Abstract
The cell of origin of hepatoblastoma (HB) in humans and mice is unknown; it is hypothesized to be a transformed hepatocyte, oval cell, or hepatic progenitor cell. In mice, current dogma is that HBs arise from preexisting hepatocellular neoplasms as a result of further neoplastic transformation. However, there is little evidence supporting this direct relationship. To better understand the relationship between hepatocellular carcinoma (HCC) and HB and determine molecular similarities between mouse and human HB, global gene expression analysis and targeted mutation analysis were performed using HB, HCC, and adjacent liver from the same animals in a recent National Toxicology Program bioassay. There were significant differences in
Keywords
Introduction
In mice, hepatoblastoma (HB) is a poorly differentiated, primitive, embryonal hepatocellular neoplasm that is often associated with preexisting hepatocellular adenomas (HCAs) or hepatocellular carcinomas (HCCs). They have a relatively late onset, occurring mainly in aged mice, and an increased incidence of HB has been associated with exposure to a number of chemicals in the National Toxicology Program (NTP 1998, 1999, 2000, 2013) carcinogenesis bioassay, including compounds such as oxazepam, diethanolamine, methyleugenol, and
The relevance of mouse HB in human hazard identification has also been a topic of great debate. Due to differences in biology between mice and humans, the use of mouse HB and other liver tumors to inform on human cancer hazard and risk has been controversial (Grisham 1997; Maronpot, Flake, and Huff 2004). In humans, HBs are predominantly observed in young children below the age of 5 years and are exceptionally rare in adults (Cruz et al. 2013; Spector and Birch 2012). In contrast, HB in mice occurs in aged animals, most often late in the course of hepatic neoplasia (Chhabra et al. 1992). HBs in the mouse most often arise in association with other hepatocellular tumors (Turusov et al. 2002); in contrast, human HBs are always solitary tumors unassociated with preexisting hepatocellular neoplasia and are thought to arise primarily
There are a number of critical questions that need to be answered regarding HBs in mice and their relationship to HCC. First, as a common model for exposure-related carcinogenicity, it is important to understand the relevance of HB in the B6C3F1 mouse in the context of HCC resulting from carcinogen exposure. If these tumors are genetically related, then an increased incidence of HB associated with HCC would suggest a progression of malignancy. If unrelated genetically, then this would suggest that HB is an independent neoplastic population resulting from ongoing genetic selection within the hepatic microenvironment, potentially targeting a multipotent hepatic stem cell. Secondly, it is critical to further understand the relevance of HB in the B6C3F1 mouse to humans in hazard identification. If these tumors are similar in humans and mice in terms of their global gene expression, then despite their differences in clinical course, an increased incidence of HB as a result of chemical exposure would provide stronger evidence that such an exposure may pose a cancer risk in humans. Given these unanswered questions, the aim of this study was to compare concurrent HB and HCC to identify relationships in global gene expression and mutation spectra of common hepatic cancer genes (
Materials and Methods
Sample Selection, Laser Capture Microdissection (LCM), RNA Isolation, and Amplification
All tissues used in this study were from animals in a single NTP chronic bioassay (bromodichloroacetic acid [BDCA], TR-583). The NTP conducts its studies in compliance with its laboratory health and safety guidelines and Food and Drug Administration Good Laboratory Practice Regulations and must meet or exceed all applicable federal, state, and local health and safety regulations. Animal care and use in this bioassay were in accordance with the Public Health Service Policy on Humane Care and Use of Animals. Samples containing HB, adjacent HCC, and associated normal (AN) liver in the same section were identified histologically in formalin-fixed, paraffin-embedded (FFPE) sections (Figure 1). Frozen samples collected from the liver of animals that corresponded to these FFPE tissues were used for RNA isolation. These frozen tissues were collected from the NTP frozen tissue repository, embedded in optimum cutting temperature freezing media, and cryosectioned for LCM in order to obtain each cell population (HB, HCC, and AN) for analysis. HB (
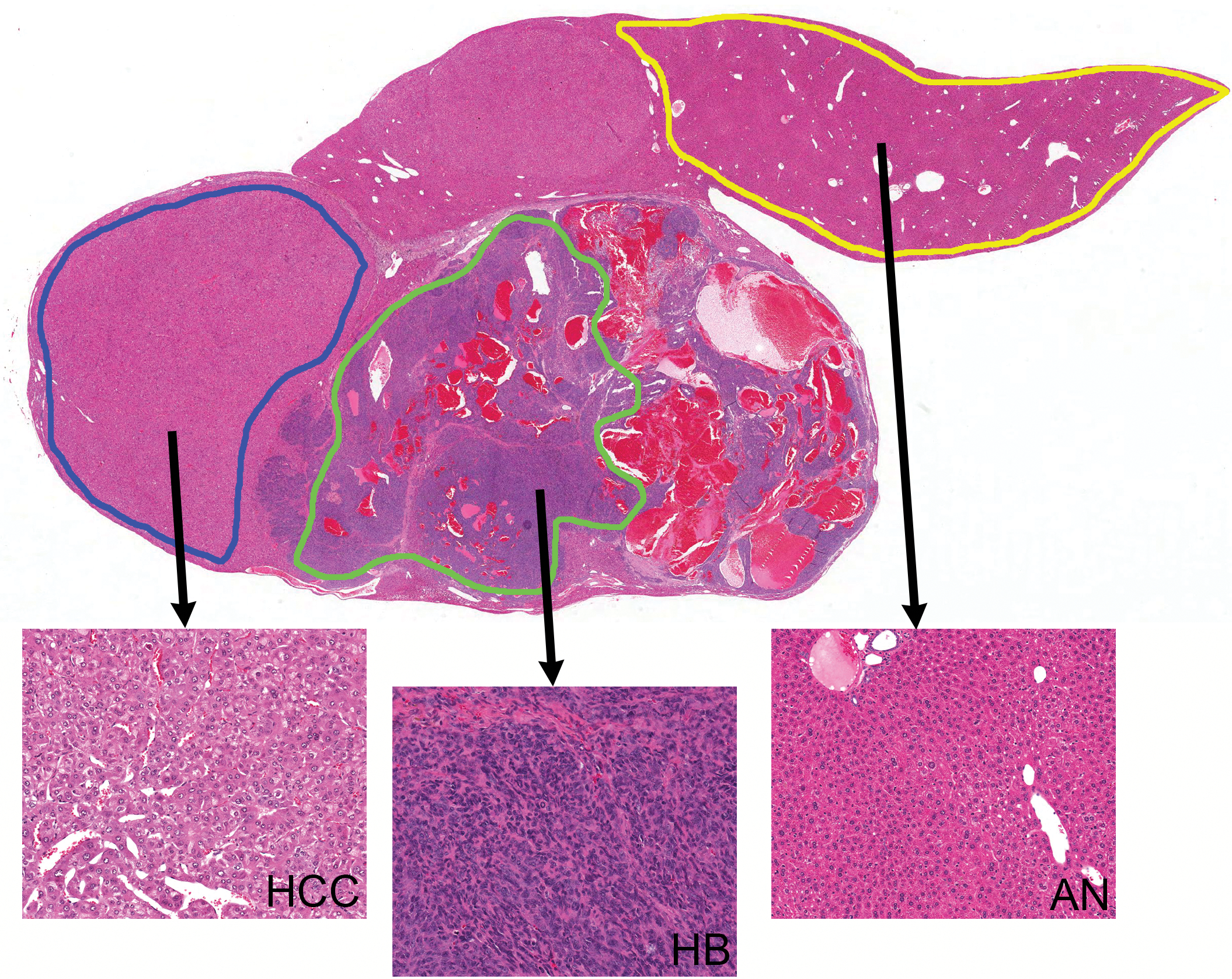
Tissue collection protocol for obtaining hepatoblastoma, hepatocellular carcinoma, and associated normal liver for each sample, using laser capture microdissection.
Mutation Analysis
Regions of the mouse
Microarray Hybridizations
Gene expression analysis was conducted on laser capture microdissected HB (
Data Processing and Identification of Differentially Expressed Genes (DEGs)
Data processing and identification of DEGs were done as previously described (Hoenerhoff et al. 2011; Pandiri et al. 2012). Briefly, array fluorescent pixel intensity measurements were acquired and gene expression data were normalized across all samples using the robust multiarray analysis (RMA) methodology (Irizarry et al. 2003). Using RMA-normalized data, for each probe set, pairwise comparisons (AN vs. HB and AN vs. HCC) were made using a bootstrap
Bioinformatics Analyses
Partek Genomics Suite, version 6.6 (release date June, 2012; Partek, St. Louis, MO), was used to perform principal component analysis (PCA) on the normalized data and to generate heat maps to compare samples for differentially expressed probe sets. PCA uses a linear transformation to find orthogonal variables (principal components [PCs]) that describe the variability in the data. The first 3 PCs that capture the majority of the variation in the data were used to visualize the spatial relationship of the HB, HCC, and AN samples.
Ingenuity pathway analysis (IPA) 9.0 (www.ingenuity.com), application build-220217, version-16542223, was used to evaluate the most statistically significant overrepresented canonical pathways. These canonical pathways are based on the ingenuity knowledge base. The significant biological canonical pathways were derived from IPA and the statistical significance was set at
The NextBio tool (www.nextbio.com) was used to compare the transcriptomic data sets of mouse HB tumor to most similar human and rodent transcriptomic data sets and evaluate pathway enrichment in both systems. NextBio is a curated and correlated repository of experimental data derived from an extensive set of public sources (e.g., ArrayExpress and GEO) that allows the user to compare patterns of gene expression in their experiment to thousands of published transcriptomic data sets. The statistical approach used by NextBio is analogous to gene set enrichment analysis (Kupershmidt et al. 2010; Subramanian et al. 2005). NextBio software was utilized to compare the differential transcriptomic changes (biogroups) common to human tumors curated by NextBio and mouse HB and HCC tumors.
Quantitative Real-time PCR
RNA extraction and amplification were performed using Ovation Pico WTA System V2 (NuGEN, San Carlos, CA) following manufacturer’s recommendation. Total RNA (10 ng) was amplified using Ovation Pico WTA System and resulted in 6 µg of amplified cDNA. Relative quantitative gene expression levels were detected using real-time PCR with the ABI PRISM 7900HT Sequence Detection System (Life Technologies, Grand Island, NY) using synergy brands (SYBR) green methodology. Primers were designed using Primer3Plus software (Rozen and Skaletsky 2000) to span exon–exon junctions with an annealing temperature of 60°C and amplification size of less than 150 bp. Briefly, 25 ng of cDNA were added to a 25-µL PCR reaction to get a final concentration of 1.00 ng/µL of cDNA. Forward and reverse primer final concentrations were 100 nM in the SYBR green assay. The reactions were performed using the Power SYBR® Green PCR Master Mix (Life Technologies, Grand Island, NY). 18S was chosen as the endogenous control gene in our quantitative PCR (qPCR) experiments. Relative quantification of gene expression changes was recorded after normalizing for 18S expression computed by using the 2−ΔΔCT method (user manual #2; ABI Prism 7700 SDS).
Immunohistochemistry (IHC)
IHC was performed on unstained FFPE sections of 3 samples containing HCC and HB using an avidin–biotin peroxidase system (Vectastain Elite ABC kit; Vector Laboratories, Burlingame, CA) according to the manufacturer’s protocol. Proteins for IHC analysis were selected based on differential gene expression between HB and HCC from the microarray data as well as the role of these proteins in HCC and HB development. The 5 µm sections were collected on charged glass slides and were deparaffinized in xylene and rehydrated through a decreasing graded ethanol series. Following antigen retrieval and blocking of endogenous peroxidases, primary antibody was applied. These included hepatocyte nuclear factor-4a (HNF4a [H-171], 1:250, rabbit polyclonal; Santa Cruz Biotechnology Inc., Santa Cruz, CA), cMYC (cMYC [AB32072], 1:75, rabbit polyclonal; Abcam Inc., Cambridge, MA), β-catenin (CTNNB1 [H-102], 1:750, rabbit polyclonal; Santa Cruz Biotechnology Inc.), and CYP2E1 ([AB1252] 1:1000, rabbit polyclonal; Millipore, Billerica, MA). Sections were incubated with avidin–biotin–peroxidase complex (Vector Laboratories), and 3,3-diaminobenzidine was used to visualize all immune reactions. Positive controls included tissues known to exhibit positive expression of proteins of interest. Negative controls received the antisera from the same animal species as the source of the secondary antibody.
Results
HB and HCC from B6C3F1 Mice Do Not Share Common Mutations in Hras and Ctnnb1
Regions of the mouse
Incidence of
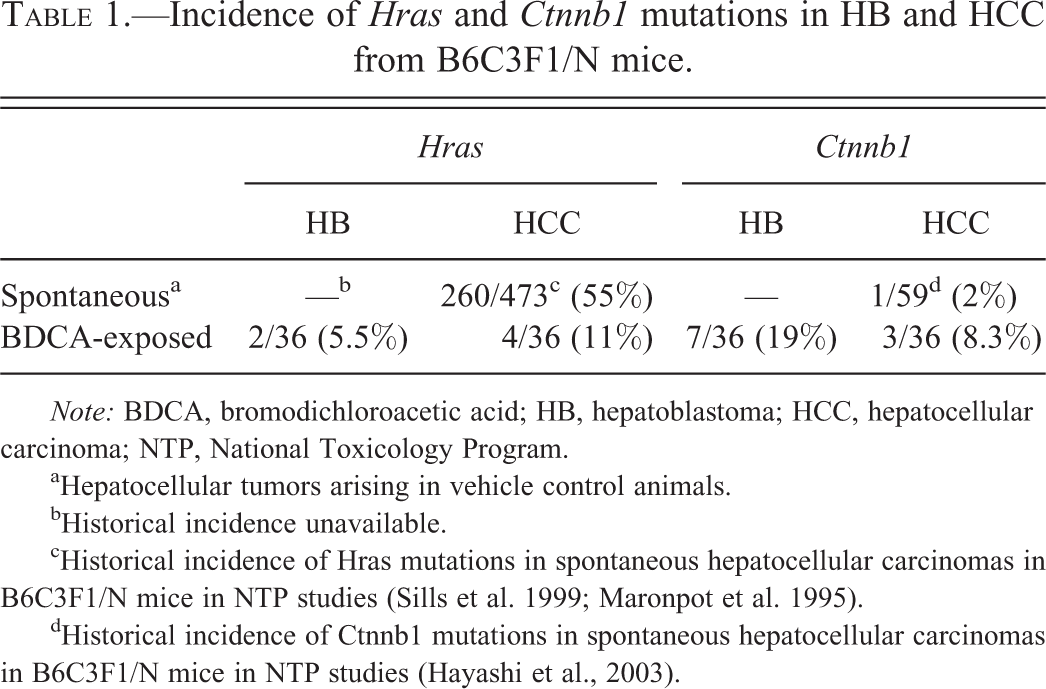

aHepatocellular tumors arising in vehicle control animals.
bHistorical incidence unavailable.
cHistorical incidence of Hras mutations in spontaneous hepatocellular carcinomas in B6C3F1/N mice in NTP studies (Sills et al. 1999; Maronpot et al. 1995).
dHistorical incidence of Ctnnb1 mutations in spontaneous hepatocellular carcinomas in B6C3F1/N mice in NTP studies (Hayashi et al., 2003).
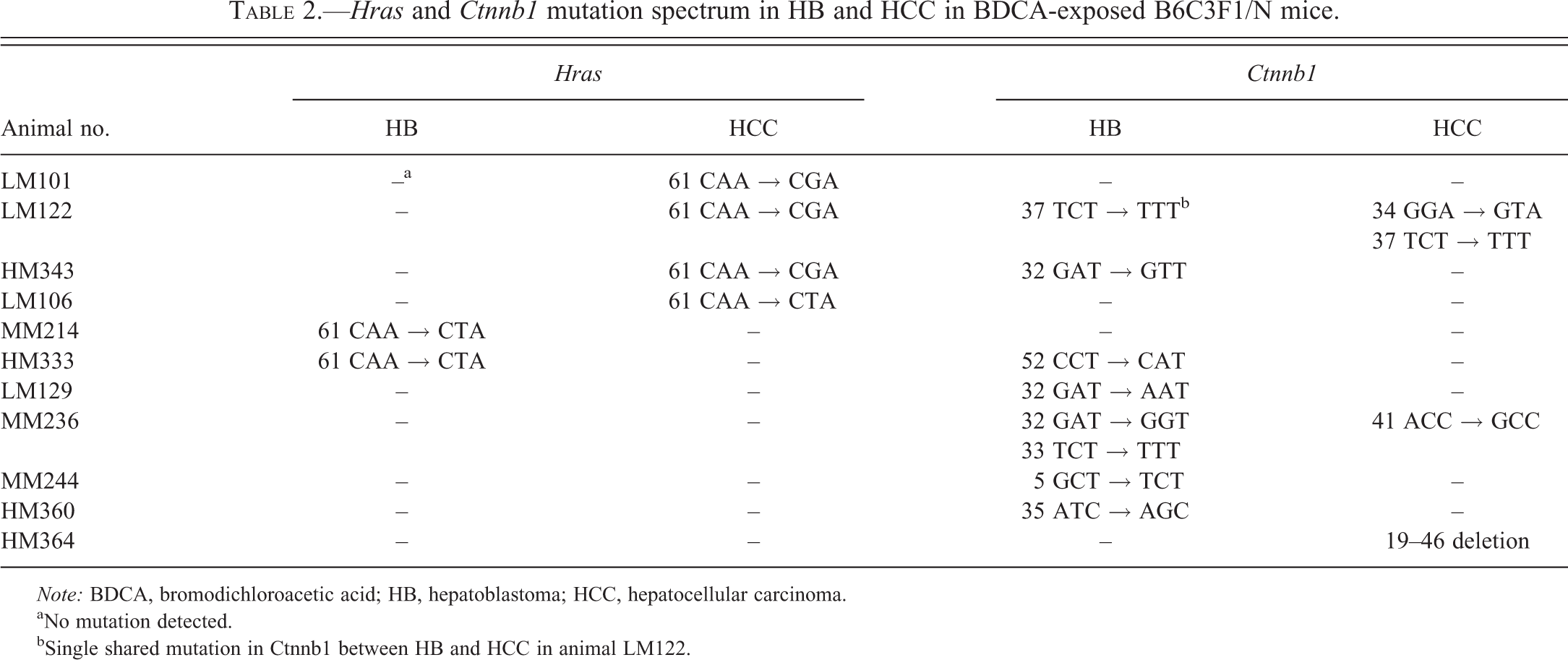
aNo mutation detected.
bSingle shared mutation in Ctnnb1 between HB and HCC in animal LM122.
HB Is Markedly Different from HCC Based on Global Gene Expression
Comparisons between global gene expression of laser capture microdissected HB, HCC, and adjacent normal liver (AN) from the same individual animals were made in order to isolate gene expression changes common and different between HB and HCC in the context of AN liver in exposed animals. This comparison was designed to minimize background chemical effect on the adjacent normal liver and tumor samples, in order to identify alterations more specific to tumorigenesis in the absence of chemical effect. While some chemical effects on tumor biology may remain at play after this comparison, this method was the most stringent to control for chemical effect while identifying significantly altered genes associated with tumorigenesis between HB and HCC. Results indicated 10,346 DEGs in HB compared to AN liver (mdFDR < 5%). In HCC, there were 1,087 genes differentially expressed (mdFDR < 5%) compared to AN liver. PCA based on all interrogated probe sets showed distinct tight clustering of HBs from HCC and AN liver (Figure 2), suggesting that there is marked variance in global gene expression that segregates HB from HCC and AN. Similarly, unsupervised hierarchical cluster analysis of DEGs segregated HB from HCC and AN (Figure 3).
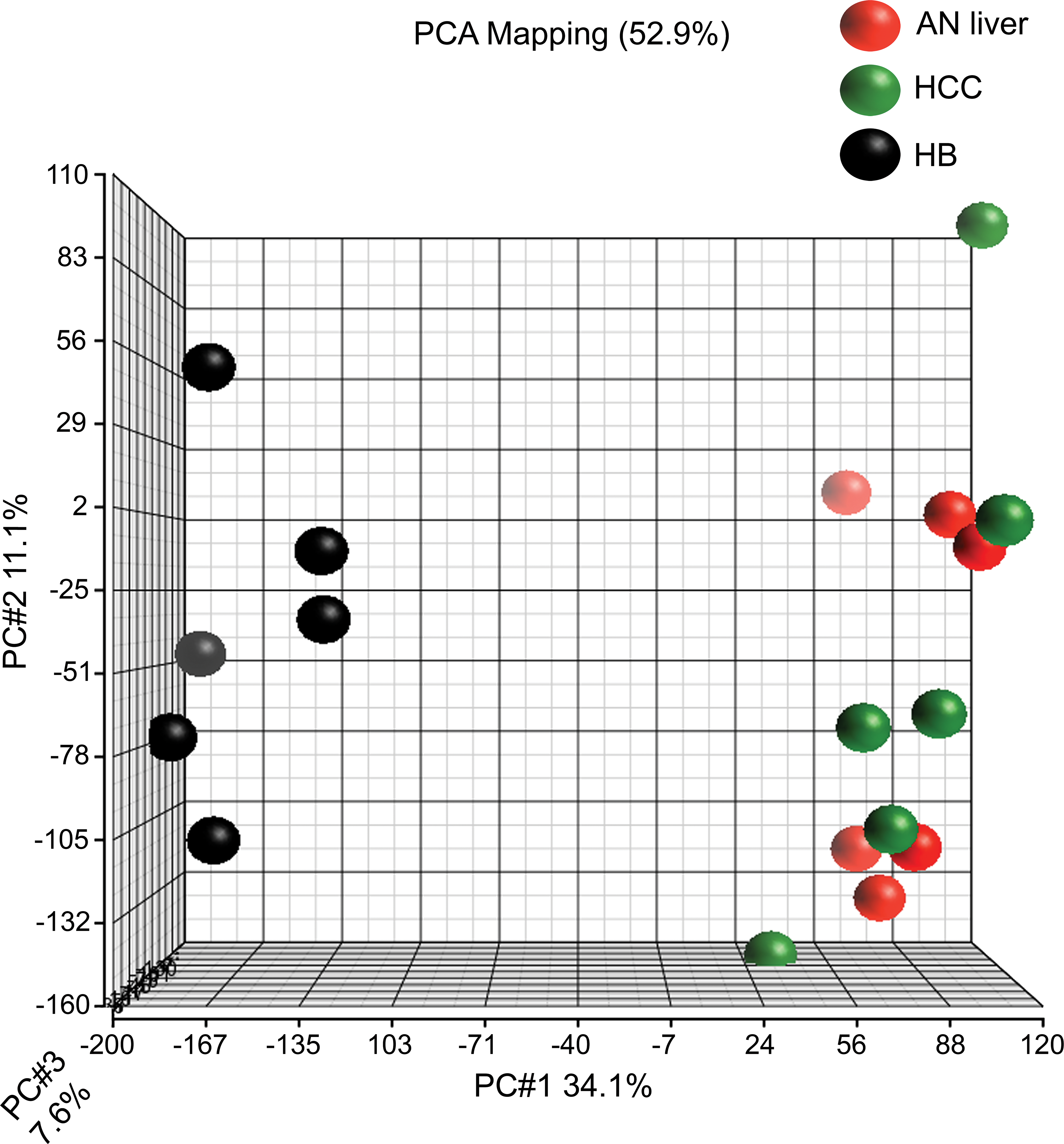
Principal components analysis comparing gene expression profiles of hepatoblastoma (HB [black]), hepatocellular carcinoma (HCC [green]), and associated normal (AN liver [red]) for differentially expressed probe sets. The plot illustrates marked differences in global gene expression in HB compared to both HCC and AN liver; overlap of HCC and AN liver samples suggests that these samples have less variance in their gene expression compared to HB, which is distinctly and markedly different.
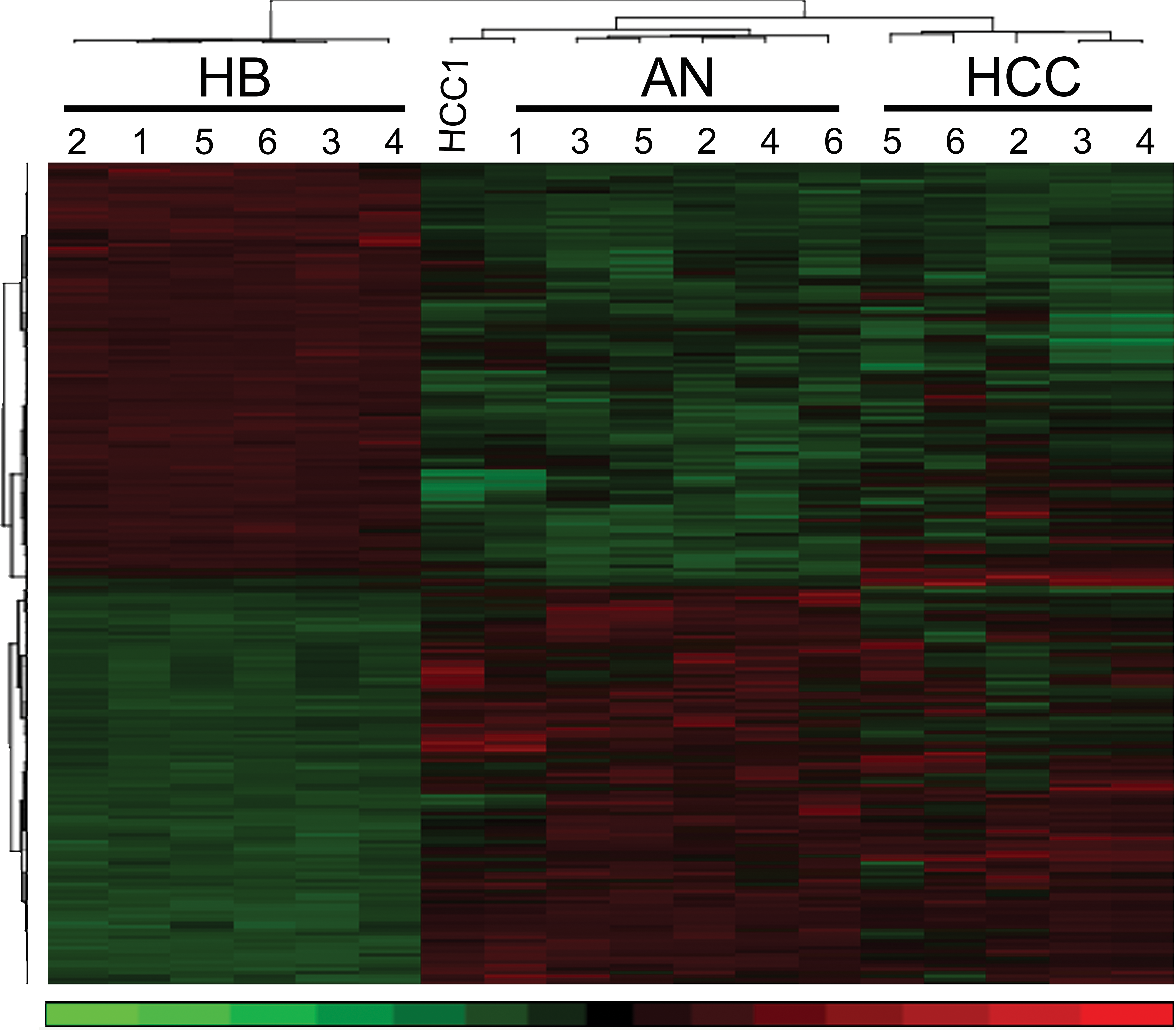
Unsupervised hierarchical cluster analysis (hepatocellular adenomas) illustrating significant differences in global gene expression of hepatoblastoma (HB) tumors compared to hepatocellular carcinoma (HCC) and associated normal (AN liver [red = upregulated genes and green = downregulated genes]). Similar to principal component analysis, hierarchical clustering shows that gene expression alterations in HCC are more similar to AN liver than HB. Alterations in significantly differentially expressed genes in HB show a marked difference when compared to HCC.
Overrepresented Pathways in Mouse HB Include Embryonic Development and Genomic Imprinting Compared to Normal Liver
IPA was performed to identify altered pathways in HB compared to AN liver. Genes targeted by
Ingenuity pathway analysis of select DEGs from overrepresented pathways in mouse HBs.
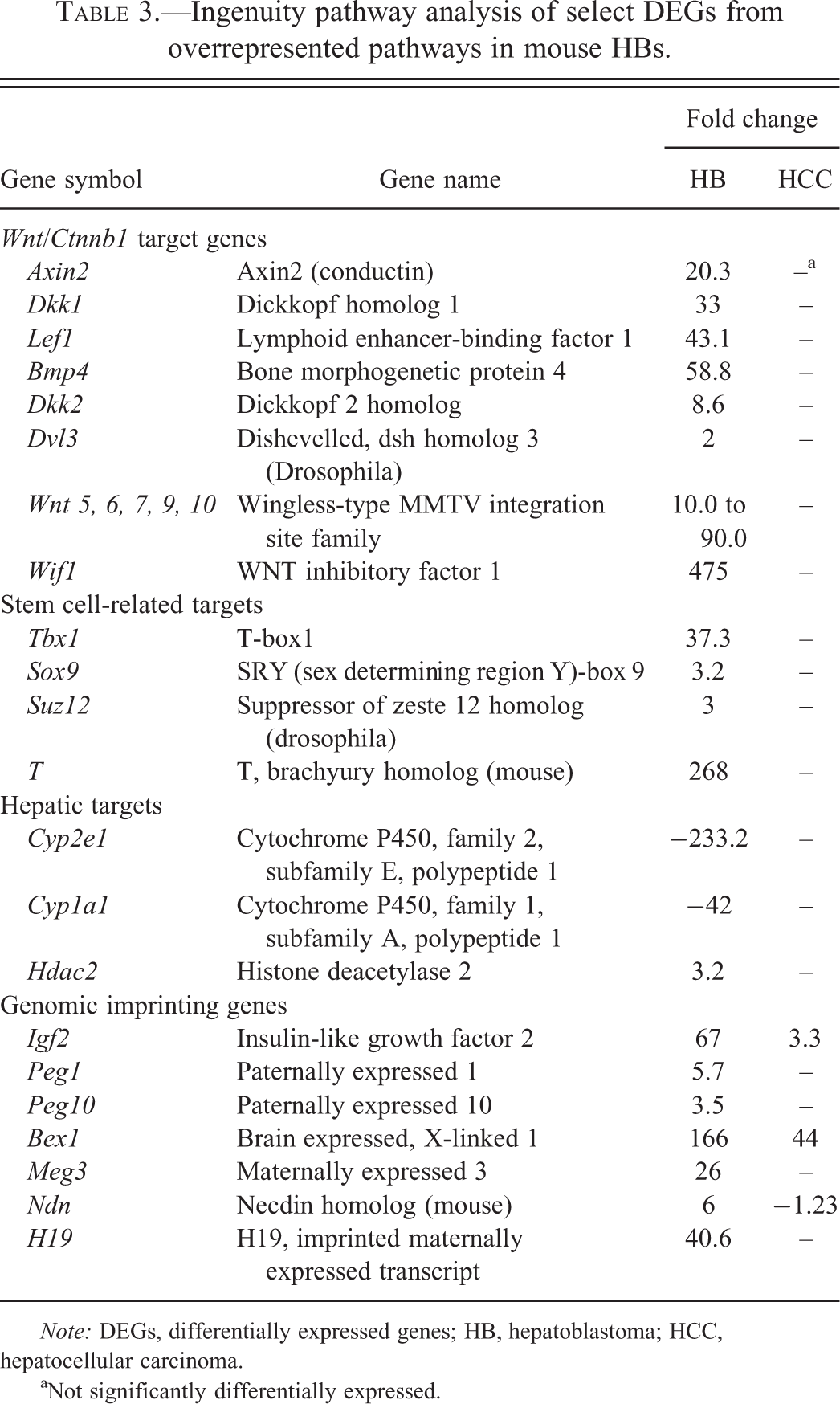
aNot significantly differentially expressed.
HBs Show Overrepresentation of Pathways Associated with Carcinogenesis and Stem/Progenitor Cell Signaling Compared to HCC
Results of IPA indicated overrepresentation of a number of biologic functions shared between HCC and HB, including cancer and cellular movement, and also identified differentially altered pathways involving protein synthesis, metabolic disease, and developmental disorders (Figure 4A). Overrepresented canonical pathways related to carcinogenesis in HB compared to HCC included EIF2 signaling, cell cycle control of chromosome replication, NRF2-mediated oxidative stress response, basal cell carcinoma signaling, and mTOR signaling (Figure 4B). Finally, there were a number of stem cell/developmental regulation pathways represented in HB compared to HCC, including Ephrin B signaling, mouse and human embryonic stem cell pluripotency, DNA methylation and transcriptional repression signaling, and sonic hedgehog (Shh) and NANOG signaling (Figure 4C). Select genes that were differentially expressed between HB and HCC were validated using qPCR (Figure 5A and B) and IHC (Figure 6). Targets for qPCR analysis included those genes observed as significantly differentially expressed between HB and HCC on microarray analysis, observed as related to genomic imprinting (

Ingenuity pathway comparison analysis of differentially expressed genes in hepatoblastoma compared to hepatocellular carcinoma in bromodichloroacetic acid–exposed B6C3F1 mice, including (A) top biological functions, (B) cancer-associated pathways, and (C) stem cell and developmental pathways overrepresented in hepatoblastoma.
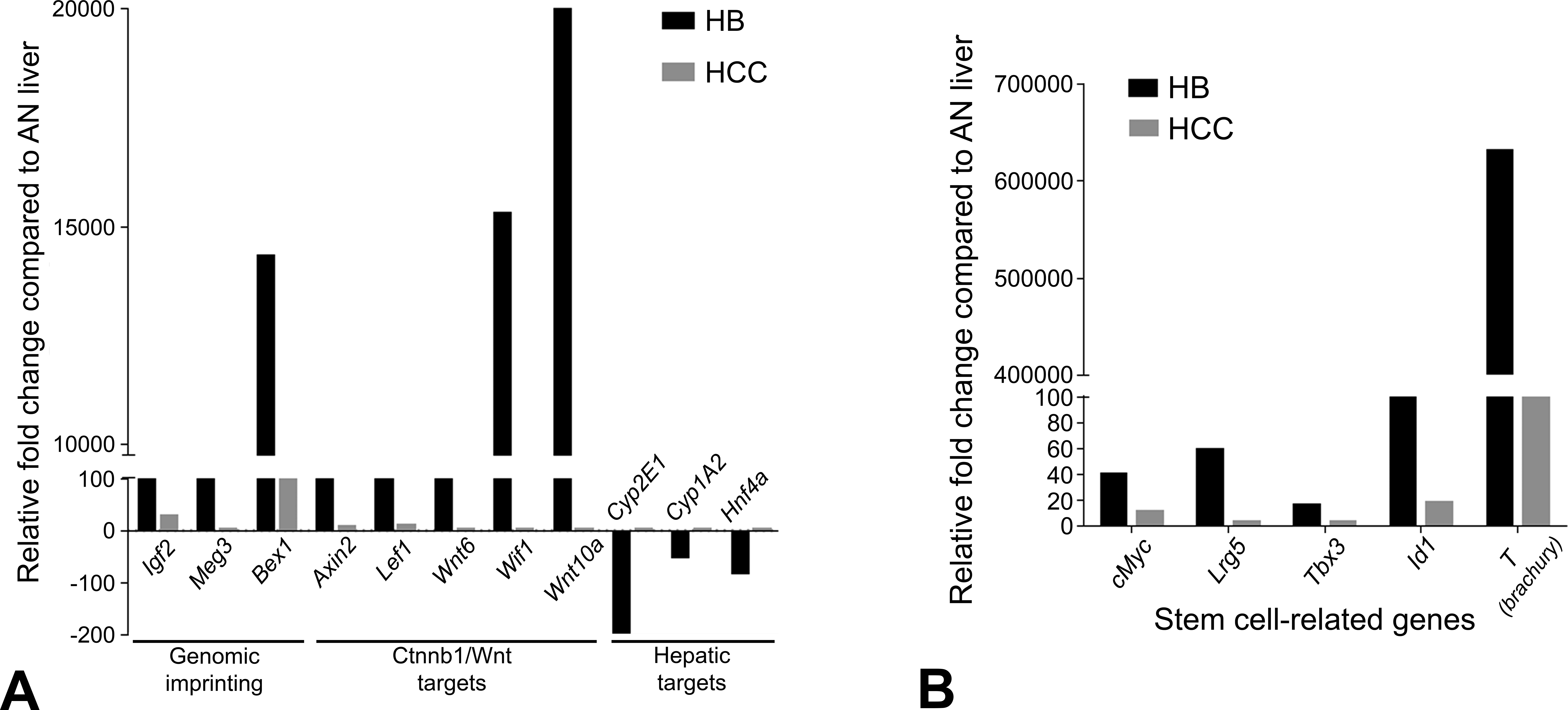
Quantitative polymerase chain reaction validation of microarray targets involved in (A) genomic imprinting, Wnt/Ctnnb1 pathway, and hepatic function and (B) stem cell genes in hepatoblastoma and hepatocellular carcinoma from bromodichloroacetic acid–exposed B6C3F1 mice.
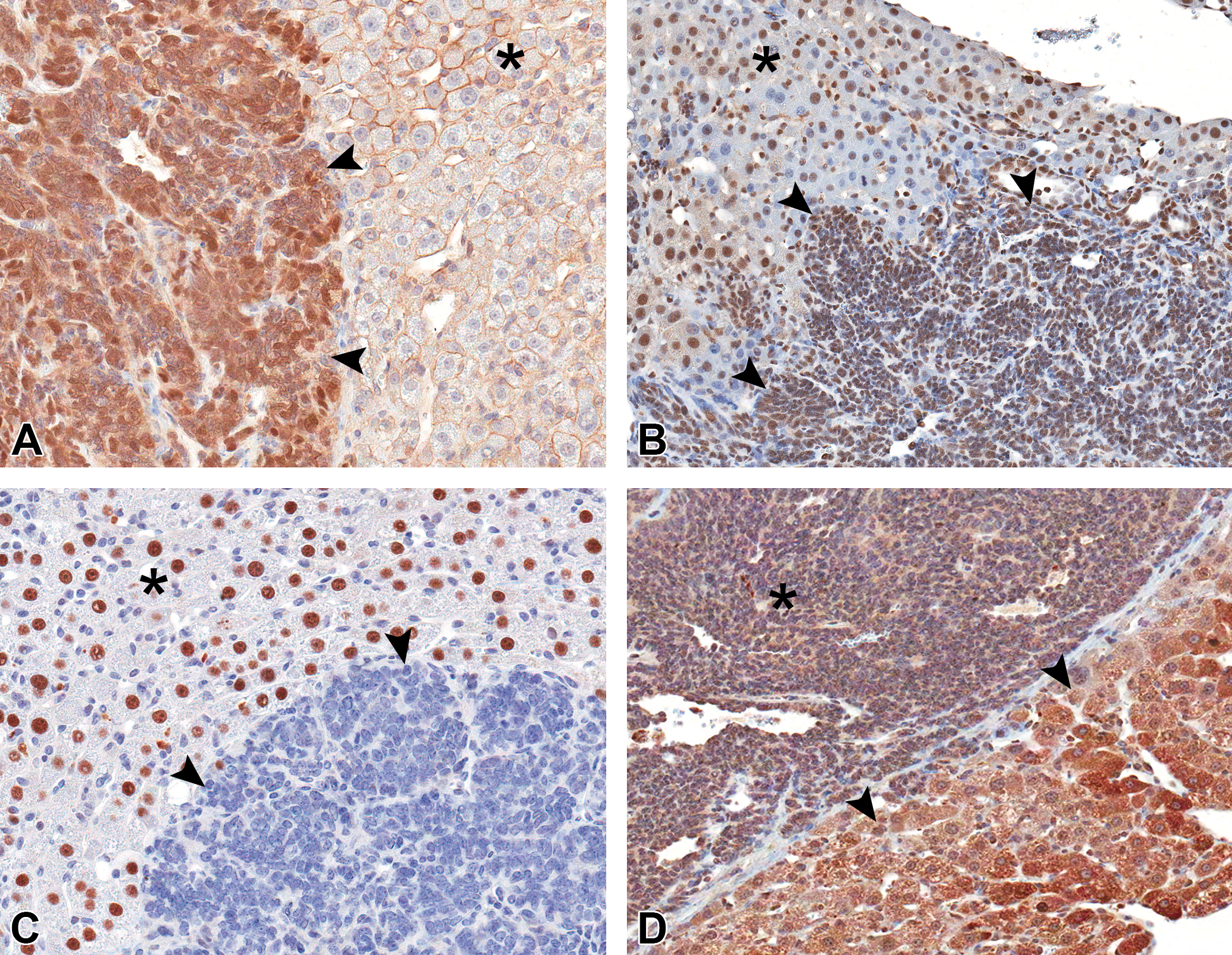
Immunohistochemistry validation of microarray targets differentially expressed between hepatoblastoma (HB) and hepatocellular carcinoma (HCC). (A) Diffuse cytoplasmic and membrane immunoreactivity to CTNNB1 antibody in HB (arrowheads) compared to mild membrane staining in adjacent HCC (asterisk) (anti-CTNNB1 antibody, hematoxylin counterstain, original objective used 40×). (B) Diffuse nuclear immunoreactivity to cMYC antibody in HB (arrowheads) compared to multifocal nuclear immunoreactivity in adjacent HCC (asterisk) (anti-cMYC antibody, hematoxylin counterstain, original objective used 20×). (C) Negative immunoreactivity to HNF4A antibody in HB (arrowheads) compared to strong and diffuse nuclear immunoreactivity in adjacent HCC (asterisk) (anti-HNF4A antibody, hematoxylin counterstain, original objective used 40×). (D) Moderate to strong cytoplasmic immunoreactivity to CYP2E in HCC (arrowheads) compared to weak cytoplasmic labeling in adjacent HB (asterisk) (anti-CYP2E antibody, hematoxylin counterstain, original objective 20×).
Gene Expression of Mouse HB Is Highly Concordant with Early Mouse Embryonic Liver and Pluripotent Hepatic Stem Cells
NextBio meta-analysis software (Kupershmidt et al. 2010) was used to compare mouse HB and HCC to other mouse and human data sets in the literature in order to identify similarities in data sets based on the total number of shared genes, and the number of genes similarly changed based on directionality of fold change in both data sets. Meta-analysis showed that, in contrast to HCC, mouse HB is transcriptomically very similar to early (E10.5) embryonic mouse liver (Chaignat et al. 2011; Otu et al. 2007) and highly concordant with hepatocyte-derived pluripotent stem cells (Lee et al. 2012; Figure 7).
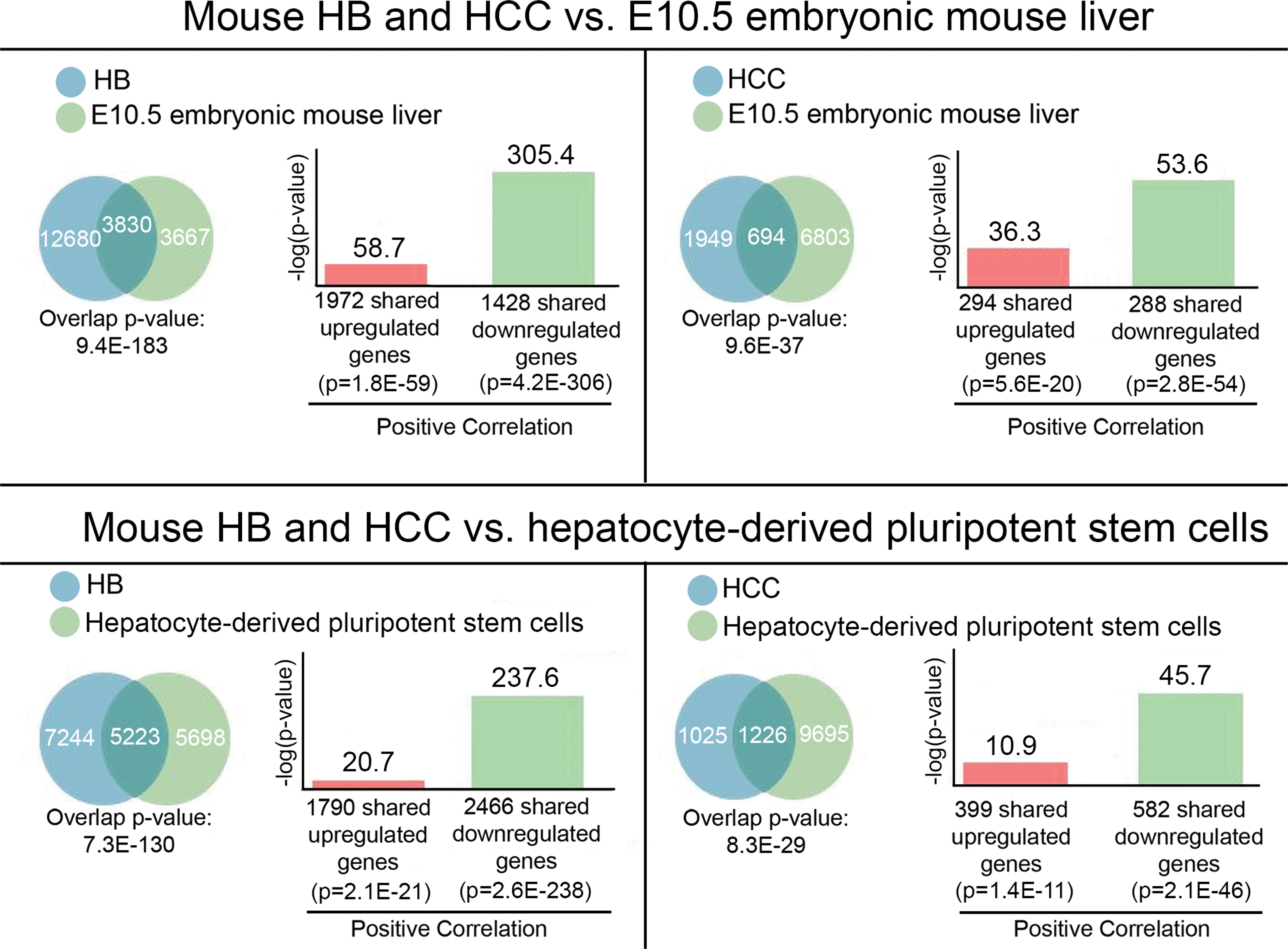
NextBio meta-analysis comparing hepatoblastoma (HB) and hepatocellular carcinoma (HCC) in bromodichloroacetic acid (BDCA)–exposed B6C3F1 mice with mouse embryonic liver (E10.5, top) and hepatocyte-derived pluripotent stem cells (bottom). There is a higher degree of concordance between HB in BDCA-exposed mice and mouse embryonic liver (3,830 total shared genes and
Discussion
The primary objectives of the current study were to better understand the underlying molecular changes that characterize the relationship between HB and HCC in B6C3F1 mice, and how these changes relate to human HB in order to better understand the relevance of the mouse HB to human health in chemical hazard identification studies. Using global gene expression profiling and mutation analysis, we have shown that HB is markedly different and quite distinct from HCC in mice. First, mutation analysis of
In the current study, numerous pathways and their respective genes involved in both mouse and human embryonic stem cell pluripotency, embryonic development, and genomic imprinting were prominent in mouse HB but not significantly represented in mouse HCC (Cairo et al. 2008; Clevers 2006; Monga 2011). Overrepresentation of genes from the hedgehog-signaling (Hh) pathway, including strong upregulation of Shh, its receptor patched 1, and the Hh ligand smoothened, was observed in HB and not HCC. This pathway is responsible for critical cell fate decisions including proliferation, apoptosis, migration, and differentiation (McMillan and Matsui 2012). It plays vital roles in tissue morphogenesis during fetal development including embryonic development of the liver and hepatic regeneration in the adult; constitutive activation of Hh signaling can lead to transformation of progenitor cells when continuously activated and lead to various cancers in humans (Beachy, Karhadkar, and Berman 2004; Omenetti et al. 2011). Activation of the Hh pathway is also important in cancer stem cell regulation and enhanced tumor initiating and self-renewal potential (McMillan and Matsui 2012), and overexpression of this pathway has been reported in about two-thirds of human HB (Eichenmüller et al. 2009; Oue et al. 2010). Significant upregulation of genomic imprinting genes primarily expressed in fetal liver was observed in mouse HB from the current study; this upregulation is similar to that observed in human HB (Cairo et al. 2008; Tomlinson and Kappler 2012). Genomic imprinting is an epigenetic mechanism in which individual alleles are silenced based on the parent of origin, and loss of imprinting is recognized in various childhood developmental disorders and in various cancers (Damaschke et al. 2013).
Genes associated with Ctnnb1/Wnt signaling and normal hepatic function were significantly differentially altered in mouse HB in this study compared to HCC. The Wnt/Ctnnb1 pathway is a master regulator of cell fate and proliferation during embryonic development, and it is essential for stem cell maintenance in a wide variety of tissues (Clevers 2006; Haegebarth and Clevers 2009). In this study, we observed dysregulation in a number of genes associated with a variety of Wnt/Ctnnb1 functions, including feedback regulation, positive effectors, antagonists of the Wnt pathway, and target genes in carcinogenesis. Many of these genes expressed at high levels in HB tumors are targets of
Meta-analysis of mouse HB indicated that the key processes of embryonic development and stem/multipotent cell regulatory pathways as the major characteristics of these tumors. There was significantly higher concondance not only with mouse embryonic liver but also with hepatic pleuripotent stem cell biosets in HB, compared to mouse HCC. This supports an embryonic phenotype of HB tumors as seen histologically and further indicates that these are distinct tumors likely arising from a primordial hepatic precursor cell. Although there are several differences between human and mouse HB, our results suggest that mouse and human HB share some important similiarities in their molecular landscape, and therefore mouse HB is a relevant lesion in toxicology and carcinogenicity bioassays and is an important indicator of potential cancer risk in hazard identification studies. Due to its marked genomic and phenotypic differences, this tumor may be considered a distinct entity from HCC.
Conclusions
Our data show that mouse HBs are profoundly different from HCC in terms of their global gene expression and mutation spectra. In addition, mouse and human HB share significant similarities in the global gene expression, including dysregulation of embryonic developmental pathways, embryonic/stem cell pluripotency pathways, and genomic imprinting. In addition, meta-analysis supports that mouse HB is highly concordant with mouse embryonic liver and hepatic pluripotent stem cells. These findings suggest that HB and HCC are very distinct and separate entities, although possibly arising from a common hepatic stem or progenitor cell. It is possible that the development of HB may require a particular microenvironment which is provided by an adjacent HCC, including one with increased genomic instability. Further studies with greater sample sizes from several bioassays as well as spontaneously occurring tumors would be warranted for further understanding of these molecular changes. Additionally, since each tumor was sampled as a whole, factors such as tumor heterogeneity may possibly have played a role in these results. Future studies sampling multiple sites within each tumor may provide more information about variable mutation spectra and differential gene expression between HB and adjacent HCC. Using newer technologies, such as exome sequencing at high read depths may provide more evidence on the conservation of mutations between HB and HCC, and cell lineage tracing experiments may provide more definitive evidence on the origin of HB.
Footnotes
Acknowledgments
The authors would like to thank the NIEHS histology and immunohistochemistry laboratories and the NTP Archives for their technical expertise, and the CMPB pathologists and NTP toxicologists for helpful discussions.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Author Contribution
Authors contributed to conception or design (SB, AP, KS, SP, and MH); data acquisition, analysis, or interpretation (SB, AP, HN, YW, JF, HH, TT, MD, KS, SP, KG, DM, RS, and MH); drafting the manuscript (SB, AP, HN, KS, SP, and MH); and critically revising the manuscript (SB, AP, YW, JF, HH, TT, MD, KS, SP, KG, DM, RS, and MH). All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
