as it is being cut 149 and other private companies also offer AI solutions for keeping images in register to supplement their image analysis software 116,138 and to aid in the creation of 3D images.
Although rendering 3D images is not necessary for all histology-based image analyses, extending histological assessment into 3D holds promise for characterizing anatomic structures with complex 3D organization but also requires a significant investment in resources. To date, 3D histology has been employed in a variety of research applications, including characterizing microvascular structure in the hind limb of C57BL/6 mice, 150 evaluating the invasion fronts in cervical carcinoma, 151 and building high-resolution anatomical atlases using Nissl-stained sections of adult mouse brain. 152
Another challenge that might be aided by AI is accurately identifying and tracking cell segmentation in formalin-fixed, paraffin embedded tissues. Correctly identifying cell and nuclear borders in mixed cell types with heterogeneous cellular morphologies and in 2D representations of sections taken through 3D structures, may result in lost nuclei and/or overlapping cell borders due to cell clumping. 153 In addition, a lack of pan-membrane markers poses additional challenges to accurate segmentation of cells, which in turn affects the analysis of molecular markers at cellular resolution. Deep learning could aid the development of robust and accurate algorithms for single-cell analysis in tissue sections that would automate image processing and analysis for cell and tissue segmentation, feature extraction and classification, and data visualization.
Multispectral Imaging
Modern image analysis software systems, built on ML algorithms, have substantially improved the identification of cell boundaries in images from multiplexed immunohistochemistry and mass cytometry experiments. 154 -158 Multispectral imaging is imaging across a wide range of wavelengths allowing the use and detections of multiple fluorescent and/other markers. Data sets generated from multispectral imaging experiments are highly multidimensional and complex; thus, they require advanced computational tools to enable pathologists to automate analysis of cell states, neighborhoods, spatial networks, and associations with clinical data. Here, ML algorithms using DNNs, 159 tSNE, 160 Phenography 161 and Citrus 162 have shown promise in dimensional reduction, thus enhancing the ability of the pathologist to examine a single cell within the context of its spatial location combined with other data.
Augmented Reality
Augmented reality uses various instruments ranging from see-through visors to smartphone screens, but in the toxicologic pathologist’s case, a microscope, to layer computer-generated, virtual objects over the viewer’s actual, physical surroundings. Augmented reality already assists clinicians during delicate surgeries by creating an overlay of anatomical structures, for example, those reconstructed from the patient’s computed tomographic (CT) data, that can be referenced by the surgeon as a guide during the procedure. 163 Similar technology is being used to train medical students and practitioners. 164 The range of AR applications has grown to include some that are useful in pathology. Like in surgery, AI can aid performing autopsies, by creating an overlay of CT data. 165 It can also aid the training of pathology residents to perform autopsies, by instructing them with real-time diagrams, annotations, and voice instructions. 166 Augmented reality can also be used to annotate areas of interest on surgical specimens in real-time, and improve identifying the precise anatomical location of important pathologic findings by overlaying previously obtained radiographs onto the samples. 166 Furthermore, AR can be used to annotate photographs taken during gross examinations, to facilitate communication between pathology technicians and pathologists. 167 Augmented reality may also improve the accuracy and efficiency of the pathologist, as in the case of microscope-integrated telepathology, 168 which allows real-time sharing of the precise field of view being seen by one individual with other experts. The technology also allows the projection of an augmented overlay of digital information, such as measurements, annotations, and information regarding the study, on top of the optical field of view. 169 Deep learning algorithms have also been developed in the AR space that has been used to assist the pathologist in detecting breast cancer metastases in lymph nodes and in classifying prostate cancer according to the Gleason scoring system. 170 These tools enable real-time image analysis highlighting the areas of interest directly into the field of view 170 allowing for the potential of real-time assessment by pathologists and ML algorithms.
Education
Overall, pathology students have positive perceptions and attitudes regarding computer-based learning environments, translating to high student satisfaction rates 171 -179 and possibly better learning outcomes, 180 -182 although students still see benefit from traditional histology laboratory training. 178,183 It is easy to envision CBIR algorithms that use ML to dynamically tutor pathology students working through digital slide sets, helping them develop cognitive expertise in image analysis. The algorithms would incorporate feedback based on a students’ search approach, visual efficiency, feature identification, and pattern matching. 20,184 Indeed, in the setting of a largely digitized histopathology viewing platform, the raw data are optimal for DL, 185,186 which we expect will become more widely adopted as a diagnostic and teaching tool in medical education in the coming years. The Pathology Informatics Essentials for Residents (PIER) program is notable for establishing a standardized, customizable informatics curriculum that can be implemented in any pathology residency program. 187 It was developed through an alliance, formed in 2013, between The College of American Pathologists, the Association of Pathology Chairs, and the Association for Pathology Informatics. The PIER working group developed 38 peer-reviewed outcome statements and released its second toolkit online in July of 2017. 188 The PIER document delineates the knowledge, skills, and competencies to be acquired by trainees and how their outcomes/achievements can be monitored. 189 A survey of alumni from pathology informatics training programs identified a strong market for formally trained informaticians, with many achieving leadership positions early in their careers. 190 On the veterinary side, the American College of Veterinary Pathologists recently designated “Shape the future practice of veterinary pathology in the age of digital pathology, AI and advanced molecular tools” as the first of 4 goals in their 2018 to 2022 strategic plan. 191 Training the next generation of expert image analysts for veterinary medicine will depend on the broader adoption of principles like those outlined in PIER by their veterinary counterparts. This is especially important as veterinary pathologists are often involved in preclinical assessments and other critical junctures in translational science.
Regulatory Environment
Regulating AI for a successful pathology future is still relatively nascent, but in the human diagnostic setting it has been growing steadily for several years. 192 Regulatory guidance on the use of AI and ML software in Good Laboratory Practice assessments is currently based on the validation principles applied to Good Laboratory Practice assays in general, such as those outlined by the International Council for Harmonization of Technical Requirements for Pharmaceuticals for Human Use (ICH) for validation of analytical procedures, 193 or those identified by the Pharmaceutical Research and Manufacturers of America committees for creating evidentiary standards for biomarkers and diagnostics. 194 In general, validated assays have well-defined parameters including assay accuracy and precision, detection and quantitation limits, linearity, range, and robustness. The assays must also be plausible, relevant to a pharmacologic target or mechanism, and linked to the outcome of a disease or toxicity. In addition to meeting the fundamental principles of assay validation, the principles of using Software as a Medical Device in clinical evaluations, as summarized by guidance from the International Medical Device Regulators Forum Medical Device Working Group, 195 should also be considered, as should maintaining Digital Imaging and Communications in Medicine (DICOM) standards for handling image data.
Digital Imaging and Communications in Medicine is the international standard for the transmission, storage, retrieval, printing, processing and displaying of medical imaging information. The DICOM Base Standard Committee (Working Group 6) meets regularly to review the output of 31 DICOM Working Groups and to identify trends and anticipate changes that might affect the integrity of medical imaging information. The group’s fundamental objective is to develop a standard that will enable the exchange of digital information between medical imaging devices and other information technology systems. With this in mind, there is an emphasis that the DICOM standard guide diagnostic medical imaging for numerous medical disciplines including pathology.
One of the most pressing issues for pathology, related to DICOM’s objective, is ensuring bidirectional communication between slide scanners and other information technology systems, regardless of vendor. 196,197 Adoption of the DICOM standard by developers and manufacturers would enable a standardized exchange of the information associated with digitized images among devices. The DICOM standard has been integrated in the design of the Philips Intellisite Open Pathology Platform. 196 This platform provides scanner interoperability using the Philips iSyntax Bridge coupled to DICOM image export systems that support data migration (including file format) to any third-party system also using DICOM compliant interfaces. An important practical outcome of using the DICOM standard is that a DICOM formatted digital image archive will allow pathology laboratories to freely exchange information between systems (eg, picture archiving and communication systems [PACS] or scanners), by avoiding the pitfalls associated with incompatible proprietary formats, 197 which could result in important time and cost savings when upgrading or replacing laboratory imaging and analysis systems. Further, adherence to DICOM standards in digital pathology could promote collaboration across disciplines, by enabling scientists and medical professionals using different systems to easily access the same information and data without reformatting it for system compatibility, which may lead to some amount of data loss. This adherence will be critical for successful AI and ML innovation. The DICOM standard includes tags with image-specific and patient information that can facilitate selecting the proper images for a specific study or protocol (preprocessing) and helps ensure that the image is identified correctly within a PACS. 197
The importance and influence of DICOM will grow as the standards group works to meet the changing needs of medical imaging as driven by new technologies and applications. As veterinary and toxicologic pathologists expand their use of digital image formats pertaining to AI and ML applications, adhering to the DICOM standard will be important for our discipline to maintain scanner and PACS flexibility while still working with image formats that support our workload, and will require close collaboration with vendors to ensure that the scanners produce images that conform to DICOM standards and that PACS users can import and view the files. 197
Professional Impact of AI and ML on Toxicologic Pathologists
Earlier, we warned that with all great technological advances come implementation challenges. Some of these challenges have been discussed, others implied; however, philosophical questions surrounding AI and ML remain. Will this burgeoning field become an enemy or an ally to the digital toxicologic pathologist? We have attempted to address this question by examining other medical imaging disciplines that have been more engaged with AI and ML, most notably radiology, in which the work of radiologists has been augmented by AI rather than replaced by it. We feel this will likely be true for toxicologic pathologists as well, at least in the short term.
At present, it is unclear how fledgling AI technologies will influence not only toxicologic pathology but also humankind. Will we face a true “technological singularity” 198 with the development of strong AI and artificial general intelligence up to the point of performing “superintelligent” 199 actions or even experiencing consciousness? Or will we contain the doomsday scenarios painted in science fiction by only developing applied, narrow, or weak AI? Although it is prudent to acknowledge the potential for future harm and mitigate accordingly, AI and ML can bring very real advantages and are already advancing the fields of toxicologic pathology, drug safety and development, and global health in general.
Amara’s Law 200 states: “We tend to overestimate the effect of a technology in the short run and underestimate the effect in the long run,” which suggests that we should focus on harnessing these technologies to make our lives easier and improve our work-life balance rather than how these technologies might negatively impact society, or eliminate the need for workers. For example, toxicologic pathologists are in high demand, a trend likely to continue into the near future.
Many toxicologic pathologists may feel overwhelmed, distracted, or underutilized, especially in relation to solving complex problems indicated by the actual test article-related changes they see. As AI and ML systems continue to develop and improve, we may be able to attain the “holy grail” of using computers to screen out “negative” slides that lack toxicologically important findings with very high sensitivity and specificity allowing pathologists to focus solely on slides with test article-related findings. This would create better work-life balance while providing more time for research, publications, and personal development. Of course, the technology will not unburden pathologists until it routinely identifies all slides with lesions (ie, a false negative rate of 0).
Conclusions
Artificial intelligence and ML are rapidly changing the face of many scientific and non-scientific fields including digital toxicologic pathology. Human-AI partnerships are already flourishing in many medical settings and collaborations between computer scientists and pathologists of all varieties will continue to make these partnerships more robust. Public and private sector organizations are working to develop ever newer ML approaches and public-private partnerships will facilitate the mining of larger and larger data sets for novel insights and predictions. Indeed, toxicologic pathology is transitioning from the analog to the digital world. It already delivers and will continue to drive exciting new ways of working with and exploring data, answering scientific questions, and potentially ameliorating global health issues. For more than a decade, the issue of AI and ML replacing the routine functions of toxicologic pathologists or replacing pathologists altogether has been raised at pathology professional meetings. The dawn of AI-only pathology still seems distant and recent studies indicate that the combined power of a trained pathologist and ML reduces diagnostic errors compared to either human or machine alone. Therefore, we encourage toxicologic pathologists to take the view that AI and ML will synergistically improve a pathologist’s individual and collective diagnostic efficacy and competency, not replace them in the future.
Footnotes
Authors’ Note
This is a review by the Society of Toxicologic Pathology, Digital Pathology, and Image Analysis Special Interest Group. As such, it does not represent the position of the entire STP.
Acknowledgments
The authors wish to acknowledge Briana Dennehey, Joi Holcomb and Christina Contreras of MD Anderson Cancer Center, Department of Epigenetics and Molecular Carcinogenesis for expert review of the manuscript and expert support of figure production and manuscript formatting.
Declaration of Conflicting Interests Statement
The author(s) declared no potential, real, or perceived conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
