Abstract
The approach undertaken to deliver a Good Laboratory Practice (GLP) validation of whole slide images (WSIs) and the associated workflow for the digital primary evaluation and peer review of a GLP-compliant rodent inhalation toxicity study is described. The contract research organization (CRO) undertook validation of the slide scanner, scanner software, and associated database software. This provided a GLP validated environment within the database software for the primary histopathologic evaluation using WSI and viewed with the database software web viewer. The CRO also validated a cloud-based digital pathology platform that supported the upload and transfer of WSI and metadata to a cache within the sponsor's local area network. The sponsor undertook a separate GLP validation of the same cloud-based digital pathology platform to cover the download and review of the WSI. The establishment of a fit-for-purpose GLP-compliant workflow for WSI and successful deployment for the digital primary evaluation and peer review of a large GLP toxicology study enabled flexibility in accelerated global working and potential future reuse of digitized data for advanced artificial intelligence and machine learning image analysis.
Keywords
Introduction
The globalization of contract research organizations (CRO) and biopharmaceutical company networks and the outsourcing of nonclinical toxicology studies has challenged the industry’s ability to collaborate effectively while controlling costs and preventing delays in drug development cycle time. Events like the 2020 coronavirus disease (COVID-19) pandemic accentuated the situation due to travel and workplace restrictions and disruption of the shipping supply chain. 1 Industries had begun investigating whole slide images (WSI) for both primary evaluation and peer review of nonclinical toxicology studies over a decade ago but progress had been slow due to multiple factors including a perceived lack of functionality relating to the storage, access and rendering speeds, and viewing of the image files. 2,3 Furthermore, there has been concern over regulatory acceptance of the process. While solutions like encrypted portable media can alleviate the issue of image access and rendering, it generally only allows for the browsing of the image files as lists of file names or thumbnails that may not display the slide label information or other helpful metadata. In effect, there has been a hesitance by toxicologic pathologists to believe that WSI, their associated software applications, and access to image files can really provide an experience comparable to the use of glass slides and a light microscope.
Overcoming these issues has generated considerable discussion within the toxicologic pathology community as to the type, extent, and nature of the end user testing that would be required to demonstrate fit-for-purpose use of WSI alongside Good Laboratory Practice (GLP) compliance. 3,4 For medical diagnostic pathology, a medical device regulatory pathway for whole slide imaging systems was achieved by undertaking the United Sates Food and Drug Administration (FDA) 510(k) compliant technical qualification of the components accompanied by diagnostic concordance studies. 5 –9 However, for nonclinical use, whole slide imaging systems have not been regarded as medical devices. This has therefore contributed to the debate within the toxicologic pathology community as to the extent of technical qualification of system components, diagnostic concordance studies, and even what WSI files actually represent in the context of nonclinical research.
One unintended risk of this debate has been the potential to introduce a level of complexity not appropriate for a specific intended use like contemporaneous peer review.
10
Recent Organization for Economic Co-operation and Development (OECD) guidance frequently asked questions (FAQs) on the use of digital histopathology in GLP studies
11
provides important concepts which are summarized below:
The concept of a “faithful reproduction” and “equivalence”: The word faithful reproduction has been carefully chosen so as to avoid categorizing WSI as true copies of glass slides or specimens. Equivalence is used in the context that WSI should be deployed to provide an equivalent experience to that of using a light microscope (functionality) as determined by the pathologist. This equivalence not only refers to color fidelity, focus, and resolution, but that systems for viewing WSI should also provide equivalent functionality in terms of providing the human readable information from the slide label and an overview of the tissues on the slide. Equivalence therefore defines the need to use fit-for-purpose software and hardware (including scanner) that allows for a faithful reproduction of the light microscope. In human surgical pathology, diagnostic concordance has been used to refer to a comparison of the diagnoses obtained when an identical set of surgical pathology biopsies are examined using glass slides versus their corresponding WSI. In human surgical pathology, publications describing diagnostic concordance have tended to focus on diagnostic outcome, rather than assessment of the functional utility of the hardware and software utilized to generate and review the WSI. We therefore believe that within the context of nonclinical toxicology, the OECD’s reference to “equivalence” is not synonymous with “diagnostic concordance.”
The concept of digitized slide integrity and the need for validation of all laboratory instruments (scanners) and information technology (IT) systems used: Integrity of the digitized slides should be ensured throughout the generation, encryption, storage, and transfer of WSI files. We believe that digitized slide integrity refers to maintaining the integrity of the unique features of WSI such as embedded metadata that provide information on the slide label and slide macro image. The trained pathologist is capable of judging whether the digitized slide integrity is preserved, and as part of the validation, the functional requirement for digital slide integrity should be tested. There is also expectation that where workflows are split between different GLP footprints, all parts of the chain involved in the generation, transfer, storage, and retrieval (viewing) of the image files are suitably validated for their intended use within each separate GLP footprint.
The concept of requirements for archiving: WSIs are only archived when they contribute to the generation of raw data and are necessary for the reconstruction of the study. They should be archived in a format that aligns with the expectations of other GLP-compliant electronic formats. For example, in contemporaneous peer review using WSI, if the final diagnosis by the study pathologist is undertaken using the glass slide, then only the glass slide is archived.
The usual principles that underpin GLP are followed: An expectation over sample integrity and traceability, chain of custody, and appropriate and documented training, procedural controls (ie, Standard Operating Procedures [SOPs]), and documentation of WSI use in study protocols.
The sponsor organization involved in this deployment of GLP-compliant peer review has been utilizing WSI as the default medium for the peer review of non-GLP studies for several years. The CRO involved has also used WSI for more than a decade to support its diverse and global pathology workforce and their clients. It was therefore a logical step to partner and expand WSI use to both the primary evaluation and peer review of GLP-compliant studies supporting first time in human submissions and ongoing clinical trials. In order to demonstrate the suitability of WSI for this purpose, distinct GLP-compliant validations were undertaken by the CRO and sponsor for the hardware, software, and procedures that would subsequently be used for a GLP-compliant primary evaluation and peer review of a rodent 3-month inhalation toxicology study.
The higher-level assumptions that underpinned the intended use and qualification, the validation process, and details of the steps undertaken by both organizations are subsequently outlined. Additionally, there are details pertaining to study conduct for the GLP study that was evaluated and peer reviewed using WSI. We believe that we have described the processes and validation needed to deliver digital primary evaluation and peer review of a GLP-compliant rodent toxicology study as exemplar for wider deployment by our industry.
Hardware and Software Validation
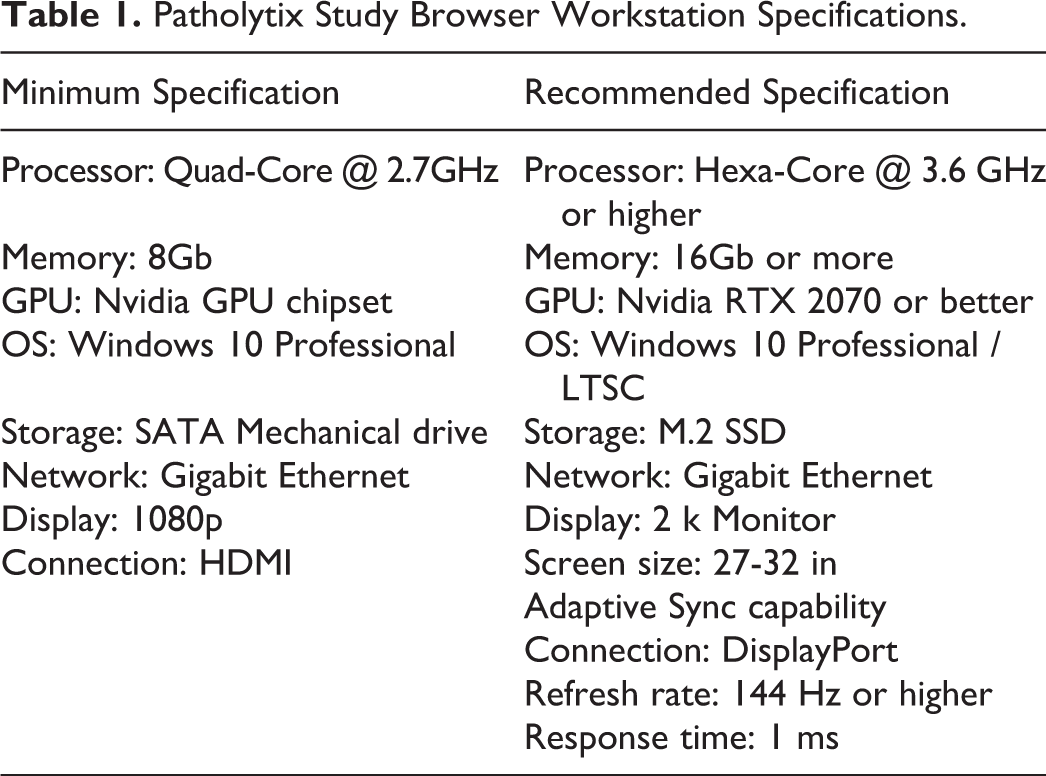
The CRO undertook a GLP validation of the Leica Aperio AT2 scanner and associated software (Scanscope Console 102.0.07005), and the Leica database software (eSlide Manager v12.4.3.7006) to provide GLP-compliant WSI that would be used for primary digital evaluation within the Leica Aperio eSlide Manager software environment using the Leica Web Viewer v12.4.3.7006. (This is subsequently referred to as the digital pathology acquisition platform). To facilitate cloud-based peer review, the CRO also undertook a GLP validation of the Deciphex Patholytix Preclinical product suite (v2.0) to include the Patholytix Core, Study Uploader, and Study Browser to enable the conversion, encryption, and transfer of the validated WSI and associated study specific metadata to the Patholytix cloud. (This is subsequently referred to as the digital pathology distribution platform). In addition, a mock peer review environment was built based on the digital pathology distribution platform recommended minimum specifications (Table 1) to support the validation of the complete CRO to sponsor workflow internally at the CRO.
Patholytix Study Browser Workstation Specifications.
The sponsor undertook a GLP validation of version 2.0 of the digital pathology distribution platform suite to cover the download and viewing of the WSI and metadata within an existing sponsor site GLP footprint in the United Kingdom. Figure 1 outlines the system architecture that covers the workflow between the 2 organizations.
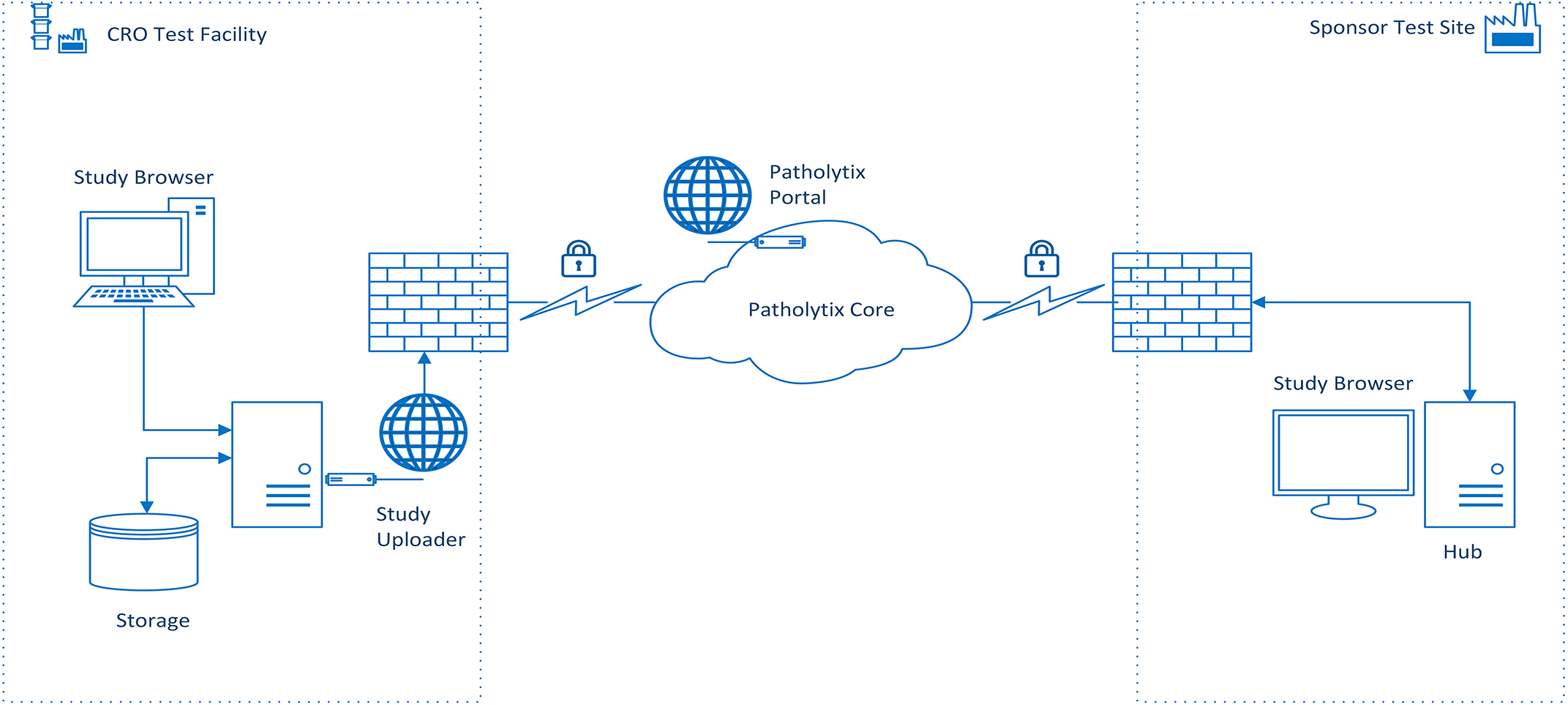
A high-level diagram that demonstrates the workflow for the transfer of WSI between the CRO and sponsor. The WSI files generated at the CRO are encrypted using the cloud-based digital pathology distribution platform and uploaded to the cloud along with the relevant associated metadata. The sponsor downloads the encrypted WSI files to a NAS drive. The downloaded files are viewed using the digital pathology distribution platform browser software installed on a Hub PC authenticated via the hub PC and browser software. CRO indicates contract research organization; NAS, network attached storage; WSI, whole slide image.
Both sponsor and CRO agreed to the overarching assumptions that would underpin the validation approach
2
: As the CRO and sponsor sat within separate GLP footprints and organizations, validation of the workflow contained within each organization would be entirely separate. As the CRO included validation of an internal mock digital peer review environment that covered upload and download of WSI and metadata from the cloud, the sponsor validation of the download and viewing of the WSI would comprise only a subset of the CRO validation activities. The sponsor would not validate upload of images to the cloud, as this feature was not required for peer review at the time of writing this manuscript. Given the acceptance and widespread use of WSI in the medical community, the process would be driven by the identification, definition, and mitigation of the specific risks relating to WSI use in a nonclinical GLP environment.
12,13
As contemporaneous peer review would not generate raw data,
10
the level of risk associated with this activity was therefore considered to be reduced compared to primary evaluation. If the scanner model used to generate the WSI has FDA 510(k) clearance, this removes the need to undertake additional specific qualification relating to the scanner technical performance. Performance testing for the scanner would thus be confined to the relevant aspects relating to installation qualification (IQ), operational qualification (OQ), and performance qualification (PQ) within the defined context of use required at the GLP test facility. The goal of validation would be to provide documented evidence that all hardware and software used to evaluate and review the WSI would consistently perform according to predetermined specifications and predefined quality attributes. These predefined quality attributes would be determined by the respective validation team, including experienced toxicologic pathologists to ensure that the quality of the WSI was suitable for primary evaluation and peer review. The underlying principle is identical to the judgment currently allowed by toxicologic pathologists when using a light microscope. Regulatory guidance has not required toxicologic pathologists to validate light microscopes but relies on the professional experience and judgment of a qualified individual to ensure the optical output meets the required standard. We believe the situation should be identical when using WSI. Additionally, as would be the case for the light microscope, annual maintenance and comparable procedurally controlled maintenance would be expected for the digital microscope (monitors, hardware/software). As the digital pathology distribution platform resulted in a workflow that would be split between 2 different organizations and their associated GLP footprints, the sponsor would include an end-to-end verification (original slide to corresponding WSI). The 2 organizations would share pertinent details of their validation plans to ensure alignment in key areas such as software versions (particularly where a software application such as the digital pathology distribution platform was used by both parties as part of the workflow).
Both the CRO and sponsor decided to base their validation plans on the principles of IQ/OQ/PQ rather than the software development life cycle model. Given that all software used by the CRO and sponsor in the WSI workflow were vendor provided production applications, we believed that the well-established IQ/OQ/PQ model would provide a robust and efficient process.
Although the approach to validation was similar between the CRO and sponsor, because the workflow involved utilized 2 separate digital pathology platforms at the CRO, and the cloud-based transfer of files to the sponsor, the authors opted to describe the validation undertaken at each organization separately.
Contract Research Organization Validation of Data Acquisition and Distribution
As a provider of the WSI and associated metadata to the sponsor, the CRO undertook the validation efforts under 2 separate streams: data acquisition (scanner, scanner software, data repository software, image management, and viewing software) and data distribution (cloud-based digital pathology platform).
The data acquisition stream included both the scanner and the software controlling the equipment, as well as the data repository and image management software. The stream was guided by a formal master validation plan and detailed risk assessment document to ensure full compliance with current regulatory expectations for Computer System Validation. 13,14 This included vendor assessment, risk assessment and associated mitigation, IQ of the hardware and software (meeting or exceeding vendor recommended specification 15 ) security and data integrity, and User Acceptance Testing (UAT) related to the quality of the images captured by the scanner. The validation cycle was conducted by a team composed of Computer System Validation experts, end users (including pathologists and technical staff), and IT personnel, with oversight by quality assurance (QA). American and/or European College of Veterinary Pathologists (ACVP, ECVP) board-certified pathologists specially trained in toxicologic pathology assessed that the WSI image was a faithful representation of the glass slide using criteria which included the presence of all tissues, appropriate focus of the image in all areas of the slide, and the color and overall quality for histopathological evaluation. This assessment was performed by 2 ACVP/ECVP diplomates reviewing WSI captured at ×20 and ×40 magnification using tissues sourced from 6 different species (dog, mini pig, mouse, rat, fish, and cat) that had undergone various staining protocols (hematoxylin and eosin, Masson trichrome, picrosirius red, toluidine blue, reticulin stain, and chromogen-based immunohistochemical stains).
Elements including audit trail and archiving were also addressed, and limitations defined such as the exclusion of some proprietary image browsing software because of lack of control over the version used (downloadable from vendor site). Quality control (QC) checkpoints were in place that were procedurally controlled and ensured image quality was as per pathologists’ expectations for the intended use. For example, the study pathologist can request rescans in the same way as a study pathologist would request recuts of nondiagnostic glass slides if any WSIs are considered to be of nondiagnostic image quality.
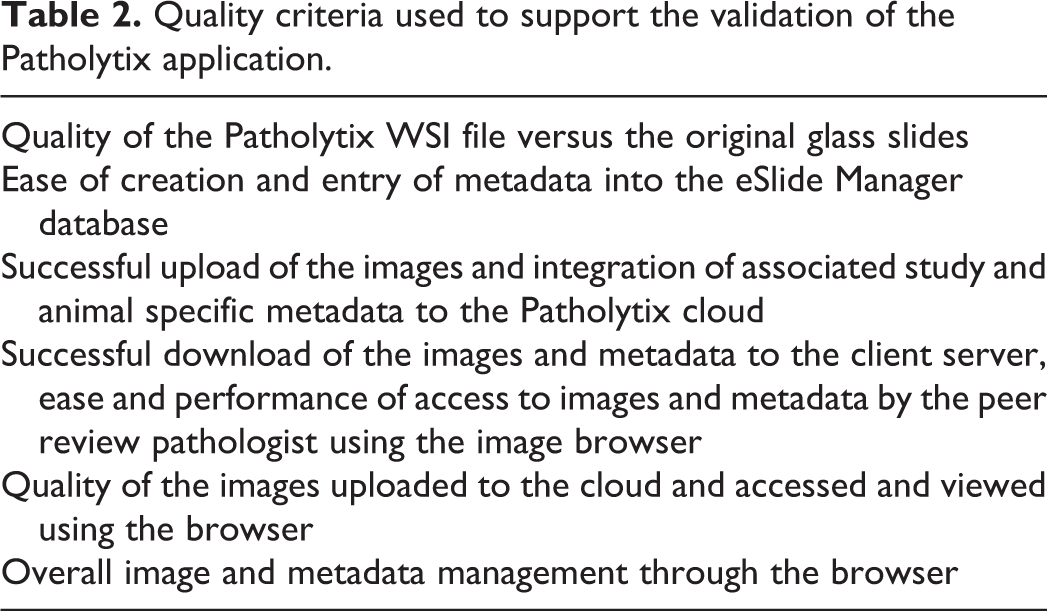
The validation of the data distribution stream (digital pathology distribution platform) was initiated after conducting a minimum of 10 pilot digital pathology studies led by ECVP/ACVP-certified veterinary pathologists, using 4 different species (minipig, non-human primate, rat, and mouse) and over 6000 different images (full tissue lists). Specific quality criteria were documented and the results internally archived. The pilot data provided confidence that the distribution digital pathology platform offered adequate image quality compared to the light microscope for the intended use of histopathological evaluation. Quality criteria used to support the use of the digital pathology distribution platform are listed in Table 2.
Quality criteria used to support the validation of the Patholytix application.

The validation of the digital pathology distribution platform, completing the workflow from data acquisition to data transmission, was initiated under version 1.4 of the product suite and completed under version 2.0, a version that provided significant enhancements to compliance elements to further support the requirements of GLP regulations. Challenges associated with the system multitier architecture of local and cloud components of the digital pathology distribution platform were addressed through detailed configuration documentation, a thorough risk assessment of the different layers and potential risks associated with the intended usage (peer review), as well as a testing strategy, again through formal IQ and UAT test cases, that included the complete workflow, including interorganization transfer (under the CRO environment in creating a “mock” sponsor environment) of images and metadata.
A process map was generated to fully assess the different roles and the functions associated with these roles that contributed to a requirement document that supported the intended use. With this mapping in mind, a detailed risk assessment was performed to ensure adequate measures were undertaken to mitigate the identified risks. Some of these included procedural controls to ensure quality of data input (images and metadata) from source to destination (peer reviewer). With the requirements compiled, accompanied by the risk assessment and validation plan, the configuration elements were also documented and confirmed through formal IQ testing. A mock “peer reviewer” environment was built based on the digital pathology distribution platform’s recommended minimum specifications (Table 1) to support the validation of the complete workflow internally at the CRO. This ensured that the entire process was validated, end-to-end, with testing of the complete cycle to support the full understanding of the application. This end-to-end validation at the CRO insured expertise to provide proper guidance and support for validation activities at the sponsor peer review pathologist’s GLP facility as well.
Testing included role definitions, trust creation, and sharing functions between uploading and receiving organizations/sites. Trust creation involved the generation and verification of site/organization information to ensure that WSI can only be shared between verified parties. Scripts evaluated image (scanned by a validated scanner and software) and metadata upload (some of it extracted directly from the laboratory information management system [LIMS] through a qualified software solution). Metadata extracted included both study and animal-related information as well as tissues and related macroscopic and microscopic findings where applicable. Other functions assessed were the integration of metadata to WSIs (including stress scenarios), image download (including file transfer integrity through checksum confirmation), and subsequent image navigation through the browser of the digital pathology distribution platform. Image quality testing was done with reference to the original glass slide by experienced pathologists to confirm that the image was a faithful representation of the slide and satisfied quality criteria for the intended use of histopathological evaluation. These criteria were identical to those used for the acquisition stream validation, allowing use of digital images for histopathological interpretation equivalent to slide evaluation.
The validation activity defined the process for both the scanning and intended use of WSI internally for primary evaluation as well as distribution of images to external partners for peer review activities. These processes were documented in written procedures and included risk mitigation registered within a risk assessment and controls for the quality of input, images, and metadata.
Most risks were categorized associated with potential quality issues or risks related to regulatory uncertainty. The risks associated with quality were mitigated through the following actions: Quality control points were placed within both the acquisition and distribution streams and involved verification of the image quality, along with the criteria and process for rescanning. There were also QC procedures targeting metadata accuracy and its availability with the images for both identification and assessment. Generation and scanning of a reference/QC multiarray slide containing samples of several different rat tissues stained with hematoxylin and eosin to confirm that the hardware and software used by study pathologist and peer review pathologist provided a faithful reproduction of the reference slide and written confirmation by peer review pathologist of the quality of the downloaded images. Qualification of the metadata extracts from the LIMS used to populate studies within the digital pathology distribution platform. Ancillary pilot studies as well as a peer-reviewed published proof of concept study
16
with cross organization input that supported the use of digital images for histopathological evaluation.
Regulatory-centric risks were mitigated with the following: A comprehensive validation approach that targeted the complete workflow against intended use (acquisition-distribution-archiving-restoration). The archiving included securing the metadata under the control of the archivist within the metadata acquisition database and transfer of the WSI to external media for physical archiving at the CRO with the study records. Standard operating procedures for all aspects of the workflow. Procedural controls (eg, ancillary forms, control over metadata files) to allow full study reconstruction through chain of custody. Verification measures to ensure the complete workflow was validated end-to-end for data chain of custody between organizations. Agreements in place to ensure proper security and management of cloud files. Clear definition of what constituted data in primary evaluation and peer review. Comprehensive audits of vendors to assess the level of compliance with regard to the appropriate quality systems in place.
At the conclusion of the 2 validation streams at the CRO site, both validation efforts were reported under a validation report, SOPs were issued, and systems were commissioned for use by test facility management.
Sponsor Validation of the Download and Viewing of WSI Files
Given that the CRO would be providing images from validated systems, the sponsors validation concentrated on OQ and PQ of the digital pathology distribution platform in the form of UAT. This activity followed established protocols used by the sponsor to validate software for GLP use. The system qualification was supported by a sponsor QA audit of the vendor and risk assessments relating to IT security. The validation team consisted of an ACVP-certified toxicologic pathologist (MJ) experienced with the use of WSI and cloud-based digital pathology software, the system owner, and IT specialists with experience in risk management, QA, system architecture, and software validation.
All relevant activities relating to the validation of the digital pathology distribution platform were captured in a validation plan that included elements to cover areas such as introduction and scope, organizational structure (personnel involved), roles and responsibilities, risk impact determination, validation strategy, test plan, acceptance criteria, document archiving, references, and relevant SOPs. The system validation also included a traceability matrix to ensure all requirements defined for the browser were met throughout the validation activities.
The validation process began with a sponsor QA audit of the software vendor which was undertaken remotely. This included confirmation that the software conformed with the expectations outlined in the International Society for Pharmaceutical Engineering Good Automated Manufacturing Practice (GAMP). GAMP 5 provides 17 a pragmatic framework for the risk-based approach to computer system validation where a system is evaluated and assigned to a predefined category based on its intended use and complexity. In addition to GAMP 5 expectations, other areas such as quality management systems, system development lifecycle, good information practice, information security, and system maintenance and support were also explored.
The vendor QA audit was accompanied by a “Risk Impact Determination: (RID) of the vendor Software as a Service as per sponsor requirements. The RID included elements to assess areas such as GAMP category, data privacy, regulatory Good Practice (GxP) status, data integrity, medical device classification, and IT security.
To avoid issues with bandwidth, it was decided to host the digital pathology distribution platform on-premise components on a workstation PC (Hub PC) in close network proximity to the end user, rather than within a standard server environment. Therefore, the software components were installed in a stand-alone Hub PC that was situated in a secure room within the existing GLP footprint of the sponsors UK site where the pathologist would undertake the peer review. The WSI files downloaded from the cloud were stored and accessed from a Network Attached Storage device (Synology DS218) situated in the same room as the Hub PC. The Hub PC contained an Intel i-7-8700 CPU with 8.0 GB of random-access memory (RAM) and an internal Intel UHD 630 graphics processor that met the vendor’s minimum specifications (Table 1). The WSIs were viewed on a Dell P2719H 27-inch 1080P 60 Hz refresh rate monitor. A 3D connexion SpaceMouse TM attached to the Hub PC was utilized to pan and zoom the images. As the CRO validation of the complete digital pathology distribution platform suite was completed using version 2.0 of the software, the sponsor also validated the cloud and image viewing components of version 2.0 of this software following its release. While the study upload feature of the digital pathology distribution platform is integral to the functionality of the platform core component, this feature was not required by the sponsor for peer review and was not therefore specifically tested in the validation activity. A representative from the vendor provided oversight of the installation of version 2.0 of the digital pathology distribution platform product suite on the GLP-dedicated Hub PC to ensure the installation process met the IQ protocol and acceptance criteria defined by the vendor. As the Hub PC was a single stand-alone installation of the vendor software, it was decided that it would represent the preproduction environment prior to the UAT and would be designated as the production environment once the UAT testing was complete, and the validation plan had been approved.
The approach to the validation of the digital pathology browser for peer review was based on the assumption that both the OQ and PQ requirements of the digital pathology browser software would be defined by a toxicologic pathologist experienced in the use of WSI. The approach used to validate the digital pathology browser consisted of combining the OQ and PQ testing in the form of UAT. The test scripts deployed for the UAT were written by a business analyst familiar with GLP system validation. Prior to the authoring of the UATs, a toxicologic pathologist familiar with the digital pathology browser (MJ), and the system owner, helped to define high level use case and user requirements (URS) for the digital pathology browser. As the digital pathology browser was a vendor application, the use cases and URS were ultimately dictated by existing digital pathology browser functionality, but these URS were based on a minimal set of specifications to cover all GLP relevant functions of the system 13 along with other features that supported an efficient workflow. GLP relevant functions included system security, user access (including audit trail), and the ability to identify GLP compliant from noncompliant studies. As the use case and URS would form the basis of the UAT scripts, we were careful to define the URS in a manner that could be practically tested and documented while undertaking the UATs. Examples of some of the high-level use cases and URS are included in Tables 3 and 4.
Examples of higher level user requirements (use cases) for a digital peer review application.
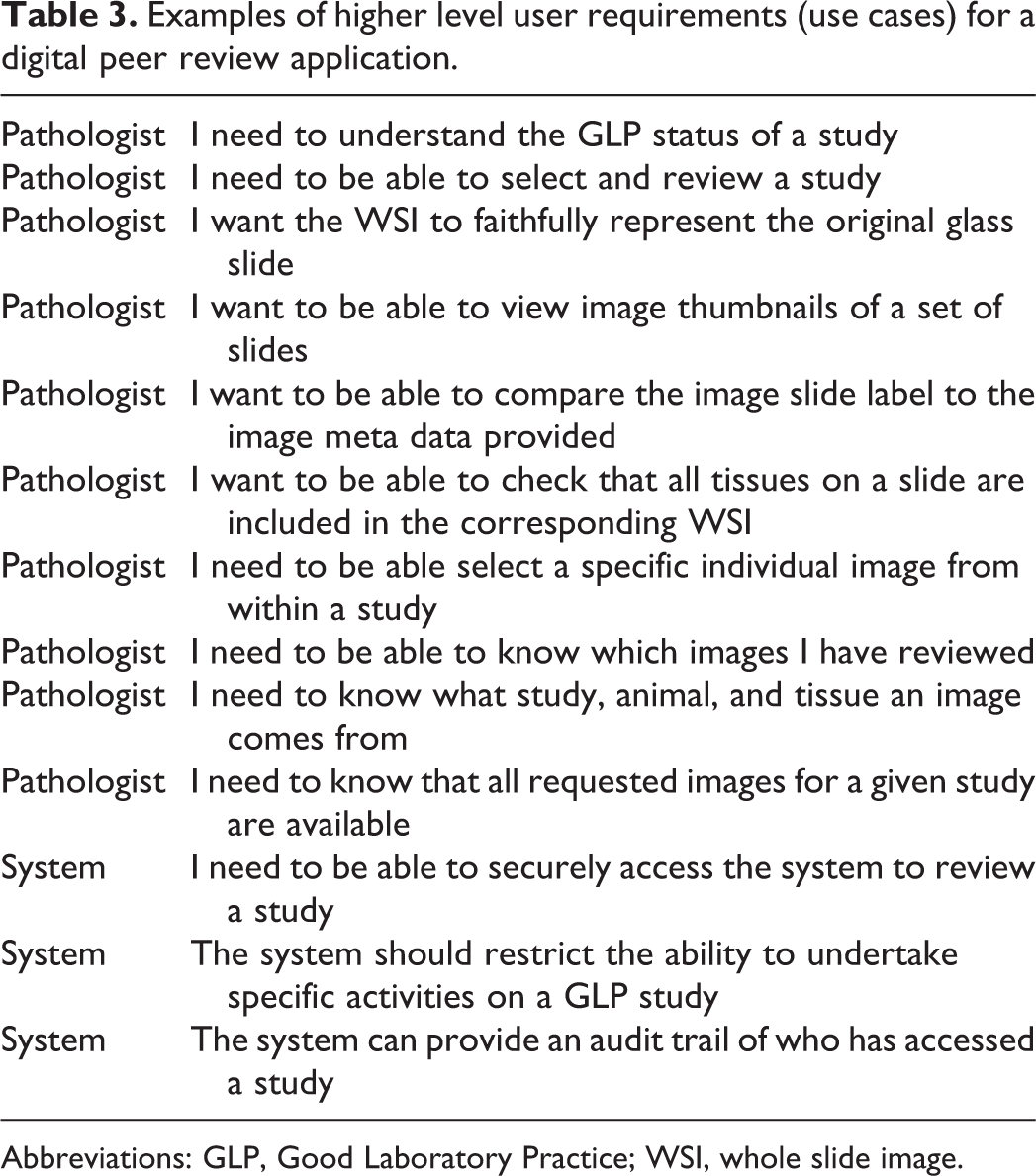
Abbreviations: GLP, Good Laboratory Practice; WSI, whole slide image.
Examples of user requirements defined for a cloud based digital peer review application.

Abbreviations: GLP, Good Laboratory Practice; WSI, whole slide image.
As the routine use of the digital pathology study browser for GLP peer review would be supported by an SOP, a new SOP was written to cover the use of the application prior to the authoring of the UAT scripts. Reference to this SOP helped the business analyst writing the UAT scripts to better understand the desired system functionality, and how the UAT scripts could effectively test this.
As the scanning of the original slides, collection of associated metadata, and upload of the WSI and metadata to the cloud were being undertaken and validated by the CRO, this process was out of scope for the sponsors validation activity. However, given that the transfer of the images from the CRO to the sponsor involved a cloud-based digital platform installed in 2 separate organizations and GLP footprints, the sponsor considered it would be prudent to include end-to-end verification of certain key attributes (glass slide to WSI) in support of the sponsor validation process. In order to provide appropriate WSI files for the UAT testing, the sponsor identified slides from a subset of animals from 2 sponsor nonclinical toxicology studies that were already archived at the CRO test facilities. Study slides included animals from different dose groups, sexes, and study numbers, in order to provide appropriate metadata to test the sort and filter functionality of the digital pathology browser. The slides identified for use in the UAT testing were scanned using GLP validated scanners at the CRO test facilities and uploaded to the cloud using a GLP-validated version of the digital pathology distribution platform. The draft UAT scripts designed to test the OQ/PQ of the digital pathology distribution platform browser underwent a “dry run” using a non-GLP installation of the browser at a separate non-GLP sponsor site, prior to their finalization and deployment for the formal testing using the GLP preproduction environment.
Formal UAT took place at the UK sponsor GLP site following the successful IQ of version 2.0 of the digital pathology distribution platform on the GLP designated Hub PC and after the system validation plan had been approved. As the sponsor only has 1 GLP-compliant footprint in the United Kingdom, and due to COVID-19 restrictions, the UAT tests were run by a single pathologist (MJ) who was familiar with the software application. The WSIs of the slide sets identified for the UAT were downloaded to the networked cache prior to performing the UAT scripts. Screenshots were captured to support the outcome of each test step (where appropriate) and were documented as supporting evidence. The supporting evidence and UAT scripts were signed off by the pathologist undertaking the UAT along with a member of IT QA.
In order to provide an end-to end QC check, the original glass slides that formed the test sets for the UAT testing were shipped to the sponsor and were used to perform a slide label and macro image QC check, which was undertaken by the pathologist running the UAT scripts. The slide label QC check involved comparing the human readable information on the glass slide label of each slide to ensure it corresponded to the thumbnail image of the slide label for each WSI file displayed in the digital pathology browser. In addition, the tissues contained within the glass slide were compared to the slide macro thumbnail of the slide in the study browser. This provided an end-to-end QC check of the workflow between the CRO, the digital pathology distribution platform cloud environment, and the sponsor. The results of this QC check were documented and used to support the UAT test scripts and other validation documents. A formal QC check of the other metadata displayed in the digital pathology distribution platform browser was not undertaken by the sponsor, as the transfer of these metadata was included in the validation undertaken by the CRO using an upload/download simulated peer review environment.
Once the UAT testing had been completed and signed off, the validation team reviewed the system validation report to ensure all requirements had been met prior to report approval and go live of the system. At system go live, the installation on the dedicated Hub PC was then designated as the production environment, with any further changes subject to standard change control practices.
Primary Evaluation and Peer Review of the GLP Toxicology Study
A 3-month rat inhalation toxicology study was chosen by the sponsor as being suitable for a pathology evaluation and peer review completed exclusively using WSI following the successful GLP validation of the hardware, software, and procedures within both organizations. Pertinent details relating to the hardware and software (model, version, etc) used for both the primary evaluation and peer review were included in the study protocol or by amendment. The study protocol was a guideline compliant full tissue inhalation toxicology study with 5 dose groups and 10/sex/group. The study slides (approximately 2400) were scanned using a validated scanner model at ×40 in 14 separate scanning runs. During each scanning run, the same tissue multiarray QC slide was also included with the study slides to provide a QC WSI for each scanner run. The tissue multiarray QC slide contained samples of several different rat tissues with a wide variety of staining intensities and cellular/morphological details stained with hematoxylin and eosin. The study pathologist (JB) only used WSI for the primary evaluation of the histopathology and accessed the WSI files remotely using a CRO server hosting the digital pathology acquisition platform software and viewed the images with the digital pathology acquisition platform’s web viewer, running on a Dell Precision 7920 Tower (Intel Xeon Gold 6130 CPU 2.10 GHz 16 Core[s] 32 GB RAM) and a Dell S2716DG Monitor (Nvidia G-Sync, 144 Hz refresh rate, QHD resolution, 2560 × 1440). The hardware used met or exceeded recommended vendor specifications. 17 Histopathology data were entered onto an LIMS system.
During the primary evaluation, the study pathologist first checked each of the tissue array slide WSI for consistency and suitability of color balance and focus. While evaluating the study WSI, checks were made of the slide label, presence of the required tissue according to the tissue list for that block, focus quality, image artifacts, and color balance. If any of these were found to be inadequate, a rescan was requested. As the WSIs were evaluated remotely from the CRO test facility, the study pathologist did not have direct access to the glass slides. However, the study pathologist could access the glass slides at the test facility. The CRO test facility SOP also stipulated that the study glass slides should be kept in a suitable location that allows the study pathologist to access them should they deem it necessary. All image metadata were archived within the local production database of the digital pathology acquisition platform with access limited to archive personnel. The WSI will be exported to external disk for physical archiving at the CRO facility archive in line with GLP expectations for electronic data. 13
To facilitate peer review, all of the original scanner derived WSI files and relevant associated metadata from the study were uploaded to the cloud environment using the digital pathology distribution platform.
The original glass multiarray QC slide used during the scanning runs was sent to the peer review pathologist to allow a QC check prior to conducting the peer review. The sponsor peer review pathologist (MJ) acknowledged safe receipt of the original glass multiarray QC slide as per GLP chain of custody expectations. The QC slide was returned to the test facility for archiving following the completion of the peer review. The full set of WSI files including the images of the glass multiarray QC slide was downloaded to the networked cache within the sponsor GLP footprint. The peer review was undertaken in person within the dedicated GLP area of the sponsors building using the validated production environment of the digital pathology distribution platform browser. The peer review pathologist viewed the WSI using the validated Hub PC (Intel i-7-8700 CPU, 8.0 GB of RAM, Intel UHD 630 graphics processor) and a Dell P2719H 27-inch 1080P 60 Hz refresh rate monitor. An ergonomic mouse attached to the Hub PC was utilized to pan and zoom the images. Prior to peer reviewing the WSI from the study slides, the peer review pathologist filtered the images to provide a set of thumbnails that represented the 14 multiarray QC glass slide scans. The pathologist was then able to use a microscope to compare the original glass multiarray QC slide with the WSI images of the corresponding scans. This provided the pathologist with an end-to-end QC check of key image quality features such as color fidelity, focus, and resolution. The successful completion of this QC check was documented by the peer review pathologist, and a signed acknowledgment was returned to the test facility for archive. Once the pathologist was confident of the quality of the scanning runs, they were free to perform the peer review as per test site SOPs. If the peer review pathologist deemed any of the study WSI to be of an insufficient quality to allow for adequate review, they would request a rescan and reupload as per the sponsor peer review SOP. The CRO and sponsor peer review SOPs did not specifically make any recommendation about what medium should be used (glass slide or WSI) in the event of a significant disagreement occurring between the peer review and study pathologist that would require escalation to a third party or peer working group. It would therefore be up to the discretion of the relevant individuals involved to decide on which medium all parties would support. For the GLP toxicology study described here, minor observations made by the peer review pathologist were resolved by both parties using the WSI available to them through their respective software applications.
Discussion
For GLP studies, the combination of achieving a realistic and timely workflow between CROs and sponsors along with concerns over regulatory acceptance has impeded the expansion of WSI use. Recent efforts by members of the European Society of Toxicologic Pathology and the Society of Toxicologic Pathology in the form of workshops and working groups have provided a better consensus regarding best practice for the use of WSI in GLP studies and provided a framework to guide validation strategies. 3 However, it is imperative that the CROs and sponsors translate these cross-industry efforts and guidance into the necessary GLP validations to enable full digital pathology workflow on GLP-compliant studies. While the basic requirements relating to GLP validation of laboratory instruments and IT systems used for digital pathology are well understood, 14 the actual validation process within a given GLP footprint is primarily driven by the intended context of use at each site, the nuances of which could vary between different CRO and sponsor scenarios and sites. With this in mind, the CRO and sponsor committed to undertake GLP validation of a WSI-based workflow for both primary diagnostic evaluation and peer review.
The potential for the division of workflow between 2 GLP footprints (CRO to sponsor) provides logistical challenges, not only in terms of the safe and efficient movement of the WSI files, but also the need to co-ordinate the approach to the GLP validation of the instruments and IT systems involved in their generation and transfer, particularly where a software platform is shared by both parties. The decision by the sponsor to request the use of WSI for a GLP-compliant primary evaluation and sponsor peer review required careful planning between all the key stakeholders, particularly as the cloud-based platform used for the peer review involved the transfer of images between 2 GLP footprints. Within the United Kingdom, there is a legal requirement that any study activity claiming GLP compliance must be undertaken in a facility that is registered with the Medicines and Health Care Products Regulatory Agency GLP compliance monitoring program. Such registered facilities must have a full GLP framework in place. It was for this reason that a full GLP validation of the digital pathology browser used at the peer review site was required in order to undertake a GLP-compliant peer review. The situation in North America is slightly different in that sponsors can claim GLP compliance for peer reviews at non-GLP sites, if certain safeguards can be demonstrated to the satisfaction of the study director.
The validation activities required a close partnership between experienced pathologists and other validation team members. Pathologists needed to carefully explain to other validation team members not familiar with WSI (IT, QA) the peculiar nuances of WSI when compared to other forms of electronic data, as these nuances often dictated the key risks and their associated mitigation strategies.
The successful completion of the GLP validation activities allowed the sponsor and the CRO to begin real-time “stress testing” of the entire digital workflow process using a large dimension GLP study. The safety study chosen for WSI use resulted in the primary evaluation and transfer of approximately 2400 image files. The workflow for the scanning and transfer of the WSI files required careful co-ordination but was achieved in a realistic time frame.
For the study described here, there were no significant disagreements between the peer review and study pathologist, and comments from the peer review pathologist were addressed by both parties referring to their respective WSI. Both the CRO and sponsor SOPs did not stipulate which medium should be used (glass vs WSI) for the resolution of significant differences that would require resorting to a third party or peer working group. Implicit in this approach is the ability of the pathologist to choose the medium they consider to be most appropriate. In the scenario where there is a significant disagreement due the presence of a finding that was or could not be observed in one or other medium, it is anticipated that the pathologists would include a review of the original glass slides as part of the remediation process. If the result of this remediation suggested that any deficit in the WSI was a contributing cause, the situation would require a thorough investigation that would include root cause and corrective and preventive action analyses.
A purely digital footprint avoided the need for travel and large-scale shipping, both problematic during the COVID-19 pandemic. With close attention to operational efficiencies and continued improvements in software and hardware supporting the digital workflow described, it is anticipated that the workflow will only advance in its potential to increase sponsor–CRO partnerships in drug development. Improvements in scanner throughput, and availability of cloud-based solutions for image evaluation and review, should increase the use of WSI on GLP studies, while providing both CROs and sponsors with flexible scenarios around scheduling pathology evaluation and reviews. At the time of writing this manuscript, the GLP study described here was in the process of finalization and has not therefore been submitted to a drug regulatory agency or been the subject of a test facility regulatory inspection.
By taking the first step in the integration of WSI for primary evaluation and peer review into a GLP study, and sharing our approach for GLP validation, we hope to encourage others to adopt WSI in a GLP-compliant manner and begin a dialogue with the regulatory inspectorate that may audit studies using WSI. As new technologies based on WSI like artificial intelligence-based machine learning emerge, a path has now been started that will open the door to exciting and innovative pathology tools within the regulatory toxicology space.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
