Abstract
This article provides guidelines for organ and tissue sampling adapted to porcine animal models in translational medical research. Detailed protocols for the determination of sampling locations and numbers as well as recommendations on the orientation, size, and trimming direction of samples from ∼50 different porcine organs and tissues are provided in the Supplementary Material. The proposed sampling protocols include the generation of samples suitable for subsequent qualitative and quantitative analyses, including cryohistology, paraffin, and plastic histology; immunohistochemistry; in situ hybridization; electron microscopy; and quantitative stereology as well as molecular analyses of DNA, RNA, proteins, metabolites, and electrolytes. With regard to the planned extent of sampling efforts, time, and personnel expenses, and dependent upon the scheduled analyses, different protocols are provided. These protocols are adjusted for (I) routine screenings, as used in general toxicity studies or in analyses of gene expression patterns or histopathological organ alterations, (II) advanced analyses of single organs/tissues, and (III) large-scale sampling procedures to be applied in biobank projects. Providing a robust reference for studies of porcine models, the described protocols will ensure the efficiency of sampling, the systematic recovery of high-quality samples representing the entire organ or tissue as well as the intra-/interstudy comparability and reproducibility of results.
Keywords
Introduction
Pigs are increasingly being used as disease models in translational medicine and as large animal model systems in surgery, transplantation research, and toxicologic pathology (Aigner et al. 2010; Gun and Kues 2014; Lunney 2007; Wuensch et al. 2014). The growing popularity of porcine models in biomedical research is due to several advantageous similarities between pigs and human beings that cannot be reproduced adequately in classical rodent models (Aigner et al. 2010). Due to the proximity to human anatomy, physiology and body dimensions, the comparably short generation interval (1 year), and high fertility rates, pigs are an ideal model organism for basic research and the study of disease mechanisms as well as model organisms for testing novel surgical and pharmacological therapeutic strategies (Aigner et al. 2010). Moreover, sound molecular biological methods for genetic modification of pigs are currently available, allowing for generation of tailored porcine large animal models for diverse human diseases (Aigner et al. 2010; Klymiuk et al. 2010; Klymiuk et al. 2012a; Kurome et al. 2015; Wu et al. 2013). Such genetically modified pig models have successfully been established for cystic fibrosis, diabetes mellitus, Duchenne muscular dystrophy, and other important human diseases (Aigner et al. 2010; Gun and Kues 2014; Klymiuk et al. 2013; Klymiuk et al. 2012b; Lunney 2007; Renner et al. 2013; Wolf et al. 2014). The potential of tailored pig models for testing targeted therapies is outlined by Klymiuk et al. in this issue of Toxicologic Pathology (Klymiuk et al. 2015).
Deriving optimal benefit from porcine animal models requires experimental study designs and examination protocols that warrant representative samples, reproducible results, and comparable analyses between different studies and investigators. Here, the applied mode of selection of biological samples, including the location, size, number, and orientation, is of great importance because it will affect the results of the subsequent investigations, ranging from histological examinations to molecular profiling analyses. In rodent models, the introduction and broad application of guidelines for the standardized generation of samples have greatly contributed to the quality as well as to the intra- and interstudy comparability of results (Kittel et al. 2004; Morawietz et al. 2004; Ruehl-Fehlert et al. 2003). Comparable sampling guidelines have not been established so far for porcine models.
However, rodent sampling protocols cannot be directly adapted to pig models because there are considerable differences in the anatomy and size of porcine and rodent organs/tissues. In addition to physical and anatomic features, several other important aspects have to be considered in the sampling strategies for porcine animal models. The considerably longer generation interval of pigs compared to rodents, as well as the significantly higher costs, time, and personnel efforts required for the generation of porcine models and for pig husbandry, limits the number of available animals.
Therefore, the individual animals of a respective porcine model and the samples generated from these pigs are particularly valuable, especially if genetically modified pigs and/or long-term experimental issues, such as prolonged disease courses, are to be examined. In the course of any study, additional experiments, which had not been scheduled at the beginning, might later turn out to be relevant. If suitable samples for such additional experiments are not available, they have to be generated from additional animals. Particularly, if aged pigs of genetically modified models are examined, the efforts that have to be deployed for the generation of additional animals are considerably higher than in corresponding rodent models.
In light of the steadily growing relevance of pig models in biomedical research and species-specific differences, the implementation of uniform and standardized protocols for sample generation from porcine organs and tissues applicable to a wide range of subsequent types of analyses is urgently needed to take full advantage of the translational value of porcine animal models. The proposed guidelines will allow the generation of comparable and reproducible high-quality specimens and might reduce the number of animals needed in a study by avoiding the unnecessary sacrifice of valuable animals for the repeated generation of samples (Tornqvist et al. 2014).
Sampling Guides for Porcine Organs and Tissues
In total, sampling protocols for ∼50 porcine organs and tissues (see Supplementary Material), adjusted to the expenditures and scopes of the following 3 different study types, are provided: Type I: Routine screenings for the detection of histopathological organ alterations in new porcine models, studies examining general gene expression patterns in organs/tissues, and general toxicity studies. Type II: Advanced examinations of distinct organs/tissues, with the generation of a sufficient number of backup samples, suitable for a wide range of diverse analyses, including analyses not specified at the time point of sampling. Type III: Biobank projects, requiring large-scale sampling procedures to generate high numbers of various different types of samples suitable for as many different types of analyses as possible, taken from a broad spectrum of different organs/tissues.
The respective protocols are designed to fit the demands of the industrial standards of the pharmaceutical industry and toxicologic pathology. They have been developed based on extensive experiences in pig toxicopathology, in pathomorphological characterization of numerous genetically modified pig models, and in porcine animal model biobanking (Abbott 2015; Aigner et al. 2010; Kemter et al. 2012; Klymiuk et al. 2013; Klymiuk et al. 2012a; Klymiuk et al. 2012b; Klymiuk et al. 2012c; Renner et al. 2010, 2012, 2013; Streckel et al. 2015; Wuensch et al. 2014). The proposed sampling protocols are intended as general guidelines but not as requirements for the sampling of tissues in any porcine model. The protocols can generally be applied to the organs/tissues of pigs weighing ∼10 to ∼400 kg and can be modified accordingly if smaller or younger animals are examined.
In studies of the first type (I) or in experiments that, in addition to a different main experimental task, a broad set of organs/tissues has to be examined in a routine, overview fashion by standard analyses methods, the applied sampling protocols allows for fast, uncomplicated and less elaborate sampling. Therefore, type I study sampling protocols include the collection of a limited number of samples per organ/tissue taken from deliberately defined locations, with uniform sample sizes and predefined orientations/cutting directions of a histological specimen. Type I sampling is considered adequate for the identification of qualitative histopathological changes and general organ-/tissue-specific gene expression patterns in routine studies. If organs/tissues display macroscopically evident pathological alterations, additional samples for histopathology, microbiology, virology, and molecular analyses are taken from the altered sites, as appropriate.
The list of porcine organs and tissues scheduled for routine examination in type I studies and lists of organs/tissues recommended for pathohistological examination in routine toxicity studies in rodent and nonrodent species by the Society of Toxicologic Pathology (STP) and by different public institutions and regulatory authorities are shown in Table 1. Except for rodent-specific organs, all organs and tissues regularly evaluated in other species are also examined in porcine models. Additionally, the generation of samples for histopathology and molecular analyses is scheduled for some porcine organs/tissues, which are not regularly included in established sampling guidelines for routine toxicity studies. These include organs and anatomical structures that are sampled because they are characteristically well developed in pigs, such as the bulbourethral gland and the palatine tonsil, or because they represent routinely examined predilection sites for pathological alterations in certain porcine diseases, such as the ileal papilla in swine dysentery. Sampling of other tissues and organs included in the type I study sampling list, such as adipose tissue, tendons, middle and inner ear structures, the urethra, the spermatic cord, the penis, and the prepuce, may be skipped in routine toxicity studies if no gross lesions are present at necropsy and no clinical findings support a histopathological examination. However, when genetically modified, “new” porcine models are necropsied for an initial, overall pathological examination, the rare opportunity to collect and examine these “uncommon” tissues/structures should be used.
List of Organs and Tissues Scheduled for Routine Examination in Type I Studies of Porcine Models and Organ Lists Recommended for Histopathological Examination in Routine Toxicity Studies by the STP and Various Public Institutions and Regulatory Authorities.
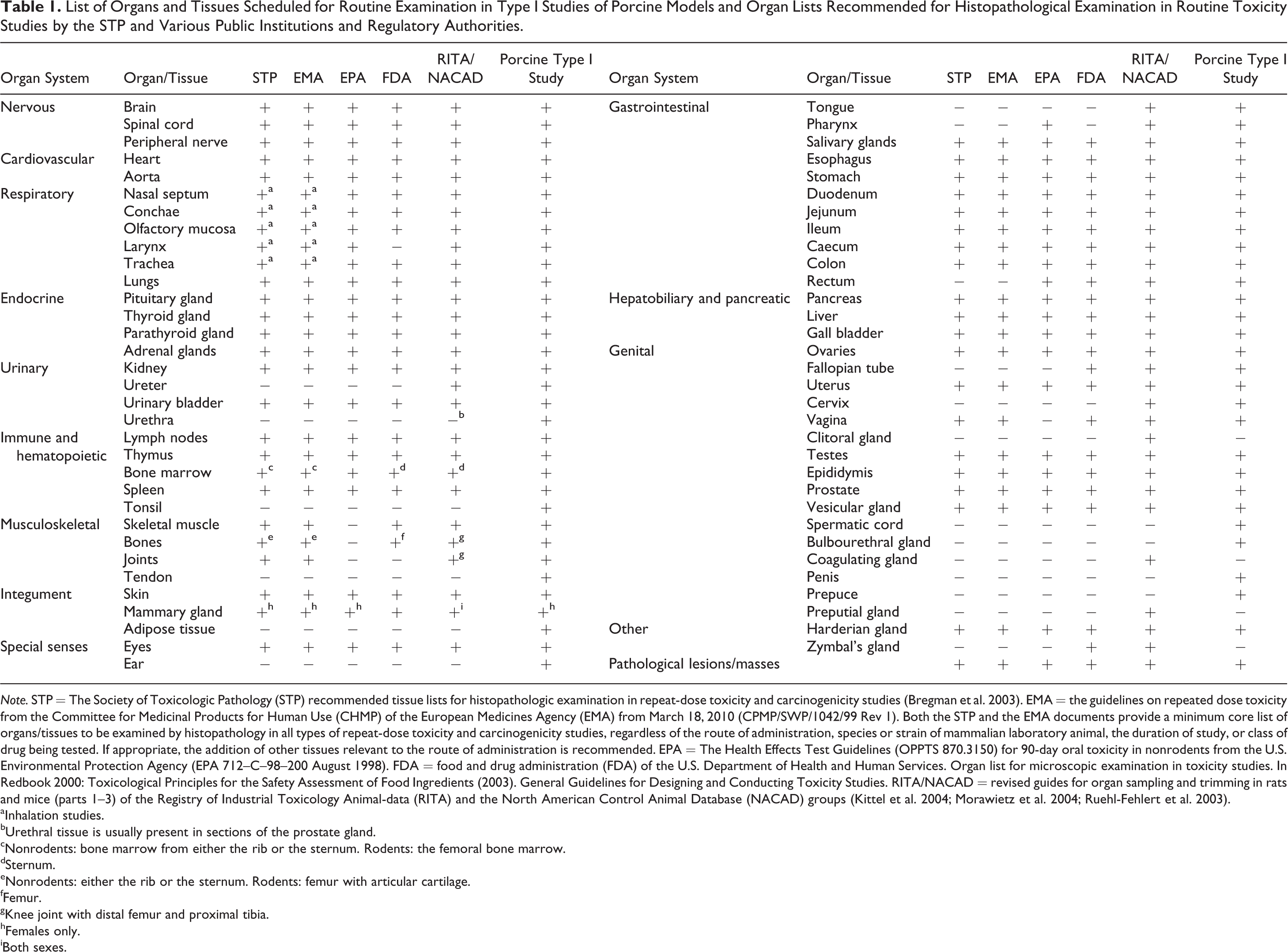
Note. STP = The Society of Toxicologic Pathology (STP) recommended tissue lists for histopathologic examination in repeat-dose toxicity and carcinogenicity studies (Bregman et al. 2003). EMA = the guidelines on repeated dose toxicity from the Committee for Medicinal Products for Human Use (CHMP) of the European Medicines Agency (EMA) from March 18, 2010 (CPMP/SWP/1042/99 Rev 1). Both the STP and the EMA documents provide a minimum core list of organs/tissues to be examined by histopathology in all types of repeat-dose toxicity and carcinogenicity studies, regardless of the route of administration, species or strain of mammalian laboratory animal, the duration of study, or class of drug being tested. If appropriate, the addition of other tissues relevant to the route of administration is recommended. EPA = The Health Effects Test Guidelines (OPPTS 870.3150) for 90-day oral toxicity in nonrodents from the U.S. Environmental Protection Agency (EPA 712–C–98–200 August 1998). FDA = food and drug administration (FDA) of the U.S. Department of Health and Human Services. Organ list for microscopic examination in toxicity studies. In Redbook 2000: Toxicological Principles for the Safety Assessment of Food Ingredients (2003). General Guidelines for Designing and Conducting Toxicity Studies. RITA/NACAD = revised guides for organ sampling and trimming in rats and mice (parts 1–3) of the Registry of Industrial Toxicology Animal-data (RITA) and the North American Control Animal Database (NACAD) groups (Kittel et al. 2004; Morawietz et al. 2004; Ruehl-Fehlert et al. 2003).
aInhalation studies.
bUrethral tissue is usually present in sections of the prostate gland.
cNonrodents: bone marrow from either the rib or the sternum. Rodents: the femoral bone marrow.
dSternum.
eNonrodents: either the rib or the sternum. Rodents: femur with articular cartilage.
fFemur.
gKnee joint with distal femur and proximal tibia.
hFemales only.
iBoth sexes.
The sampling protocols designed for type II and III studies are particularly designed for the examination of genetically modified pig models and allow for the generation of samples that are quantitatively and qualitatively suitable for a large(r) range of possible subsequent analyses. Aside from the generation of samples for the analyses actually scheduled in the experimental design of a specific study, sampling protocols for type II and III studies also provide the opportunity to generate sufficient numbers of differentially processed backup samples for additional types of analyses in advance. Furthermore, these protocols allow for the provision of a comprehensive biobank collection of redundant, adequately processed samples from any organ or tissue of potential interest (Abbott 2015). The spectrum of possible downstream analyses may include descriptive and quantitative histopathological analyses, such as histological examinations of differentially fixed samples and samples embedded in different embedding media, including paraffin or plastic resin as well as frozen-section histology, immunohistochemistry, in situ hybridization, electron microscopy, and quantitative stereological analyses. Additionally, clinical laboratory diagnostic analyses as well as DNA, RNA, and protein analyses including holistic OMICS profiling of frozen, and of otherwise preserved, sample materials might be performed.
Wherever applicable, type II and III sampling protocols schedule volume-weighted systematic random sampling procedures (Gundersen and Jensen 1987) for several organs, including the liver, spleen, kidneys, adrenal and thyroid glands, pancreas, salivary glands, thymus, and lungs. In this instance, the sampling positions and numbers of samples to be taken are based on stochastic parameters and depend on anatomic–morphological and functional properties of the respective organ/tissue. These sampling regimes ensure sampling of representative specimens, avoid systematic sampling biases, reduce experimental variability, and efficiently increase the precision of the overall experiment (Howard and Reed 2005). From each of the systematically randomly determined sampling positions, multiple samples are harvested and differentially processed according to the respective scheduled subsequent analyses. This time- and labor-saving principle may easily be adjusted to the individual sample-number and sample-type demands of a specific study. An exception to the general systematic random sampling approaches, where the entire organ is sampled, was made in organs/tissues with numerous and/or complexly structured morphologic components, such as the central nervous system or the heart, where the necessary number of sampling sites determined by systematic random sampling over the total organ would be exceedingly disproportionately high and therefore impractical. Therefore, in these organs, samples are taken from defined locations, such as defined brain areas, or distinct cardiac structures that are of interest in a specific experiment. If appropriate, the excised tissue regions of interest are then subjected to a subsequent random sampling procedure to generate representative subsamples for different downstream analyses. In practice, the workload, the personnel requirements, and the time frame and temperature conditions of a distinct sampling procedure must be compatible with the requirements of the study design and the scheduled analyses. Thus, unless the primary scientific scope of a study necessarily requires systematic random sampling of the entire organ/tissue, taking samples from defined anatomical locations appears sufficient for most qualitative histopathological and molecular–biological analyses in tissues, such as the mammary glands, adipose tissue, skin, and skeletal musculature.
For selected organs, study type II and III sampling protocols additionally present appropriate methods for the determination of the total organ (i.e., the reference compartment) volumes by Cavalieri volumetry or via the determination of the specific density of the tissue (Howard and Reed 2005; Scherle 1970). Moreover, the generation of backup specimens suitable for quantitative histomorphological analyses requiring isotropic uniform random–sectionable and vertical uniform random–sectionable samples is routinely scheduled in type II and type III study sampling protocols for several organs/tissues. These samples enable the assessment of a wide range of quantitative stereological parameters that might yet emerge to be of interest in later courses of a study and could not be adequately determined without the respective specimen (Gundersen et al. 2013; Howard and Reed 2005). Further information on the practical application of systematic random sampling, volumetry, and sample processing for quantitative stereological analyses is provided in the Supplemental Material and in the pertinent literature cited there.
Sampling Protocol Instructions and Illustrations
The sampling protocols proposed in the Supplemental Material provide detailed descriptions of applicable sampling procedures (type I–III studies) for different organs/tissues and various different downstream analyses, as illustrated by schematic drawings, macroscopic images, and histological images. The initial section presents different sampling strategies applicable to porcine organs and tissues, the determination of the specific density of porcine tissues, organ volumetry, estimation of embedding-related tissue shrinkage, and the generation of samples for quantitative stereological analyses. The sampling guides for the different organs/tissues usually cover particular information on the following topics:
Relevant pig-specific anatomic features and practical recommendations regarding the preparation of different organs/tissues.
Cutting directions and orientations of samples for histopathological examinations. The symbols used to indicate different cutting directions, sample orientations, and section planes in schematic drawings and photo images are explained in Supplemental Figure S14.
Sample numbers/locations. For routine screenings in type I studies, the anatomic location of the samples to be taken from the respective organs/tissues is indicated. The number of samples that has to be taken by systematic random sampling generally depends on the size of the organ/tissue, the size of the tissue sample pieces, the statistical properties of the investigated parameters, such as interindividual/biological and interspecimen variances, as well as the type and extent of the scheduled subsequent analyses. Therefore, the sample numbers indicated in the sampling guidelines merely represent recommended guidance levels, which should generally be sufficient for most analyzed parameters. Depending on the investigated parameter, the actual number of necessary sampling positions per organ/tissue might, thus, be lower for a specific experiment.
Individual sample sizes and specific tissue processing methods for different downstream analyses. The maximal size of an individual sample is limited by different factors, including the size of the respective organ, the number of samples to be harvested, and the specific conditions of the subsequent processing of the sample, such as the maximal penetration depth of fixatives and the size of embedding cassettes and test tubes. The dimension of individual samples designated for molecular analyses is approximately 3 × 3 × 3 mm. These samples are frozen on dry ice and then stored at −80°C until further analysis. The size of samples to be processed for histological examination is usually up to 2 × 2 × 0.5 cm for paraffin-embedded specimens, ∼1 × 1 × 0.5 cm for plastic resin (glycol methacrylate/methylmethacrylate [GMA/MMA])-embedded specimens, ≤1 × 1 × 0.5 cm for cryohistology samples, and ≤2 × 2 × 2 mm for the glutaraldehyde-fixed specimen. The standard fixatives used in the present guidelines are 10% formalin (4% neutrally buffered formaldehyde solution), fixation for 24 hr at room temperature (RT); methacarn solution (60% absolute methanol, 30% chloroform, 10% glacial acetic acid v/v), fixation for ≥24 hr at RT, samples are then rinsed in 70% ethanol and tissue processing for paraffin embedding is started in 70% ethanol; and glutaraldehyde solution (2.5% or 6.25% in Sorensen’s buffer, as indicated). For cryosectioning, tissue specimens are embedded in Tissue-Tek® O.C.T.™-blocking medium (Sakura Finetek Europe B.V., the Netherlands), frozen in liquid nitrogen-cooled isopentane and stored at −80°C until sectioning. Alternatively, the samples can be frozen in dry ice-cooled isopentane without a blocking medium to include methods such as the mass spectrometry imaging in the spectrum of possible downstream analyses (Goodwin et al. 2011). Formalin-fixed bone specimen are decalcified, using a slow-acting, acid-based decalcification solution (DC3; Labonord, Germany) for 3–30 days at RT. The embedding media for samples used for histological and quantitative histopathological analyses are paraffin, glycidyl ether (Epon) resin, and GMA/MMA (Hermanns, Liebig, and Schulz 1981). Nonstandard materials used for sampling/tissue processing are specified in the descriptions of the respective sampling protocols. For a simplified presentation, the different downstream analysis types are indicated by pictograms (Supplemental Figure S15).
A comparison of the study type I sampling protocols with established recommendations for histopathological examinations in routine toxicity studies in rodents. For each organ/tissue, the similarities and differences between the type I study sampling protocols for porcine models, standard guidelines for organ sampling and trimming in rats and mice (Kittel et al. 2004; Morawietz et al. 2004; Ruehl-Fehlert et al. 2003), and applicable “Best Practices” guides of Working Groups of the STP are discussed (Bolon et al. 2013; Haley et al. 2005; Reagan et al. 2011).
The recommended cross grid sizes for systematic random sampling procedures.
Schematic illustrations are provided where appropriate and are drawn in gray scales to preserve the recognizability of image details in black and white printouts.
Histological images. In the type I study sampling guidelines, HE-stained histological images of the indicated section planes are provided, with the relevant tissue structures indicated.
Estimates of the expected time and personnel efforts for sample collection in type II and III studies. References to the most relevant literature.
Conclusions
Because of the increasing importance of porcine animal models in biomedical research, the application of consistent and appropriate sampling procedures for tissue evaluation will increase the quality of these studies. The sampling protocols provide a basis for the generation of representative, high-quality samples using standardized procedures, which will contribute to the validity of inter- and intrastudy comparisons in porcine studies. The proposed sampling procedures and indicated sample numbers and sizes are intended as guidelines for sampling organs/tissues in pigs but not as requirements that must be met in any research project involving porcine animal models. The different sampling protocols for the type I, II, and III studies can be combined and individually adjusted to the protocol-defined objectives of a specific research project. The proposed “forward-looking” sampling strategies ensure that all necessary samples are correctly collected and processed for a given study so that they might contribute to a reduction in the number of animals needed in a study (Tornqvist et al. 2014).
Footnotes
Acknowledgments
The authors thank L. Pichl, J. Grieser, M. Handl, and A. Hinrichs for excellent technical assistance and Dr. M. Leipig, Dr. B. Keßler, and P. B. Uhl for their conceptual input. LP, JG, MH, and ML are affiliated with the Institute of Veterinary Pathology, Center for Clinical Veterinary Medicine, Ludwig-Maximilians-Universität München, Munich, Germany. AH and BK are afilliated with Minitüb GmbH, Tiefenbach, Germany. PBU is affiliated with the Institute of Animal Physiology, Department of Veterinary Sciences, Ludwig-Maximilians-Universität München, Munich, Germany.
Author Contribution
Authors contributed to conception or design (BA, SH, CB, ES, SR, FS, EW, RW, AB); data acquisition, analysis, or interpretation (BA, SH, CB, ES, SR, FS, EW, RW, AB); and drafting the manuscript (BA, EW, RW, AB). All authors critically revised the manuscript, gave final approval, and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved. AB crafted the illustrations.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: C.B.-R., E.S., S.R., A.B., R.W., and E.W. declare that they have no competing interests. This study was supported by the Federal Ministry of Education and Research (Leading-Edge Cluster m4—Personalized Medicine and Targeted Therapies; German Center for Diabetes Research). B.A. is an employee of Minitüb GmbH, Tiefenbach, Germany. F.S. is an employee of AstraZeneca RD, Mölndal, Sweden. S.H. received a PhD scholarship from the Hanns-Seidel-Stiftung e. V., Munich, Germany. The authors of this study are members of the EU COST Action BM1308 “Sharing advances on large animal models—SALAAM.”
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
