Abstract
Long-term exposure to piperonyl butoxide (PBO) induces multiple nodular masses along with hepatocellular tumors in the liver of mice. The histopathological features of the nodules led to our diagnosis of nodular regenerative hepatocellular hyperplasia (NRH). However, because of the lack of data on the biological characteristics of NRH, whether this lesion is truly nonneoplastic remains unknown. In this study, the molecular characteristics of NRH were compared with those of hepatocellular adenoma (HCA) by global gene expression analysis. Six-week-old male ICR mice were fed a diet containing 6,000 ppm PBO for 43 weeks to induce NRH and HCA development. Complementary DNA microarray analysis was performed using messenger RNA extracted from NRH and HCA frozen sections collected by laser microdissection. Hierarchical cluster analysis showed that all NRH samples clustered together but were separate from the HCA cluster. Pathway analysis revealed activation of the cell cycle and Delta–Notch signaling in both lesions, but the latter was more upregulated in HCA. Downregulation of cytochrome p450 enzymes was observed in NRH, but not in HCA. These results imply that NRH differs from HCA in terms of not only morphological but also molecular characteristics.
Keywords
Long-term exposure to piperonyl butoxide (PBO) induces hepatocellular tumors in mice (Takahashi et al. 1997). Previously, we reported that multiple nodular masses in addition to hepatocellular tumors are induced in the liver of mice by treatment with PBO (Tasaki et al. 2014). Macroscopically, these masses exhibit a partially pedunculated shape and, microscopically, they are composed of nodules compressing the surrounding tissue (Tasaki et al. 2014). These findings suggest that this nodular mass may be proliferative. Almost all of the nodules are composed of various phenotypes of hepatocytes such as basophilic, eosinophilic, and vacuolated hepatocytes. However, the lobular architecture remains mostly intact despite the presence of hepatic lobes with distortion of the hepatic cords and immature central veins and portal triads. In addition to these morphological features, the occasional existence of focal necrosis in the nodules indicates that this lesion results from compensatory response due to hepatocellular damage and allows us to speculate that this lesion is regenerative. Based on these histopathological findings, we diagnosed this proliferative lesion as nodular regenerative hepatocellular hyperplasia (NRH; Tasaki et al. 2014). Thus, although morphological distinction of NRH from neoplastic lesions has been proposed, the biological characteristics that distinguish NRH from hepatocellular tumors remain unclear. Determining the molecular characteristics of NRH would help confirm the properties of nonneoplastic and sustained proliferative hepatocellular lesions.
A number of studies using surgical or genetic engineering approaches have been carried out to elucidate the molecular processes involved in liver regeneration based on global gene expression profiles (Locker et al. 2003; Kakizaki et al. 2007; Togo et al. 2004; Chiba et al. 2011). However, almost all of these studies focused on transient regenerative reactions and the early phase of liver regeneration. On the contrary, there have been few studies on chronic liver regeneration. In terms of the causal relationship between chronic hepatitis and hepatocellular tumors (Bishayee 2014), clarifying the molecular pathological differences between nonneoplastic and neoplastic lesions, which are both characterized by sustained cell proliferation, is of obvious importance.
In the present study, to elucidate the molecular differences between NRH and hepatocellular tumors, we compared the global gene expression profiles of NRH with those of hepatocellular adenomas (HCA) induced by long-term exposure of mice to PBO.
Materials and Methods
Animals
Following 1 week of acclimatization, 25 male ICR mice were randomized by body weight into the basal diet (10 animals) and PBO treatment (15 animals) groups. They were housed in polycarbonate cages with hardwood chips for bedding in a conventional animal facility, air-conditioned to 23°C ± 2°C with 55% ± 5% humidity, on a 12-hr light–dark cycle. PBO (Wako Pure Chemical Industries, Osaka, Japan) was added to the CRF-1 basal diet (Charles River Japan, Kanagawa, Japan) at a concentration of 6,000 ppm, and the prepared diet was fed to the animals ad libitum for 43 weeks. This dose was selected based on previous studies (Takahashi et al. 1997; Tasaki et al. 2014). At the end of the experiments, all animals were euthanized under deep anesthesia. At necropsy, the livers were immediately removed, and nodular mass tissues detected by macroscopic examination were collected individually. The collected nodular mass tissues were cut in half and fixed in 10% buffered formaldehyde or frozen after embedding in OCT compound (Sakura Finetek Japan, Tokyo, Japan). The protocol for this study was approved by the Animal Care and Utilization Committee of the National Institute of Health Sciences (Tokyo, Japan).
Histopathological Examination and Sample Collection
Nodular mass tissues fixed in 10% buffered formaldehyde were embedded in paraffin, sectioned, and stained with hematoxylin and eosin (H&E). The frozen masses embedded in OCT compound were also sectioned using cryostats and stained with H&E. The nodular mass tissues in these sections were diagnosed as NRH or HCA in accordance with diagnostic criteria reported previously (Tasaki et al. 2014) and International Harmonization of Nomenclature and Diagnostic Criteria for Lesions in Rats and Mice Project (Thoolen et al. 2012) and divided into four groups: the control (nonproliferative tissue in mice fed a basal diet) and nonproliferative tissue, NRH and HCA in mice treated with PBO. Each area of serial frozen sections was microdissected using a laser microdissection system (Leica Microsystems, Wetzlar, Germany). For HCA sampling, we randomly selected 3 mice from 4 mice with HCA and corrected an HCA per animal. For NRH sampling, we randomly selected 3 mice from 9 mice with NRH and corrected an NRH per animal. Nonproliferative area in PBO treated group was corrected from liver tissue surrounding NRH in the mice using NRH sampling.
RNA Isolation
Total RNA was isolated from microdissected samples using RNeasy Mini Kits (Qiagen GmbH, Hilden, Germany). The concentration and quality of the total RNA were analyzed using a UV–VIS spectrophotometer (Nanodrop ND-1000, NanoDrop Technologies, Wilmington, DE) and an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA).
Global Gene Expression Analysis by complementary DNA (cDNA) Microarray
Total RNA were labeled with cyanine 3 dye using the Quick Amp Labeling Kit (Agilent Technologies). RNA concentration, dye incorporation, and quality were analyzed using a UV–VIS spectrophotometer and the Agilent 2100 Bioanalyzer. Fluorescently labeled cRNA was hybridized to Agilent 4 × 44 K whole mouse genome expression microarrays following the manufacturer’s protocol (Agilent Technologies). Hybridized microarrays were then scanned using the Agilent Microarray Scanner (Model G2565BA, Agilent Technologies). Feature Extraction Software (Agilent Technologies) was employed for image analysis and data extraction processes. Normalization of gene expression data and filtering of probe sets by expression levels, flags, and errors were performed using GeneSpring software (Agilent Technologies). Significant differences in gene expression among the control and PBO-treated nonproliferative, NRH and HCA tissues were analyzed by analysis of variance (p < .05; multiple testing correction: Benjamini–Hochberg false discovery rate). Genes exhibiting expression changes between NRH or HCA and nonproliferative tissues in PBO-treated mice were detected by Tukey’s honestly significant difference test (p < .05, fold change >2.0) in order to clarify the differences between NRH and HCA. The identified genes were analyzed by hierarchical clustering, principal component analysis (PCA), gene ontology (GO) analysis, and single experiment analysis (SEA) using GeneSpring software.
Results
Histopathological Examination
Histopathological examination revealed nodules compressing the surrounding tissues in the liver of mice treated with PBO. The nodules were composed of hepatocytes, but the lobular architecture remained intact despite some immature central veins (Figure 1A–C). These lesions were classified as NRH according to previous reports (Tasaki et al. 2014). In addition to NRH, there were well-demarcated nodules from the surrounding liver, which were composed of hepatocytes with varying size and cellular atypia such as pleomorphic nuclei and sinusoid compression, and did not involve central veins or portal triads (Figure 1D–F); these nodules were classified as HCA. The incidences of NRH and HCA in the PBO-treated group were 9/15 and 4/15, respectively. There were no NRH and HCA in basal diet group.
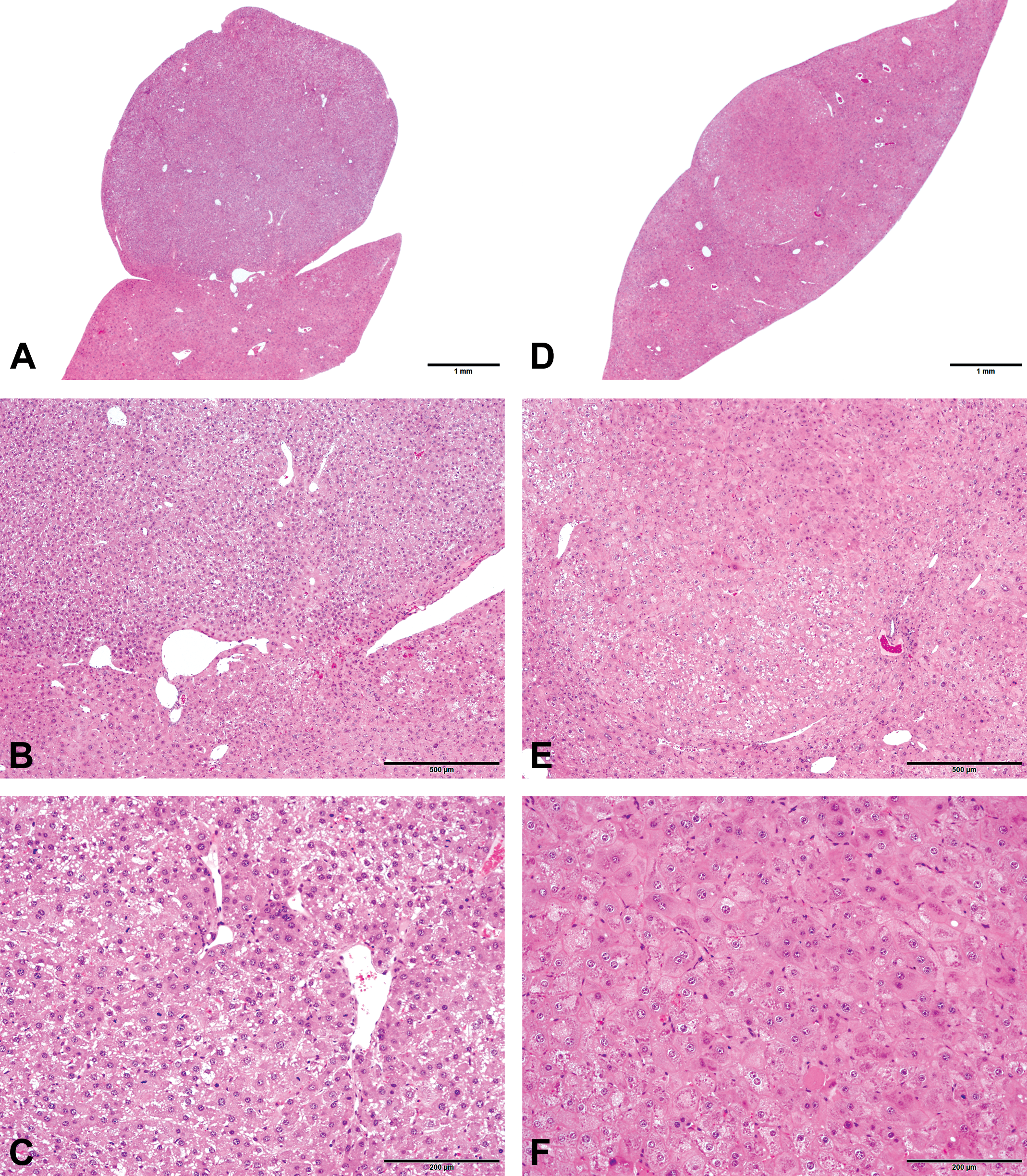
H&E-stained images of NRH (A–C). The nodules exhibited a pedunculated shape (A), compressing the surrounding tissue (B). The lobular architecture such as the central veins remained intact (C). H&E-stained images of HCA (D–F). The nodules compressed the surrounding tissues (D and E) and were composed of hepatocytes with cellular atypia without an intact lobular architecture (F). H&E = hematoxylin and eosin; NRH = nodular regenerative hepatocellular hyperplasia; HCA = hepatocellular adenoma.
Global Gene Expression Analysis Using cDNA Microarrays
Filtering, which was performed to minimize errors, generated 17,581 entities. The expression of 1,241 or 847 genes was significantly altered among all four groups or among the three PBO-treated groups, respectively. When comparing NRH or HCA with nonproliferative tissue in PBO-treated mice, 646 and 444 genes with significantly (P < .05) altered expression in the NRH and HCA, respectively, compared with the nonproliferative tissue in PBO-treated mice, and 470 (210 upregulated and 260 downregulated) or 344 (158 upregulated and 186 downregulated) genes, respectively, were significantly (P < .05) altered in expression by at least 2.0 fold. A total of 242 (124 upregulated and 118 downregulated) genes were commonly altered in both lesion types.
Hierarchical Clustering and PCA
The 847 genes that were significantly altered among the three PBO-treated groups were subjected to hierarchical cluster analysis and PCA. Hierarchical clustering results showed that all three NRH samples clustered together, separately from the HCA samples. The principal bifurcation in the clustering occurred between the NRH/HCA samples and the control/nonproliferative tissues in PBO-treated mice (Figure 2A). PCA was applied on genes significantly altered among the three PBO-treated groups to identify those genes that could distinguish between nonproliferative tissue (PBO), NRH, and HCA (Figure 2B).
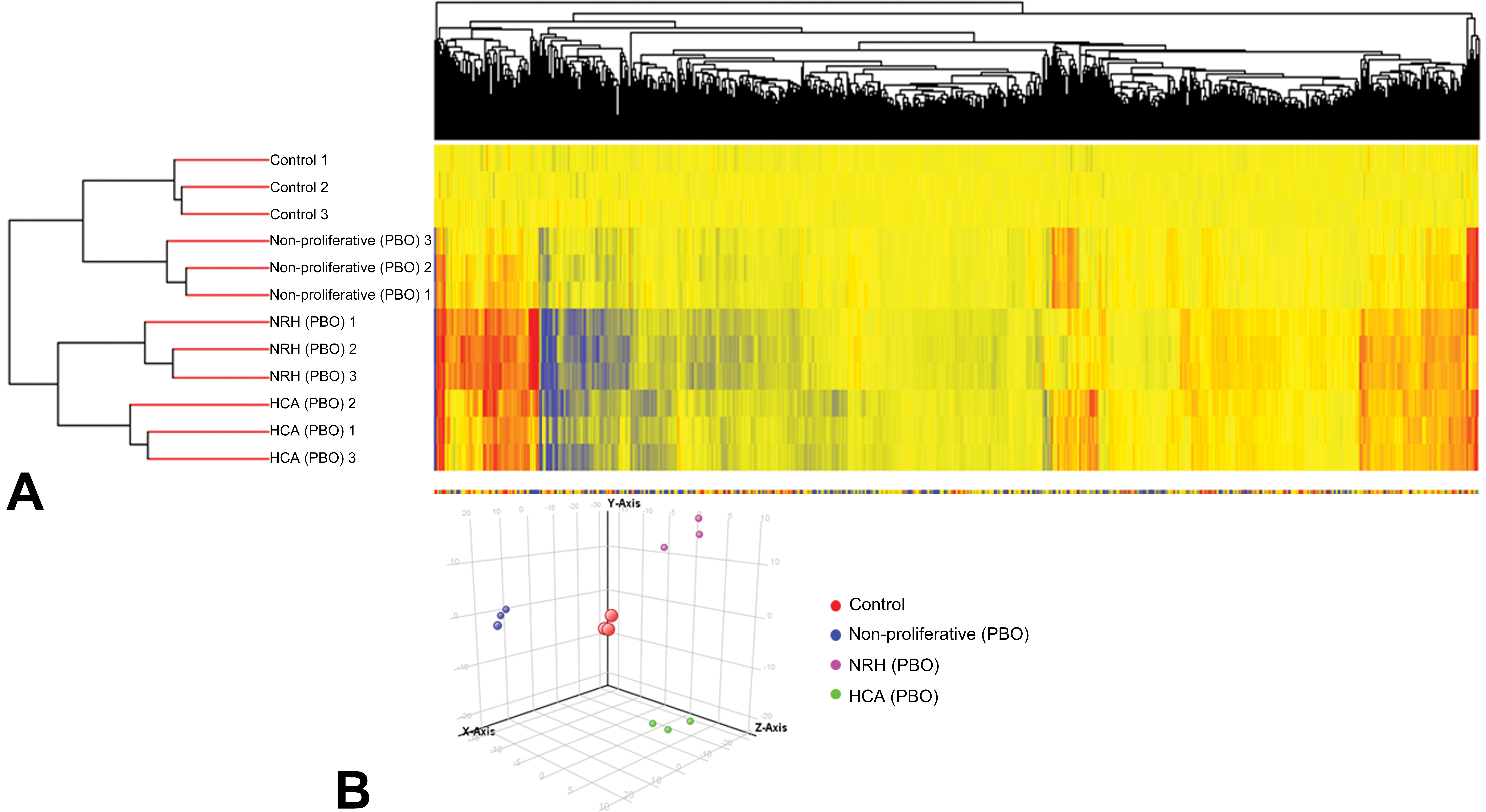
Hierarchical cluster analysis illustrated significant differences in global gene expression between NRH and HCA as well as between nonproliferative tissues and proliferative lesions including NRH and HCA (A). PCA analysis results of gene expressions that differentiate between control, nonproliferative tissue (PBO), NRH, and HCA (B). NRH = nodular regenerative hepatocellular hyperplasia; HCA = hepatocellular adenoma; PCA = principal component analysis; PBO = piperonyl butoxide.
GO Analysis
GO analysis showed that the genes upregulated in both the NRH and HCA groups were enriched mainly in cell cycle and mitotic process (Figure 3A). The genes downregulated in the NRH group were enriched in metabolic processes and catalytic activity, and those in the HCA group were associated with components of the plasma membrane (Figure 3B and C). No terms were significantly enriched among the genes downregulated in both the NRH and HCA groups or among those upregulated in either the NRH or HCA group only.
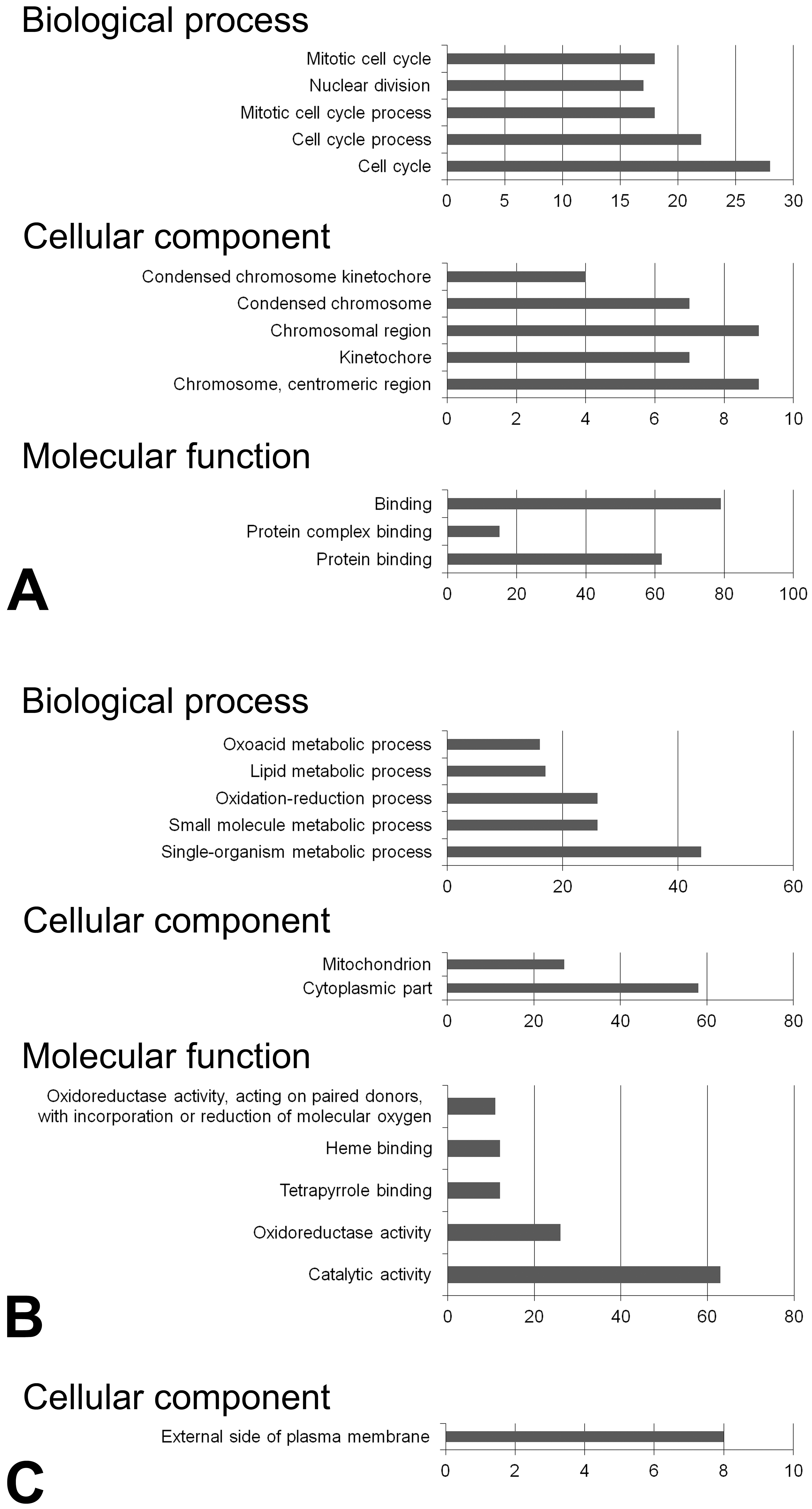
Top five most enriched GO terms among the genes upregulated in NRH and HCA (A) and among those downregulated in NRH (B) or HCA (C). The horizontal axis represents the number of enriched genes. The vertical axis represents the GO terms, increasing in p value from bottom to top. GO = gene ontology; NRH = nodular regenerative hepatocellular hyperplasia; HCA = hepatocellular adenoma.
SEA
SEA identifies matching pathways within the experiment and pathways related to the differentially expressed genes. The 646 and 444 genes with significantly altered expression in the NRH and HCA groups, respectively, compared with the nonproliferative tissue group were analyzed. The genes matched 23 or 15 pathways in the NRH or HCA groups, respectively (Table 1). Of these pathways, 12 were common to both NRH and HCA. These common pathways included cell cycle as well as protein–protein interactions. The genes associated with cell cycle were mainly upregulated (Table 2). Eleven pathways were associated with NRH only and not HCA, including oxidation by cytochrome P450, metapathway biotransformation, and peroxisome proliferator-activated receptor (PPAR) signaling. These pathways involve cytochrome p450 enzymes, including Cyp1a2, Cyp26a1, Cyp27a1, and Cyp7b1, which were downregulated in the NRH group only (Table 3). Three matching pathways were observed in the HCA group only, including the Delta–Notch signaling pathway. This pathway involves Notch4, Dll4, and Jag2, which were upregulated in the HCA group (Table 4). These genes had a tendency to be upregulated in NRH compared with nonproliferative tissue in PBO-treated mice, although the extent of upregulation was more prominent in the HCA group (Table 4). The expression level of ephrin-related genes also showed a gradual increasing tendency from nonproliferative tissue to NRH and HCA tissues (Table 5). However, signaling pathways of hepatocyte growth factor (HGF), epidermal growth factor (EGF), tumor necrosis factor (TNF)-α, and interleukin (IL)-6, which are important factors underlying acute liver regeneration, were not significantly enriched in NRH and HCA (Table 6).
Pathways Altered in NRH and HCA Identified by Single-experiment Analysis.
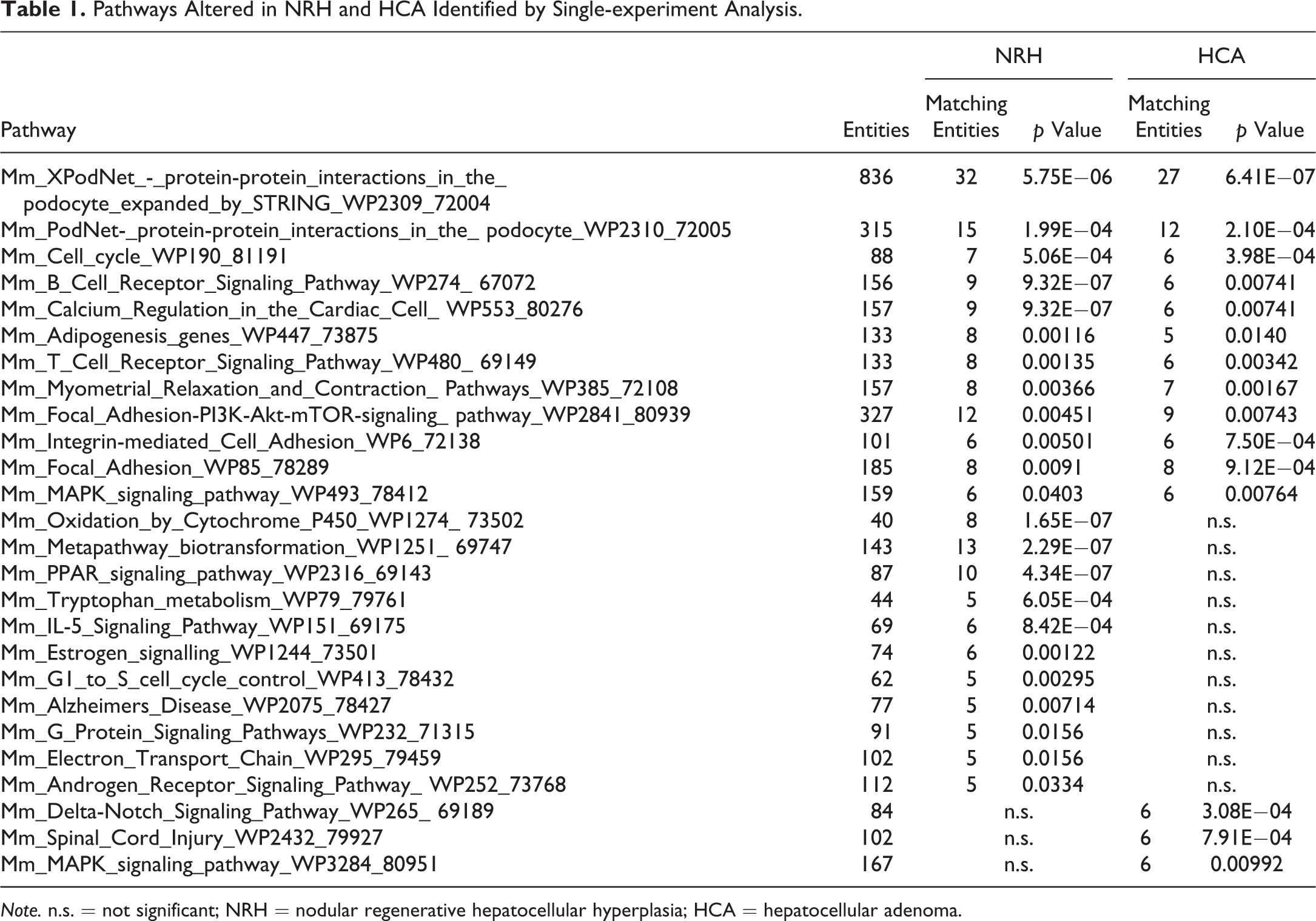
Note. n.s. = not significant; NRH = nodular regenerative hepatocellular hyperplasia; HCA = hepatocellular adenoma.
List of Matching Genes Involved in Cell Cycle Pathways.
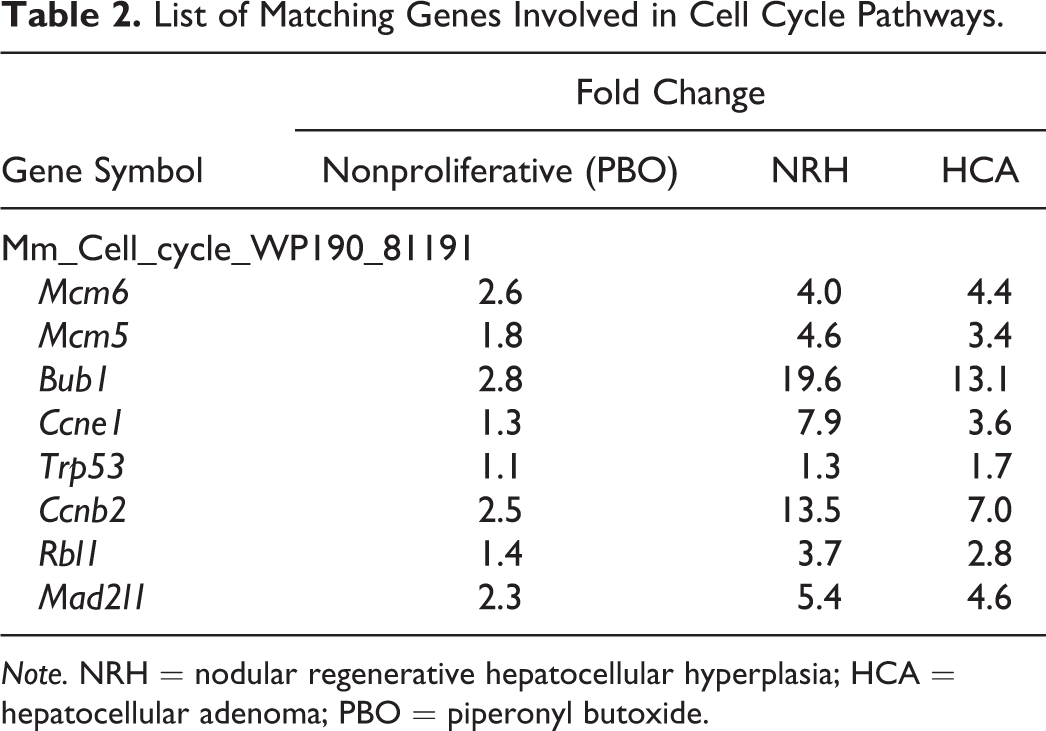

Note. NRH = nodular regenerative hepatocellular hyperplasia; HCA = hepatocellular adenoma; PBO = piperonyl butoxide.
List of Matching Genes Involved in Oxidation by Cytochrome P450 and Metapathway Biotransformation Pathways.
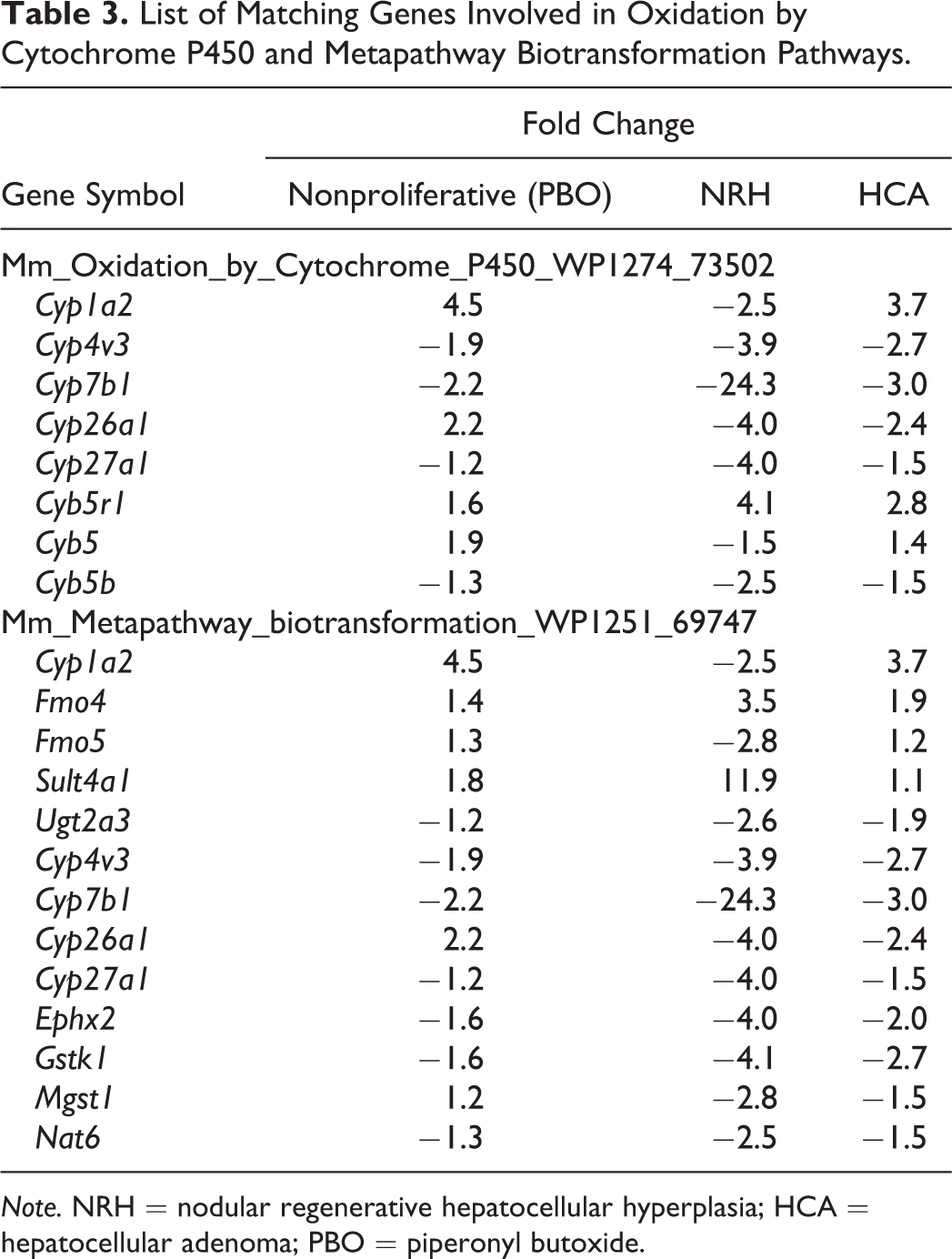
Note. NRH = nodular regenerative hepatocellular hyperplasia; HCA = hepatocellular adenoma; PBO = piperonyl butoxide.
List of Matching Genes Involved in the Delta–Notch Signaling Pathway.
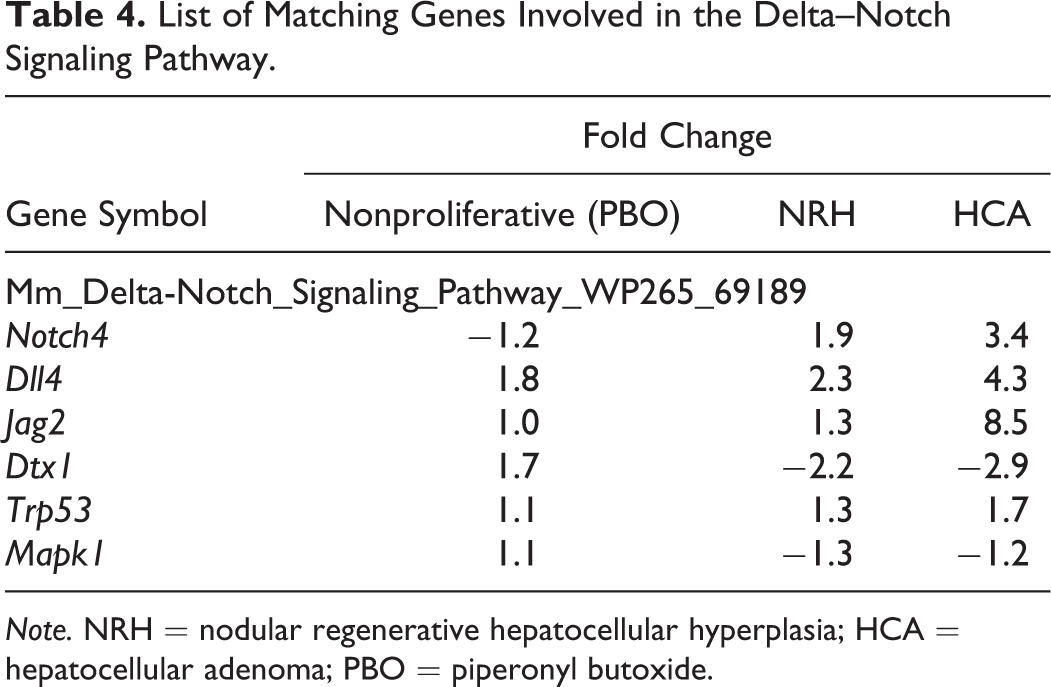
Note. NRH = nodular regenerative hepatocellular hyperplasia; HCA = hepatocellular adenoma; PBO = piperonyl butoxide.
Fold Changes in Efnb2 and Epha4 Expression in Nonproliferative, NRH, and HCA Tissues in PBO-treated Mice.
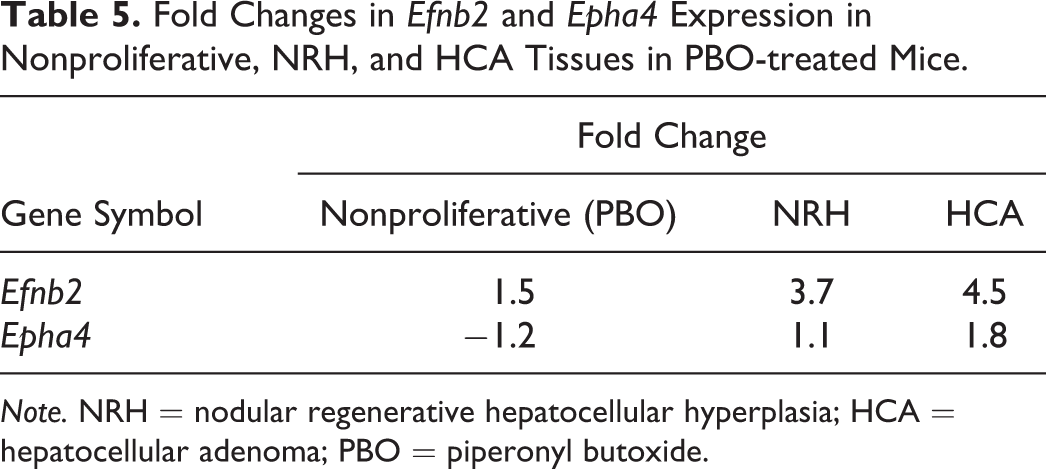

Note. NRH = nodular regenerative hepatocellular hyperplasia; HCA = hepatocellular adenoma; PBO = piperonyl butoxide.
HGF, EGFR1, TNF-α, and IL-6 Signaling Pathway Enrichment in NRH and HCA.
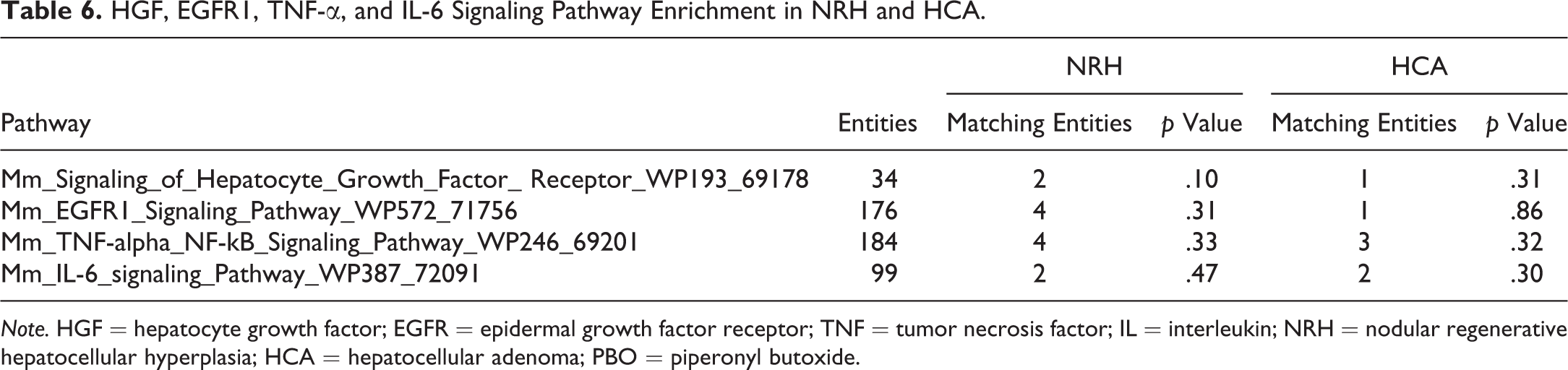
Note. HGF = hepatocyte growth factor; EGFR = epidermal growth factor receptor; TNF = tumor necrosis factor; IL = interleukin; NRH = nodular regenerative hepatocellular hyperplasia; HCA = hepatocellular adenoma; PBO = piperonyl butoxide.
Discussion
In the present study, we investigated the molecular characteristics of NRH by comparative analyses of global gene expression with that of nonproliferative or HCA tissues. Grouping by hierarchical clustering showed that NRH clustered most closely with HCA compared with the other two groups. GO analysis showed that genes involved in cell proliferation processes such as cell cycle, mitosis, and nuclear division were upregulated in both NRH and HCA. In addition, SEA revealed that genes related to cell cycle were upregulated in both NRH and HCA compared with normal tissues in PBO-treated mice. These results supported our diagnosis based on morphological characteristics that NRH is a proliferative lesion.
Pathway analysis revealed significant activation of Delta–Notch signaling in HCA. Notch proteins are a family of transmembrane proteins that interact with ligands to play a key role in the development of tissues and cells, via diverse effects on differentiation, survival, and proliferation (Andersson and Lendahl 2014; de la Pompa et al. 1997; Yamamoto, Schulze, and Bellen 2014). In mammals, the Notch signaling pathway involves four receptors (Notch1–4) and five ligands (Jagged1, Jagged2, Delta1, Delta3, and Delta4). Notch4 and Dll4, which were upregulated in HCA in the present study, are reportedly expressed specifically in the vasculature (Uyttendaele et al. 1996; Shutter et al. 2000). Constitutive expression of active Notch4 induced dramatic vascular alterations in mice (Carlson et al. 2005; Murphy et al. 2008; Miniati et al. 2010). Therefore, Notch4/Dll4 is considered important in many aspects of vascular biology (Kume 2012). Furthermore, in carcinogenesis, coordinated activation of the DLL4/Notch4 and ephrin B2 pathways plays a key role in abnormal remodeling of tumor vessels (Hainaud et al. 2006). In the present study, in addition to upregulation of Notch4 and Dll4, ephrin B2 and Eph receptor A4 tended to be upregulated in HCA. Therefore, it is likely that HCA induced by long-term exposure to PBO acquires neoplastic properties as a consequence of concomitant activation of Notch and ephrin proteins. Interestingly, Delta–Notch signaling and ephrin-related genes also had a tendency to be upregulated in NRH compared with nonproliferative tissues in PBO-treated mice, although their upregulation was more prominent in HCA. DLL4 and VEGF are known as the only genes whose haploinsufficiency results in vascular abnormalities (Gale et al. 2004). Hainaud et al. (2006) also showed that expression of Notch4, Dll4, and ephrin B2 was temporally increased with progression from a hyperplastic lesion to carcinoma. Therefore, further activation of Delta–Notch signaling might be a driving force to transform NRH into HCA. Further studies for expression, lobe specific differences, and genetic alternation involved in Notch pathway in NRH and HCA might be needed.
The characteristic alteration observed in NRH exclusively was decreased expression of Cyps. Pathway analysis revealed significant alterations in metapathway biotransformation, PPAR signaling, and oxidation by cytochrome P450 pathways in NRH only. These pathways involve Cyp1a2, Cyp7b1, Cyp26a1, and Cyp27a1, which were downregulated. Inhibition of Cyp expression in regenerative liver with high proliferative activity after partial hepatectomy in rats was reported previously (Marie et al. 1988). For many decades, the mechanisms underlying liver regeneration following resection or moderate damage have been well described (Forbes and Newsome 2016). The initial step of liver regeneration following partial hepatectomy is the activation of a combination of signaling pathways involving mitogenic growth factors, such as HGF and EGF, and nonmitogenic cytokines, such as IL-6 and TNF-α (Michalopoulos 2013). Furthermore, IL-6 and TNF-α also play important roles in promoting liver regeneration following acetaminophen-induced injury (Chiu et al. 2003; James et al. 2003). In the present study, however, significant enrichment of IL-6, TNF-α, HGF receptor, or EGF receptor 1 signaling pathways was not found in NRH, although downregulation of Cyps was observed. In contrast to acute liver regeneration models such as partial hepatectomy or chemical-induced injury, NRH was induced by chronic exposure to PBO in this study. Upstream regulation between acute and chronic regenerative stimuli may be quite different despite common Cyp downregulation. In terms of intracellular signaling, nuclear translocation of Notch intracellular cytoplasmic domain (NICD) increases within 15 min after partial hepatectomy (Köhler et al. 2004). The activation of Notch signaling contributes to regulate cell cycle in proliferating hepatocytes through NICD/Akt Akt/Hif-1α pathway in liver regeneration (Zhang et al. 2018). In addition, Notch and recombination signal-binding protein immunoglobulin kappa J regulates biliary regeneration (Lu et al. 2016). Likewise, β-catenin protein increases during the first 5 min with nucleus translocation following partial hepatectomy, indicating the contribution to induce cell proliferation and target gene expression (Monga et al. 2001). In present study, Delta–Notch signaling was upregulated in NRH. This finding indicated that there were differences in signaling pathway between acute and chronic liver regeneration. It is likely that Delta–Notch signaling is common pathway in acute and chronic liver regeneration.
Although the present result showed no significant changes in Cyp expression in HCA, aberrant CYP expression has been previously reported in hepatocellular tumors (Chen, White, and Eaton 1993; Degawa et al. 1995; Wastl et al. 1998; Ashida et al. 2018). Furthermore, aberrant Cyp expression in hepatocellular tumors has been reported to be involved in status of gene mutation such as β-catenin and Ha-ras (Loeppen et al. 2005; Hailfinger et al. 2006). These findings suggested that regulation of Cyp expression in hepatocellular tumors might be involved in not only proliferative nature but also status of gene mutation or types of carcinogen. Also considering mutation status is important to classify proliferative lesions into neoplastic or nonneoplastic lesions, further studies for genetic alternation of cancer-related genes in NRH and HCA might be needed.
In conclusion, we examined global gene expression profiles in NRH and HCA induced by long-term exposure to PBO in mice. Activation of Delta–Notch signaling was detected in these lesions, but to a greater extent in HCA, indicating that NRH is a proliferative, but not a neoplastic, lesion. In addition, NRH induced by chronic regenerative stimuli involves Cyp downregulation, as in regenerative lesions induced by acute stimuli, but the upstream regulation causing these Cyp alterations may differ between the two lesion types. The present results may help elucidate the molecular relationship between chronic liver regeneration and liver tumor development.
Footnotes
Author Contributions
Authors contributed to conception or design (ST, YY, YI, AK, KO, TU); data acquisition, analysis, or interpretation (ST, YY, YI, AK, KO, TU); drafting the manuscript (ST, YY, AK, TU); and critically revising the manuscript (ST, YI, KO, TU). All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of Conflicting Interests
The author(s) declared no potential, real, or perceived conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by JSPS KAKENHI Grant Number JP 15K07732.
