Abstract
Introduction
GBE was nominated for study by the National Cancer Institute (NCI) as part of a review of botanicals being used as dietary supplements in the United States. GBE was selected for review because (1) GBE and its active ingredients, the flavonoids and ginkgolides, have demonstrated biological activity, (2) there is widespread exposure to GBE and it may potentially be consumed in large amounts for prolonged periods of time, (3) there are insufficient studies to evaluate for potential carcinogenicity after prolonged use, and (4) some ingredients in GBE are known
As with all herbals in the NTP testing program, selection of a test article is challenging due to the complexity of the materials and the breadth of products represented in the marketplace. GBE, in particular, is a complex mixture with many diverse constituents. The terpene lactones and flavonol glycosides are generally regarded as the active constituents responsible for the positive biological activity ascribed to GBE (van Beek and Montoro 2009), while the ginkgolic acids have been associated with cytotoxicity and mutagenicity
Hepatocellular carcinoma (HCC) in humans is an extremely complex disease, involving dysregulation of numerous growth and oncogenic pathways, chromosomal aberrations, and genetic mutations. The disease accounts for greater than 90% of liver cancer in humans and is the third leading cause of cancer mortality worldwide (Altekruse, McGlynn, and Reichman 2009). The cause and pathophysiology of human HCC is multifactorial, associated with various carcinogens, infectious agents (hepatitis B virus, hepatitis C virus), toxic agents (aflatoxin B1), genetic disease, and lifestyle factors such as chronic excessive alcohol intake (El-Serag and Rudolph 2007). HCCs occur spontaneously as a background lesion in
Materials and Methods
Ginkgo biloba Extract Test Material
The
Hepatocellular Neoplasms
Male and female B6C3F1 mice were exposed to 0, 200, 600, and 2,000 mg/kg GBE by corn-oil gavage, 5 days a week for 2 years (Table 1; NTP 2012). The statistical trend analysis from the incidences of spontaneous and GBE-treated hepatocellular tumors were evaluated statistically using the poly-3 test. Husbandry and experimental procedures were in compliance with requirements set forth by the Public Health Service’s
Incidences of select hepatic lesions in B6C3F1 mice treated with GBE by gavage in subchronic (90-day) and chronic (2-year) National Toxicology Program studies.
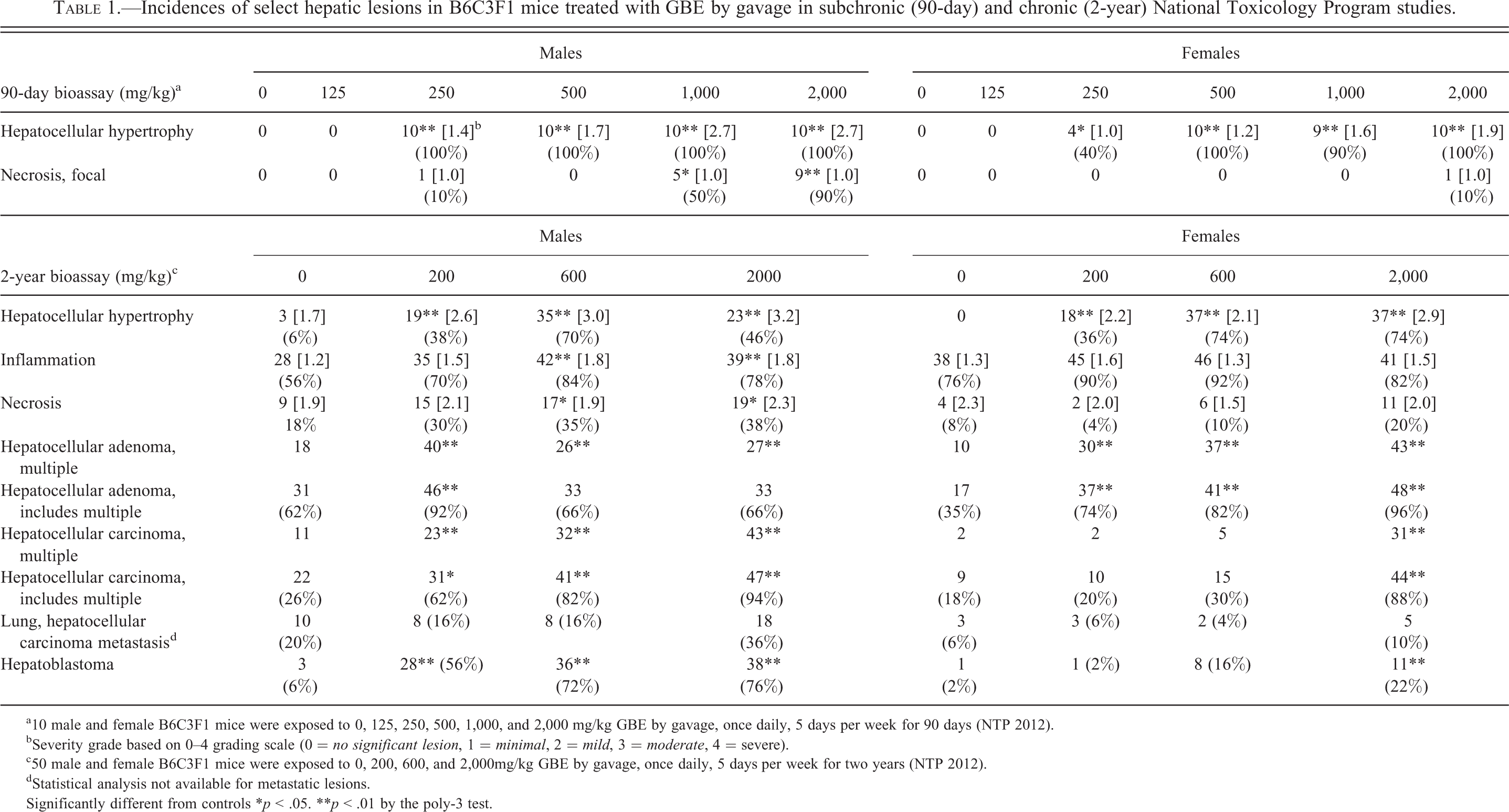
a10 male and female B6C3F1 mice were exposed to 0, 125, 250, 500, 1,000, and 2,000 mg/kg GBE by gavage, once daily, 5 days per week for 90 days (NTP 2012).
bSeverity grade based on 0–4 grading scale (0 =
c50 male and female B6C3F1 mice were exposed to 0, 200, 600, and 2,000mg/kg GBE by gavage, once daily, 5 days per week for two years (NTP 2012).
dStatistical analysis not available for metastatic lesions.
Significantly different from controls *
In Vitro Genotoxicity
GBE was assessed for mutagenic potential in a bacterial mutagenicity assay that employed
DNA/RNA Isolation, Polymerase Chain Reaction (PCR) Amplification, and Autosequencing
Sixty GBE-treated and 20 spontaneous HCCs were evaluated for mutations in exon 2 (codons 5-57) of
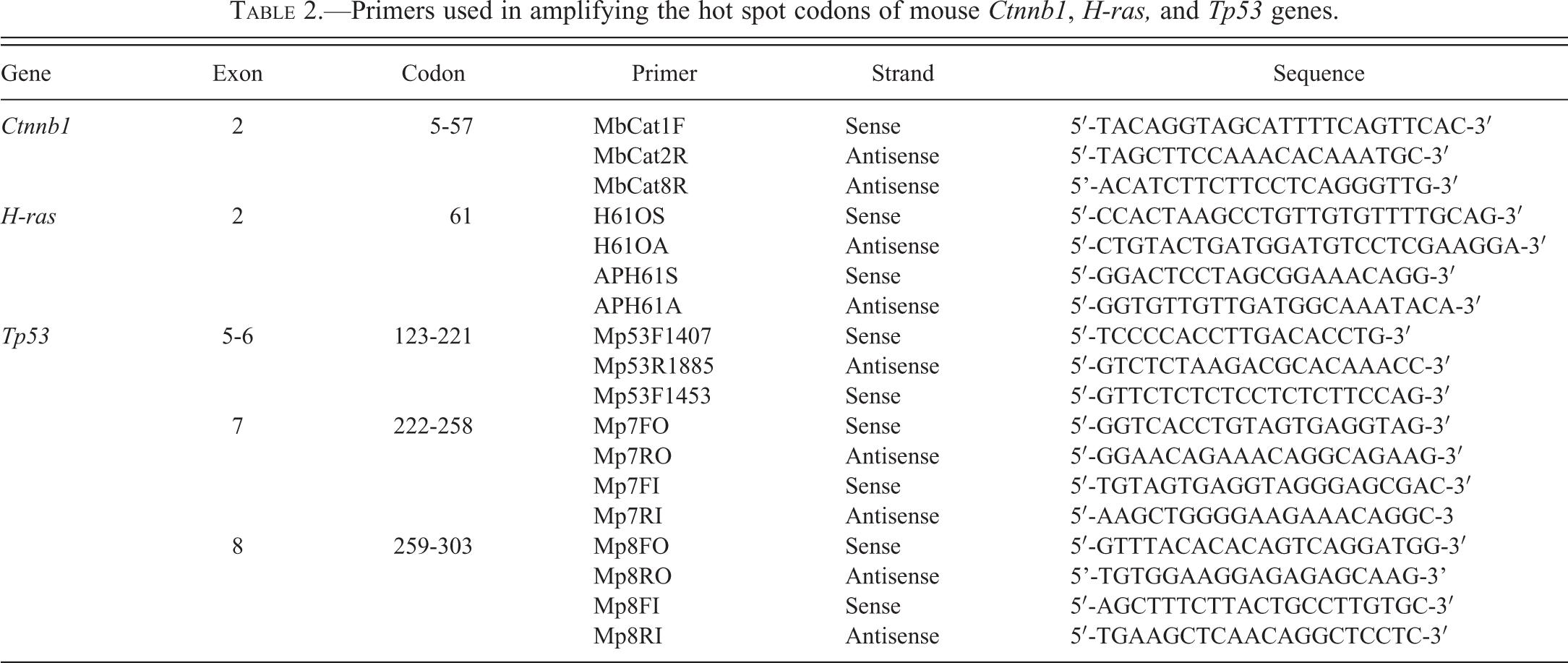
Primers used in amplifying the hot spot codons of mouse
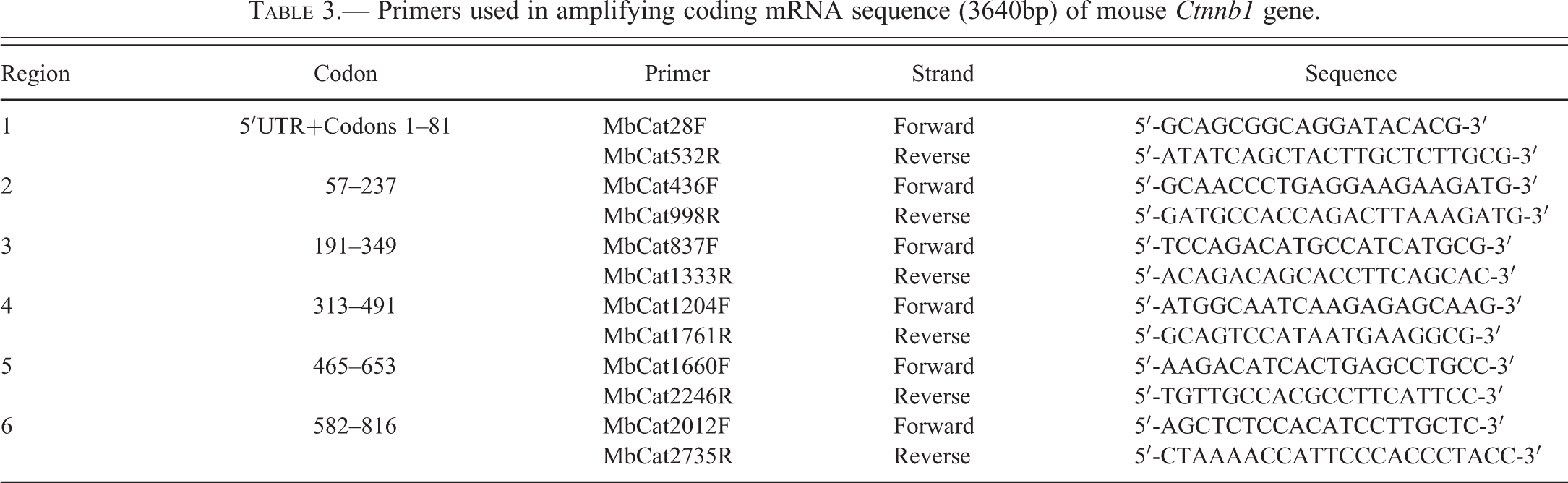
Primers used in amplifying coding mRNA sequence (3640bp) of mouse
Microarray Analysis and Quantitative Real-time (QRT)-PCR Validation
Global gene expression analysis was used to examine differential gene expression across normal, spontaneous, and GBE-treated HCCs. Affymetrix Mouse Genome 430 2.0 GeneChip arrays (Affymetrix, Santa Clara, CA) were used to assess gene expression. Amplification of 1 µg total RNA was performed as instructed in the Affymetrix One-Cycle cDNA Synthesis protocol. For each array, 15 µg of amplified biotin-cRNAs were fragmented and hybridized to each array for 16 hour at 45°C in a rotating hybridization oven using the Affymetrix Eukaryotic Target Hybridization protocol. Array slides were double stained with streptavidin/phycoerythrin and washed using the EukGE-WS2v5 protocol from the Affymetrix Fluidics Station FS450 for antibody amplification. Arrays were scanned in an Affymetrix Scanner 3000, and GeneChip® Command Console Software (AGCC; Version 1.1) was used to obtain the data. Fluorescent pixel intensity measurements were processed using the MAS5 algorithm (Hubbell, Liu, and Mei 2002), and signals were background subtracted and averaged across probes within a probeset using a mean Tukey biweight function. Control probes on the array were removed and then the gene expression data from the remaining probsets were normalized across the groups of samples using the Robust Multi-array Analysis (RMA) methodology (Irizarry et al. 2003) in the R statistical software.
QRT-PCR was performed with the ABI PRISM 7900HT Sequence Detection System (Applied Biosystems) and TaqMan MGB probes (FAM™ dye labeled). Primers and probes were obtained from Applied Biosystems Assays-on-Demand Gene Expression products. For amplification, cDNA was combined with a reaction mixture containing TaqMan universal PCR Master Mix (Applied Biosystems, Catalog No. 4304437) according to manufacturer's instructions. Technical duplicates for each sample were analyzed, and a sample without RT was included in each plate to detect genomic DNA contamination. Amplification steps included cycles at 50°C for 2 min for uracil-N-glycosylase incubation, denaturation at 95°C for 10 min, and denaturation and amplification at 95°C for 15 s, then 60°C for 30 s, for 40 cycles. Fold changes in gene expression in spontaneous and GBE-treated HCC were determined by quantification of target samples relative to vehicle control liver. The 18S RNA gene was used as the endogenous control for normalization of RNA levels. To determine this normalized value, 2−(ΔΔCt) values were compared between tumor and control samples, where the changes in crossing threshold (ΔCt)=CtTarget gene−Ct18S RNA, and ΔΔCt=ΔCtcontrol−ΔCttarget.
Bioinformatics Data Processing and Statistical Analysis
Arrays were scanned in an Affymetrix Scanner 3000, and data were obtained using the GeneChip® Command Console Software (AGCC; Version 1.1), and Partek Genomics Suite (6.4) was used to perform principal component analysis (PCA) on the normalized data and to generate heat maps to compare samples as previously described (Hoenerhoff et al. 2011). The methodology described by Guo, Sarkar, and Peddada (2010) was used to control the false discovery rate at
Incidence of
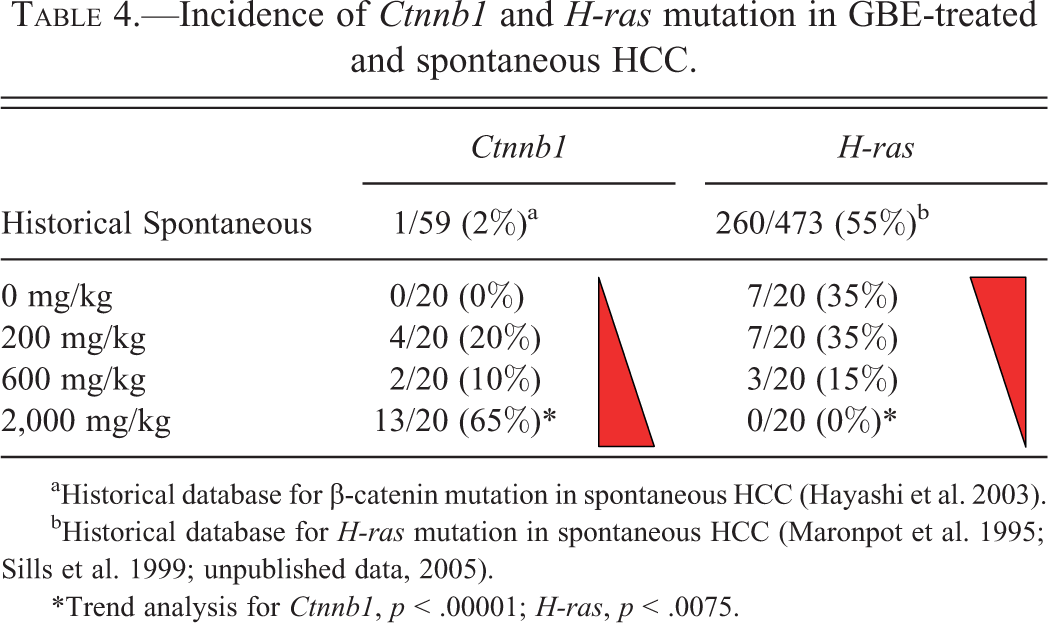
aHistorical database for β-catenin mutation in spontaneous HCC (Hayashi et al. 2003).
bHistorical database for
*Trend analysis for
Immunohistochemistry and Western Blot Analysis
For immunohistochemical analysis, following deparaffinization, rehydration, antigen retrieval, and quenching of endogenous peroxidase activity, polyclonal or monoclonal primary antibodies were applied. These included goat polyclonal CTNNB1 (sc-1496, 1:50; Santa Cruz Biotechnology, Santa Cruz CA), mouse monoclonal E-cadherin (CDH1; C20820, 1:50; Transduction Labs, Lexington KY), and rabbit polyclonal Glutamine synthetase (GLUL; ab49873, 1:10,000; Abcam, Cambridge MA) antibodies. Negative controls were obtained by substitution of the primary antibody with normal serum of the species that the secondary antibody was made in. Following washing, the labeled streptavidin ABC technique was employed for detection of primary antibody binding. For visualization, 3,3-diaminobenzidine (DAB) was applied and counterstained with Mayer’s hematoxylin. The sections were dehydrated through graded alcohols, immersed in xylene, and mounted with coverslips. Detailed immunohistochemistry protocols can be found at http://www.niehs.nih.gov/research/atniehs/labs/lep/path-support/core-support/immuno/protocols/. For Western blotting, spontaneous and GBE-treated tumor and normal liver lysates were prepared as previously described (Hoenerhoff et al. 2011), and the following antibodies were used: N-terminus goat polyclonal CTNNB1 (C-18; sc-1496, 1:100; Santa Cruz Biotechnology, Santa Cruz, CA); C-terminus rabbit polyclonal CTNNB1 (E247; ab32572, 1:2500; Abcam, Cambridge MA); rabbit polyclonal Calpain 1 (H-240; sc-30064, 1:500; Santa Cruz Biotechnology) and Calpain 2 (ab39165, 1:250; Abcam); and rabbit polyclonal CYP2B (generous gift from Tatsuya Sueyoshi, Pharmacogenetics Group, NIEHS, 1:50,000).
CTNNB1 Immunoprecipitation and Protein Microcharacterization
Tissue homogenates were prepared in radioimmunoprecipitation assay (RIPA) buffer containing protease and phosphatase inhibitors. The supernatants obtained after centrifuging the homogenates at 12,000 × g for 15 min at 4°C were used for analysis. Protein concentration was measured using the bicinchoninic acid (BCA) assay (Pierce, Rockford, IL). Samples containing 500 µg of protein were incubated with 10 µl of agarose-conjugated CTNNB1 antibody (sc-1496 AC; Santa Cruz Biotechnology, Santa Cruz, CA) at 4°C with constant rotation. The samples were centrifuged at 12,000 rpm for 2 min at 4°C, the supernatant was removed, and the agarose-protein immune complexes were washed 3 times in PBS. The pellet was resuspended in 50 µl of 2× loading buffer, and the samples were fractionated by SDS-PAGE. After electrophoresis, the gel was cut in half. One half of the gel was stained with SimplyBlue SafeStain (Invitrogen, Carlsbad, CA). Briefly, the gel was rinsed 3 times for 5 min each in deionized water and stained in SimplyBlue SafeStain for 1 hr at room temperature with gentle shaking. Following staining, the gel was washed in water for 1 hr and submitted for protein microcharacterization. The other half of the gel was transferred to a nitrocellulose membrane (Invitrogen, Carlsbad, CA), and nonspecific binding sites were blocked using 10% nonfat dry milk. The membranes were incubated with an antibody to CTNNB1 (1:100 dilution; sc-1496; Santa Cruz Biotechnology, Santa Cruz, CA) for 1 hr at room temperature with gentle agitation. Immunoreactivity was detected using horseradish peroxidase (HRP) conjugated IgG (1:5,000 dilution; sc-2020; Santa Cruz Biotechnology, Santa Cruz, CA), ECL Plus Western Blotting reagents (Amersham, Piscataway, NJ), and film autoradiography.
For protein microcharacterization, the area of interest (as determined by Western blot analysis) was digested with trypsin (Promega, Madison, WI) for 8 hr using a ProGest robotic digester (Digilab, Holliston, MA). Briefly, minced gel bands were incubated 2 times in 25 mM ammonium bicarbonate, and 50% (v/v) acetonitrile for a total of 30min. The gel was dehydrated in acetonitrile for 20 min, dried under a nitrogen stream, and incubated with 250 ng of trypsin for 8 hr at 37°C. The resulting peptides were extracted using 5% (v/v) formic acid, 50% (v/v) acetonitrile and lyophilized. The lyophilized samples were resuspended in 0.1% formic acid, loaded onto an Agilent C18 chip, and washed with 5% acetonitrile, 0.1% formic acid. NanoLC-ESI-MS/MS analysis was performed using the Agilent 1100 nanoLC system and the Agilent XCT Ultra ion trap mass spectrometer with a mass range of 200 to 2,200 m/z, an ionization potential of 2.1 kV, an Ion Charge Control (ICC) smart target of 10,000 or 200 ms of accumulation, and 1.0 volt fragmentation amplitude. Peptides were eluted from the Agilent chip by applying a linear gradient of acetonitrile (5 to 95%) and 0.1% formic acid. MS/MS data were acquired using the data extractor feature of the SpectrumMill software (Agilent, Santa Clara, CA) with an ion size limit 300 to 5,000 Da and retention time of 10 to 60 min. The resulting spectra containing sequence tag information of greater than 2 residues were submitted for database searching in the NCBI database with the following restrictions: trypsin specificity with one missed cleavage, a precursor ion mass tolerance of 2Da, a product ion mass tolerance of 1.0 Da, variable methionine oxidation, and a minimum matched spectral intensity of 70%. Proteins identified with an MS/MS search score greater than 25 were tabulated. The false positive rate was 0% as determined by a reverse sequence database search and manual sequence validation.
Results
Oral exposure to GBE results in a dose-dependent increase in the incidence of hepatocellular necrosis and centrilobular hypertrophy
Exposure of male and female B6C3F1 mice to 125, 250, 500, 1,000, and 2,000 mg/kg GBE over a 90-day period resulted in a treatment related increase in hepatocellular hypertrophy and focal hepatocellular necrosis (Table 1 and Figure 1A). Hepatocyte hypertrophy was characterized by centrilobular to midzonal enlargement of hepatocytes, with increased amounts of homogenous to finely granular eosinophilic to amphophilic cytoplasm and enlarged nuclei. Focal necrosis (Figure 1A) was characterized by loss of hepatocyte architecture, hypereosinophilia, and nuclear pyknosis, karyorrhexis, and karyolysis. In the 2-year bioassay, treatment with 200, 600, and 2,000 mg/kg GBE resulted in a dose-dependent increased incidence and severity of centrilobular hepatocellular hypertrophy in male and female mice. These lesions were also accompanied by a dose-related increase in hepatocellular necrosis in all treated males and mid- and high-dose females, and inflammatory cell infiltrates composed predominantly of lymphocytes, plasma cells, neutrophils, and rare macrophages (Figure 1B and Table 1). These findings indicate that oral exposure to GBE at the indicated doses results in hepatocellular hypertrophy and damage that increases in severity over time with continued exposure.
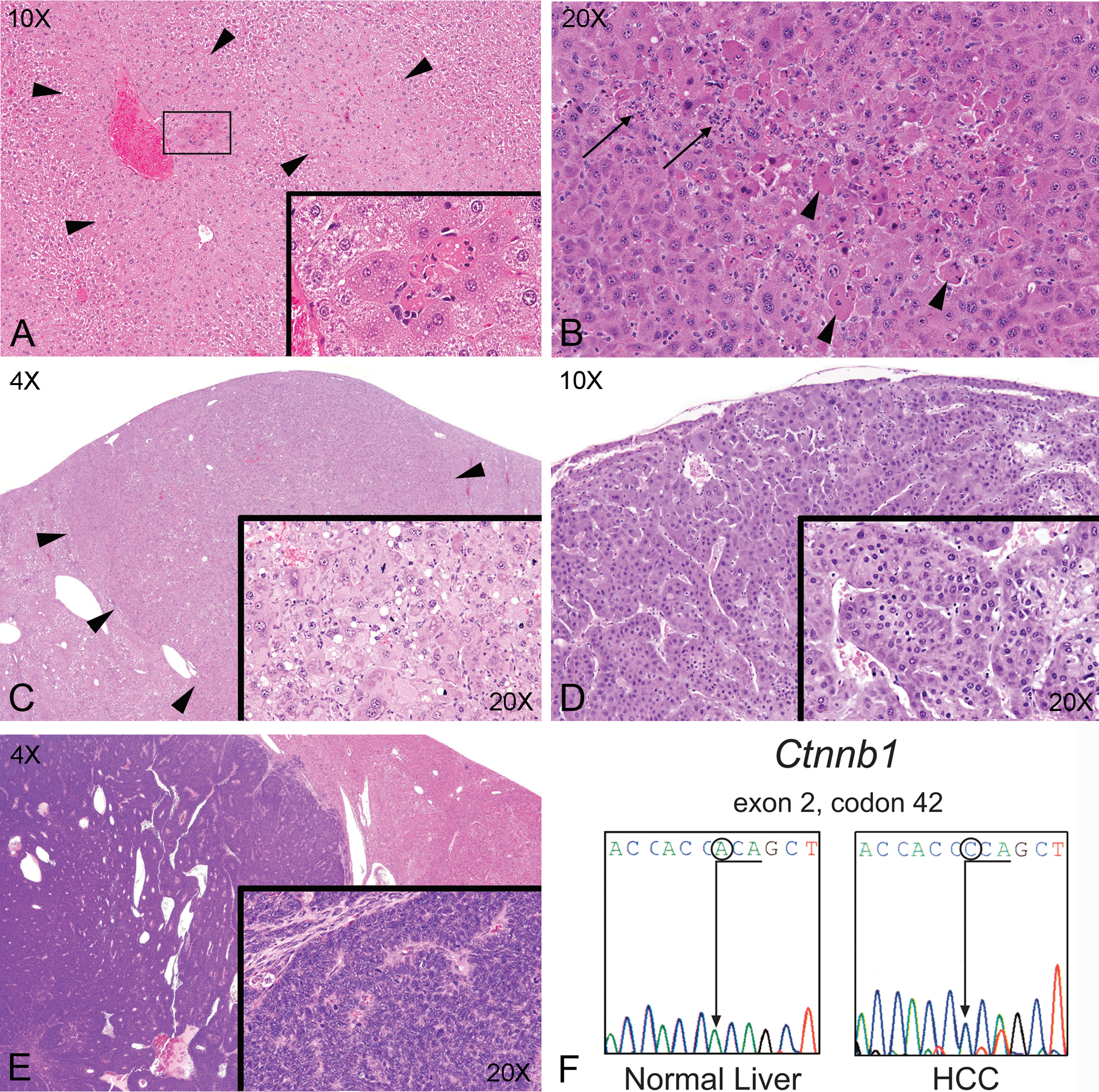
Hepatic lesions in B6C3F1 mice exposed to GBE for 90 days and 2 years by gavage; 90-day GBE exposure induced marked centrilobular hepatocellular hypertrophy (A) (arrowheads, 10×) and focal hepatocellular necrosis (box, 10× and inset, 40×). At 2-year exposure, hepatocellular hypertrophy was accompanied by minimal to mild necrosis (arrowheads) and inflammatory infiltrates (B) (arrows, 20×) composed predominantly of neutrophils, lymphocytes, and rare macrophages in all male treated groups and mid- and high-dose females groups. Two-year GBE exposure resulted in a dose-dependent increase in hepatocellular adenoma (C), carcinoma (D), and hepatoblastoma (E). Hepatocellular adenomas (C) were expansile and solid proliferations of sheets of hepatocytes (arrowheads, 4×), some of which had atypical features of enlarged cells with abundant eosinophilic cytoplasm, large nuclei and areas with numerous mitotic figures (inset, 20×). Hepatocellular caricnomas (D) (10×) were characterized by invasive cords, trabeculae, and sheets of atypical hepatocytes (inset, 20×). Hepatoblastomas (E) invariably arose within hepatocellular carcinomas and were composed of solid lobules and sheets of densely packed ovoid to round cells with deeply basophilic cytoplasm and dense, angular to ovoid nuclei, with frequent mitoses (inset, 20×). Hepatocellular carcinomas in B6C3F1 mice exposed to GBE were associated with a marked increase in point mutations in exon 2, codon 42 of
Oral exposure to GBE results in a dose-dependent increase in the incidence of hepatocellular tumors by 2 years
Exposure of male and female B6C3F1 mice to GBE over a 2-year period resulted in a dose-related increase in the incidence of hepatocellular adenomas and carcinomas (single and multiple) in male and female mice (Table 1). Histologically, hepatocellular adenomas and carcinomas associated with GBE treatment were similar to spontaneously arising tumors in vehicle control groups. Hepatocellular adenomas were generally well-circumscribed proliferations of variably sized, well-differentiated hepatocytes with variable tinctoral characteristics and displayed loss of the normal hepatic lobular architecture (Figure 1C). Adenomas varied from solid growth patterns to cords of cells 1 to 3 cell layers thick. Some adenomas were atypical and consisted of large cells with abundant eosinophilic cytoplasm, large nuclei, and areas with numerous mitotic figures (2–3 per 40× field; Figure 1C, inset). Fatty change (lipidosis) and eosinophilic intracytoplasmic inclusions were noted in some adenomas.
HCCs (Figure 1D) were poorly demarcated with irregular borders and focal invasion into the surrounding parenchyma. HCCs tended to present as larger masses than adenomas, often replacing nearly the entire hepatic lobe. Cellular atypia and mitotic figures were common. Nucleoli were often enlarged and multiple. Cells had variable tinctorial appearances from eosinophilic, basophilic, or vacuolated, to a combination of these phenotypes. Carcinomas exhibited multiple growth patterns, including trabecular, solid, and pseudoglandular patterns, often within the same tumor as previously described (Hoenerhoff et al. 2011). Tumors with a trabecular pattern were composed of cords of atypical hepatocytes three or more cell layers thick, separated by dilated vascular spaces (Figure 1D). When the solid growth pattern was present, the cells tended to be anaplastic, characterized by large size, large hyperchromatic irregular nuclei or double nuclei, 2 or 3 nucleoli, and abundant eosinophilic cytoplasm, and numerous mitoses (2 or 3 per 40× field). Although tumor phenotypes did not differ considerably between spontaneous and GBE-treated HCC, tumor multiplicity (in all treated males and high dose females) and pulmonary metastasis (in high-dose males) was more pronounced in GBE-treated mice, and often zones of marked hepatocellular hypertrophy merged imperceptibly with hepatocellular adenomas or carcinomas in these tumors.
Generally, a dose-related increase in hepatoblastomas (single and multiple) was also noted in treated males, and in mid- and high-dose females (Figure 1E). Hepatoblastomas were characterized by expansile irregular proliferations of compacted basophilic neoplastic cells arranged in sheets, often palisading around vascular spaces (pseudorosettes). Nuclei were generally oval to round to irregular, with scant basophilic cytoplasm, and mitoses were frequent. Blood-filled cystic spaces, necrosis, and hemorrhage were common components of hepatoblastomas. These tumors frequently arose within HCCs.
GBE Is Genotoxic at High Doses and GBE-treated HCCs Are Associated with Alterations in Ctnnb1 and H-ras Mutation Spectra
In the bacterial gene mutation assay (Ames assay), GBE was mutagenic at high doses (generally >2,000 µg/plate) in
GBE-treated HCCs Are Associated with Modification of CTNNB1 Protein and Alterations in the Expression of Other Wnt Mediators
Since there was a statistically significant increasing trend in
Recent studies in breast and prostate cancer have identified a similar 75kDa protein cleavage product that is associated with a more metastatic phenotype (Benetti et al. 2005; Rios-Doria et al. 2004). In these studies, cleavage of CTNNB1 by the calpain enzyme system occurs at the N-terminus, resulting in loss of the GSK3β phosphorylation site responsible for CTNNB1 degradation, resulting in a constitutively active protein fragment that retains transcriptional activity. To determine if the observed 75kDa band in GBE-treated HCC in this study was a result of such post-translational modification, an N-terminus CTNNB1 antibody was used to evaluate for changes in this region of the protein. Results of Western blotting using the N-terminus antibody showed loss of the second 75-kDa band, indicating N-terminal CTNNB1 cleavage (Figure 1G). To assess expression of the Calpain system,
Since there was a significant increase in
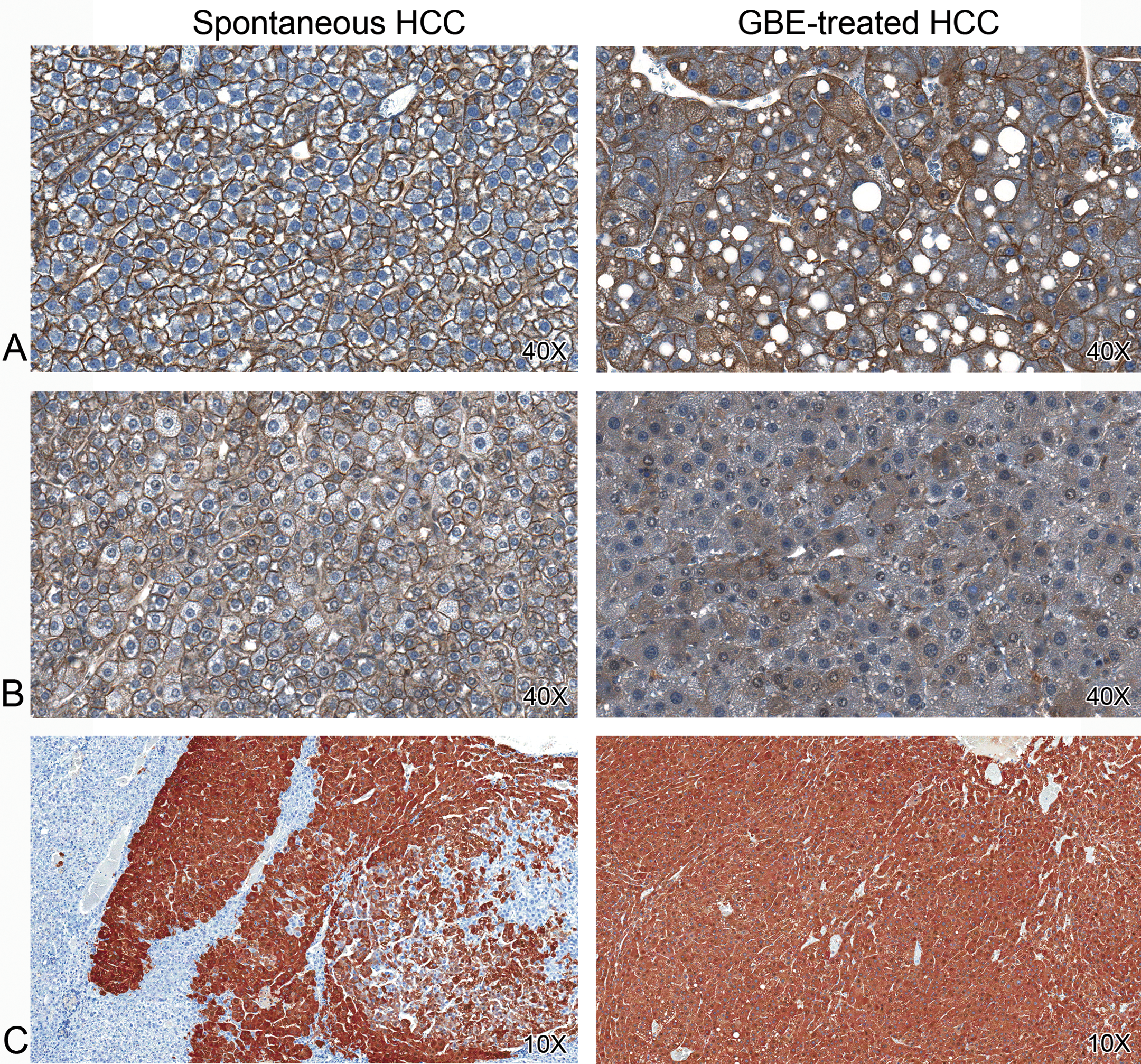
Immunohistochemistry of WNT mediators in spontaneous and GBE-treated hepatocellular carcinomas. In GBE-treated HCC, there was overexpression and cytoplasmic accumulation of CTNNB1 (A) and overexpression, cytoplasmic accumulation, and loss of membrane immunoreactivity for CDH1 (B) (20×). GBE-treated tumors overexpressed GLUL (C), a downstream transcriptional target of CTNNB1 (10×).
Differential Gene Expression Reveals Marked Differences in Global Gene Expression between Spontaneous and GBE-treated HCC
To elucidate other mechanisms that may be involved in hepatocarcinogenesis in GBE-exposed animals, global gene expression profiling was performed using age-matched vehicle control liver, spontaneous HCC, and GBE-treated HCC. From analysis of 34,000 genes on the array, a set of 2,864 significantly differentially changed (up- or downregulated) genes (
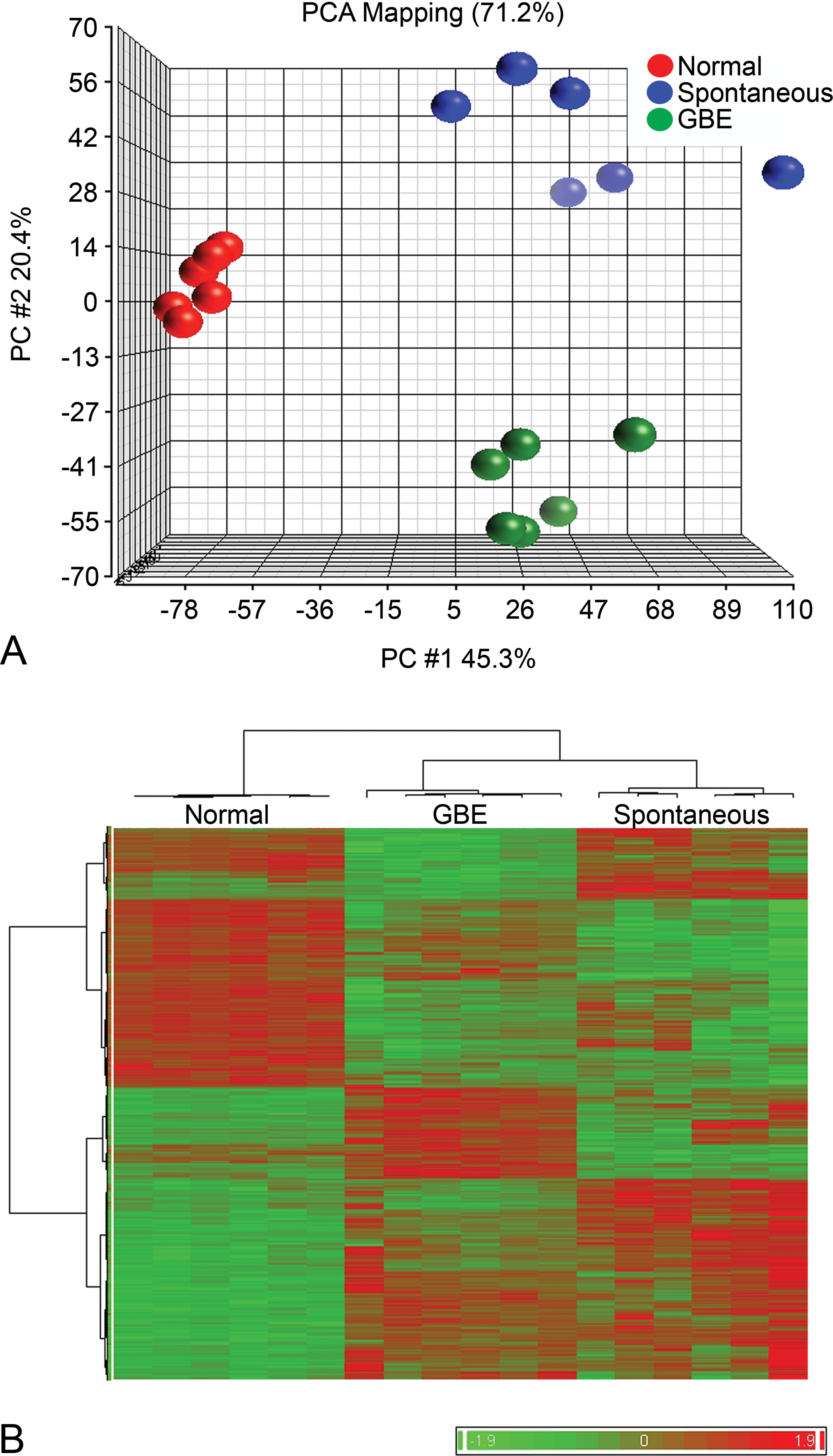
Differential gene expression profiling of spontaneous and GBE-treated HCC. Principal component analysis (PCA) (A) demonstrated significant clustering of normal liver (red), spontaneous HCC (blue), and GBE-treated HCC (green), based upon global gene expression. Hierarchical cluster analysis (HCA) (B) illustrated significant differences in global gene expression between normal liver, spontaneous HCC, and GBE-treated HCC (red = upregulated genes, green = downregulated genes).
Comparison Analysis of Differentially Expressed Genes Identifies Overrepresentation of Genes Associated with Cancer Signaling, HCC Development, and Chronic Oxidative and Xenobiotic Stress in GBE-treated HCC
Since global gene expression profiling indicated significant differential gene expression between spontaneous and GBE-treated HCC, we performed a comparison analysis between tumor types (relative to vehicle control normal liver) using Ingenuity Pathway Analysis (IPA) software. Results of comparison analysis indicated overrepresentation of genes associated with pathways involved in (1) cancer signaling, (2) HCC development, and (3) chronic xenobiotic and oxidative stress (Figure 4). Gene categories associated with cancer signaling included “cell cycle,” “cell proliferation,” “hepatic disease,” “cellular movement,” “inflammatory disease,” “immunologic disease,” and “cell death” in GBE-treated HCC compared to spontaneous tumors (Figure 4A). Gene categories associated with human HCC development (Figure 4B) included “liver inflammation,” “liver hyperplasia,” “liver cholestasis,” “liver damage,” “liver steatosis,” “liver necrosis,” “liver degeneration,” and “liver fibrosis.” Finally, overrepresented gene categories associated with chronic xenobiotic and oxidative stress included “aryl hydrocarbon receptor signaling,” NRF2-mediated oxidative stress,” “PPAR signaling,” “xenobiotic signaling,” “glutathione signaling,” and “production of nitric oxide and reactive oxygen species” (Figure 4C).
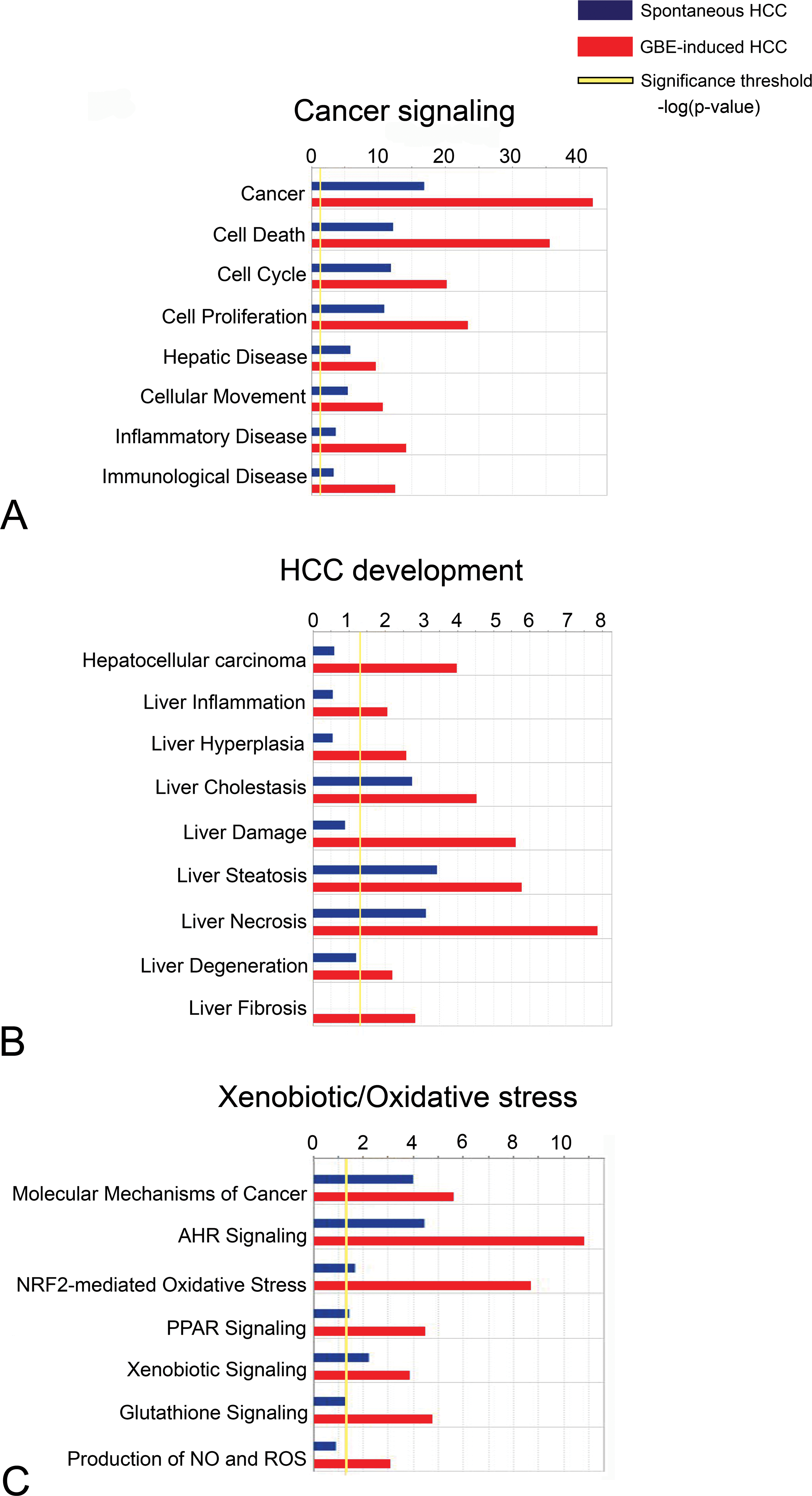
Ingenuity pathway analysis (IPA) comparison analysis of spontaneous and GBE-treated HCC. In GBE-treated tumors, there was significant overrepresentation of genes associated with biologic functions of human cancer (A), toxicologic functions associated with human hepatocellular carcinoma development (B), and pathways related to chronic xenobiotic and oxidative stress (C).
Within overrepresented pathways of cancer signaling and HCC development, there was overexpression of several
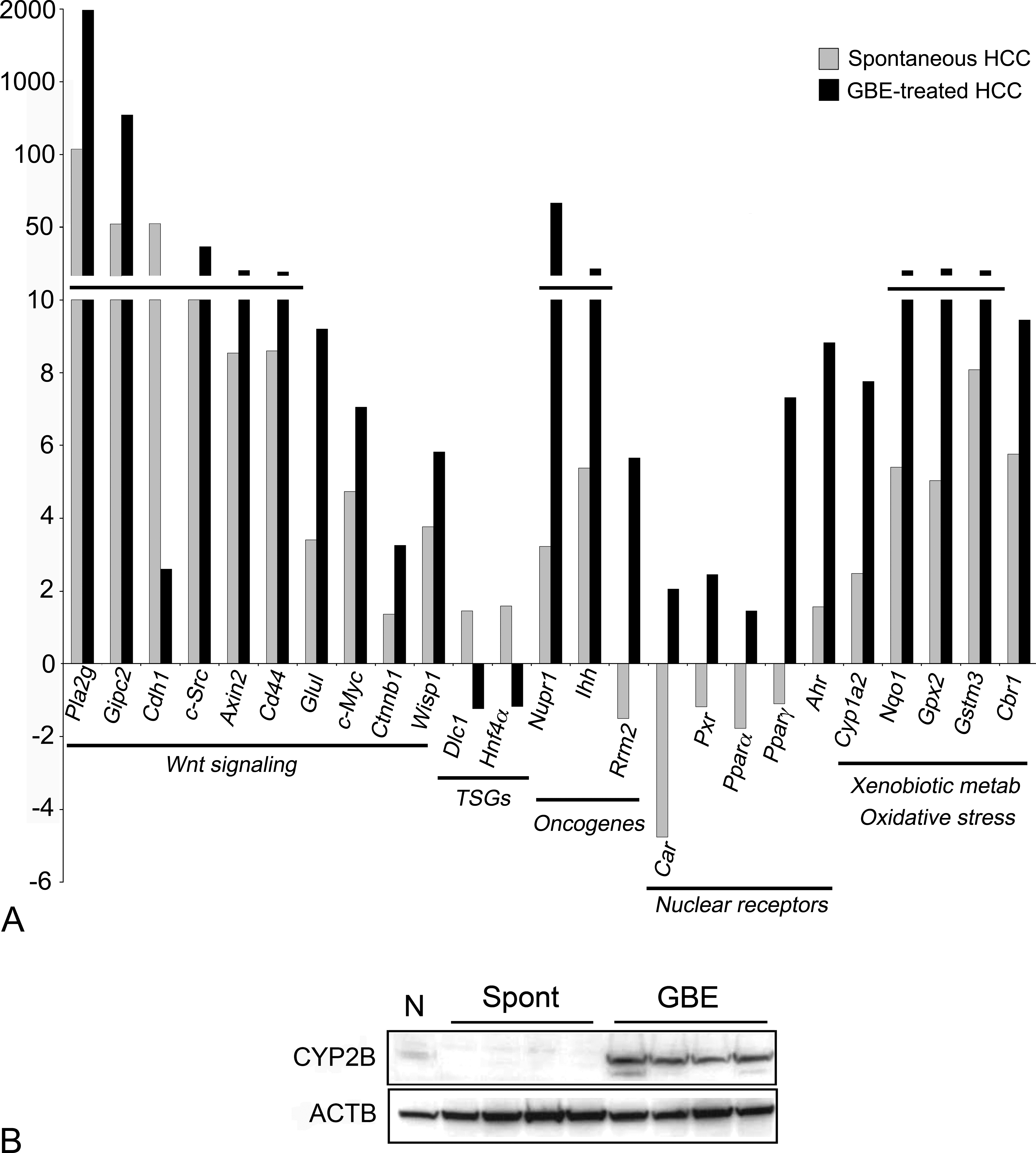
Validation of differential gene expression changes between GBE-treated and spontaneous HCC, including QRT-PCR quantification illustrating (A) upregulation of
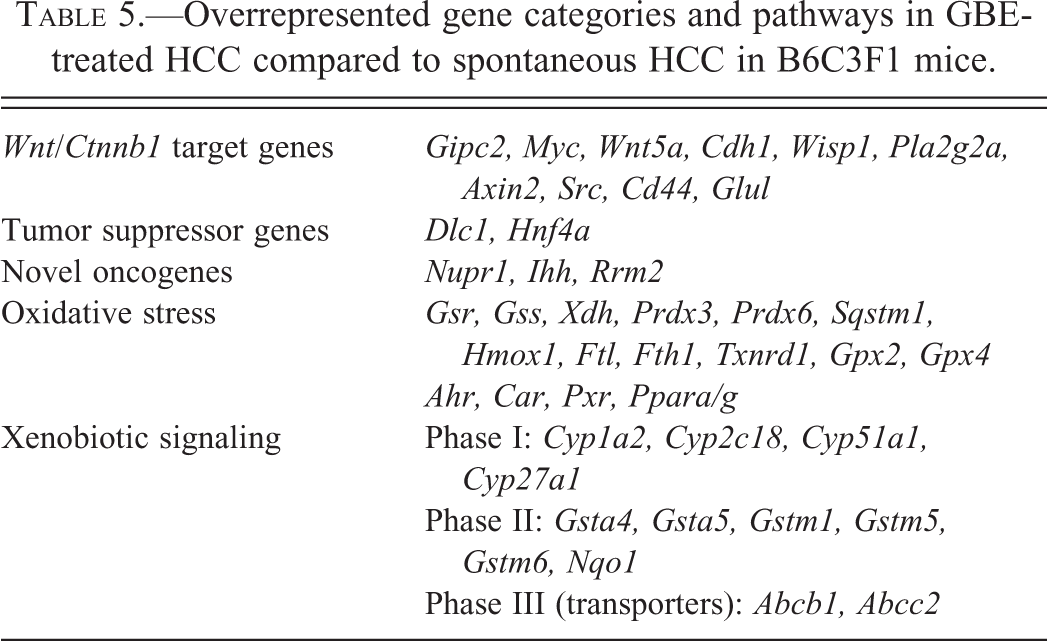
Overrepresented gene categories and pathways in GBE-treated HCC compared to spontaneous HCC in B6C3F1 mice.
Discussion
While spontaneous and GBE-treated HCC in this study were morphologically very similar, in terms of their gene expression and mutation spectra, these tumors are actually quite different. HCCs in mice exposed to GBE were characterized by dose-dependent
In addition to the genetic alterations in
We have shown that the extract was genotoxic at high doses
In order to extrapolate the doses of GBE used in the NTP bioassay to exposure in humans, we calculated potential human exposure based on simple comparisons to mass (mg/kg) and surface area (mg/m2; Reagan-Shaw, Nihal, and Ahmad 2008). Based on administered mass dose alone, the low dose (200 mg/kg) in mice is 50 times the normal recommended human dose of 240 mg/d. According to the Food and Drug Administration (FDA)’s guidelines on surface area adjustment (mg/m2), the low dose is 4 times higher. Given the unregulated nature of this compound, exposure to increased concentrations of this extract in the human population over long periods of time is possible.
In conclusion, while spontaneous and GBE-treated HCC in B6C3F1 mice are very similar at the morphologic level, we have shown that the molecular alterations in GBE-treated tumors are very different from those seen in spontaneous tumors. These include unique alterations in
Footnotes
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The authors received no financial support for the research, authorship, and/or publication of this article.
Abbreviations:
Acknowledgments
We would like to thank Kevin Gerrish and Laura Wharey in the NIEHS Microarray Core and Pierre Bushel in the Biostatistics Branch for their assistance with global gene expression experiments. We would like to thank the NIEHS Histology and Immunohistochemistry Laboratories, Protein Microcharacterization Core, and DNA Sequencing Core for their technical expertise, and the NTP Toxicogenomics Faculty for thoughtful discussions.
This work was supported by the National Institutes of Environmental Health Sciences (NIEHS), National Institutes of Health, and The Division of the National Toxicology Program. This article may be the work product of an employee or group of employees of the National Institute of Environmental Health Sciences (NIEHS), National Institutes of Health (NIH); however, the statements, opinions, or conclusions contained therein do not necessarily represent the statements, opinions or conclusions of NIEHS, NIH, or the United States government.
