Abstract
As the use of digital techniques in toxicologic pathology expands, challenges of scalability and interoperability come to the fore. Proprietary formats and closed single-vendor platforms prevail but depend on the availability and maintenance of multiformat conversion libraries. Expedient for small deployments, this is not sustainable at an industrial scale. Primarily known as a standard for radiology, the Digital Imaging and Communications in Medicine (DICOM) standard has been evolving to support other specialties since its inception, to become the single ubiquitous standard throughout medical imaging. The adoption of DICOM for whole slide imaging (WSI) has been sluggish. Prospects for widespread commercially viable clinical use of digital pathology change the incentives. Connectathons using DICOM have demonstrated its feasibility for WSI and virtual microscopy. Adoption of DICOM for digital and computational pathology will allow the reuse of enterprise-wide infrastructure for storage, security, and business continuity. The DICOM embedded metadata allows detached files to remain useful. Bright-field and multichannel fluorescence, Z-stacks, cytology, and sparse and fully tiled encoding are supported. External terminologies and standard compression schemes are supported. Color consistency is defined using International Color Consortium profiles. The DICOM files can be dual personality Tagged Image File Format (TIFF) for legacy support. Annotations for computational pathology results can be encoded.
Keywords
Introduction
The use of digital and computational techniques throughout the field of toxicologic histopathology is expanding in academic institutions, government, the pharmaceutical industry, and contract research organizations. Challenges related to efficiency, scalability, and interoperability arise as slide scanning devices, image management platforms, image viewers, and analysis tools from different commercial and open sources proliferate. The equipment and software used for the examination and analysis of small animal research tissue typically overlaps that used for human anatomic pathology. Consequently, the solutions developed for clinical use can be applied. This article will focus particularly on the needs of whole slide imaging (WSI) and virtual microscopy (VM), though similar approaches are applicable to imaging of gross specimens, electron microscopy, and other related techniques.
Interoperability
Interoperability may not be a factor when a single very high throughput use case can be identified, for which a single supplier can provide an end-to-end solution for acquisition, management, and result production and dissemination. A significant expense for a dedicated solution used for no other purpose may be justifiable. More typically though, a laboratory is challenged by a multitude of different, heterogeneous tasks of variable complexity requiring different sources of tissue, different preparation steps (including sectioning, fixation, embedding, and staining), image acquisition with different resolution, illumination and throughput requirements, and varying combinations of human interpretation and manual measurement, classical image processing and quantification, and application of artificial intelligence, machine learning, and computational pathology (CP) techniques. 1 Rapidly evolving technology applicable to each of these suboperations means that using a single solution from a single supplier may be impractical. A best-of-breed approach that integrates multiple components from multiple suppliers is needed. This presents a challenge for interoperability at each of the boundaries between successive components.
Aside from immediate operational considerations, interoperability is also relevant to regulatory requirements and reproducibility concerns. No matter which regulations in what jurisdiction apply to a laboratory and its software and devices, authorities typically recognize the benefits of adoption of well-known industry standards. The preservation of records in an interoperable form not only satisfies regulatory requirements but also facilitates the reusability of data and reproducibility of experiments. Quite apart from added business value, increasingly, the general need to adopt Findability, Accessibility, Interoperability, and Reusability principles 2 is understood, and in the clinical realm, interoperability of health records has risen to the level of specific legislation and regulation. 3 In the context of toxicologic pathology, which may be subject to Good Laboratory Practice (GLP) regulations, use of a standard, open, nonproprietary format helps address archival requirements for data as well as long-term accessibility of required electronic records compromised by proprietary system obsolescence. 4 A cynical but realistic additional benefit of interoperability is that during corporate acquisitions and mergers, whether of suppliers, service providers, or customers, integration, combination, or replacement of disparate but interoperable solutions is greatly simplified.
Of more immediate contemporary relevance, the ongoing COVID-19 pandemic has resulted in the displacement of the workforce from on-premise to offsite and emphasized the importance of telepathology solutions to enable remote work. 5 Even though previously deployed infrastructures may not have been ideal, expedient solutions and the deployment of research-only tools for clinical use through the application of waivers from burdensome regulatory requirements 6 , as well as expedited review and validation, have proven popular. In the long term, the increasing experience with and availability of telepathology will inevitably lead to an increase in demand for interoperability. This will be particularly so when heterogeneous types and sources of imaging data are involved, and third parties are able to provide scalable and secure archival, distribution, and viewing, as well as integrate a multitude of analysis tools to provide a consistent solution for on-premise and remote use.
Interoperability in this context is defined in the IEEE Standard Computer Dictionary to mean “the ability of two or more systems or components to exchange information and to use the information that has been exchanged.” 7
A simple model of the components involved and the interoperability boundaries between them are illustrated in Figure 1. The slide scanner device acquires digital whole slide images; the image archive receives, stores, and distributes them; the image manager tracks and organizes the images and indexes their metadata; viewers display them on the screen and allow for human-operated annotation and quantification tools; and analysis systems automatically produce derived categorical or quantitative information from acquired images.
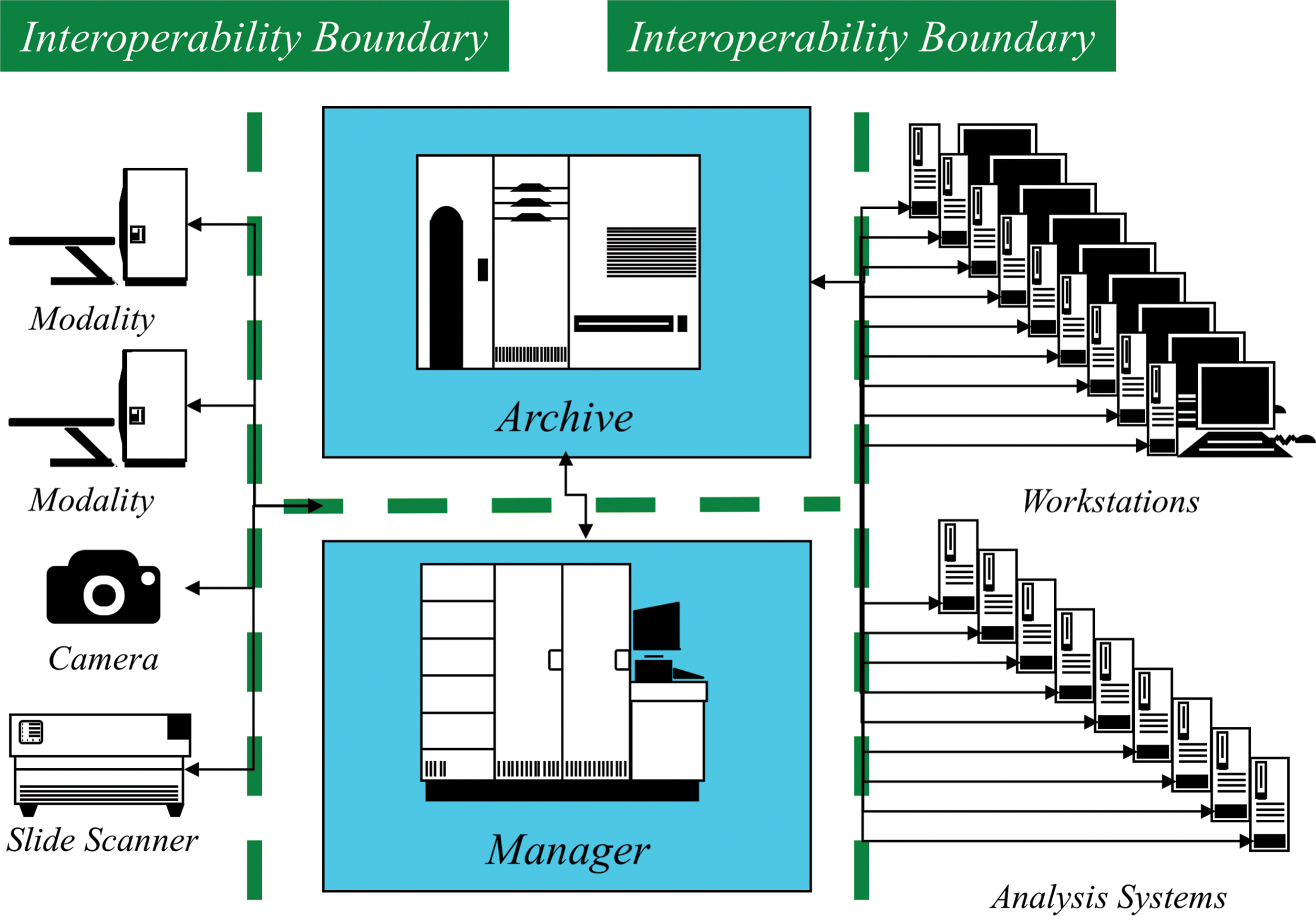
A simple model of interoperability boundaries between whole slide imaging (WSI) components.
Information is exchanged across each interoperability boundary in the form of an object, such as an image in a file format supported by both sides, and a protocol that manages the exchange, again one understood by both sides. In the simplest case, the management of the exchange can be performed by a human, for example, a user can manually move an image file from one file system to another. This is not scalable and is error-prone, so more typically an automated protocol is used, ranging from shared file systems with watched folders through generic industry and consumer-oriented file transfer protocols to dedicated biomedical imaging protocols with application-specific features. More complex transactions across the interoperability boundaries can address requirements beyond transfer, such as query and retrieval, reliability (including storage commitment), and annotation (labeling, tagging). Underpinning the information exchange may be security-related requirements for confidentiality, integrity, and availability.
There are essentially two approaches to interoperability between a system that is a source of information and the other a recipient: the source defines its own objects and protocol, and every recipient understands all possible source objects and protocols and the source uses a standard object and protocol, and every recipient needs only to understand that standard.
Both approaches can be used relatively successfully on a small scale. Only the second approach, the use of standard objects and protocols, is scalable in the long term.
Proprietary Format Reading Libraries
When WSI scanners came to market, each vendor developed their own proprietary format. Consequently, as these machines entered use in academic research environments, tools and libraries were developed to read those proprietary formats. Some were shared as open source software projects, notably OpenSlide 8 and Bio-Formats. 9 The emphasis was initially on reading the proprietary file formats, and manual intervention was used instead of an automated interchange protocol. Proprietary, but open, platforms were developed for storage and management, such as OMERO, 10 and proprietary, but open, protocols and application programming interfaces (APIs) were added to facilitate access by viewers and analysis tools. Fortunately, many of the proprietary file formats were based on well-known generic image file formats used in other industries, such as the Tagged Image File Format (TIFF), 11 so generic software components, libraries, and tools were reusable. Indeed, the TIFF standard was extended to satisfy the requirements of very large images for WSI, resulting in BigTIFF, 12 support for which was assured by extending the well-known libtiff implementation. 13 Unfortunately, not all formats were based on TIFF, thus complicating matters. Nor was there agreement on how sections of the images, tiles or strips, should be encoded, overlapped (or not), and compressed. As scanners and updated software versions and capabilities proliferate, so too do the variations in the proprietary file formats. Both open source and commercial developers are thus faced with the need to constantly revise, extend, and update their implementations, chasing the variations in the input format, coupled with the need to reverse engineer undocumented formats and variations. Further complicating deployment, user applications for viewing and analysis have dependencies on updated versions of the libraries they use and constantly need to update their copy of the libraries and perform regression testing to assure nothing has been broken by an update. The addition of fundamental new features, such as z-stacks or fluorescence channels, may trigger the need for a revised library API, further complicating the integration of conversion libraries with downstream software.
In short, though the use of proprietary formats and closed single-vendor platforms has been prevalent until now, such an approach is dependent on the availability and maintenance of multiformat conversion libraries. Though expedient for small deployments, this is not sustainable at industrial scale, as is evident by the stagnation of development of, or even the formal announcement of end of support for new variants in, well-known libraries. 14
Interoperability Standards
Rather than every scanner producing a proprietary format, every scanner could theoretically output a single standard format that is usable by every recipient. In practice, though there are subtle variations in the patterns of the acquired and reconstructed pixel data, in almost all cases, a single encoding approach, or very small subset of approaches, is sufficient for the vast majority of use cases.
Though cynics would suggest that the development of a suitable standard would require a definition of yet another new image standard,
15
in practice, there are essentially two candidates for consideration as the basis for biomedical WSI application standards: TIFF Digital Imaging and Communications in Medicine (DICOM)
As has already been mentioned, TIFF, which is the basis of many proprietary formats, had been extended to support WSI pixel data encoding as BigTIFF. Further, some WSI scanners and management software have the ability to export so-called “generic TIFF” images. Support for reading and writing TIFF and BigTIFF files is common in many general-purpose image display and processing tools, though dedicated WSI support in microscopy-specific tools is usually needed to handle tiled images, multiple resolution layers, and overview and label images. Though the TIFF documentation 11 describes the use of strips or tiles for images, as well as the potential presence of images flagged as subresolution images, there is no formally defined profile to describe how a pyramidal tiled multiresolution needed for WSI and VM should be encoded. This has resulted in some variation in choices by different implementers and the detailed specification of implementation choices by some library producers. 16,17 Regardless, the use of TIFF for WSI is widespread among commonly used open source and commercial histopathology research software viewing and analysis tools. Further, the open source and commercial whole slide image reading and conversion libraries support generic tiled pyramidal TIFF. The TIFF supports various different lossless and lossy compression schemes, including baseline Joint Photographic Experts Group (JPEG), which is very commonly used, and proprietary TIFF extensions introduced to support other standard schemes, like JPEG 2000. Note that TIFF is only a file format and does not define a protocol for exchanging and managing the images. The official TIFF documentation 11 defines a basic set of metadata for describing characteristics of the images, as well as a means of adding new application and product-specific tags for metadata. In practice, WSI TIFF implementers have eschewed the use of TIFF tags for metadata in lieu of other alternatives, such as their own semi-structured plain text or eXtensible Markup Language (XML)-encoded information, either buried inside a TIFF tag or in a separate accompanying file. 18
The DICOM standard, 19 on the other hand, defines both formats for image pixel data and metadata encoding and protocols for exchange and management. It is a dedicated biomedical standard, though it has been applied in other industries such as nondestructive testing of parts 20 and security screening. 21 It is ubiquitous in radiology and increasingly commonly used for almost every medical imaging specialty that has made the digital transition within the healthcare environment. 22 Just as TIFF needed to be extended to BigTIFF to encode large images and to have conventions defined to support tiled pyramidal multiresolution WSI, so too DICOM needed to be extended to support WSI pixel data as well as to provide application-specific metadata including specimen description. This work was performed over a decade ago 23,24 and has long since been incorporated into the current DICOM standard. 19 An ongoing maintenance process by the still active digital pathology working group (WG 26) assures that appropriate clarifications, corrections, explanations, and extensions are incorporated as necessary. 25
Why is DICOM ubiquitous in clinical image management rather than alternatives like TIFF, or other consumer-oriented formats like JPEG or Portable Network Graphics (PNG)? Various factors account for this. The DICOM standard’s rich biomedically specific information model provides clear definitions of metadata for both identifying and describing application-specific subject and acquisition information, which is common across all specialties and modalities. The inclusion of protocols for exchanging and managing the objects in an automated manner is not provided by any other standard in this space. Like TIFF, DICOM is a completely open standard, free to obtain, read and implement without license fees; it is also available in a machine-readable format that simplifies updating of toolkits and libraries. High-quality open source reference implementations of the standard are available, as well as commercially supported toolkits, for most common platforms and programming languages. Commoditization of all manner of products has occurred through the use of DICOM, including scanners, archives, workstations, viewers, and toolkits for products, testing, analysis, and research. Conformance and interoperability testing venues and opportunities abound. A core principle of DICOM is backward compatibility, such that changes add new features but do not invalidate the installed base of systems, software, and archived images. Finally, there is its history, in that DICOM filled a void at a time when there was no serious competition.
Today, 35 years after it was conceived, DICOM’s huge installed base ensures that off-the-shelf scalable enterprise and cross-enterprise open source and commercial solutions for almost every conceivable imaging application are available. Additionally, for environments in which the use of formal international standards is required or preferred, DICOM has been adopted as ISO 12052. 26
Radiology was among the first of the medical imaging fields to recognize that images would be shared electronically beyond the confines of an individual device and that communication between acquisition, storage, management, and display devices would be necessary. 27 At the first formal conference about what would become known as Picture Archiving and Communications Systems (PACS), the need to standardize such communication was recognized and a whole track of papers on this subject was included. 28 During this period, DICOM was established as a collaboration between academic and industry radiology partners encouraged by regulators in the early 1980s, though it didn’t really become successful until the release of the current form of the standard in 1993. 29
Despite its age, DICOM has evolved over time, not only with new information models, metadata, and pixel data encoding and compression mechanisms to support novel applications and modalities but also with new protocols to support workflow and new access mechanisms, such as the http-based DICOMweb protocols, 30 as well as new representations to augment the traditional binary file encoding, including XML and JavaScript Object Notation (JSON) transformations of the metadata. The DICOM standard has included explicit support for microscopy since 1999 and WSI since 2010.
As such, DICOM is a suitable interoperability standard to enable the operationalization of digital pathology imaging on an industrial scale, leveraging the same tools that are used throughout the enterprise for all other types of medical imaging. This applies equally to animal toxicological pathology as it does to human clinical histopathology imaging, particularly to the extent that the same scanning devices, archiving and management systems, viewing software, and analysis platforms can be shared, leading to economies of scale and mitigation of niche provider dependencies. The remainder of this article will address the specifics of the use of DICOM for WSI in toxicological pathology, as well as describe how TIFF and DICOM tools and images may peacefully coexist in the same deployment environment.
Tiled Pyramidal Multidimensional Pixel Data
The base (highest) resolution layer of WSI pixel data is typically very large. For example, a typical 20 mm × 15 mm piece of tissue scanned at 0.25 µm per pixel (nominal 40×) is 80,000 × 60,000 pixels, 4.8 billion pixels. With one byte used for each of three red, green, and blue (RGB) channels, that is 14.4 GB, uncompressed. Even with significant lossy compression applied, the bulk data size is still large. Since the pathologist does not view the entire slide at full resolution all at once, some means of rapidly accessing the subset of pixels in the desired subregion, at the desired level of resolution, is necessary to support the VM viewing paradigm. When adding support to DICOM for WSI, WG 26 was faced with two alternatives to address this: use of a pyramidal multiresolution, tiled representation, in which individual tiles in different regions at different resolutions are separately compressed and use of a full image transformation and compression scheme with inherent multiresolution decomposition, together with a means of accessing different regions at different resolutions.
In essence, the debate revolved around using the relatively simple and expedient TIFF-like approach that had been popularized in various proprietary formats, as opposed to the full image wavelet-based approach used in JPEG 2000 with JPEG Interactive Protocol used to address subregions. 31 There were vociferous protagonists of each alternative. In the end, it was concluded that only one approach would be selected for inclusion. The nominally simpler TIFF-like tiled representation with separate encoding of tiles and resolution layers was chosen, despite the wavelet approach allowing for greater compression, absence of tile boundary artifacts, and an encoding that did not require extra layers to be transmitted. The estimated 30% overhead for this choice was deemed a reasonable tradeoff against complexity. The use of JPEG 2000 for intratile compression can still be used in DICOM, though the more common choice is baseline JPEG.
Though DICOM could have encoded each tile in a separate file, since they would number in the hundreds of thousands, this was not deemed practical, so a multiframe encoding was chosen in which each tile is a frame in a DICOM file. Though DICOM could have encoded all resolution layers in a single file, it was decided to send each layer as a separate image and further to separate label and overview images into additional files. Thus, a whole slide image with say, five resolution layers, would consist of five separate DICOM files, with additional label image and overview image files. It is typical, though not required, to group all the images from a single scan into a single DICOM series and not to mix images from different scans in the same series. Note that the DICOM design differs from the most commonly used TIFF-based approach to encoding the multiple resolution layers and associated images in a single file, each described by a separate entry in the TIFF Image File Directory (IFD).
As mentioned above, baseline JPEG is typically used for RGB bright-field WSI. It is only one of the various different compression schemes (Transfer Syntaxes) supported by DICOM, however, and numerous cross-industry-standard lossless (reversible) and lossy (irreversible) methods are defined, including JPEG 2000 and JPEG-LS. The DICOM standard supports not only 8 bits per channel RGB representations but also 16-bit depth as well as single and multichannel data, such as is necessary to represent multispectral and fluorescence images. The decision to use lossy compression or not in GLP regulated environments needs careful consideration with respect to the integrity of the source data, just as for digital photographs. However, this decision is separable from the question of the format chosen, TIFF or DICOM, since both support lossless and lossy schemes.
Not only multiresolution images but multidimensional data beyond a single focal plane, including the use of Z-stacks (multiple focal planes), as well as multichannel, multispectral data, are supported. Hence, additional structural metadata is needed to describe the tile pattern and the location and ordering of tiles. The physical size and physical layout of tiles on the slide are considered important not only for the viewer experience but also for size measurement and other quantitative analysis applications. The physical size information for each resolution layer is explicitly defined, as opposed to being implicit as in some proprietary formats. The DICOM standard defines a normative slide-relative coordinate system that defines both the orientation and origin of physical coordinates, as well as a means of describing the total pixel matrix imaged and each tile’s relative and absolute position within it. Tiles may be encoded sparsely, each with attributes defining an explicit location. Alternatively, a raster scan pattern predefined by the standard may be used, such that all tiles of an entire rectangular region are encoded, with the location of tiles being computable; this makes the structural metadata shorter and simpler to parse and the tile index for viewing faster to build.
Having selected a tiled approach, the conventional DICOM pixel data encoding mechanisms were used for each tile (each frame), just as for all other modalities. All of the existing DICOM protocols and mechanisms for transport, storage, management, query, and retrieval of objects can be reused unchanged, including those for accessing metadata and retrieving selected frames. Obviously though, a WSI-aware application is needed to view the images in a meaningful way, since a traditional radiology viewer that is expecting single independent images or slices will not provide the VM experience, which requires an understanding of tiles and resolution layers.
Metadata
One of the primary strengths of DICOM is its information model and its definition of metadata. Like all image-based formats, some structural metadata is necessary to describe the encoding of the pixel data, such as the size of rows and columns, number of tiles, type of pixel (grayscale or color), the number of bits per channel, and in what manner the pixel data are compressed. Both TIFF and DICOM are very similar in this respect, having evolved at the same time, ostensibly with some cross-fertilization of ideas between their developers.
Where DICOM and other formats differ significantly, however, is in the definition and inclusion of both identifying and descriptive metadata related to the subject of the image, under what circumstances it was acquired, and in what manner it was acquired. This metadata helps to address attributability requirements in a GLP environment. The DICOM standard defines a largely hierarchical information model that contains various entities, such as patients, specimens, and equipment, and various organizational groupings such as imaging studies and series (Figure 2).
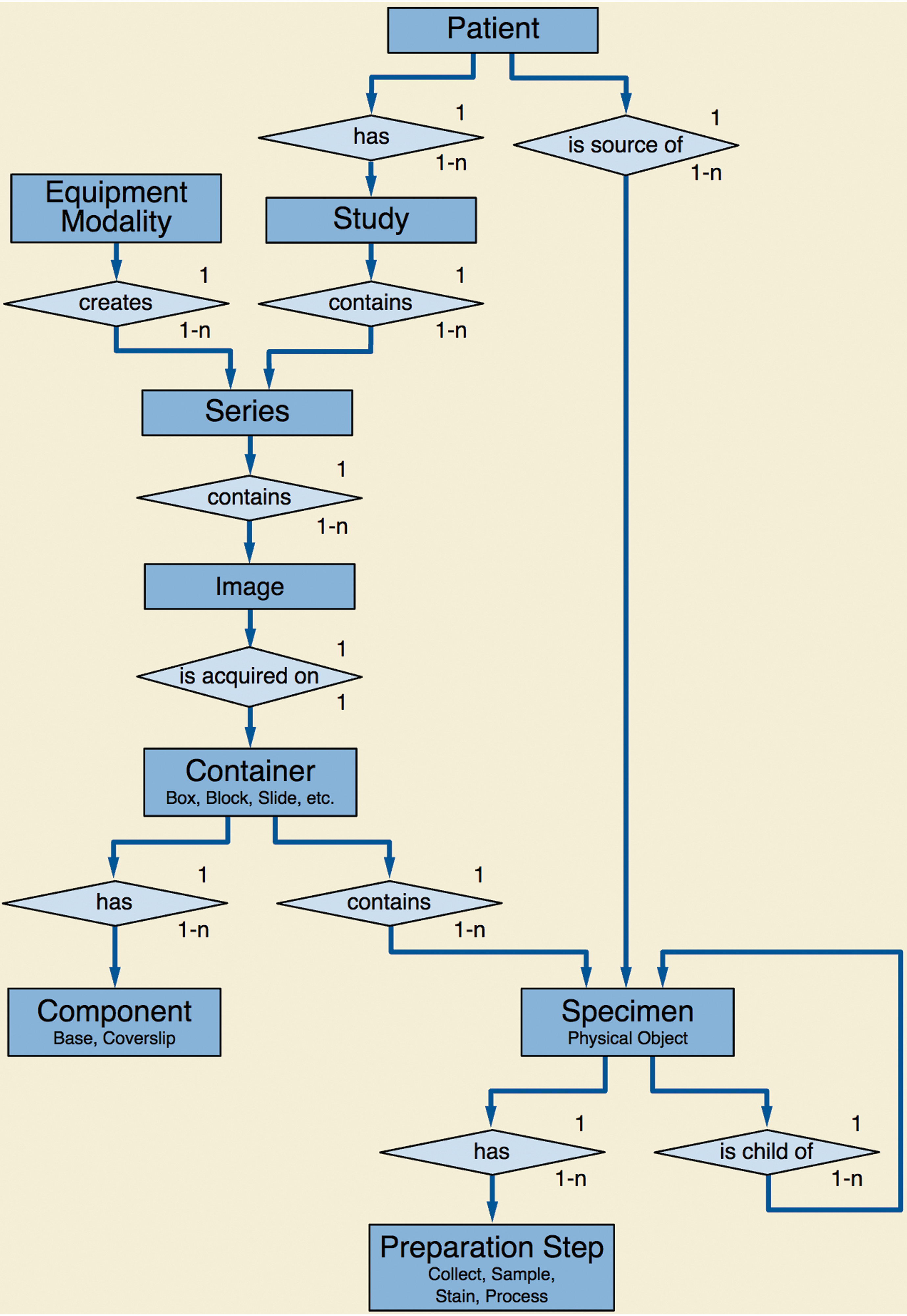
Digital Imaging and Communications in Medicine (DICOM) model of the real world—Specimens.
To the extent that whole slide images are the same as any other image of a patient (or research subject), a significant commonality exists across different modalities. For example, patients are identified by attributes such as name and ID, and described by attributes such as age and sex, using the same data elements regardless of whether the image is for WSI, or a computed tomography (CT) or magnetic resonance (MR), or a skin lesion photograph. This commonality of the higher level context of an image allows for reuse of the same protocols, storage, query and retrieval mechanisms, databases, and archives, regardless of the image type. In an enterprise-wide system that is modality and application agnostic, whole slide images can peacefully coexist with all other types of images, and the user can select those that are relevant to view or analyze. No other standard provides this level of interoperability and cross-modality functionality.
The imaging subject need not be a human, and even though the term “patient” is used, can be a research subject. Species (or other taxon) of the subject may be encoded, as well as greater detail, including breed and strain. In nonclinical research applications involving small animals, other modalities than pathology often image multiple subjects at the same time and encode them in the same image (eg, an MR, CT, or positron emission tomography “mouse hotel”). 32 Accordingly, DICOM includes a model for describing multiple subjects in one image masquerading as a nominal single “patient,” as well as a mechanism for describing the location in the image pixel data of specific animals and a mechanism for referencing original images and locations from derived images in which individual subjects have been separated (split). The same mechanism could be applied to WSI of tissue microarrays (TMAs) that include samples from different subjects, though there has been little experience with this to date.
To the extent that specimen imaging, or WSI in particular, is different, application- and modality-specific modules define appropriate identifying and descriptive attributes. For example, specimens and their subparts, including individual labeled slides, are identified by specimen- and container-specific human-readable identifiers and machine-readable unique identifiers, as well as being described by text and coded attributes. Specimen preparation is addressed in detail, such that one can provide text or coded information to describe such things as sampling, fixation, embedding, and staining.
The acquisition equipment can be identified and described, using both common cross-modality attributes (such as manufacturer, model, software version, serial number, and universal device identifier) and microscopy-specific information including lenses, illuminants, and filtration. Different optical channels may be described, and individual frames matched to a specific optical channel. This mechanism is intended to support multispectral and fluorescence imaging. It is also possible to describe samples that have been multiply stained (eg, with different antibodies and fluorophores) and imaged simultaneously or cleared, restained, and reimaged, though experience with such multiplexed imaging is limited so far, and there may be opportunities to extend and improve the model and the cross-referencing between the specimen preparation and optical path descriptions.
In all cases where types and descriptions of recognized well-defined entities, activities, and processes are needed, standard codes can be used instead of plain text. The need to use controlled terminology, standard lexicons, and standard ontologies for both common biomedical and domain-specific applications such as histopathology is well recognized and has been embraced by DICOM from early on. In particular, DICOM has a close relationship with Systematized Nomenclature in Medicine (SNOMED) International, which allows for license-free and fee-free global use of a relevant subset of codes. Most of the WSI-related codes, including those for anatomy, specimen preparation, and imaging technique, are drawn from SNOMED Clinical Terms. Other sources of codes are used as necessary, and specialty-specific codes can also be either mapped or substituted as appropriate. A relevant example might be the International Harmonization of Nomenclature and Diagnostic Criteria for Lesions in Rats and Mice (INHAND) or the Standard for Exchange of Nonclinical Data (SEND) standard codes for toxicological pathology, 33 –35 to the extent that they contain concepts related to context or acquisition, as opposed to image findings.
In addition, identifying and descriptive metadata is acquired for image management and provenance. Not only is it critical to capture the identity of the imaging subject but also to be able to relate the image to the physical assets (such as the slide identifier) but also the request for the procedure (such as via an accession number). Appropriate date and time information also needs to be recorded and persisted.
This description of metadata begs the question of its source and its relevance. The traditional DICOM approach expects the acquisition device to populate a rich set of descriptive and identifying metadata. In the early days of CT and MR scanning, some of this information was entered by the human operator at the console and the result automatically generated by the device from implicit knowledge of the imaging technique. Nowadays, all high-volume DICOM image acquisition in radiology, and most in related fields, involves the integration of the acquisition device with reliable primary sources, such as the departmental information system. In radiology, this is the Radiology Information System, and in human clinical histopathology, it is the Anatomical Pathology Laboratory Information System (AP-LIS). To the extent that similar metadata is required for toxicological pathology, integration with similar systems using similar standards for messages and protocols can be envisaged. In radiology, DICOM Modality Worklist 36 is usually used, and this was originally proposed for pathology too. 37 However, since AP-LIS vendors generally produce HL7 V2 messages routinely, integrations between WSI scanners and existing AP-LIS generally use that approach. There are ongoing efforts to standardize the necessary HL7 V2 messages in IHE. 38 There have also been suggestions that the HL7 FHIR 39 may be a suitable messaging protocol to use in future, since it is more amenable to queries. The selection of the relevant set of metadata from the primary source is made by either manual or automated provision of an appropriate key, such as a barcode read from the label, in the case of a microscope slide.
Regardless, it is highly desirable to somehow fully populate the DICOM image metadata before distributing the images. The approach of integrating the scanner with the AP-LIS is preferable, but another alternative is to improve the DICOM image produced by the scanner before propagating it. Figures 3 and 4 illustrate the two alternatives of upstream integration and a downstream processor that augments the metadata retrospectively. Other variants can be envisaged, but it is extremely important that the image archive not be populated with DICOM images that are devoid of useful metadata, which significantly undermines the value proposition for using the standard.
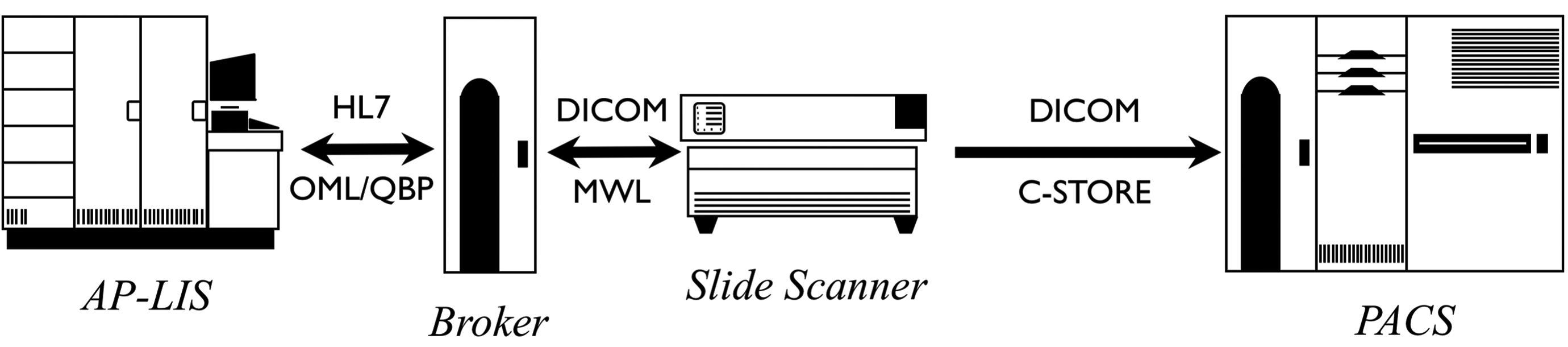
Sources of metadata—Upstream integration of metadata from Anatomical Pathology Laboratory Information System (AP-LIS) into images.
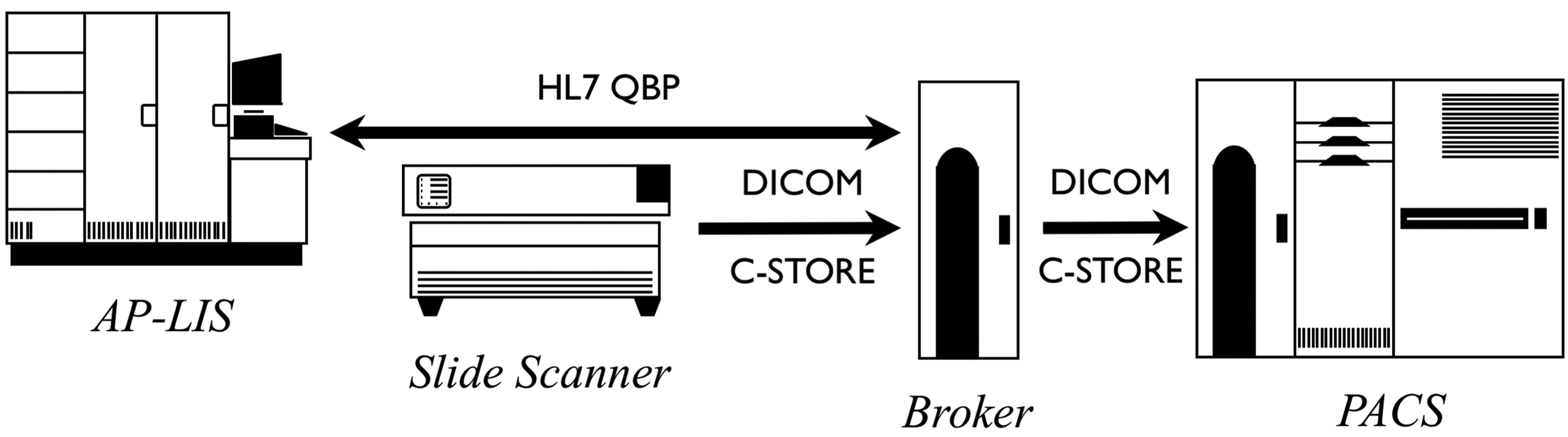
Sources of metadata—Downstream addition of metadata from Anatomical Pathology Laboratory Information System (AP-LIS) into images.
The importance of the DICOM metadata is apparent when the images are viewed or are detached from the system in which they are stored. Consider an image viewer that receives a set of DICOM (or any other format) image that is devoid of identifying and descriptive metadata. How does the recipient know which person or animal is the subject of the image, what part of the specimen it is, what specimen processing has been applied, and so on? The answer would be that the viewer has to be integrated with a separate source of this information, that is, integrated with the AP-LIS, as well as possessing some key that can be used to query for it (typically the barcode value populated in the DICOM metadata, or in the case of proprietary formats, embedded in the file or folder name per some mutually agreed convention). Integrating every downstream viewer and analysis tool with the AP-LIS does not scale as well as integrating the limited number of sources, and dealing with the problem earlier, on creation or ingestion.
Another consideration is when the image is detached from its environment, such as when it is sent for outside review, consultation, or referral. The recipient is unlikely to have any means of accessing the source AP-LIS (which is undoubtedly protected from access by outsiders), so any relevant metadata will have to be sent out of band somehow (eg, in a document or spreadsheet), likely precluding its automated use and display by the receiving viewer or analysis tool. The rich metadata routinely incorporated in DICOM images for all other modalities has proven tremendously valuable for clinical image sharing, enabling routine interchange of images on off-line interchange media (such as CD and memory sticks) as well as network-based solutions. Though de-identification is required for interchange for research and clinical trial use, standards are well defined 40 and tools are readily available, which also support WSI. Indeed, clinical research involving imaging for therapeutic response assessment would be nearly impossible without the use of DICOM images and their rich metadata, and similar benefits may be anticipated for DICOM use in both clinical anatomical pathology and toxicological pathology fields, for operational, primary research as well as secondary reuse applications.
From a security perspective, though DICOM is often deployed unencrypted on small closed networks that are physically protected, generic security mechanisms, such as encryption in transit using Transport Layer Security (TLS), are available, as well as mechanisms for communicating identity for authorization.
Conversion
In the long term, the widespread adoption of the DICOM standard, or some similar standard, for all digital pathology deployments is inevitable. Operation on an industrial scale is simply not achievable using the current hodgepodge of proprietary formats, customized integrations, and error-prone manual workarounds. In the interim, however, given the proliferation of proprietary format images in archives, and the installed base of scanners that cannot or will not be upgraded to support DICOM image production, a conversion-based approach can be deployed. There are both open source and commercial providers of converters that can create DICOM from proprietary format images, often using the same libraries that support viewing of different proprietary formats. Some of these can be deployed in high-throughput situations to either migrate an entire archive of existing images, or inserted inline in a production scanning environment to automate the conversion, thus sending only DICOM images downstream.
Care should be taken when deploying such tools to assure that rich metadata is included as described. Some customization may be required to integrate primary metadata sources with the conversion tool.
Dual Personality DICOM-TIFF
It is possible to produce DICOM files that are at the same time valid TIFF files and readable by generic TIFF software. With careful attention to the structuring of the corresponding DICOM and TIFF content, a full pyramidal multi-resolution tiled image can be created that can be read by either type of recipient, without having to duplicate the entire compressed pixel data, which is shared. Since DICOM uses separate files for each resolution layer, addition of hidden layers to represent lower resolution content does result in modest expansion (about 30%), but this may be offset by the accessibility benefits. This is not necessary if only the base resolution layer is required for analysis, but is a common requirement for VM viewers. Space precludes discussion of the details, which are described elsewhere. 41
DICOMweb
Traditionally, DICOM systems have made use of a dedicated Internet Protocol (IP)-based network protocol that requires DICOM-aware software tools to implement. More recently, DICOM has added hypertext transfer protocol (HTTP)-based protocols, collectively referred to as DICOMweb, 30 to simplify implementation and use with off-the-shelf web servers, browsers, and tools. For WSI in particular, this has considerably simplified the addition of DICOM support to existing or new VM WSI viewers, since DICOMweb provides a simple uniform resource locator–based means of searching for the appropriate images, retrieving the metadata that describes their structure in JSON or XML format, and then retrieving only the necessary tiles (frames) to display on the screen, in a browser-friendly consumer-industry format like JPEG (or PNG, when lossless transport is required). DICOMweb transactions are often deployed with network encryption (TLS, ie, HTTPS), and conventional authentication and authorization mechanisms can be used (such as OpenID Connect with OAUTH2).
Color Management
It is recognized that there is considerable variability in staining procedures and preferences between laboratories and individual pathologists and that true color may be difficult to define. There have been attempts at color calibration, color normalization, and substance quantification. 42 For general use, beyond following the manufacturers’ calibration procedures, users may only be able to achieve consistency of display downstream from the scanner, as opposed to being able to achieve some nominal “truth” in color.
Consistency of color appearance in the viewer is generally considered important for diagnostic interpretation, though there is debate about this. 43 –45 Indeed, regulatory approvals for third-party viewers based on technical rather than clinical performance have dwelt on this aspect. 46 –48
From an informatics perspective, and with respect to DICOM in particular, the state of the art in achieving consistency of appearance on different displays depends on the use of International Color Consortium (ICC) profiles. The DICOM standard requires that every WSI contains an ICC for each color optical path. Gray scale channels (used for multispectral and fluorescence) do not require an ICC profile unless a color palette used for pseudo-coloring is present. In this manner, the scanner vendor is responsible for specifying the intended color space to be used (even if it is only a default standard RGB (sRGB) or similar profile), and once specified, all recipients are expected to use their platform’s color management software to map the colors to the calibrated display. Consistency of color appearance is then achievable if displays are actually calibrated, displays have a similar gamut and luminance range, and the viewing environment is similar.
Note that though some vendors perform color calibration to a default profile (like sRGB), others include sophisticated (and bulky) ICC profiles for each scan. Failure by the viewer (or analysis tool) to make use of the supplied ICC profile may produce unsatisfactory results.
The manner of application of color management before displaying the images may require careful consideration. It may be appropriate to leave color management to the display platform, for example, by embedding the ICC profile in the JPEG image sent to the web browser, or perform it manually on the client side, or have it performed on the server side, rendering the image to a well-known color space. The DICOMweb retrieve rendered image services have been extended with specific color management parameters to support these alternatives.
Note also that the ICC profile–based color management support in DICOM is not specific to WSI. The same mechanisms are used for all color imaging in DICOM, so that common libraries and tools can be used.
Annotations and Derived Information
The majority of our discussion has focused on DICOM use for encoding scanned images, but the standard also supports the encoding of derived data in the form of both derived images and image-like objects, as well as other forms of annotations, labels, tags, measurements (quantitative), and categorical (qualitative) information. These are particularly relevant for research applications and toxicological pathology, for which the production of quantitative and categorical information is important. In many cases, aggregate-derived data need to be passed to other nonimaging systems, such as databases, data management systems, and statistical software. For these, simple text-based formats (such as comma-separated value (CSV) files) or simply structured data (in XML or JSON format) are appropriate, rather than DICOM. However, when an intimate relationship needs to be maintained between the derived data and the images from which it was derived, DICOM supports the appropriate mechanisms. Though space precludes a thorough description, in summary, DICOM provides, among other mechanisms: Structured report (SR) objects, which support encoding of trees (and directed acyclic graphs) of coded, textual, numeric, and coordinate-based vector graphic data related to 3D or 2D images, such as to define lines segments or contours defining of regions of interest (ROIs) Segmentation (SEG) objects, which support one or more single-bit plane or 8-bit continuous probabilistic or fractional occupancy classification of pixels into coded segment classes (defined by anatomy, category, and type), which may also be referenced by SRs as the definition of ROIs Parametric map (PM) objects, which support numeric encoding of image-like (rectangular arrays) of scaled integer or floating point data that represent some physical quantity or model parameter and be referenced by SRs as the source of some derived measurement
These mechanisms allow encoding the output from a broad variety of annotations and categorization tasks, varying in level of granularity:
patient, case, or specimen
imaging study (set of slides for one or more specimens and sections)
series or acquisition (one or more scans of a single slide, including resolution layer, z-depth, or fluorescence channel)
entire image or subset of frames (tiles)
ROI including patches and super pixels
single points (single pixels or voxels or subpixel resolution)
Examples of specific use cases relevant to contemporary CP applications include:
encoding of human-generated truth values for ROIs representing subcellular structures, such as nuclei
recording of quantitative measurements, metrics, or scores of a particular immunohistochemical staining process, together with ROI or image references illustrating the source of the derived values
recording of categorical observations and conclusions, including classification and diagnosis
storing heat maps, identifying which pixels had greater influence on the behavior of a CP algorithm (eg, saliency maps), which can be pseudo-colored and superimposed on the WSI for display to the user
In all such cases, where generic DICOM objects previously lacked WSI-specific tiled multiresolution features, such support has been added. For example, the SEG and PM objects have both been extended with tile organization and position support.
It is recognized that the SR mechanism, though flexible, is too bulky to encode vast numbers of contours of objects (eg, the outlines of all nuclei on an entire slide). For situations in which it is not appropriate to use the pixel-wise segmentation object encoding, a new WSI-specific bulk annotation object is under development. It is expected that this will use large binary floating point arrays of contour points, with integer indices into these arrays for shapes that have a variable number of points (open polylines and closed polygons). The intent is that the architecture will be backward compatible with the existing SR structures used for smaller numbers of human-generated annotations but more compact and more quickly constructed and parsed.
All of the DICOM annotation-related objects share exactly the same information model as all the image objects in the standard. Hence, they are identified and described at the patient, study, series, and instance level with the same metadata and can be indexed and stored in the same archives and management systems as the images and listed in the same browsers. Additional effort is of course needed to interpret and apply the annotations for display, as well as to create them in the first place, and to allow editing and tracking of changes and provenance (the last being particularly important in regulated industries). Given the current lack of interoperability of annotations used in proprietary WSI viewers, workstations, and analysis tools and the relative ease of converting to and from a standard representation, it is expected that the benefits of using DICOM for annotations and other derived information will significantly enhance the value of using DICOM as a WSI image format.
Reality Check and Connectathons
It is no secret that the adoption of digital pathology in the real-world has been sluggish. The business case for digitizing slides when the glass slide still has to be made is not always obvious. The availability of CP applications to automate tedious manual tasks or to perform tasks hitherto impossible is thought to be a strong driver toward digitization.
So too has the adoption of the DICOM standard, or indeed any standard for WSI, been correspondingly sluggish. The ability of an interoperability standard to generate sufficient return on the nontrivial investment required will likely not occur until high-throughput uses cases involving heterogeneous sources and consumers dominate. It is encouraging that several scanner vendors have recently released DICOM implementations, or plan to shortly. In the interim, it is possible to lower the barrier to entry by the reuse and extension of tools for implementation, deployment, and testing that have been used for DICOM in other specialties. As previously mentioned, a DICOM WSI, though large, is just like any other DICOM object, hence can be stored in, and regurgitated from, off-the-shelf open source and commercial archives. Query and retrieval can be improved by extension of such archives with indices of WSI-relevant metadata. DICOMweb support is already widely available in such archives, and the requirements for WSI are not much different than for any other image on the server side. Customized VM viewing clients are still required, but the DICOMweb access patterns are not dissimilar to proprietary tile-based APIs, so all that is required is computation of a tile-to-frame and resolution layer image index, and existing VM viewers can be readily adapted.
On the scanner side, the conversion of proprietary formats to DICOM is relatively straightforward if the tiles are already suitably organized. Reorganizing strips or tiles that are in a different pattern can be computationally intense and may require a separate processor than is normally resident in the scanner. In a production environment, the creation of (or conversion to) DICOM and transport off the scanner must be sustainable and keep up with rate of scanning to prevent a backlog and to allow for immediate quality control (whether manual or automated). Over time it is expected that scanners will evolve from needing proprietary format conversion to creating DICOM natively, that is, to have DICOM “inside”; this was the transition that radiology scanners made some quarter of a century ago. Integration of the AP-LIS at the scanner in order to include rich identifying and descriptive metadata is crucial, for the reasons previously emphasized, but remains challenging with the current crop of products.
In order to promote the adoption of DICOM for WSI, and to demonstrate to product managers and customers alike that DICOM is appropriate and feasible, DICOM WG 26 organized a series of “Connectathons” over the last few years. Held at scientific meetings in the United States and Europe, these have been very successful in attracting participation by many vendors, both small and large. They have also served to provide a neutral forum for experimentation and testing, and the resulting feedback has resulted in numerous small and significant improvements to the standard as well as greater consensus on what choices to make when there are multiple alternatives. 49 The WSI files used during the Connectathons, even if imperfect, are publicly shared to facilitate testing. 50
It is fair to say though, that so far interoperability has not been as high a priority for many customers as other factors, and pure DICOM WSI deployments are relatively few in number. Worst case, if a desirable vendor cannot provide DICOM in today’s delivered product, a customer can contractually insist on the near future availability of a DICOM upgrade, including a conversion or migration strategy for previously acquired proprietary format images and annotations. The DICOM standard is now being required for some local and national WSI sharing projects. 51
Unfortunately, far too many suppliers and sites today are taking a path of least resistance, accepting, storing, and regurgitating all formats encountered, and passing the format reading and AP-LIS metadata integration problems downstream for the viewer or analysis tool to deal with. A better approach would be to convert to DICOM with rich metadata on ingestion, so that downstream clients have only one format to deal with and further a format detachable from the system without loss of functionality.
In the realm of toxicological pathology, both in-house tools and third-party service provider platforms need to be enhanced to support DICOM. An informal survey of service provider format support claims online would suggest that support is patchy at best, 52 –54 but some have already begun the transition, which is encouraging, though some have also expressed skepticism. 55,56
Conclusions
Adoption of DICOM for WSI is inevitable in the long term, if for no other reason than that scanner vendors will need to support it to be compatible with the enterprise imaging systems used by every other department in the hospital. For toxicologic pathology, other constraints apply, but the benefits of selecting a single interoperable standard outweigh any disadvantages. Interim measures such as the use of dual personality DICOM-TIFF images may serve to mitigate short-term software support concerns. The ability to encode quantitative results and annotations in a standard DICOM format is important for clinically deployed DP but is particularly attractive for toxicologic pathology research given regulatory record retention and reproducibility requirements and the routine use of third-party service providers.
Footnotes
Authors’ Note
This paper addresses research involving animals only in generic terms and no animals were used in the production of this material. This paper addresses research involving human subjects only in generic terms and no human subjects were used in the production of this material.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: The author is the editor of the DICOM Standard, under contract to the National Electrical Manufacturers Association (NEMA).
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article, except that the article processing charge was reimbursed by Janssen Pharmaceutica.
