Abstract
An increasing number of studies have focused on the role of microRNAs in liver fibrosis/cirrhosis. miR-214 has recently attracted more attention as a fibrosis-related factor; however, the molecular mechanisms in hepatic fibrogenesis remain largely unknown. Here, we investigate the pathological role of miR-214 during progression of liver cirrhosis in rats. Rats were injected intraperitoneally with thioacetamide at a dose of 100 mg/kg body weight, twice a week. The liver was collected at post first injection weeks 5, 10, 15, and 20. Hepatic expression of miR-214 was analyzed by real-time polymerase chain reaction,
Introduction
Liver fibrosis is a reversible scarring response characterized by an excessive accumulation of extracellular matrix (ECM), occurring in chronic liver diseases such as viral hepatitis and alcoholic and nonalcoholic steatohepatitis (Friedman 2008). Liver fibrosis leads to cirrhosis and eventually hepatocellular carcinoma. Hepatic stellate cells (HSCs), located in the space of Disse, play a central role in liver fibrosis (Bataller and Brenner 2001). By hepatic damage and/or exposure to profibrotic cytokines such as transforming growth factor (TGF)-β1, HSCs move into injured regions and then are differentiated into myofibroblasts, the principal producer of ECM components.
microRNAs (miRNAs) are endogenous small noncoding RNAs that are involved in the posttranscriptional regulation of gene expression. miRNAs are associated with pathophysiologic events as well as fundamental cellular processes such as development, differentiation, apoptosis, and proliferation (Ambros 2004; Bartel 2004). Growing evidence shows that miRNAs are key modulatory molecules in liver diseases, including liver fibrosis (Chen 2009; Wang, Heegaard, and Orum 2012). Recent studies indicate that miR-29 modulates the regulation of liver fibrosis, and its downregulation is mediated by TGF-β in HSCs (Roderburg et al. 2011). miR-29b is downregulated in livers of patients and mice with fibrosis and suppresses activation of HSCs (Sekiya et al. 2011). Other studies indicate that expression levels of miR-199 and miR-200 families are positively correlated to the progression of liver fibrosis and that expression of fibrosis-related genes in HSCs are significantly increased by overexpression of these miRNAs (Murakami et al. 2011). Recent studies have reported differing roles of miR-214 in fibrosis: 3 in the kidney (Zarjou et al. 2011; Denby et al. 2011, 2014), 1 in the liver (Chen et al. 2014), and 1 in the heart (Aurora et al. 2012). However, the pathophysiological roles of miR-214 in liver fibrosis remain largely unknown.
Here, we report the upregulation of miR-214 during progression of liver fibrosis in a rodent model. Our results suggest that miR-214 is a negative regulator of liver fibrosis, which could be used as a potential therapeutic target for chronic liver diseases.
Materials and Methods
Animals
Six-week-old male F344/DuCrlCrlj rats were obtained from Charles River Japan (Shiga, Japan). The rats were injected intraperitoneally with thioacetamide (TAA), dissolved in saline at a dose of 100-mg/kg body weight, twice a week. The dose was determined based on our previous study (Ide et al. 2002). Four rats were euthanized under deep isoflurane anesthesia at each of post first injection weeks 5, 10, 15, and 20. Control rats received equivalent volumes of saline and were sampled at week 5. TAA and isoflurane were purchased from Wako (Osaka, Japan). Experimental procedures were conducted in agreement with the guidelines on animal care and use at Osaka Prefecture University.
Histopathology
Liver samples from the left lateral lobe were fixed in 10% neutral-buffered formalin, and samples from the right medial lobe were embedded immediately in Tissu Mount (Chiba Medical Co, Saitama, Japan) and stored at −80°C before use. Formalin-fixed tissues were embedded in paraffin, sectioned at 3–5 μm and stained with Sirius red for detecting collagen deposition. Histopathologic examination was performed by a Japanese Society of Toxicologic Pathology-certified toxicologic pathologist. The area positive for Sirius red was measured by an image analyzing software (WinRoof version 7.4; Mitani Corp., Fukui, Japan). Data are presented as percentage of positive area per total area.
miRNA Expression Arrays
Liver samples from the right medial lobe were immediately soaked in RNA later (Qiagen, Duesseldorf, Germany) and stored at −80°C. Total RNA was isolated using an SV Total RNA Isolation System (Promega, WI) according to the manufacturer’s instruction. Expression profiles of miRNAs were analyzed by miRNA microarrays, manufactured by Agilent Technologies (SurePrint Rat miRNA Array kit; CA).
Real-time Reverse Transcription-Polymerase Chain Reaction (RT-PCR)
The total RNA from the right medial lobe was reverse-transcribed by TaqMan MicroRNA Reverse Transcription kit (Life Technologies, CA). Real-time RT-PCR was performed with TaqMan Universal PCR Master Mix (Life Technologies) and TaqMan MicroRNA Assays (Life Technologies) in PikoReal real-time PCR system (Thermo Scientific, MA). Data were calculated using the ΔΔCt method. U87 or U6 ncRNA was used as an internal control.
In Situ Hybridization (ISH) for miRNA
Formalin-fixed, paraffin-embedded sections of the liver were cut at 4 μm. Before hybridization, sections were pretreated as follows: (1) 4% paraformaldehyde in phosphate buffered saline for 15 min, (2) 20 μg/ml proteinase K (Life Technologies) at 37°C for 20 min, (3) 0.16 M 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide in 1-methylimidazole buffer (0.13M 1-methylimidazole, 33nM HCl, and 300 nM NaCl) for 1 hr, and (4) 0.25% acetic anhydride in 0.1 M triethanolamine (pH 8.0) for 10 min. miR-214 probe (miRCURY-LNA detection probe; Exiqon, Vedbaek, Denmark), scramble-miR control probe (negative control; Exiqon) were diluted at 1:1,000 in hybridization buffer (50% formamide, 10 mM Tris-HCl, pH 8.0, 200 μg/ml yeast tRNA, 10% dextran sulfate, 1× Denhardt's solution, 600 mM NaCl, 0.25% sodium dodecyl sulfate [SDS], and 1 mM ethylenediaminetetraacetic acid [EDTA]) and were placed on slides. Sections were then coverslipped and incubated at 65°C overnight. After hybridization, sections were rinsed in 2× sodium saline citrate (SSC), containing 50% formamide at 60°C for 30 min and rinsed in 2×, 0.2×, and 0.1× SSC (each at 60°C for 15 min). RNA hybrids were immunostained with alkaline phosphatase-conjugated anti-digoxigenin antibody (Roche, Basel, Switzerland) and visualized using 5-bromo-4-chloro-3-indolyl-phosphate/nitro blue tetrazolium substrate (Roche).
Laser Microdissection (LMD)
Ten micromiter frozen sections of the right medial lobe were cut onto polyethylene naphthalate membrane slides using a cryostat. Sections were fixed in ethanol containing 5% acetate and stained with toluidine blue. Hepatic parenchyma and fibrotic area at week 20 were captured by LMD7000 (Leica, Wetzlar, Germany). Samples were soaked in QIAzol lysis buffer (Qiagen) and total RNA was extracted using miRNeasy Micro Kit (Qiagen). Reverse transcription and real-time RT-PCR were performed as described previously.
Cell Culture
MT-9 cells are derived from rat malignant fibrous histiocytoma and reported to have myofibroblastic phenotypes after TGF-β1 stimulation (Yamate et al. 2001, 2007). Cells were cultured in minimum essential medium (MEM, Life Technologies), containing 10% fetal bovine serum (FBS; Wako) in a humidified 5% CO2 atmosphere at 37°C.
miRNA Transfection
MT-9 cells were grown to 50% confluency and transfected with miR-214 mimic (for overexpression; Qiagen) using HilyMax reagent (Dojin Kagaku, Kumamoto, Japan) at a concentration of 0 to 10 nM. The cells were sampled 24 hr after transfection. For transfection after TGF-β1 stimulation, MT-9 cells were grown to 50% confluency in MEM with 2% FBS. Then, the cells were incubated in MEM with 2% FBS and rhTGF-β1 (10 ng/ml; #240-B, R&D Systems, Minnesota). After 24 hr, the cells were incubated in MEM with 2% FBS and miR-214 mimic at a concentration of 0 to 10 nM. The cells were sampled 12 hr or 24 hr after transfection.
Western Blot
Cells were lysed in radioimmunoprecipitation assay buffer (20 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 1 mM ethylene glycol tetraacetic acid, 1% NP-40, 0.1% deoxycholate, 0.1% SDS, 1 mM NaF, 100 mM Na3VO4, 1 mM phenylmethylsulfonyl fluoride, and protease I cocktail; Nakarai Tesque, Kyoto, Japan) for 30 min on ice. Supernatants were collected after centrifugation at 15,000 rpm at 4°C for 10 min. The protein concentration was determined by an absorption spectrometer using protein assay (Bio-Rad Laboratories, CA). The supernatants were incubated at 95°C for 5 min with SDS sample buffer (Cosmo Bio, Tokyo, Japan), containing 10% beta-mercaptoethanol. The samples were run on 10% polyacrylamide gel and transferred to polyvinylidene difluoride membrane (Bio-Rad Laboratories). The membranes were reacted with anti-α-smooth muscle actin (SMA) antibody (1:500; Dako, Gskimlostrup, Denmark) or anti-cleaved caspase-3 antibody (1:1,000; Cell Signaling, Massachusetts) overnight at 4°C. The membranes were reacted with peroxidase-conjugated secondary antibody (Histofine simple stain MAX PO; Nichirei Biosciences, Tokyo, Japan) and were incubated with ECL Prime Western Blotting Detection Reagent (GE Healthcare, NJ). Signals were detected by using LAS-3000 imaging system (GE Healthcare).
Cell Viability Assay
Relative cell number was determined by WST-8 Cell Counting Kit-8 (water-soluble tetrazolium; Dojindo, Kumamoto, Japan) according to the manufacturer's instructions. Absorbance was measured at 450 nm using a microplate reader. Experiments were done in triplicate.
Statistical Analysis
Data were presented as mean ±
Results
miR-214 Is Highly Upregulated in the Fibrotic Lesions during Cirrhosis
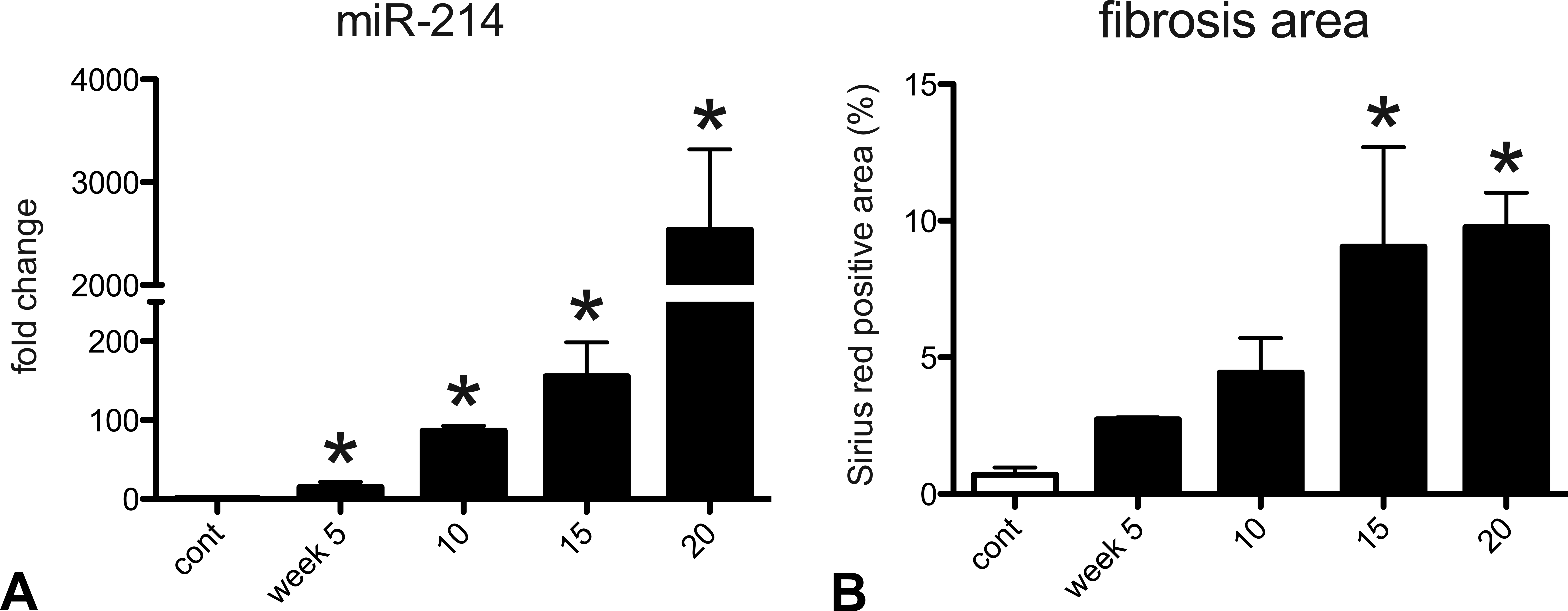
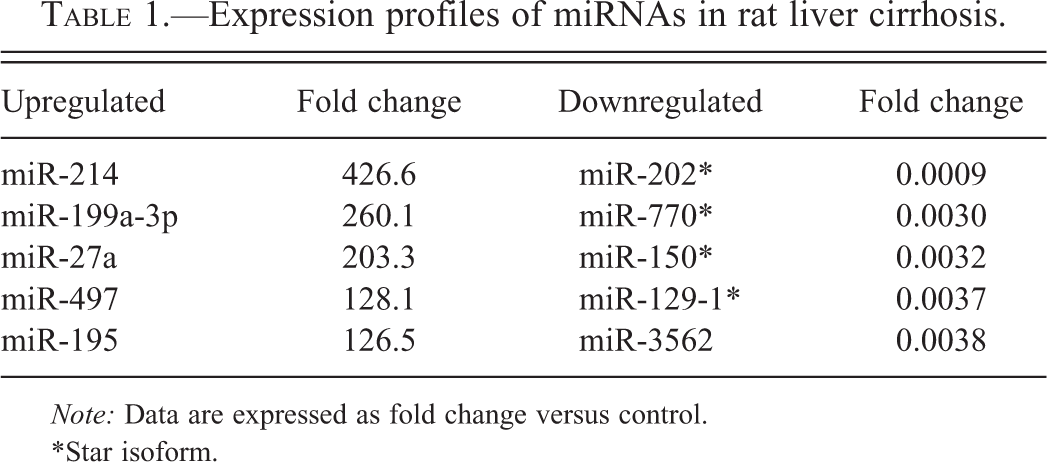
Control livers did not show any histopathological changes. At week 5, collagen deposition, stained red by Sirius red, started to appear in the perivenular areas, forming thin fibrous septa (Figure 1A; arrow). At week 10, the fibrous bridges were completely formed, thus separating the hepatic lobules into pseudolobules (Figure 1B; surrounded by arrows). At week 20, the fibrotic lesions became more thickened and micronodules of various sizes were seen (Figure 1C; surrounded by arrows). Microarray analysis identified 63 miRNAs that were significantly changed in cirrhotic liver at week 10; 39 miRNAs were upregulated (fold change > 2) while 24 were downregulated (fold change < 0.5). The top 5 of upregulated or downregulated miRNAs are listed in Table 1. Of the 63 miRNAs, the change in miR-214 expression was most prominent; miR-214 was highly upregulated (Figure 2A) in parallel with the cirrhosis progression (Figure 2B).
Collagen deposition during thioacetamide-induced liver cirrhosis. Liver sections were stained with Sirius red at weeks 5 (A), 10 (B), and 20 (C) after the first injection. Fibrous septa is indicated by arrow (A), and pseudolobules are surrounded by arrows (B, C). Bar = 100 μm. Expression pattern of miR-214 in thioacetamide -induced liver cirrhosis (A). Hepatic miR-214 expression was determined by real-time reverse transcription-polymerase chain reaction. Data are normalized by internal control (U87 ncRNA) and are expressed as fold change from control. Positive area for Sirius red (B) was calculated by an image analyzing software (WinRoof). Data are presented as percentage of positive area per total area. * Expression profiles of miRNAs in rat liver cirrhosis.
*Star isoform.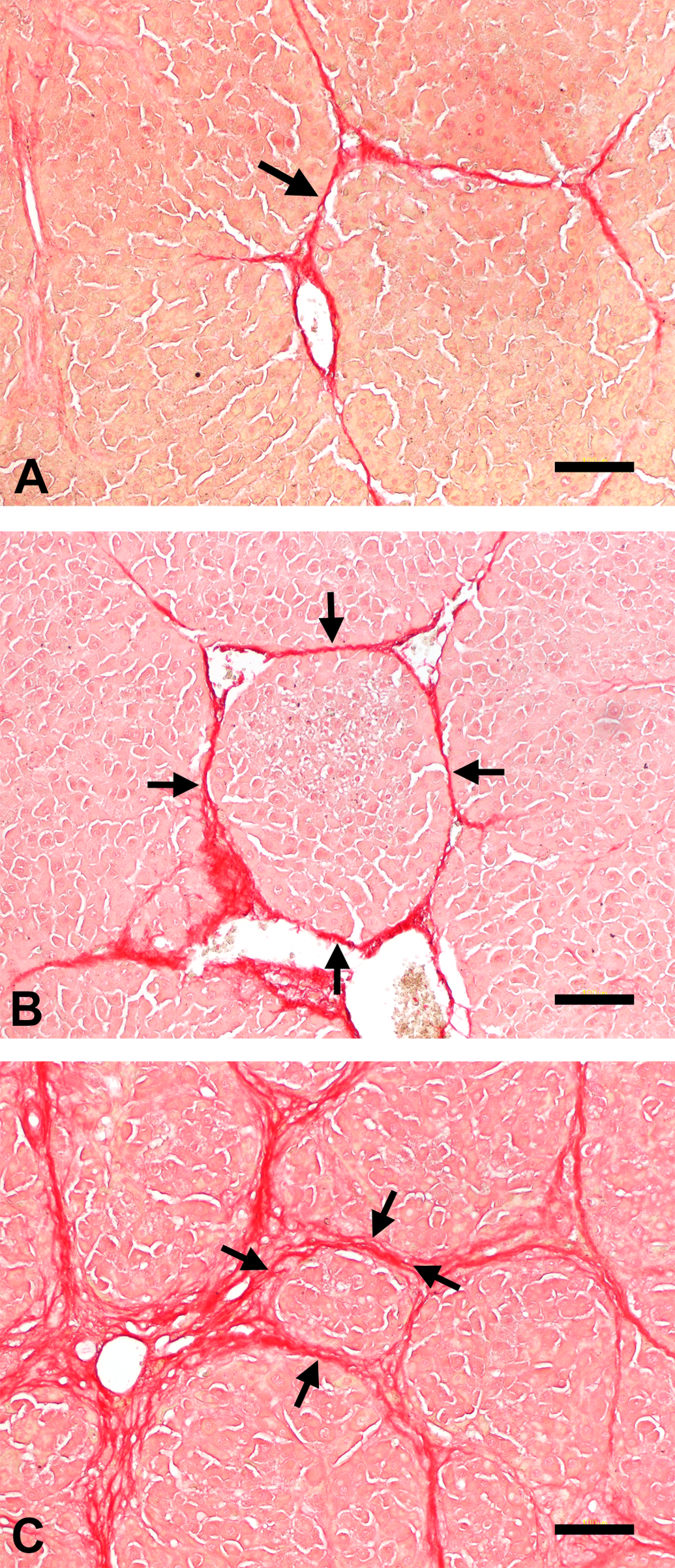
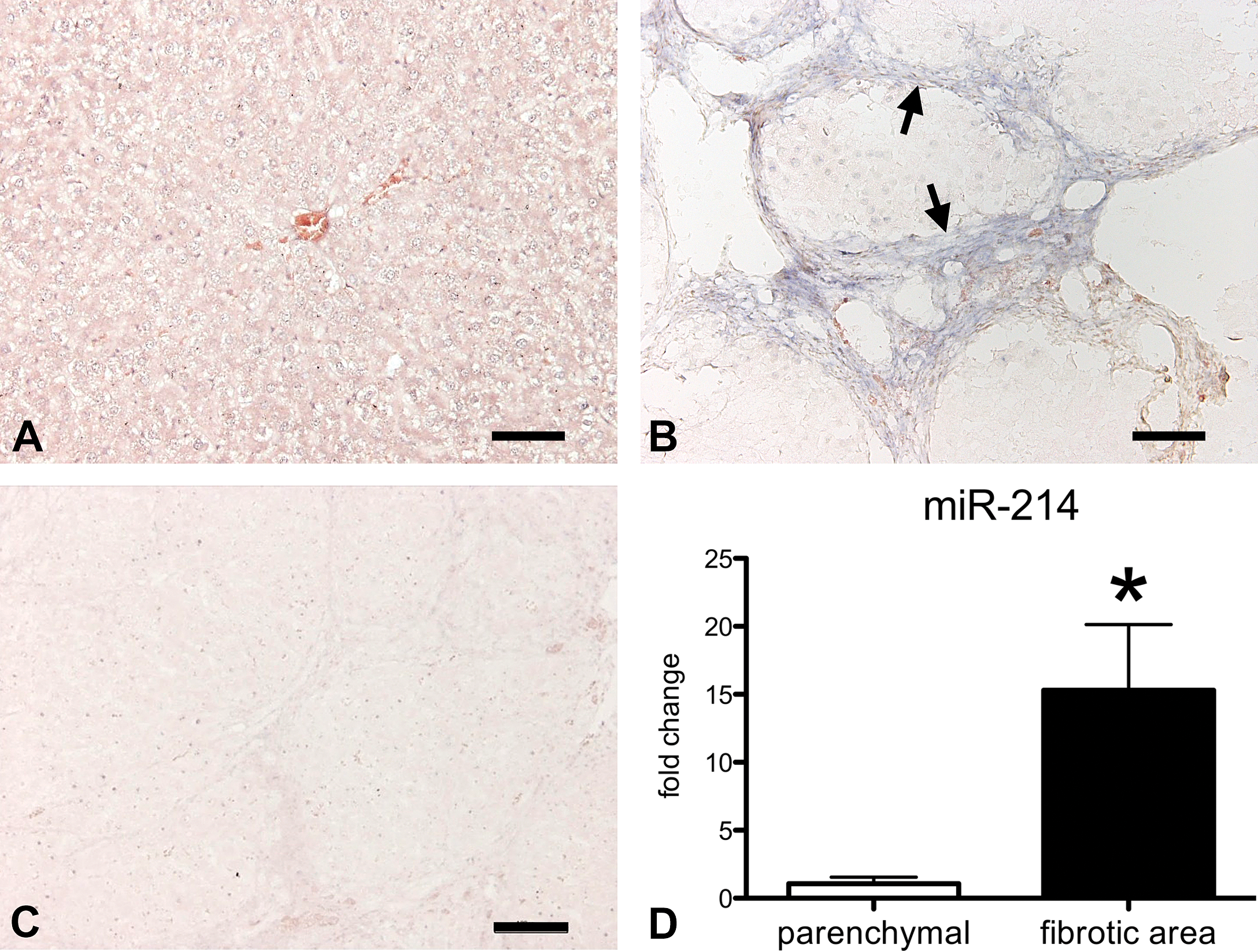
ISH was performed to identify the localization of miR-214 in liver cirrhosis. miR-214 expression was undetectable in control liver (Figure 3A), whereas it was mainly found in the fibrotic area at week 20 (Figure 3B). No signal was detected with the probe of scramble RNA (Figure 3C). To further verify the miR-214 localization, LMD followed by real-time RT-PCR was performed. miR-214 expression was 15.3-fold higher in the fibrotic area compared with that in the hepatic parenchyma at week 20 (Figure 3D). These results suggest that the miR-214 upregulation is closely associated with fibrosis and that miR-214 can be expressed by fibrogenic cells such as fibroblasts or myofibroblasts.
miR-214 Promotes Apoptosis and Inhibits Myofibroblast Differentiation under TGF-β1 Stimulation
To elucidate the pathological role of miR-214 in fibrogenesis, miR-214 was overexpressed in fibrogenic MT-9 cells by transfection. miR-214 expression was upregulated by miR-214 mimic treatment in MT-9 cells (100-fold at 10 nM; Figure 4A). Protein expression of α-SMA did not change after transfection (Figure 4B). Relative cell number did not show any significant change after transfection (Figure 4C).
Effect of miR-214 overexpression on MT-9 cells. miR-214 expression was determined by real-time reverse transcription-polymerase chain reaction (A). Data are normalized by internal control (U6 ncRNA) and are expressed as fold change from 0 nM. Protein expression of α-SMA was examined by Western blot using chemiluminescence (B). Relative cell number was determined by WST (water-soluble tetrazolium) assay (C). Data are expressed as relative absorbance.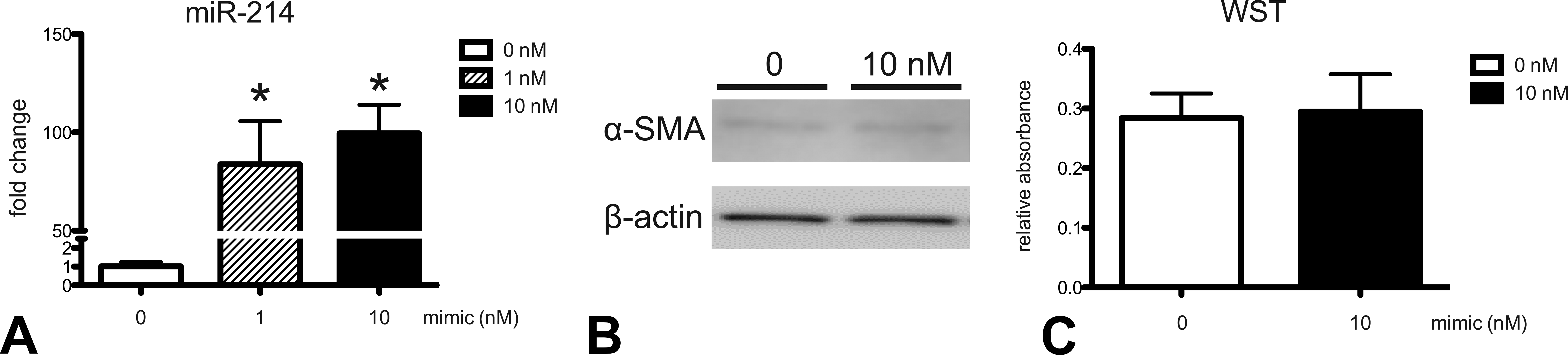
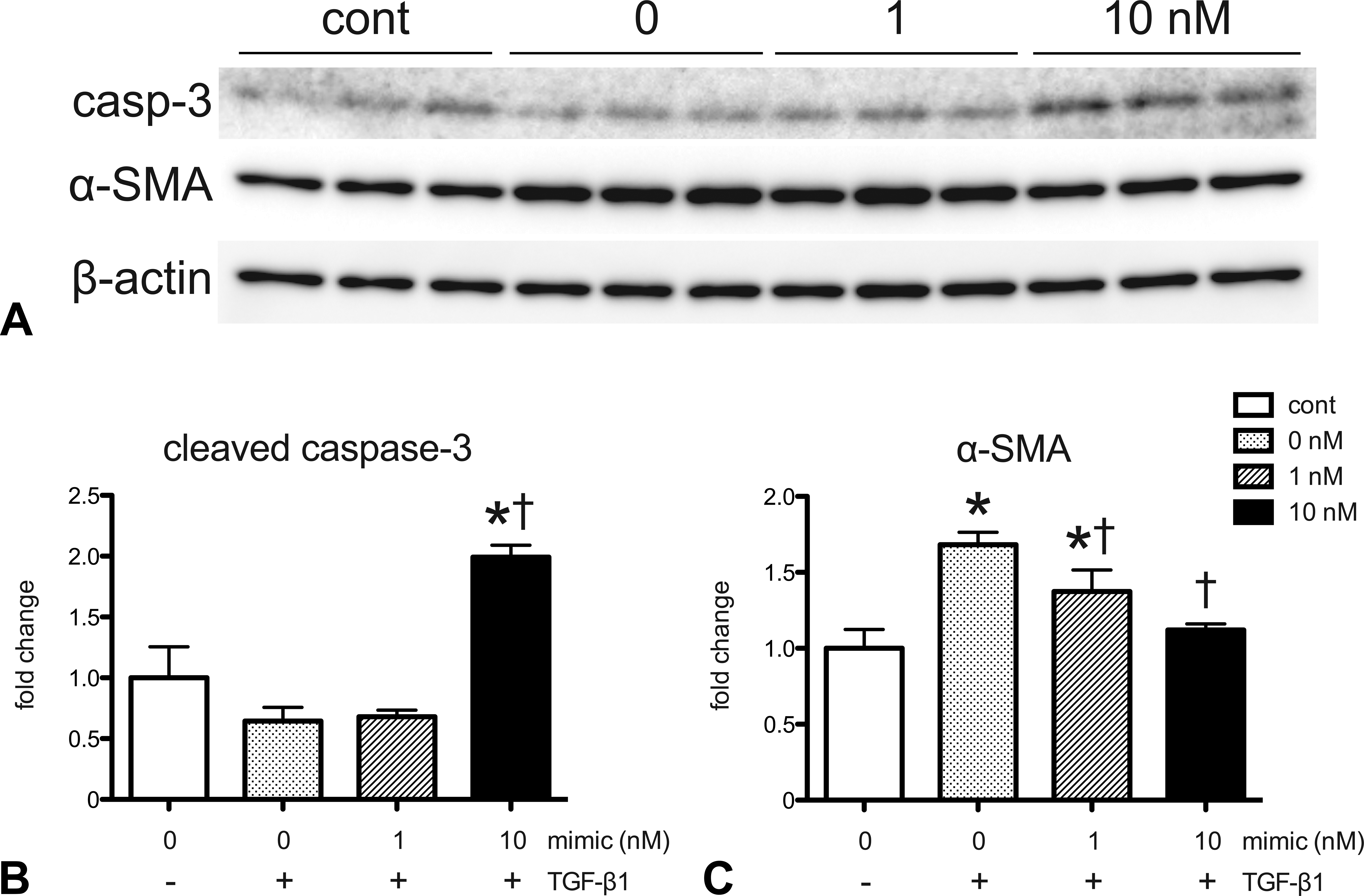
In TAA-induced liver cirrhosis, expression of TGF-β1, a central fibrogenic cytokine, is persistently increased (Ide et al. 2002; Wijesundera et al. 2014). Therefore, we investigated the role of miR-214 on MT-9 cells under TGF-β1 stimulation. miR-214 expression did not change by TGF-β1 stimulation without transfection of miR-214 mimic in MT-9 cells, whereas it was significantly increased by transfection after TGF-β1 stimulation (Figure 5A). In contrast to the single transfection (Figure 4C), relative cell number was decreased in MT-9 cells after transfection with miR-214 mimic under TGF-β1 stimulation (Figure 5B). Expression of cleaved caspase-3 was significantly increased by treatment with 10 nM of miR-214 mimic under TGF-β1 stimulation (Figure 6A and B). Expression of α-SMA was increased by TGF-β1 stimulation, which was suppressed by miR-214 overexpression (Figure 6A and C). These results suggest that miR-214 overexpression promotes apoptosis of fibrogenic cells and inhibits their differentiation into myofibroblasts under TGF-β1 stimulation.
Effect of miR-214 overexpression under transforming growth factor-β1 stimulation on MT-9 cells. miR-214 expression was determined by real-time reverse transcription-polymerase chain reaction (A). Data are normalized by internal control (U6 ncRNA) and are expressed as fold change from control. Relative cell number was determined by WST assay (B). Data are expressed as relative absorbance. * Effect of miR-214 overexpression on protein expression under transforming growth factor-β1 stimulation in MT-9 cells. Representative image of Western blot for cleaved caspase-3 and α- smooth muscle actin (SMA) (A). Expression levels of caspase-3 (B) and α-SMA (C) were determined by densitometry. Data are normalized by internal control (β-actin) and are expressed as fold change from control. *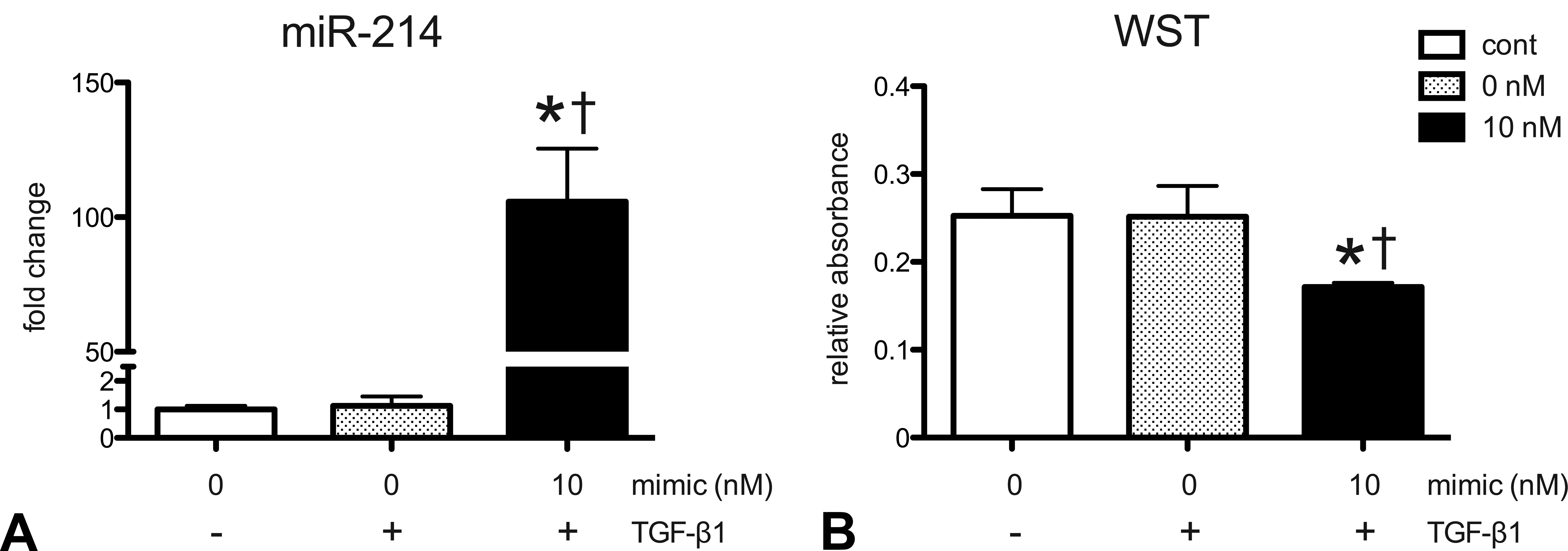
Discussion
Recently, miR-214 has attracted attention as a modulator of fibrogenesis. In the liver, miR-214 expression is decreased in the early stage of liver fibrosis in mice treated with carbon tetrachloride, TAA, and ethanol (Chen et al. 2014). In this study, miR-214 expression is markedly increased in the advanced stage of liver cirrhosis. In the kidney, expression of miR-214 is increased in unilateral ureteral obstruction (UUO)-induced fibrosis (Zarjou et al. 2011). These findings suggest that miR-214 expression may differ between organs and between stages of fibrosis/cirrhosis. In the TAA model, miR-214 was highly expressed in the fibrotic area, where activated HSCs are predominant (Ide et al. 2002; Tennakoon et al. 2015). HSCs play a central role in fibrosis through activation and differentiation into myofibroblasts (Bataller and Brenner 2001). Thus, it is considered that HSCs function in the cirrhotic liver is modulated by miR-214 expression.
Our results demonstrated that miR-214 overexpression under TGF-β1 stimulation induces apoptosis and prohibits myofibroblast differentiation in fibroblastic MT-9 cells, suggesting an anti-fibrotic role of miR-214 in liver fibrosis/cirrhosis. Apoptotic myofibroblasts are seen in the lesions of TAA-induced cirrhosis (Tennakoon et al. 2015). This hypothesis is supported by a recent study showing that miR-214 downregulates connective tissue growth factor, thereby inhibiting myofibroblast differentiation and ECM production in humans and mice (Chen et al. 2014). On the other hand, genetic deletion of miR-214 in mice significantly attenuated renal fibrosis induced by UUO (Denby et al. 2014). In the heart, genetic deletion of miR-214 causes increased apoptosis of cardiomyocytes and increased fibrosis through repression of sodium/calcium exchanger 1 (Ncx1), a key regulator of Ca2+ influx (Aurora et al. 2012). These findings suggest that the pathological role of miR-214 in fibrosis may differ between the organs.
Recent studies show that miR-214 inhibits cell apoptosis via targeting phosphatase and tensin homolog (PTEN)/Akt signaling pathway involved in cell cycle progression and prevention of apoptosis (Chang et al. 2003; Paez and Sellers 2003; Yang et al. 2008; Li et al. 2011). PTEN protein is a lipid phosphatase that antagonizes phosphoinositide 3-kinase function and consequently inhibits downstream signaling such as Akt. In this study, overexpression of miR-214 had no influence on the protein expression of PTEN in MT-9 cells (data not shown). In this TAA model, miR-214 is mainly expressed in the fibrotic area, but to some extent by hepatocytes. A recent study demonstrates upregulation of miR-214 in human embryonic stem cells–derived hepatocytes, suggesting that miR-214 may be involved in hepatocyte differentiation (Kim et al. 2011). It is possible that miR-214 may contribute to the formation of pseudolobules via modulating function of hepatocytes in cirrhosis.
In conclusion, our study demonstrated that miR-214 upregulation is associated with progression of TAA-induced liver cirrhosis and that miR-214 promotes apoptosis and inhibits myofibroblast differentiation of fibroblastic cells, suggesting its anti-fibrotic role in liver fibrosis/cirrhosis. However, despite the marked increase in miR-214, the liver cirrhosis progresses with time. Recent studies suggest that genetic deletion of miR-214 in mice attenuates renal fibrosis and exacerbates cardiac fibrosis (Aurora et al. 2012; Denby et al. 2014). Another study showed that miR-214-5p, the star sequence of miR-214 (miR-214-3p), is involved in the activation of HSCs and the progression of liver fibrosis (Iizuka et al. 2012). These findings provide a new insight into the role of miR-214 as a potential therapeutic target in liver fibrosis/cirrhosis. Further study (i.e., knockout animal,
Footnotes
Author Contribution
Authors contributed to conception or design (TI, TH); data acquisition, analysis, or interpretation (TI, TH, MH, MK, JY); drafting the manuscript (TI, TH); and critically revising the manuscript (MH, MK, JY). All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) received no financial support for the research, authorship, and/or publication of this article.
