Abstract
Objective
Orthodontic tooth movement (OTM) relies on mechanotransduction and inflammatory responses within the periodontal ligament (PDL). This study aimed to elucidate the role of transient receptor potential vanilloid 4 (TRPV4) in mediating mechanical force-induced pyroptosis in human PDL (hPDL) cells and its functional impact on osteoclast activation during OTM.
Methods
hPDL cells were isolated from premolars extracted for orthodontic reasons and characterized for stem cell properties (CD73+/CD90+/CD105+; CD34−/CD45−). An in vitro OTM model was established by applying compressive force (0–2.0 g/cm2). Pyroptosis was assessed via qRT-PCR, Western blot, and immunofluorescence for NLRP3, caspase-1, GSDMD, and IL-1β. Osteoclast differentiation was evaluated in hPDL-PBMC co-cultures using TRAP staining, ELISA (RANKL/OPG), and gene expression analysis (CTSK/TRAP). TRPV4 activation was modulated using agonist GSK1016790A.
Results
Compressive force (1.5 g/cm2, 24 h) significantly upregulated pyroptosis markers (NLRP3, cleaved caspase-1, N-GSDMD, IL-1β) and inflammatory mediators (COX2, TNF-α, IL-6, and IL-8). TRPV4 activation was mechanosensitive and enhanced by GSK1016790A, which amplified pyroptosis and subsequent RANKL secretion. In co-cultures, hPDL pyroptosis promoted osteoclast differentiation, evidenced by increased TRAP + multinucleated cells and elevated CTSK/TRAP expression.
Conclusion
Mechanical force activates TRPV4 to drive NLRP3/caspase-1-dependent pyroptosis in hPDL cells, which accelerates osteoclastogenesis via RANKL signaling. Targeting TRPV4-pyroptosis axis may represent a novel strategy to optimize orthodontic treatment efficiency.
Introduction
Orthodontic Tooth Movement (OTM) is a critical process in orthodontic treatment that involves the continuous application of mechanical forces to facilitate the movement of teeth within the alveolar bone, thereby improving occlusion, function, and aesthetics.1,2 The periodontal ligament (PDL), a specialized connective tissue between teeth and bone, serves as the primary mechanotransduction hub during OTM.3,4 The PDL plays a vital role in tooth support, force distribution during mastication, neural feedback, tooth eruption regulation, and OTM itself. 5 It comprises various cell types, including fibroblasts, endothelial cells, osteoblasts, odontoblasts, and osteoclasts, which collectively maintain the tissue characteristics and functions of the PDL. 6 During OTM, the PDL undergoes a series of biomechanical and biochemical responses that guide tooth position changes. 7 Given the prolonged duration of orthodontic treatment, there is a risk of periodontal tissue damage and discomfort. 8 Therefore, exploring novel biological pathways to accelerate tooth movement while mitigating periodontal tissue damage has become a focal point in modern orthodontic research.
During orthodontic treatment, PDL cells respond to changes in mechanical forces by promoting inflammatory responses, matrix remodeling, and cell death, thus participating in the remodeling of alveolar bone and tooth movement.9,10 While inflammation and apoptosis in PDL cells during OTM are well-documented,8,11 the role of pyroptosis—a highly inflammatory form of programmed cell death characterized by gasdermin-D (GSDMD) cleavage, membrane pore formation, and IL-1β/IL-18 release 12 —remains poorly defined. Recent evidence suggests pyroptosis may accelerate bone remodeling by amplifying local immune responses, 10 yet its mechanistic regulation by orthodontic forces is unknown. This knowledge gap limits the development of targeted strategies to optimize OTM efficiency. Notably, a recent study has highlighted that mechanosensitive signaling pathways can bridge mechanical stimuli with pyroptotic cell death in periodontal tissues, emphasizing the need to dissect specific molecular mediators involved in this process. 13
Transient receptor potential (TRP) channels are key mechanosensors in PDL cells. 14 Among them, Transient Receptor Potential Vanilloid 4 (TRPV4) is a non-selective cation channel widely expressed in various cell types, including PDL cells.15,16 TRPV4 channels exhibit high sensitivity to multiple stimuli, such as mechanical forces, temperature changes, and osmotic pressure, and have been confirmed to play a critical role in cellular mechanosensation and physiological responses. 17 TRPV4 was prioritized over other mechanosensitive channels (e.g. Piezo1/2) due to its established role in PDL mechanotransduction, its sensitivity to compressive forces in periodontal tissues, and its direct link to inflammasome activation in inflammatory pathways.18–21 Activation of TRPV4 not only regulates cellular mechanosensation but may also promote tissue remodeling by modulating cell survival, apoptosis, and immune responses. 22 Although previous studies have indicated the role of TRPV4 in orthodontics, 20 the specific mechanisms, particularly whether it accelerates tooth movement through pyroptosis, remain unclear.
Therefore, this study aims to investigate the effects of pyroptosis on PDL cells and the relationship between TRPV4 channel activation and the promotion of pyroptosis in PDL cells, as well as to assess the impact of this process on tooth movement during orthodontic treatment. In this research, the TRPV4 agonist GSK1016790A 11 will be used for the first time in the context of pyroptosis-related studies during OTM. We hope that this research will provide a new biological basis for orthodontic therapy and potential therapeutic strategies for accelerating tooth movement in clinical practice.
Materials and method
Isolation of human PDL (hPDL) cell
hPDL tissue was obtained from the middle third of the roots of premolars extracted from three volunteers (2 females, 1 male; ages 18–25 years; premolars extracted for orthodontic reasons), whose teeth were free of decay. The clinical procedure was approved by Changsha Stomatological Hospital (CSH-2022-038), and written informed consent was obtained from all participants. This study was conducted in accordance with the Helsinki Declaration. Briefly, the periodontal ligament scraped from the root surfaces was digested at 37°C with a mixture of 3 mg/mL type I collagenase and 4 mg/mL dispase II for 1 h to facilitate cell dissociation. A single-cell suspension was then obtained and cultured in α-MEM supplemented with 20% fetal bovine serum (FBS) and 1% penicillin/streptomycin. After 3 days of adherence, the primary cells were digested and further cultured in six-well plates for subsequent experiments. The culture medium for the isolated cells was replaced with DMEM containing high glucose, 10% FBS, and 50 mg/L vitamin C, maintained in a humidified environment at 37°C with 5% CO2. This study utilized cells from passages 3 to 5. In the study of TRPV4 signaling pathway, the TRPV4 agonist GSK1016790A (10 μmol/L, MCE, 942206-85-1) was applied to hPDL for 1 h, followed by mechanical loading (1.5 g/cm2) for 24 h. 13 For experiments involving GSK1016790A, control groups indicated hPDL cells with no treatment.
Mechanical loading on hPDL cells in vitro
The previously isolated hPDL cells were digested with trypsin and centrifuged at 300 g. A total of 90,000 cells were seeded into six-well plates and cultured for 4-day until they reached 90% confluence. After 24 h of incubation, compressive loading was applied using glass layers and 50 mL plastic tube caps containing weighted metal balls.23,24 Compressive forces of 1.5 g/cm2 were applied to the PDL progenitor cells at different time points (0, 24, and 48 h), or varying compressive forces (0, 0.5, 1.0, 1.5, and 2.0 g/cm2) were applied to the hPDL cells for 24 h.13,20 Accordingly, the control groups were defined as follows: for the time-course experiments, cells harvested at the 0 h time point under 1.5 g/cm2 served as the control; for the dose-response experiments, cells cultured under identical conditions but without the application of any compressive force (i.e. 0 g/cm2) constituted the control. The hPDL cells were then collected for subsequent analyses. Cellular morphological changes were observed using an Olympus IX73 inverted optical microscope (Olympus Corporation, Tokyo, Japan) equipped with an Olympus DP74 digital camera. Images were acquired using Olympus CellSens Dimension software (version 1.18).
Flow cytometry analysis
Flow cytometry was employed to identify the expression of specific stem cell antigens in hPDL cells. Briefly, 2.5 × 105 hPDL cells were treated with trypsin and resuspended in FACS buffer, followed by centrifugation at 300 g for 5 min at 4°C. The HPDL cells were then incubated with primary antibodies against CD34 (1:11550, abcam, ab81289), CD45 (Abcam, ab40763, 1:15), CD73 (Abcam, ab317462, 1:50), CD90 (Abcam, ab307736, 1:500), and CD105 (Abcam, ab2529, 1:10) for 30 min. After incubation, the cells were centrifuged again at 300 g for 5 min. Analysis was performed using BD CellQuest Pro software.
QRT-PCR
The hPDL cells were processed with TRIzol reagent (Invitrogen, CA, USA) to extract total RNA. Following the manufacturer's instructions for isolation, RNA quantification was performed using NanoDrop, measuring absorbance at 260 and 280 nm to ensure an A260/A280 ratio between 1.8 and 2.0. Subsequently, cDNA was synthesized using the PrimeScript® RT reagent kit (Takara, Shiga, Japan). QRT-PCR was conducted using the ABI Prism 5700 Sequence Detection System (Applied Biosystems). GAPDH served as the internal control gene. Relative gene expression was calculated using the comparative CT (2−ΔΔCT) method. The primers are listed in Table 1.
The primer sequences.

ELISA assays
According to the manufacturer's instructions, the ELISA kits (Jianheng Bioengineering, Nanjing, China) were used to measure the concentrations of COX2, TNF-α, IL-6, IL-18, and RANKL in the cultured supernatants. Results were determined by comparing the samples with the standard curve generated by the kit. All samples and standards were tested in triplicate.
Western blot
The hPDL cells were lysed on ice for 30 min using a protein extraction buffer containing RIPA, NaF, and PMSF. The collected proteins were denatured in a water bath at 95°C for 10 min and then centrifuged at 12,000 r/min for 10 min at 4°C. The supernatant was transferred to a new tube. Equal amounts of protein were loaded onto a gel for separation by SDS-PAGE. The proteins were then transferred to a PVDF membrane (Millipore) and blocked with 5% non-fat milk or bovine serum albumin, followed by incubation with primary (incubated overnight at 4°C) and secondary antibodies (incubated for 1 h at room temperature). The primary antibody used was as follows: NLRP3 (Abcam, ab263899, 1:1000), Caspase-1 (Abcam, ab207802, 1:1000), Cleaved Caspase-1 (Cl-Casp-1, Invitrogen, # PA5-99390, 1:500), GSDMD (Abcam, ab210070, 1:1000), N-terminal GSDMD (N-GSDMD, Abcam, ab215203, 1:1000), IL-1β (Abcam, ab283818, 1:1000), Cleaved IL-1β (Cl-IL-1β, Invitrogen, PA5-105048, 1:1000), RANKL (Abcam, ab45039, 1:1000), OPG (Abcam, ab73400, 1:1000), TRPV4 (Abcam, ab307444, 1:1000), GAPDH (Abcam, ab8245, 1:1000). Protein levels were normalized using glyceraldehyde-3-phosphate dehydrogenase (GAPDH) as the internal control. After electrophoresis and transfer, bands were visualized using enhanced chemiluminescence, and densitometric analysis was performed using ImageJ software (NIH). The relative expression of target proteins was calculated as the ratio of the gray value of the target protein band to that of the GAPDH band in the same sample.
Osteoclast activation assay and TRAP staining
Human peripheral blood mononuclear cells (hPBMCs) were isolated from peripheral blood samples collected from healthy, consenting volunteers (with no active systemic or periodontal diseases). To account for donor variability, hPBMCs from each individual donor were processed and analyzed separately in independent experiments; cells were not pooled across donors. The hPDL progenitor cells, at a concentration of 5 × 10^3 cells/mL, were seeded into 24-well plates and co-cultured with hPBMCs at 1 × 10^6 cells/mL. The culture medium was supplemented with macrophage colony-stimulating factor (MCSF) at 30 ng/mL and soluble receptor activator of nuclear factor-κB ligand (sRANKL) at 50 ng/mL, lasting for 14 days. After a 14-day co-culture period, the cells were fixed and stained for tartrate-resistant acid phosphatase (TRAP) activity using an acid phosphatase assay kit (387A-1KT; Sigma). Briefly, cells were fixed in 4% paraformaldehyde (PFA) for 10 min, followed by fixation in a 50:50 mixtures of acetone and ethanol for 1 min. The cells were then air-dried and stained for TRAP according to the previously described protocol. 24 The control groups for the co-culture system indicated hPBMCs co-cultured with hPDL cells without mechanical loading.
Immunofluorescence assay
The hPDL cells were fixed in 4% paraformaldehyde and permeabilized at room temperature with 0.1% Triton X-100 for 10 min. Subsequently, the cells were incubated with 5% bovine serum albumin (BSA) at room temperature for 1 h, followed by incubation with TRPV4 antibody (Abcam, ab39260, 5 µg/ml) and Caspase-1 antibody (Invitrogen, PA587536, 1:50) overnight at 4°C. After thorough washing, the cells were incubated with fluorophore-conjugated secondary antibodies in the dark at room temperature for 1 h. The nuclei were counterstained with DAPI (Invitrogen, D1306, 1:1000). The caspase-1 immunofluorescence staining intensity was quantified using ImageJ software (NIH). Following imaging with a laser scanning microscope (Zeiss), three random fields per sample were selected for analysis. The mean fluorescence intensity (MFI) of caspase-1 within the cytoplasmic region of individual cells was measured after background subtraction, with DAPI counterstaining used to define cellular boundaries. Imaging was performed using a Zeiss LSM 880 laser scanning confocal microscope (Carl Zeiss AG, Oberkochen, Germany) equipped with a Zeiss Plan-Apochromat 63×/1.4 oil immersion objective. Fluorescence signals were detected using a Zeiss GaAsP detector, and images were acquired and processed using Zeiss ZEN 3.2 (Blue Edition) software.
Statistical analysis
Statistical analysis was conducted using GraphPad Prism 9.0 software. The data are expressed as mean ± SD. Specified 3 biological replicates per experiment were performed. Differences between two independent groups were evaluated using a two-tailed, unpaired Student’s t-test. Differences between multi-group comparisons were evaluated using a One-way AVOVA with Tukey's post hoc test. A p < .05 was considered statistically significant.
Results
Extraction and isolation of PDL cells for in vitro modeling of OTM
PDL stem/progenitor cells are the primary cells responding to mechanical forces that lead to OTM. Therefore, we first isolated stem cells from primary human periodontal ligaments (hPDL) and cultured them in vitro. Flow cytometry was used to assess the proportion of specific stem cell markers on the surface of hPDL cells, evaluating their stemness. All donor cells expressed specific markers (CD34−, CD45−, CD73+, CD90+, and CD105+), which are characteristic of pluripotent adult mesenchymal stem cells

Isolation of hPDL cells for in vitro modeling of OTM. (A) Flow cytometry was used to detect specific stem cell antigens on the surface of primary human PDL (hPDL) cells. (B) The mRNA levels of cell inflammation-related factors COX2, TNFα, IL6, and IL18 at different time points (0–48 h). (C) Changes in cell secretion of inflammation-related factors COX2, TNF-α, IL-6, and IL-18 at different time points (0–48 h). N = 3 biological replicates, technical triplicates per condition. Data analyzed by one-way ANOVA with Tukey's post hoc test. *p < .05, **p < .01, ***p < .001 between groups.
The hPDL cells exhibit pyroptosis under compressive forces
The induction of pyroptosis in hPDL progenitor cells under mechanical stimulation was further investigated. Compression force was applied to hPDL progenitor cells in vitro. The qPCR detection revealed that the mRNA expression of pyroptosis markers (NLRP3, Caspase-1, GSDMD, IL1β) significantly increased over time (Figure 2(A), p < .001). Similarly, WB detection of protein expression of pyroptosis markers (NLRP3, Caspase-1, Cl-Casp-1, GSDMD, N-GSDMD, IL-1β, Cl-IL-1β) showed a comparable trend (Figure 2(B), p < .001). Next, the effects of different compressive force (0, 1.0, 1.5, and 2.0 g/cm2) on pyroptosis in hPDL cells were examined. As shown in Figure 2(C), WB analysis demonstrated that the expression of pyroptosis markers increased with pressure, reaching a peak at 1.5 g/cm2. Likewise, real-time PCR analysis of pyroptosis marker mRNA levels under varying mechanical pressures revealed a similar trend (Figure 2(D), p < .001). Hence, a force of 1.5 g/cm2 was applied for 24 h as the subsequent compressive forces modeling condition. Furthermore, optical microscopy (OM) images showed blurred cell contours, large vacuoles, and swollen, flattened cells, characteristic of pyroptotic morphology in compressive forces (CF) modeling group (Figure 2(E)). Overall, these findings reveal that compressive force induces pyroptosis in hPDL cells in vitro.
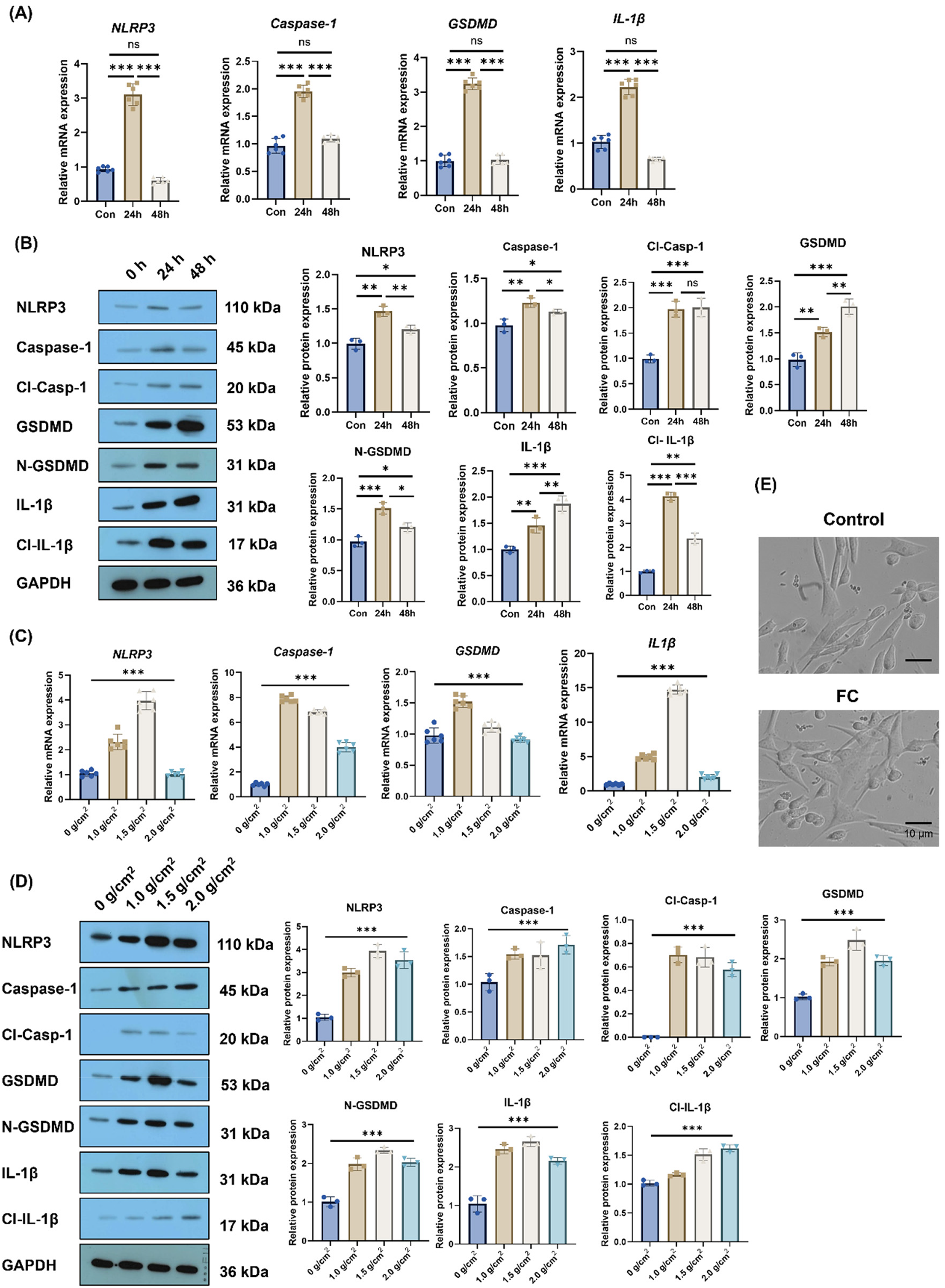
The hPDL cells exhibit pyroptosis under compressive forces. (A) PCR was used for detection of mRNA levels of pyroptosis markers (NLRP3, Caspase-1, Cl-Casp-1, GSDMD, N-GSDMD, IL1β, Cl-IL1β), which increased with time (0, 24, and 48) after modeling. (B) WB was used for detection of protein level of cell pyroptosis markers (NLRP3, Caspase-1, GSDMD, and IL-1β) over time (0, 24, and 48) after modeling. (C) PCR detection of mRNA levels of pyroptosis markers with changes in pressure after modeling (0, 1.0, 1.5, and 2.0 g/cm2). (D) WB was used for detection of protein level of cell pyroptosis markers, which increased with pressure after modeling (at 0, 1.0, 1.5, and 2.0 g/cm2). (E) Microscopic observation of morphological changes via optical microscopy (OM) in control group and compressive forces (CF) group (1.5 g/cm2, 24 h). Scale bar = 10 μm (original magnification: 400×). N = 3 biological replicates, technical triplicates per condition. Differences between the two groups were evaluated using a two-tailed independent Student's t-test. Differences between multi-group comparisons were evaluated using a one-way AVOVA with Tukey's post hoc test. *p < .05, **p < .01, ***p < .001 between groups.
Force-induced hPDL cells pyroptosis promotes osteoclast activation
To investigate the relationship between pyroptosis and osteoclast activity, hPDL progenitor cells were co-cultured with peripheral blood mononuclear cells (PBMCs), and osteoclast markers or morphological changes were observed. As shown in Figure 3(A), compared to the control group, the expression of the osteoclast activation marker RANKL mRNA and the RANKL/OPG ratio was significantly increased in the OTM modeling group under compressive force (1.5 g/cm2) (p < .001), while OPG expression remained unchanged. Similarly, protein expression level of RANKL was significantly upregulated in the OTM group versus the control group (Figure 3(B), p < .001). Furthermore, ELISA assays showed increased secretion of RANKL in the OTM modeling group (Figure 3(C), p < .001). Notably, the number of TRAP + osteoclasts was significantly higher in the OTM modeling group (Figure 3(D), p < .001). Additionally, mRNA expression level of cathepsin K (CTSK) and TRAP was significantly elevated in osteoclasts (Figure 3(E), p < .001). These data suggest that mechanical force enhances osteoclast activity in hPDL progenitor cells.
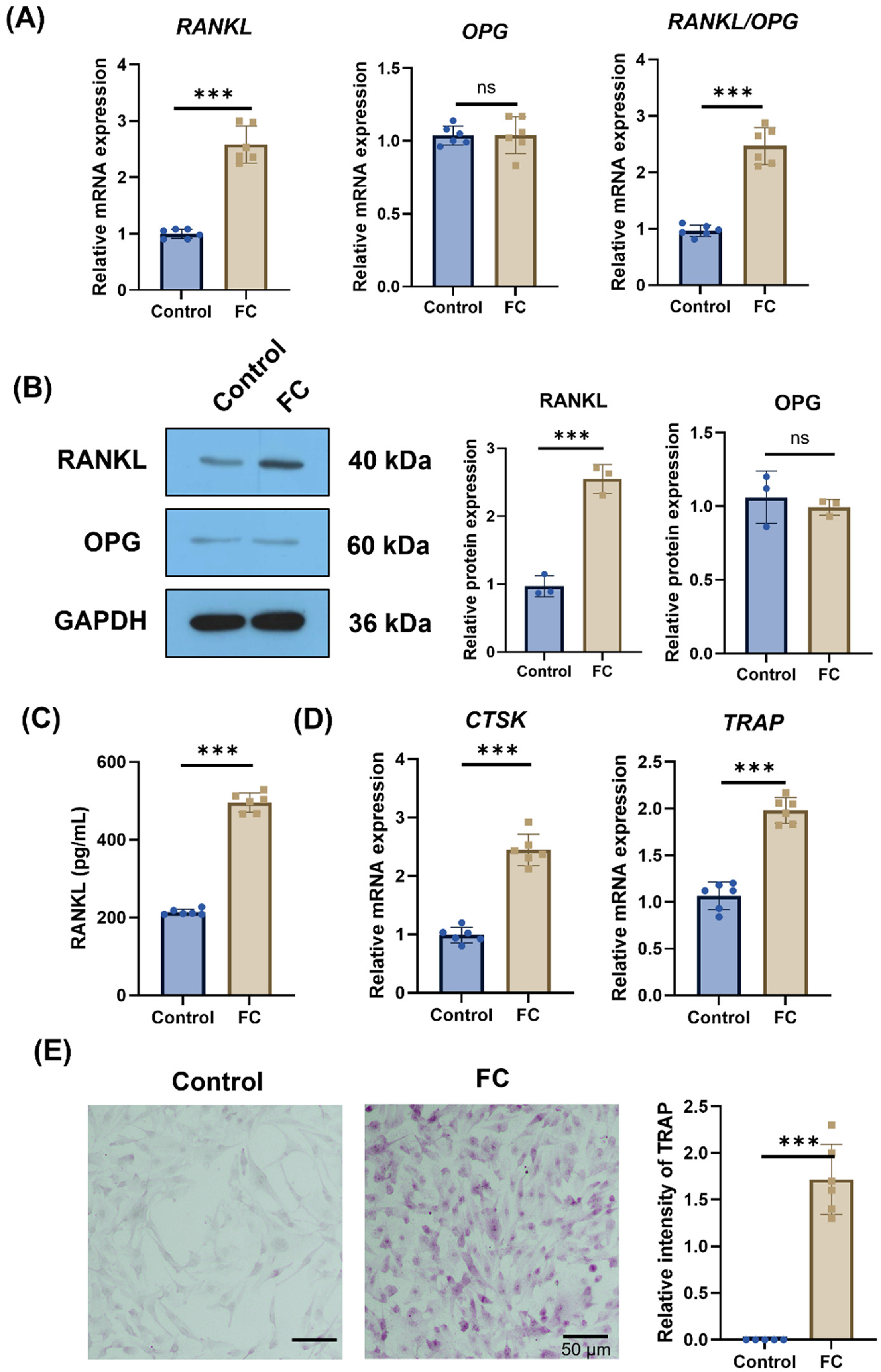
Force-induced hPDL pyroptosis promotes osteoclast activation. (A) PCR was used to detect changes in mRNA levels of RANKL, OPG, and RANKL/OPG in hPDL cells. (B) WB was used to detect changes in RANKL/OPG protein, a marker of osteoclast activation, in both control and modeling groups. (C) ELISA detection of RANKL secretion concentration. (D) PCR detection of mRNA expression changes of TRAP and CTSK in PBMC cells. (E) TRAP staining was used to detect the number of osteoblasts in the control and modeling groups. Scale bar = 50 μm (original magnification: 200×). N = 3 biological replicates, technical triplicates per condition. Differences between the two groups were evaluated using a two-tailed independent Student's t-test. n.s.: not significant, *p < .05, **p < .01, ***p < .001 between groups.
GSK1016790A activates TRPV4 pathway to participates in pyroptosis of hPDLcells
TRPV4 channels can be activated under mechanical force. A significant increase in TRPV4 protein expression in the OTM modeling group versus control group was observed (Figure 4(A), p < .001). Immunofluorescence staining showed a significant increase in Caspase-1 + TRPV4 + hPDL progenitor cells under mechanical stress (1.5 g/cm2) (Figure 4(B)), which indicating that the TRPV4 pathway was activated in hPDL cells undergoing pyroptosis under mechanical force stimulation. To further validate this conclusion, the TRPV4 agonist GSK1016790A was used to treat hPDL cells to confirm that activation of the TRPV4 signaling pathway induces increased pyroptosis. WB showed that treatment with GSK1016790A significantly upregulated the expression of key proteins of pyroptosis (NLRP3, Caspase-1, Cl-Casp-1, GSDMD, N-GSDMD, IL-1β, Cl-IL-1β) in hPDL cells, especially under mechanical force stimulation (Figure 4(C), p < .001). Optical microscopy revealed more pronounced cell swelling and membrane vesicle formation after GSK1016790A treatment (Figure 4(D)). These results demonstrate that GSK1016790A activates the TRPV4 pathway to participate in pyroptosis in hPDL progenitor cells.

GSK1016790A activates TRPV4 pathway and participates in pyroptosis in hPDL cells. (A) WB was used to detect TRPV4 proteins in control and CF modeling groups. (B) Immunofluorescence assay was used to detect the co-expression of TRPV4 and Caspase-1 in hPDLcell. Alexa Fluor 488 (green) for TRPV4 and Alexa Fluor 594 (red) for Caspase-1. Nuclei were counterstained with DAPI (blue). Scale bar=20 μm (original magnification: 630×). (C) WB experiment was used to detect key proteins (NLRP3, Caspase-1, Cl-Casp-1, GSDMD, N-GSDMD, IL-1β, Cl-IL-1β) involved in cell pyroptosis. (D) Observation of cellular morphological changes under an optical microscopy. Scale bar = 10 μm (original magnification: 400×). N = 3 biological replicates, technical triplicates per condition. Differences between the two groups were evaluated using a two-tailed independent Student's t-test. n.s.: not significant, *p < .05, **p < .01, ***p < .001 between groups.
Activation of TRPV4 signaling pathway by GSK1016790A promotes osteoclast activation under compressive forces
The effect of TRPV4 pathway activation on osteoblast activation was further investigated. The hPDL progenitor cells were pretreated with GSK1016790A to activate the TRPV4 pathway, followed by co-culturing with PBMCs. QPCR analysis of RANKL/OPG mRNA levels showed a significant increase in the osteoclast activation marker RANKL/OPG mRNA levels (Figure 5(A), p < .001). WB analysis of RANKL/OPG protein levels showed consistent changes with the mRNA levels (Figure 5(B), p < .001). Additionally, after GSK1016790A pretreatment, an increase of RANKL secretion was observed versus the control group (Figure 5(C), p < .001). In PBMCs with TRPV4 pathway activation, mRNA expression of TRAP and CTSK was significantly elevated versus control group (Figure 5(D), p < .05

GSK1016790A can promote osteoclastogenesis under compressive forces. (A) PCR was used to detect changes in mRNA levels of RANKL, OPG, and RANKL/OPG ratio in hPDL cells. (B) WB was used to detect changes in RANKL/OPG protein, a marker of osteoclast activation, in both control and CF groups. (C) ELISA detection of RANKL secretion concentration. (D) PCR detection of mRNA expression changes of TRAP and CTSK. (E) TRAP staining was used to detect the number of osteoclasts in the control group, CF group and CF with GSK1016790A treated group. Scale bar=50 μm (original magnification: 200×). Three biological replicates per experiment were performed. Differences between the two groups were evaluated using a two-tailed independent Student's t-test. n.s.: not significant, *p < .05, **p < .01, ***p < .001 between groups.
Discussion
OTM is a complex biomechanical process involving the movement of teeth within the alveolar bone. 25 The periodontal ligament connects the dentin to the alveolar bone surface, transmitting mechanical forces from the teeth to the alveolar bone, making it an ideal location for cells responsible for bone remodeling. 26 Teeth lacking PDL do not respond to orthodontic forces. 27 Therefore, PDL cells play a critical role in OTM, responding to mechanical forces through processes such as cell proliferation, differentiation, and tissue remodeling. 28 Stem cells in the PDL exhibit characteristics similar to mesenchymal stem cells (MSCs), expressing MSC surface markers such as CD105, CD90, and CD73, while lacking hematopoietic markers like CD45 and CD34. 29 These findings are consistent with our flow cytometry results following isolation, indicating successful separation and culture of hPDL cells, thereby providing a foundation for subsequent exploratory experiments. Inflammation often occurs during orthodontic treatment. 30 Our study demonstrates that compressive force activates TRPV4 channels in hPDL cells, triggering NLRP3 inflammasome assembly, caspase-1 activation, and gasdermin-D (GSDMD) cleavage, ultimately leading to pyroptosis. This mechano-pyroptotic pathway significantly promotes osteoclast differentiation and activation via enhanced RANKL secretion, providing a novel mechanistic link between mechanical force and bone remodeling during OTM.
The finding that TRPV4 activation drives pyroptosis in hPDL cells under mechanical loading extends current understanding of mechanotransduction in periodontal tissues. Pyroptosis is primarily initiated through the activation of caspase-1 by inflammasomes. 31 Inflammasomes are multiprotein complexes that recognize pathogen-associated molecular patterns (PAMPs) and damage-associated molecular patterns (DAMPs). Upon detection of these danger signals, inflammasomes activate caspase-1, which subsequently cleaves Gasdermin-D (GSDMD) protein. The cleavage of GSDMD generates a pore-forming N-terminal domain that inserts into the cell membrane, leading to alterations in osmotic pressure, cell swelling, and eventual rupture, resulting in pyroptosis.32,33 While TRPV4 has been implicated in PDL mechanosensing,20,34 its specific role in regulating pyroptosis had not been previously established. Our results align with emerging evidence suggesting that mechanical forces can induce pyroptosis in hPDL cell. 13 In this study, we found that the key pyroptosis proteins NLRP3, caspase-1, and N-GSDMD exhibited significantly increased expression under mechanical stress, demonstrating a pressure gradient dependence. The observed upregulation of these pyroptosis markers following TRPV4 activation by GSK1016790A further supports the involvement of this channel in inflammasome signaling.
Studies indicate that stem cells in the PDL can differentiate into osteoblasts and osteoclasts in response to mechanical stimulation, contributing to bone formation and resorption, respectively.35,36 This differentiation process begins within hours of orthodontic force application and is associated with the local synthesis and release of various cytokines and growth factors. 37 Osteoblasts also play a vital role in bone remodeling during OTM. Under mechanical stress, these mediators guide bone resorption and formation, facilitating tooth movement. 36 RANKL are recognized as key factors in the OTM process, particularly in osteogenesis. 38 Upon binding of RANKL to RANK, a series of intracellular signaling pathways are activated, contributing to the remodeling of the alveolar bone and accelerating the OTM process. On the pressure side, RANKL-RANK-mediated osteoclast activation leads to bone resorption, while on the tension side, mechanical stimuli promote osteoblast activity and bone formation. 39 Additionally, OPG, a decoy receptor for RANKL that binds to RANKL, preventing its interaction with RANK, thereby inhibiting osteoclastogenesis and activity. 40 The equilibrium between RANKL, RANK, and OPG is crucial for regulating the balance of bone resorption and formation during OTM. 41 Notably, our data reveal that pyroptosis-mediated inflammatory signaling (evidenced by IL-1β/IL-18 release) creates a pro-osteoclastogenic microenvironment, primarily through increased RANKL/OPG ratio. This finding bridges the gap between mechanical stimulation and osteoclastogenesis, suggesting that pyroptosis contributes to coupling bone resorption with orthodontic force application. However, this inflammatory response may represent a double-edged sword: while promoting tooth movement, excessive or prolonged pyroptosis could potentially exacerbate root resorption or periodontal tissue damage—a concern not sufficiently addressed in previous studies.
Transient receptor potential (TRP) channels are typical mechanosensitive channels that detect a variety of stimuli in different tissues and cells. 42 The TRPV4 channel can be activated by various physicochemical stimuli, regulating mechanosensation, inflammation, and energy homeostasis. 20 Studies on inflammatory diseases have shown that excessive activation of TRPV4 may promote the assembly of inflammasomes and the activation of caspase-1, leading to the cleavage of GSDMD and the occurrence of pyroptosis. This process releases a significant amount of inflammatory mediators, exacerbating the inflammatory responses. 43 Previous research has indicated that the stem cell properties of PDL are influenced by TRPV4 activation under mechanical stimulation, facilitating alveolar bone remodeling and ultimately affecting tooth movement. 13 In this study, we verified that pressure stimulation can activate the TRPV4 signaling pathway in an in vitro model of OTM, with TRPV4 activation triggering pyroptosis. The calcium influx induced by TRPV4 activation may influence intracellular signaling pathways, linking it to the activation of inflammasomes or the regulation of pyroptosis-related proteins.
GSK1016790A, an effective and selective TRPV4 activator, induces Ca2+ influx and elevates intracellular calcium levels.11,44 Our study is the first to investigate the effects of GSK1016790A on TRPV4 activation in hPDL cells concerning pyroptosis. Experimental results demonstrate that GSK1016790A-induced TRPV4 activation leads to pyroptosis in hPDL cells, which is further enhanced under pressure stimulation. We discovered that activation of the TRPV4 signaling pathway by GSK1016790A further enhances osteoblast activation, deepening our understanding of the molecular signaling mechanisms involved in osteoblasts during OTM. This research provides new insights into the regulation of osteoclastogenesis under mechanical stimulation, suggesting that targeting caspase-1-dependent pyroptosis may be a promising strategy to accelerate OTM.
Compared to existing literature established the role of TRPV4 in force-induced pyroptosis in PDL cells, our study provides substantial mechanistic and therapeutic advancements: First, while prior research identified TRPV4 activation under mechanical loading, we specifically demonstrate that TRPV4 acts as a decisive mechanosensor triggering pyroptotic cell death via NLRP3 inflammasome assembly and GSDMD cleavage, refining the molecular pathway from force sensing to inflammatory death. Second, moving beyond descriptive association, we clearly show that pyroptosis-derived mediators (e.g. IL-1β) actively drive osteoclastogenesis by markedly increasing RANKL/OPG ratio and TRAP + multinucleated cell formation, establishing a functional link between pyroptosis and bone remodeling. Most notably, third, we extend the paradigm by using pharmacological activation via GSK1016790A to potentiate TRPV4-mediated pyroptosis, resulting in significantly enhanced osteoclast differentiation—an approach not yet explored in prior studies and presenting a novel therapeutic strategy for accelerating OTM.
Despite these advancements, we acknowledge several limitations that temper the clinical implications of our findings. First, the co-culture model utilized exogenous M-CSF and sRANKL supplementation to standardize osteoclast differentiation, which may not fully replicate the paracrine dynamics of the native periodontal microenvironment; although necessary to ensure consistency and mitigate variability in primary cell models, this approach may oversimplify the physiological context and potentially amplify the observed effects. Moreover, the suggestion that targeting caspase-1 could accelerate OTM requires cautious interpretation, as pyroptosis is inherently inflammatory and its uncontrolled induction might lead to adverse effects including increased root resorption, exacerbated inflammation, and possible tissue damage [PMID: 34998415]. Finally, the exclusive use of in vitro models limits direct extrapolation to clinical OTM scenarios, and future studies should employ conditioned media from mechanically stimulated hPDL cells to better model paracrine signaling, alongside in vivo validation using animal models or clinical samples to confirm the translational potential of targeting TRPV4-pyroptosis signaling for accelerating OTM.
Conclusion
In sum, this study shows that compressive force induces inflammatory response and pyroptosis (correlated with pressure intensity) in hPDL cells, and this force-induced pyroptosis promotes osteoclast activation critical for alveolar bone remodeling during OTM. Mechanical force activates TRPV4 in hPDL cells, and the TRPV4 agonist GSK1016790A further enhances hPDL cell pyroptosis and osteoclast activation, thereby accelerating OTM. These findings clarify the molecular mechanism of OTM, identify TRPV4 as a key regulator linking mechanical stimuli to hPDL pyroptosis and bone remodeling, and suggest targeting TRPV4 or caspase-1-dependent pyroptosis as a potential strategy to accelerate clinical OTM.
Footnotes
Abbreviations
Acknowledgments
No artificial intelligence (AI) tools were used in the refinement or enhancement of the manuscript's text, the creation of images, or the generation/modification of any content. All scientific data, analyses, and written content were generated, analyzed, and prepared solely by the authors. The authors fully confirm the accuracy, originality, and integrity of all submitted material and take full responsibility for its content.
Ethical considerations
This study was conducted in accordance with the Helsinki Declaration. The protocol of the study was approved by the Ethics Committee of Changsha Stomatological Hospital on March 6, 2022 (Changsha, Hunan Province, China) (CSH-2022-038). Written informed consent to participate was obtained from all of the participants in the study.
Consent to participate
Written informed consent was obtained from all participants.
Authors’ contributions
He and Tian designed and performed the experiments, analyzed data, and drafted the manuscript. Xingyu Liu participated in cell isolation, mechanical loading experiments, and flow cytometry analysis. Yige Duan conducted qRT-PCR, ELISA assays, and statistical analysis. Mengshan Chen performed Western blot, TRAP staining, and immunofluorescence assays. Zhenyan Luo conceived the study, supervised the entire project, interpreted results, revised the manuscript critically, and provided funding support. All authors read and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Program of Hunan Provincial Health Commission (202208032486), Research Project of Changsha Health Commission (KJ-B2023086) and Research Project of Changsha Health Commission (KJ-B2023092).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
