Abstract
The measurement of plasma microRNAs (miRNAs) and messenger RNAs (mRNAs) is the most recent effort to identify novel biomarkers in preclinical safety. These genomic markers often display tissue-specific expression, may be released from the tissues into the plasma during toxic events, change early and with high magnitude in tissues and in the blood during specific organ toxicities, and can be measured using multiplex formats. Their validation as biomarkers has been challenged by the technical difficulties. In particular, the concentration of miRNAs in the plasma depends on contamination by miRNAs originating from blood cells and platelets, and the relative fraction of miRNAs in complexes with Argonaute 2, high-density lipoproteins, and in exosomes and microvesicles. In spite of these hurdles, considerable progress has recently been made in assessing the potential value of miRNAs in the clinic, especially in cancer patients and cardiovascular diseases. The future of miRNAs and mRNAs as biomarkers of disease and organ toxicity depends on our ability to characterize their kinetics and to establish robust collection and measurement methods. This review covers the basic biology of miRNAs and the published literature on the use of miRNAs and mRNAs as biomarkers of specific target organ toxicity.
Introduction
MicroRNAs (miRNAs) are short (approximately 19–23 nucleotides) noncoding ribonucleic acid (RNA) molecules that bind to complementary sequences on target messenger RNA (mRNA) causing translational repression or target degradation and gene silencing. They were first discovered in
A biomarker is anything—diagnostic or surrogate—that may be used as an indicator of a particular biologic endpoint. In the field of toxicologic pathology, the term biomarker is applied to proteins, nucleic acids, or endogenous metabolites derived from body fluids that correlate with specific tissue damage, pathway modulation, or drug efficacy. It is implied that novel biomarkers or biomarker panels should perform or enable the panel as a whole to perform with increased specificity, sensitivity, predictivity, more robustly, or with increased correlation across species than current established biomarker/biomarkers for specific conditions/toxicities. For example, cardiac troponins are excellent translational biomarkers of acute myocardial necrosis (Clements et al. 2010; Berridge et al. 2009; Engle et al. 2009); therefore, the likelihood of identifying better markers than cardiac troponins for myocardial necrosis is limited. Indeed, the plasma miRNAs proposed as biomarkers of cardiac damage perform well or less well than cardiac troponins and yet have to go through rigorous validation (Meder et al. 2011; Widera et al. 2011; Gidlof et al. 2011). In contrast, serum creatinine and blood urea nitrogen are less sensitive renal biomarkers because they fail to identify early kidney damage. As a result, multiple candidate biomarkers of renal damage have been proposed and are currently being investigated (Sistare et al. 2010). Circulating miRNAs represent an opportunity to overcome some of the technologic limitations of current biomarkers.
miRNAs and miRNA clusters are often expressed in a tissue-, development-, or disease-specific manner and modify the expression of many genes, acting as a “master switch.” Their innate characteristics afford potential miRNA biomarkers a high level of sensitivity and specificity. Quantitation of miRNAs employs the sensitivity of the real-time PCR which eliminates the matrix effects and circumvents the need to develop specific ELISA-based methods for each marker. Also, circulating miRNAs offer the advantage over circulating mRNAs of being more stable, and therefore their measurement may be more reliable while providing data of similar mechanistic and translational value. Finally, the high level of conservation of miRNAs across species suggests that the identification of miRNA signatures of tissue damage will be conserved across species although comprehensive tissue distribution data are still lacking.
An important fraction of miRNAs and measurable amounts of mRNA circulate in plasma either freely or in exosomes. Exosomes are RNA- and protein-containing vesicles secreted by mammalian cells. Exosomal RNA is protected from nucleases and can be shuttled from cell to cell, which results in silencing specific mRNAs in recipient cells. While considerable progress has been made to identify plasma miRNA and mRNA signatures of cancer and disease (Mendrick 2011; Zen and Zhang 2010), advances in the identification of drug-induced genomic signatures of organ toxicity have been slower. Here, we review the current literature about the biology of miRNAs and discuss the use of circulating miRNAs and mRNAs as candidate biomarkers.
Nomenclature of miRNAs
Names for novel miRNA genes are provided by the miRBase Registry, an organization hosted and maintained in the Faculty of Life Sciences at the University of Manchester with funding from the Biotechnology and Biological Sciences Research Council (http://www.mirbase.org/). The names of miRNAs are assigned in numerical order after a manuscript describing their discovery is accepted for publication, and these new names are used in the final version of the manuscript (Griffiths-Jones et al. 2006).
miRNA genes are preceded with a three- to four-letter prefix to identify the species of origin (Table 1). For example, the prefix “rno” identifies
Nomenclature of microRNAs.
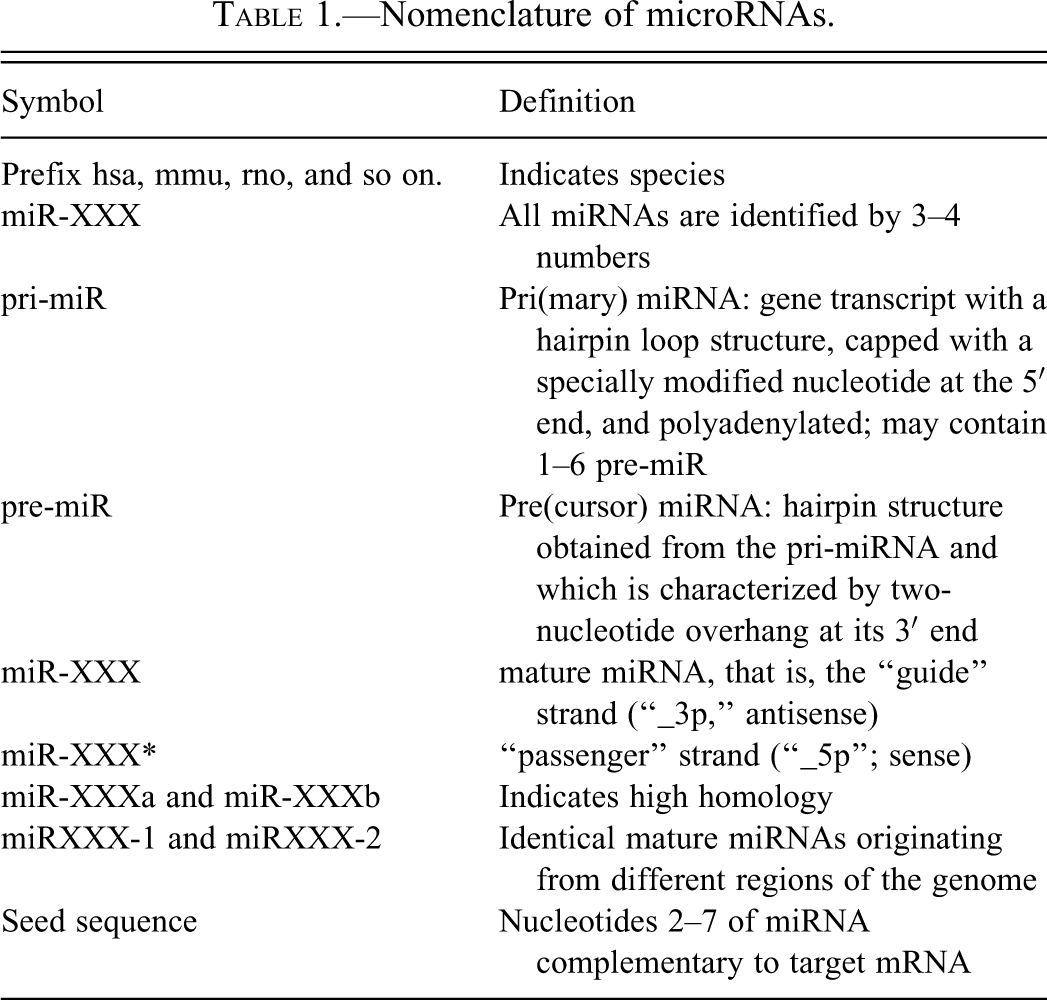

Letter suffixes identify mature miRNAs with high homology. Number suffixes identify mature miRNAs that originate from different regions of the genome but have the same sequence. The seed region is the oligonucleotides 2–7of the mature miRNA that are complementary with the target mRNA.
There are a few exceptions to the nomenclature. For example, the Let7 family was named Let for “Lethal” because knock-outs
miRNA Biosynthesis
miRNA genes are present in all human chromosomes except the Y chromosome (Kim and Nam 2006). To date, over 1,500 miRNA genes have been identified in man, 700 in mouse, and 400 in rat (http://www.mirbase.org/). Thus, miRNAs are one of the most abundant classes of regulatory genes in mammals. Based on computational methods, miRNAs were predicted to regulate 92% of mRNAs (Miranda et al. 2006).
The molecular events leading to the biosynthesis of miRNAs have been extensively reviewed (Figure 1; Faller and Guo 2008; Krol et al. 2010b; Murchison and Hannon 2004; Winter et al. 2009). Most miRNAs are coded by regions of the genome that are distant from known genes, which implies that they derived from independent transcription units. However, some miRNAs are intronic to specific genes, a situation that is best exemplified by the network of cardiac-specific miRNAs (van Rooij et al. 2006, 2009). Also, approximately half of miRNA genes comprise multiple exons with up to six mature miRNAs, and many miRNA genes are organized in “clusters” with arrangements and expression patterns implying transcription as a primary transcript with multiple exons. Examples of such clusters include the cluster miR-17-92, also known as oncomir-1, which was implicated in promoting proliferation, angiogenesis, cell survival, neoplastic transformation, and inhibition of differentiation during development (Olive, Jiang, and He 2010; Tanzer and Stadler 2004). Some of these clusters have tissue-specific expression, such as the cluster miR-183/96/182, which is expressed in the retina (Xu et al. 2007), and the cluster miR-208/499, which is expressed in the muscles (van Rooij et al. 2006, 2009).
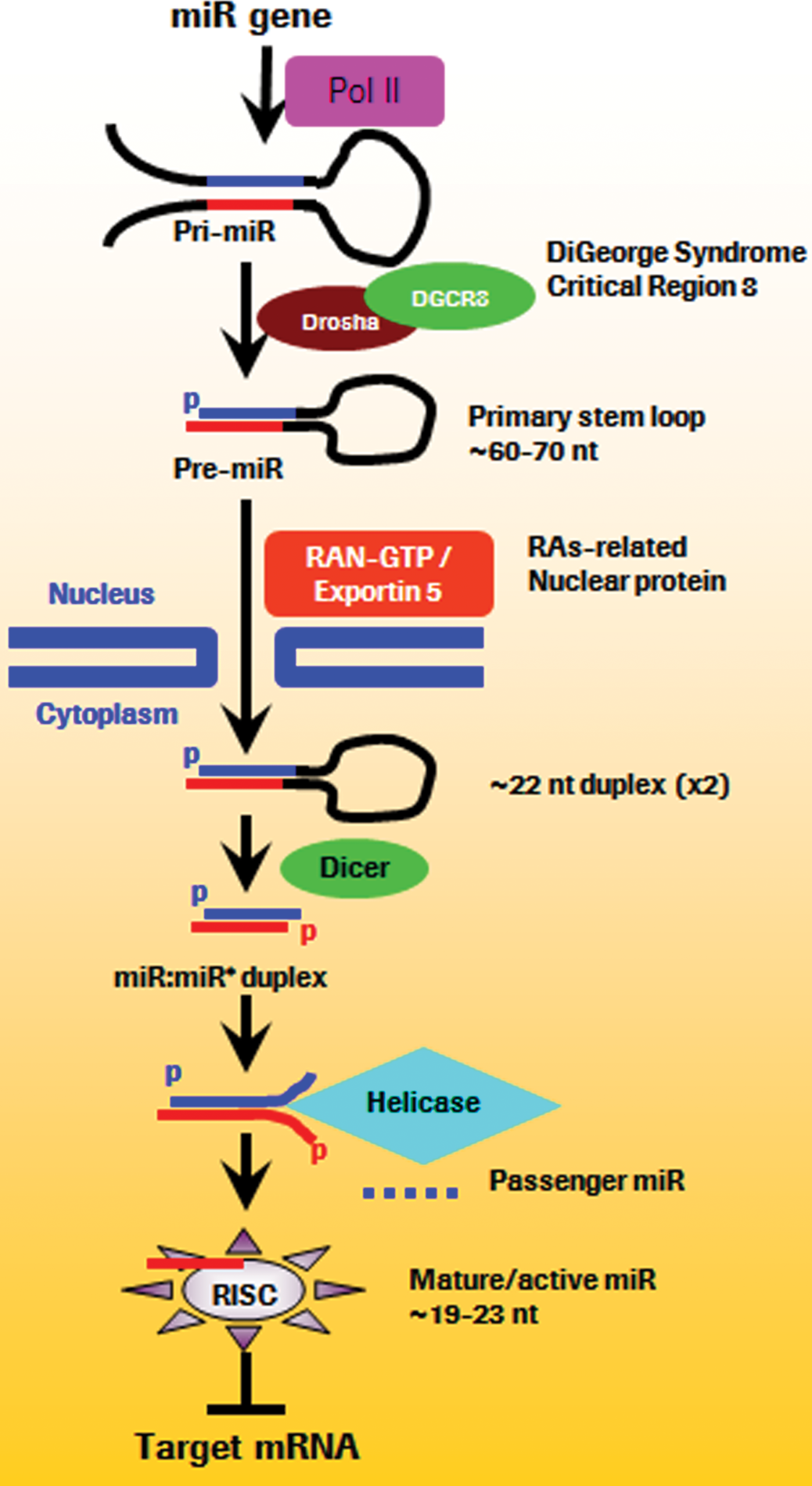
microRNA biosynthesis: microRNAs are transcribed by the RNA polymerase Pol2, and results in a pri-miR, which is modified similarly to protein-coding transcripts by addition of a 5′ cap and a 3′ poly-A tail. Within the nucleoli, pri-miRNA is cleaved by the endonucleases Droscha and DGCR8 in the pre-miRNA. Through the energy-dependent transporter Ras-related nuclear protein coupled with exportin 5, the pre-miR is exported into the cytosol where it undergoes further maturation. First, Dicer, an RNA (III) endonuclease, recognizes the pre-miRNA, which results in cleavage of the loop, leaving the 5′phosphate and a 2-3 nucleotides 3′ overhang. The miRNA duplex is then unwound into the mature single-stranded form, called the guide strand, which is antisense, and which integrates the RNA-induced silencing complex (RISC) where complementary mRNAs are degraded. The complementary miRNA strand, also called the passenger strand and designated “miR*,” which has a sense orientation, is degraded, a process which most likely occurs in the RISC and depends on the thermodynamic instability and weaker base-pairing of miR* relative to the mature miR. (Modified from Faller and Guo 2008; Krol et al. 2010b; Murchison and Hannon 2004; Winter et al. 2009.)
miRNA Biological Functions
miRNAs bind to complementary sequences on target mRNA transcripts within the RNA-induced silencing complex (RISC) causing translational repression or target degradation and gene silencing. The RISC is a multiprotein complex that uses a strand of miRNA to recognize and subsequently degrade complementary mRNA. The RISC may reside in the P-bodies (processing bodies; Liu et al. 2005) or in the GW bodies (Gibbings et al. 2009) named because they are the preferential site of localization of the 182 kDa glycine-tryptophan protein (GW182), which acts as downstream effectors in the repression. P-bodies and GW bodies may represent a single structure with many subtypes defined by the accessory proteins associated with the RISC (Kulkarni, Ozgur, and Stoecklin 2010). Although most of the biologic activities of miRNAs are considered to be cytoplasmic, the substantial amounts of miRNAs and Argonaute 2 (Ago2) present in the nuclei of mammalian cells may identify a role for miRNAs in transcriptional silencing (Kim et al. 2008). Ago2 is the catalytic component of RISC. It has two major domains: a PAZ domain that binds the 3′ end of the mature miRNA, and a PIWI domain that binds the 5′ end (Wang et al. 2010c). The second important component is DICER, which is responsible for cleavage and de-adenylation of the target mRNA.
The guide strand interacts with the mRNA in the RISC under three main modalities (Figure 2; [Bartel 2009; Brennecke et al. 2005]): (1) canonical recognition is based on perfect pairing between the target mRNA and the seed region of the miRNA, which is the oligonucleotides 2–7; (2) supplementary pairing occurs through complementarity in the 13 to 16 oligonucleotide region of the miRNA, which results in stronger affinity binding between the miRNA and the target mRNA; and (3) compensatory pairing may occur for slightly mismatched target mRNA and seed regions of the miRNA. Also, there is mounting evidence that miRNA–mRNA pairing occurs under other modalities, including modalities independent of the 3′ untranslated region of the mRNA and of the seed region of the miRNA (Thomson, Bracken, and Goodall 2011). This process of mRNA recognition and degradation by miRNAs in the context of the RISC is likely energy-dependent because it occurs in association with the mitochondria and the microtubule network (Huang et al. 2011). Multiple databases which are mostly based on computational data provide information on the predicted mRNAs targets of known miRNAs (Table 2). Efforts are in progress to integrate these databases into a single database (Mestdagh et al. 2011).
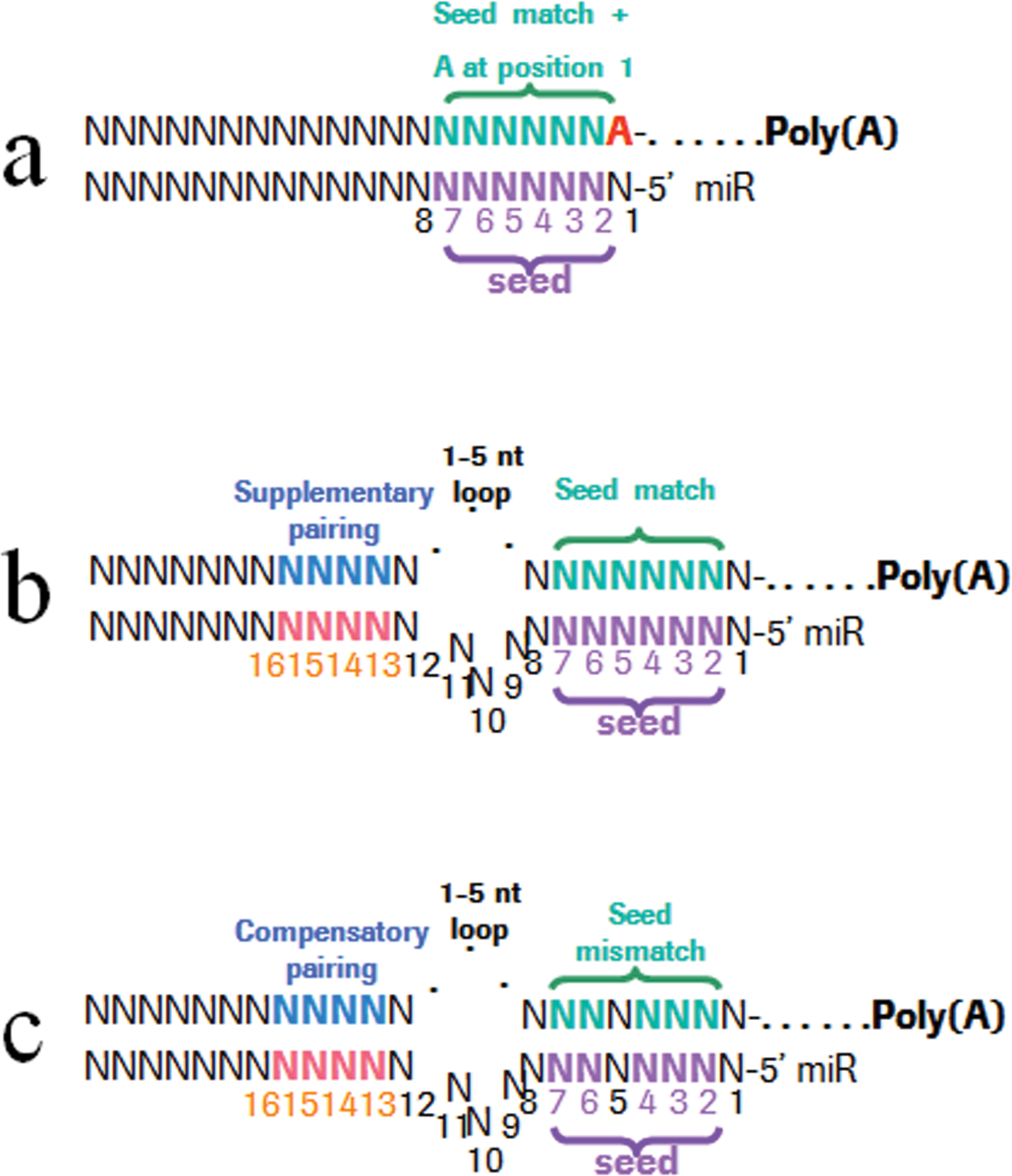
Recognition of target mRNA species by miRNAs occur under canonical (A), marginal (not illustrated here), supplementary (B), and compensatory (C) pairing modalities. The oligonucleotides 2–7 of the miRNA, which represent the seed region, are critical to the pairing. Additional affinity may be gained through pairing of the oligonucleotides in the 3′ region of the miRNA with the target mRNA. (Modified from Bartel 2009.)
Online and downloadable databases to predict mRNA targets of microRNAs.

In addition, miRNAs hybridize with mRNAs outside of the RISC complex, with the result of “withholding” specific mRNAs from the translation process. As a result of these multiple modalities of sequence recognition, an average miRNA may have hundreds of target mRNAs (Brennecke et al. 2005), which implies that the regulation of a mRNA target by a specific miRNA not only depends on the relative concentration of the target mRNA and of the effector miRNA but also depends on the concentration and availability of many other mRNAs which are less complementary. In summary, the relationship between miRNAs and mRNAs is a highly dynamic multicompartment kinetic model which results in spatiotemporal fine-tuning of multiple biologic processes. In addition to their presence in the cytosol, miRNAs were recently identified in mitochondria (Barrey et al. 2011; Bandiera et al. 2011). This finding suggests miRNAs may be involved in the energy balance of the cell at the mitochondrial level.
miRNAs are generally considered to be stable molecules, with half-lives of hours to days in the liver and the heart (Krol et al. 2010b). However, miRNAs in the retina and the neurons have considerably shorter half-lives with half-lives of 1 hr or less (Xu et al. 2007; Krol et al. 2010a). The degradation of miRNAs may be dependent on 5′-to-3′ exonuclease XRN-2 and the family of exoribonucleases encoded by the Small RNA Degrading Nuclease genes (Ramachandran and Chen 2008; Kai and Pasquinelli 2010), and the presence or absence of these proteins may provide another level of complexity contributing to the spatiotemporal dynamics of miRNA. In addition, target mRNA availability affects the release of miRNAs from Ago2, and therefore the availability of miRNAs for degradation by XRN2 (Kai and Pasquinelli 2010).
Biology of miRNAs and mRNAs in Body Fluids
Until recently, it was accepted that the half-life of RNA in the plasma was short due to the presence of potent ribonucleases (El-Hefnawy et al. 2004). Indeed, circulating RNAses degrade 99% of the exogenous RNA within 15 sec (Tsui, Ng, and Lo 2002). However, miRNAs remain stable after being subjected to severe conditions, which is likely the result of their packaging in macromolecular complexes (Arroyo et al. 2011; Turchinovich et al. 2011), their association with high-density lipoproteins (Vickers et al. 2011), and their distribution to circulating microparticles (CMPs) where they are protected from plasma endonucleases (Wang et al. 2009). In addition to protecting against degradation, the high-density lipoproteins function to upload and deliver operational miRNAs capable of effecting changes in gene expression in specific-recipient cell types.
Plasma CMPs originate from viable and degenerating nucleated and nonnucleated cells, including red blood cells and platelets. These CMPs are membrane-bound and measure 30 to 100 nm for exosomes, 0.1 to 1 µm for microvesicles which are released during sublethal cell damage (Hickman et al. 2010; Cocucci, Racchetti, and Meldolesi 2009), and 0.05 to 2 µm in diameter for apoptotic vesicles (Halicka, Bedner, and Darzynkiewicz 2000). Exosomes derive from the intracellular multivesicular bodies. In contrast, microvesicles and apoptotic vesicles are released from the surface of the plasma membranes. It is important to note that this nomenclature is not rigorously applied throughout the literature, but the key features to focus on are the origins, contents, and mechanisms of release of these cellular fragments (Thery, Ostrowski, and Segura 2009). The populations of CMPs may be separated based on their density using ultracentrifugation in sucrose gradients (Thery, Ostrowski, and Segura 2009), surface markers (Thery, Ostrowski, and Segura 2009; Orozco and Lewis 2010), microfluidic immunoaffinity (Chen et al. 2010), ExoQuick precipitation (Taylor, Zacharias, and Gercel-Taylor 2011), and microfiltration (Merchant et al. 2010). These different fractions of CMPs host different populations of mRNAs (Wetmore et al. 2010), and likely miRNAs as well. Differences in isolation methods are likely to account for some of the inconsistencies in the published literature about candidate biomarker miRNAs.
Among the CMPs, exosomes are of special interest as potential source of biomarker miRNAs because one of their functions includes intercellular transfer of miRNAs (Gibbings et al. 2009). Exosomes are membrane-bound vesicles of endocytic origin that are secreted by cells into the extracellular environment (Kosaka et al. 2010b; Cocucci, Racchetti, and Meldolesi 2009). Exosomes were identified in the early 1980s (Trams et al. 1981; Taylor, Homesley, and Doellgast 1980). Their biologic relevance was questioned until their release was associated with specific processes (Beaudoin and Grondin 1991). They were then primarily implicated in antigen presentation, as they often express the cell adhesion and costimulation proteins ICAM-1, CD86, CD63, and CD82, MHC class I and MHC class II (Thery, Zitvogel, and Amigorena 2002), and TNFR1 (Cocucci, Racchetti, and Meldolesi 2009). In addition, their lumen contains Annexin 2 and cytokines, and their membrane contains lysobisphosphatidic acid, cholesterol, and ceramide (Cocucci, Racchetti, and Meldolesi 2009).
The role of exosomes as carriers of mRNA and miRNAs was proposed in the mid-1980s (Bastida et al. 1984), but was broadly recognized only recently (Gibbings et al. 2009; Kosaka et al. 2010b). In experiments with mast cells, exosomes were shown to contain 1,300 mRNA species, many of which were not present in the cytoplasm of the donor cells (Valadi et al. 2007). The transfer of these exosomes to recipient cells results in translation of the mRNAs of the donor cells. The export of miRNAs from the cytoplasm of cells to the plasma in the form of exosomes is controlled by a specific ceramide-dependent mechanism (Kosaka et al. 2010b). These results imply that exosomes are involved in intercellular communication at the local level, a hypothesis which is gaining momentum with recent supportive evidences in infectious (Singh et al. 2011; Pegtel et al. 2010) and inflammatory diseases (Mittelbrunn et al. 2011), and in cancer (Mohla et al. 2011; Lee et al. 2011). RNA undergoing exosome-mediated transfer between cells has been denoted “exosomal shuttle” RNA (Valadi et al. 2007; Lotvall and Valadi 2007). Exosomes have been identified in numerous body fluid where they may function in intercellular signaling and may serve as diagnostic markers (Lasser et al. 2011; Keller et al. 2011). This suspected role of exosomes in intercellular communication has led to their investigation as potential delivery systems for small interfering RNAs and miRNAs (Quesenberry and Aliotta 2010; Zhang et al. 2009), including as anticancer agents (Morse et al. 2005; Chaput et al. 2003).
Little is known about the half-life and the mechanisms of clearance of exosomes and other CMPs. These mechanisms may rely on nonspecific clearance and specific uptake. The precedence of clearance of apoptotic cells and senescent red blood cells and platelets suggests that nonspecific clearance of exosomes may involve the organs with high clearance capabilities, such as the spleen, liver, kidney, lungs, and bone marrow, and the degree of sialylation, glycosylation, or phospholipid oxidation of their membrane (Gorgani and Theofilopoulos 2007; Pereira et al. 1999; Kiefer and Snyder 2000). However, specific mechanisms of uptake exist because exosomes shed by platelets interact with macrophages and endothelial cells but not with neutrophils (Losche et al. 2004). These specific mechanisms of uptake of the exosomes are energy-dependent (Escrevente et al. 2011). Upon uptake by a target cell, the exosome may fuse with the membrane of the target cell or may be endocytosed, which may result in their degradation in lysosomes or the release of their content in the cytosol of the target cell (Cocucci, Racchetti, and Meldolesi 2009).
In addition to packaging of m/miRNA in secretory vesicles, circulating miRNAs have been shown to be closely associated with proteins involved in mRNA degradation (Arroyo et al. 2011; Turchinovich et al. 2011). Following verification that purified miRNA was rapidly degraded in plasma but was protected by prior inhibition of RNase activity, further investigation utilizing differential centrifugation and size-exclusion chromatography revealed that most circulating miRNA is not present in exosomes but rather exists in complexes with the RISC-component protein Ago2. The RISC is necessary for cleavage of the target mRNA sequence, and integration of the miRNA target into this molecular complex also appears to protect the miRNA from degradation. While current research indicates that most circulating miRNA is likely found in the Ago2 RISC complex, it is likely that other proteins may also serve a protective/transport function for exported RNA molecules (Wang et al. 2010b).
Sample Collection, Processing, and Analysis Methods
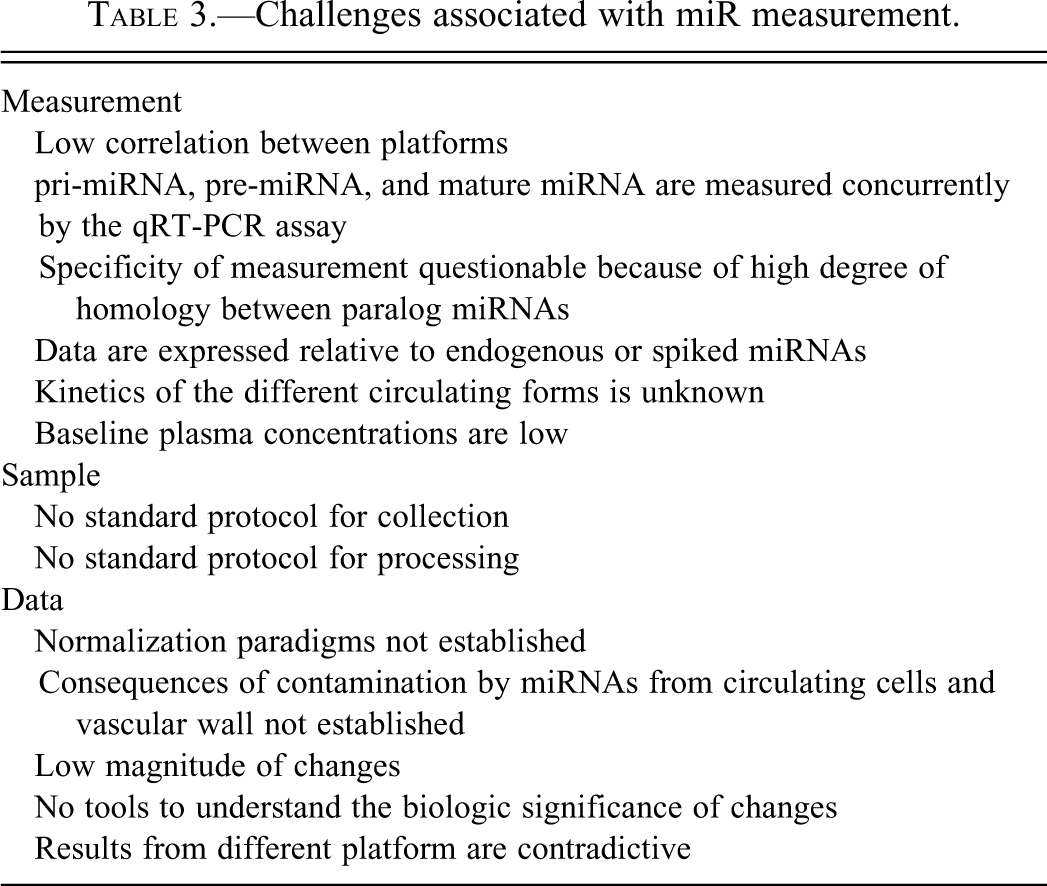
The reliability, quality, and standardization of the sample collection, extraction, and quantitive methods are critical toward the establishment of miRNA and mRNA biomarker signatures (Becker et al. 2010). It is important to recognize and understand the variables affecting the genomic expression being measured and minimize these technical variables (Table 3). These technical variables include differences in blood sampling methods, anticoagulants used, centrifugation, pipetting, sample preservation, sample processing, method of quantitation, normalization, and analytical methods.
Challenges associated with miR measurement.

Published information is lacking to help guide the sampling method. In our experience at Roche, mRNA measured on whole blood from retro-orbital bleeds is often contaminated by cells of the conjunctiva and the lachrymal glands. Similarly, concern may be raised about contamination of the sublingual bleeds by the oral mucosa and the salivary glands; of the jugular bleeds by the skin, adipose tissue, and vascular wall; and of the aortic bleeds by cells of the wall of the aorta. Specific experiments are needed to evaluate the impact of the sampling method on the circulating mRNA and miRNA profile.
miRNAs and mRNAs are present in all body fluids (Kosaka et al. 2010b; El-Hefnawy et al. 2004; Bianchi, Maron, and Johnson 2010; Li et al. 2004b; Weber et al. 2010). Although miRNA and mRNA profiles from whole blood have been published (Hausler et al. 2010; Tan et al. 2009; Keller et al. 2009; Li et al. 2004a; Wong et al. 2008), circulating RNAs in whole blood samples are impacted by the ratio of cells contained in the whole blood and by the time of storage of the whole blood. For example, miRNAs let-7b, miR-16, miR-7, and miR-145 are differentially expressed by platelets in response to storage at room temperature (Zimmerman and Weyrich 2008; Kannan et al. 2009). Similarly, white blood cells may become activated during storage (Hammerle-Fickinger et al. 2010). Therefore, the whole blood is not a promising biofluid to measure miRNA and mRNA profiles.
The most commonly used biofluids to measure circulating genomic biomarkers are the plasma and the serum. Plasma is generally preferred over serum which contains microparticles released during platelet activation (Heijnen et al. 1999; Sims et al. 1988; Kannan et al. 2009; Zimmerman and Weyrich 2008). However, plasma and serum miRNA profiles are highly correlated (Kroh et al. 2010). EDTA and citrate are the anticoagulants of choice because heparin inhibits the reverse polymerase transcriptase (Yokota et al. 1999). Erythrocytes contain significant amounts of miRNAs, and the presence of miR-20a, miR-106a, miR-144, and miR-185 in the serum indicates hemolysis (Sangokoya, LaMonte, and Chi 2010; Chen et al. 2008).
The conditions of storage and time elapsed between blood collection and plasma aliquot collection, and speed of centrifugation should be controlled and documented precisely because these factors may influence the release of microparticles from the circulating blood cells which are a major contributor to the circulating miRNAs (Chen et al. 2008) or may result in activation of the white blood cells and platelets (Hammerle-Fickinger et al. 2010; Duttagupta et al. 2011). Also, care must be taken during pipetting of the plasma not to contaminate the plasma supernatant with the cell pellet. In our experience at Roche, the plasma is obtained from blood collected in EDTA-coated tubes by a 10-min centrifugation at 1,900
A key component in obtaining accurate circulating genomic information is to ensure adequate preservation of the RNA. miRNAs are stable compared to mRNA, especially during storage in severe conditions such as boiling, very low or high pH, extended storage, and freeze thaws (Jung et al. 2010; Chen et al. 2008; Mitchell et al. 2008; Gilad et al. 2008). The stability of miRNAs is such that samples remain adequately preserved for analysis unrefrigerated in dried serum blots at room temperature for 5 months and at 37 degrees for 4 weeks (Kannan et al. 2009). Conversely, the stability of mRNA is less and differs between mRNA species (Tsui, Ng, and Lo. 2002; Holford et al. 2008). In-house and published information indicate that miRNAs and mRNAs are most stable when stored at −70°C to −80°C (Mraz et al. 2009), although this was questioned in one publication (Bravo et al. 2007).
The choice of the extraction kit to isolate miRNA and mRNA from serum/plasma will affect the experimental outcome. Kits/methods are optimized for the extraction of small RNAs either in conjunction with full length total RNA or as a fraction enriched for small RNAs. Currently available kits include mirVana™ miRNA isolation kit (Life Technologies, Austin, TX), miRNAeasy kit (Qiagen, Valencia, CA), Trizol® LS method (Life Technologies, Carlsbad, CA), the QIAamp® UltraSens™ Virus kit (Qiagen), and Exiqon’s Locked Nucleic Acid™ microarray platform (Vedbaek, Denmark; Mraz et al. 2009; Dovc-Drnovsek et al. 2008). The method used in our laboratory combines Trizol LS with column-based RNA isolation. Once the phases are separated with Trizol LS, the aqueous phase is placed over a mirVana miRNA isolation column and total RNA containing small RNA is isolated using the mirVANA™ Total RNA Extraction Kit followed by DNase treatment. Total RNA containing small RNA is then precipitated, and then quantitated using the Quant-iT™ RiboGreen® low range assay (Life Technologies). cDNA synthesis is performed using the Taqman® MicroRNA Reverse Transcription Kit (Life Technologies) followed by preamplification with Taqman® PreAmp Master Mix (Life Technologies). The yield of RNA from serum/plasma is typically low and is approximately 20 ng of total RNA from 200 µL of serum. The Ribogreen® method is preferred to ensure accurate total RNA quantitation. Other laboratories use spiked-in oligonucleotides to adjust for RNA recovery when concentrations are too low for spectrometry reading (Kroh et al. 2010). As previously stated, the RNA yield is low from serum samples which makes sample to sample and cross platform comparisons difficult.
Normalization and control of the variance are key when analyzing miRNA data sets and help control for factors that adversely affected the RT-PCR performance. The reliability, reproducibility, and interpretation of miRNA experiments can be improved by including a normalizing reference miRNA gene or set of genes in the assay (Benes and Castoldi 2010; Bissels et al. 2009). With that being said, there are currently no universally accepted miRNA reference genes or guidelines for normalizing miRNA expression data. This is concerning as inappropriate normalization can result in erroneous conclusions (Peltier and Latham 2008). U6B is one of the most common normalizing reference standards for miRNA expression studies in tissues but has not been validated for all tissue types and experimental treatments. In the plasma and the serum, miR-16 and miR-223 may be used for normalization because they are expressed at high levels and were relatively invariant across a large number of samples (Kroh et al. 2010). It is thought that normalizers for miRNA expression data are tissue- and species-specific just like normalizers for mRNA expression data, which makes choosing an appropriate reference gene difficult (Becker et al. 2010). There is a need to validate the expression stability of proposed normalizing reference candidates for each experiment to ensure that the data generated are accurately reflecting the expression change in the sample.
Many methods are available for measuring miRNAs, each with its advantages and disadvantages. Those methods include qRT-PCR, microarrays, Northern blots,
RT-PCR is attractive because it is specific for mature miRNAs, can discriminate among related miRNAs that differ by as little as one nucleotide, and is a sensitive method when incorporating a preamplification step (Chen et al. 2005). The cost- and labor-intensiveness of RT-PCR may be partly overcome using multiplex technologies such as the Fluidigm microfluidic technology (Jang et al. 2011). Using microarray technology allows for high-throughput global miRNA profiling but unlike RT-PCR is not quantitative and suffers from a lower sensitivity. The use of Northern blots for studying miRNA expression was the method first used to study miRNAs. Like microarrays, this method suffers from sensitivity issues and is not quantitative.
Data analysis for miRNAs is constantly evolving because the precise functions of miRNAs are not fully established and may differ among species, cell types, and development stages (Burchard et al. 2009; Rosa, Spagnoli, and Brivanlou 2009). Multiple web-based resources are available for miRNA data interpretation, including commercial databases (Table 2). In our experience, the same input will yield different results in these different databases vouching for the need of integrating multiple biological endpoints when developing novel biomarkers. These endpoints may include mRNA, immunohistochemistry, and
Considerations of miRNAs Evaluation as Biomarkers for Anticancer Drug Development
A key aspect of miRNAs is their potential to function as noninvasive biomarkers for a wide variety of diseases. Recognizing their positive attributes in stability, sensitivity, and specificity especially when compared to preexisting biomarkers, many investigators have examined the utility of miRNAs as predictor biomarkers to anti-cancer therapy. There is strong evidence that plasma miRNA profiles are powerful cancer biomarkers. Additionally, due to the highly conserved nature of miRNAs among mammals, it is reasonable to speculate that miRNAs found to be of value in preclinical species would have significant translational advantages in human beings relative to current antibody-based biomarker detection systems. However, while numerous studies have shown positive associations between altered expression of particular miRNAs and specific diagnostic and prognostic endpoints in cancer, much work needs to be done to verify these findings, as the history of “novel” cancer biomarkers is replete with high expectations followed by disappointing results (Diamandis 2010).
miRNAs are well positioned to play a role in the neoplastic process, as they are the key players in the fundamental cellular processes commonly dysregulated in neoplastic transformation such as progression through the cell cycle, programmed cell death, proliferation, and differentiation (Miska 2005). In support of a possible role of miRNAs in cancer progression, miRNA genes are commonly located in chromosomal regions often subject to rearrangements, deletions, and amplifications in cancer cells (Calin et al. 2004). miRNAs may function in cancer as oncogenes (so-called oncomirs) or as tumor suppressor genes. While the typical expectation is for increases in miRNA to result in downregulation of the mRNA target/targets, the converse has occasionally been shown to be true. This process occurs through miRNAs targeting gene regulatory sequences and is termed RNA activation (Portnoy et al. 2011).
Measurement of circulating miRNA has identified specific plasma miRNA profiles for a variety of neoplasms and important clinical endpoints such as survival (Brase et al. 2010; Heneghan et al. 2010; Lodes et al. 2009; Mitchell et al. 2008; Shen et al. 2011; Resnick et al. 2009; Hausler et al. 2010; Luo et al. 2011; Roth et al. 2011). The occurrence of specific miRNA profiles associated with tumor grade (Brase et al. 2011) and decreases in the circulating concentrations of specific miRNAs following treatment (Kosaka, Iguchi, and Ochiya 2010a; Ng et al. 2009) lend credit to the value of circulating miRNAs as biomarkers. Thus, plasma miRNAs may be envisioned as biomarkers to assess response to treatment in orthotopic and transgenic models of internal malignancies.
Specific plasma miRNA profiles have been associated with the formation of foci of hepatocellular alteration and hepatocellular carcinomas in the rat model of carcinogenesis induced by N-nitrosodiethylamine (Sukata et al. 2011). These results need to be confirmed in other models, especially since the miRNAs identified in this study were not those typically associated with liver cancer in man.
miRNAs as Biomarkers of Cardiovascular and Vascular Injury
Endothelial cells and plasma are in direct contact. Therefore, it is reasonable to assume that endothelial damage and the ensuing release of endothelial cell fragments will result in broad peripheral distribution of endothelial-specific miRNAs compared to damage to sequestered cells such as the neurons of the brain, germ cells of the testis, tubular cells of the kidney, or epithelial cells of the intestine. Similarly, miRNAs enriched in vascular smooth muscle cells are potential biomarkers for vascular damage (Cheng et al. 2009; Zhang 2009; Liu et al. 2009; Cordes et al. 2009).
In support of the hypothesis that miRNAs may be used as markers of vascular damage, the plasma concentration of many endothelial-specific miRNAs is decreased in patients with coronary artery disease (Fichtlscherer et al. 2010), diabetes mellitus (Zampetaki et al. 2010), and arteriosclerosis obliterans (Li et al. 2011). miR-126 is the most promising candidate biomarker of vascular injury and its functions have been extensively studied. They include regulation of leukocyte adherence and vascular inflammation, endothelial cell-to-cell contacts, and angiogenesis during development and ischemic conditions (Kuhnert et al. 2008; Wang et al. 2008; Schmidt et al. 2007; van Solingen et al. 2009; Harris et al. 2008; Wienholds et al. 2005). Other miRNAs, including let-7f, miR-15a, miR-21, miR-26a, miR-27b, miR-28-3p, miR-29b, miR-130a, miR-143, miR-210, miR-221, miR-222, miR-233, and the cluster miR-17-92 are central to angiogenesis, are highly expressed by endothelial cells or vascular smooth muscle cells, or have been described as dysregulated in the plasma of patients with vascular disease (Chen and Gorski 2008; Camps et al. 2008; Fasanaro et al. 2008; Kuehbacher et al. 2007; Bonauer, Boon, and Dimmeler 2010; Daubman 2010; Zampetaki et al. 2010). Some of these miRNAs may have value as biomarkers, too. Also, the specific miRNA profiles of specific endothelial cell types may be utilized to identify novel biomarkers (McCall et al. 2011).
In preclinical species, dysregulation of endothelial-specific miRNAs was confirmed in the Lepob mouse model of diabetes (Zampetaki et al. 2010). In addition, miR-134, miR-409-3p, and miR-511 have been identified as candidate biomarkers in the fenoldopam and dopamine rat model of mesenteric injury (Thomas et al. 2012).
miRNAs as Biomarkers of Specific Organ Toxicity
The possible use of miRNAs as biomarkers of specific organ toxicity relies on a limited number of publications. The promise of these publications resulted in research throughout the pharmaceutical industry to confirm and expand on these results. Part of this research is coordinated by the Predictive Safety Testing Consortium (https://pstcworkspace.c-path.org/; Roberts 2009) and the Health and Environmental Science Institute (http://www.hesiglobal.org/).
The first efforts to identify circulating miRNAs as biomarkers of specific organ toxicity were applied to the liver. In the first published study on this topic, miR-122 and miR-192 levels were increased in liver and plasma of mice 24 hr after the administration of acetaminophen (Wang et al. 2009). These two miRNAs were more sensitive and more specific for liver injury than ALT and were also increased in human patients with acute liver injury (Starkey Lewis et al. 2011). These results were confirmed with the CCl4 and ethanol models of hepatotoxicity (Laterza et al. 2009; Zhang et al. 2010). Most of the miRNAs with decreased expression in liver were increased in plasma, a process that was attributed to the cellular damage in the liver. In addition, miRNAs thought at the time to be specific of the brain, heart, lung, and kidney were increased in the plasma, which the authors attributed to acetaminophen toxicity in these organs, too. In another publication, urinary miRNA changes in rats administered acetaminophen and CCl4 were attributed to primary liver toxicity (Yang et al. 2012).
The specificity of miRNAs was also evaluated (Table 4; Laterza et al. 2009). miR-122, miR-133a, and miR-124 are specific to the liver, muscle, and brain, respectively. The study supported the hypothesis that miR-122 is more specific than ALT and more sensitive than histopathology to identify liver damage. These results were confirmed in another study where miR-122 was identified as a sensitive and specific marker of liver damage in man with hepatitis B infection or alcohol abuse, and in the mouse models of liver injury caused by
Plasma microRNAs and specificity for organ damage.
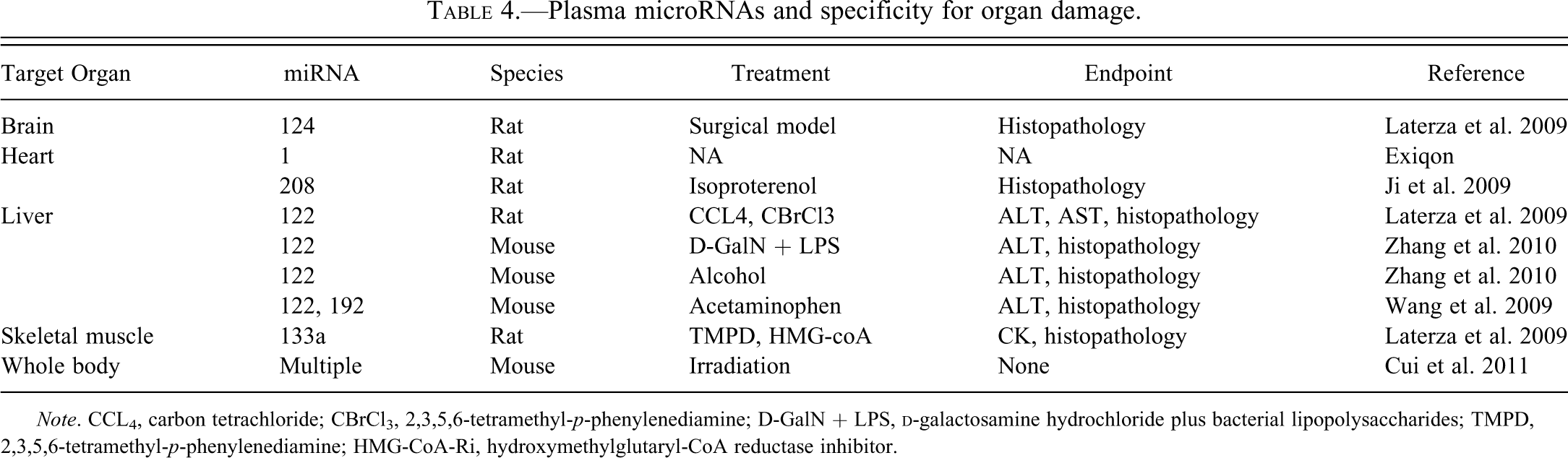
Finally, circulating miRNA profiles that accurately identified the level of irradiation were identified 6 and 24 hr after acute irradiation in mice (Cui et al. 2011). Unfortunately, hematology and histopathology were not assessed to identify possible correlates.
Circulating mRNAs as Biomarkers of Specific Organ Toxicity
The field of circulating mRNAs as candidate biomarkers was muddied by the “sentinel principle,” that is, the concept that blood cells can act as sentinels of disease and their mRNA profile could be used for the diagnosis/prognosis of disease (Liew et al. 2006). Based on the “sentinel principle,” a considerable amount of work was expanded to identify peripheral whole blood mRNA biomarkers in patients and in preclinical species. Yet, most of these efforts in man have been fruitless resulting in a limited number of often flawed publications on this topic (reviewed in Mendrick 2011). In particular, the proposed mRNA and miRNA markers of myocardial infarction are unlikely to compare favorably with cardiac troponins, which are sensitive and specific markers of cardiac damage (Muller et al. 2011; Liangos et al. 2010). The proposed whole blood signatures for autoimmune diseases, Alzheimer’s disease, and sepsis are either preliminary results or did not lead to the development of accepted clinical biomarkers (Reynier et al. 2010; Batliwalla et al. 2005; Tan et al. 2006; Grunblatt et al. 2009; Booij et al. 2011; Sutherland et al. 2011). In our laboratory, the attempt to identify a whole blood transcriptomic signature of endothelial damage in four rat models was inconclusive because of the preponderance of mRNA shifts reflecting the shifts in the major populations of white blood cells (unpublished data).
The inability to identify whole blood mRNA biomarkers was a disappointment because the breadth of knowledge for mRNAs is considerable compared to that for miRNAs, and the availability of mRNA biomarkers would provide insights into the pathobiology of diseases. Possible causes for this inability to identify whole blood mRNA biomarkers are many, and include the presence of large numbers of potent circulating ribonucleases resulting in the rapid clearance of most circulating mRNAs and DNA; contamination of the circulating mRNA by mRNA from circulating and vascular wall cells; and the abundance of globin mRNA and mRNAs from the circulating cells.
These possible issues were elegantly circumvented by our Pfizer colleagues by measuring mRNA in the plasma and in circulating microvesicles in three rat models of liver toxicity. The work of Kudo et al. identified increases in plasma albumin mRNA in the bile duct ligation model (Kudo et al. 2008). The work of Wetmore and Vrees (2010) identified hepatic microvesicles and their transcriptomic profile as early biomarkers in the acetaminophen model and the
Conclusion
The difficulty to establish reliable standardized methods for sample collection and processing has hampered our ability to identify and validate miRNA and mRNA biomarker profiles. A few publications have paved the route to success, which promises to be labor-intensive (Wetmore et al. 2010). In this endeavor, the collaborative consortia across the pharmaceutical industry and academia have a critical role (Roberts 2009).
Footnotes
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The authors received no financial support for the research, authorship, and/or publication of this article.
