Abstract
Background
Direct oral anticoagulants (DOACs) are first-line medications for stroke prevention in non-valvular atrial fibrillation (AF). However, variability in drug response poses risks of hemorrhagic or thromboembolic events.
Objectives
Although genetic influences on DOACs safety are increasingly recognized, robust evidence directly linking specific polymorphisms to bleeding risk remains limited.
Design
Multi-center observational case–control study including exome-wide association analysis of 196 non-valvular AF patients treated with rivaroxaban or apixaban, comprising 97 with bleeding complications and 99 without.
Methods
DOAC plasma concentrations, urinary 6-β-hydroxycortisol and cortisol levels were measured for CYP3A4 phenotyping. Sequencing was performed on the DNBSEQ G-400 platform. Single-nucleotide variant (SNV) associations with bleeding risk were assessed using logistic regression with additive, dominant, and recessive genetic models. Polygenic risk scores (PRSs) were calculated to evaluate cumulative genetic effects.
Results
No SNVs reached Bonferroni-corrected significance under any model. PRSs showed weak predictive ability for bleeding with apixaban. For rivaroxaban, regression indicated that ln Css min/D + 1 index increased with PRS, age, and 6-β-hydroxycortisol/cortisol ratio, but decreased with higher 6-β-hydroxycortisol and coronary heart disease presence. No statistically significant differences were found for the PharmGKB Level 3 variants rs1045642 (rivaroxaban) and rs2231142 (apixaban). Trends toward statistical significance were observed for the rs2472304-G variant in rivaroxaban users, rs6977165-C in apixaban users, and for the CYP3A4*1/*36 diplotype.
Conclusion
Residual equilibrium concentration of DOACs, including dose-adjusted, did not independently predict bleeding risk in non-valvular AF patients. Variants rs2472304 and rs6977165 may warrant further investigation as potential contributors to bleeding risk.
Keywords
Introduction
Direct oral anticoagulants (DOACs) remain the first-line, guideline-recommended medications for the prevention of stroke in patients with non-valvular atrial fibrillation (AF) 1 and are a cornerstone of treatment for patients with venous thromboembolism. 2 Traditional drugs of the group of vitamin K antagonists (VKAs, e.g. warfarin) and low-molecular-weight heparins had been the gold standard of anticoagulant therapy for decades. However, these agents are associated with challenges such as complex dosing regimens, narrow therapeutic range, the need for frequent international normalized ratio (INR) monitoring, and a high rate of drug–drug interactions. In current clinical practice, there has been a shift toward the use of DOACs, which directly inhibit factor IIa (dabigatran etexilate) or factor Xa (apixaban, rivaroxaban, edoxaban). Compared with VKAs, DOACs offer notable advantages: fixed dosing, no requirement for routine INR monitoring, less dependence on food intake for bioavailability, and a lower likelihood of drug–drug interactions. These benefits have driven the widespread adoption of DOACs. For instance, in England, DOACs accounted for 61.8% of all oral anticoagulant prescriptions in 2019, a substantial increase from 16.4% in 2015. 3 Similarly, in Germany, DOACs constituted less than 0.1% of all anticoagulant prescriptions in 2011 but surged to 49.9% by 2016. 4 In the United States, the proportion of patients receiving DOACs through the Medicare Part D program rose from 7.4% in 2011 to 66.8% in 2019, corresponding to an increase from 200,000 to 3.5 million patients. 5 Notably, apixaban ranked among the top three most-prescribed medications in the United States by 2022. 6
Despite their advantages, DOACs are associated with significant interindividual variability in drug response, which can lead to hemorrhagic or thromboembolic events. Bleeding is one of the most serious adverse reactions associated with anticoagulant therapy. Approximately 2–4% of patients receiving DOAC therapy experience major bleeding annually, while 10–12% report clinically relevant non-major bleeding events. 7 The increased use of DOACs has been paralleled by a rise in bleeding-related medical encounters. In the United States, the annual rate of such encounters increased by an average of 27.9% between 2016 and 2020. 8 In the United Kingdom, from 2011 to 2016, each additional 10% of DOAC prescriptions among all anticoagulant prescriptions was associated with a 0.9% increase in the incidence of hemorrhagic complications. During this period, the number of hospitalizations related to DOAC-induced bleeding complications totaled 4929 cases. 9 These findings underscore the critical need for continued efforts to optimize anticoagulant prescribing and monitoring.
The variability in DOAC response and the associated risk of adverse events are influenced by a range of clinical and demographic factors, including sex, age, comorbidities, renal and hepatic function, and concomitant medications. Recently, the genetic determinants of DOAC safety profiles have garnered increasing attention. Pharmacogenetic studies have highlighted associations between sequence variants involved in drug metabolism and biotransformation—including CES1, ABCB1, CYP3A4, CYP3A5, and ABCG2—and variability in DOACs.10–13 However, definitive evidence linking specific polymorphisms to an elevated risk of bleeding remains lacking. Further research is needed to clarify the genetic factors that influence interindividual variability in treatment response.
The objective of the current investigation was to identify novel pharmacogenetic biomarkers associated with the risk of bleeding in patients with non-valvular AF using DOACs, specifically apixaban and rivaroxaban. This article is based on an earlier version previously posted as a preprint on Preprints.org. 14
Materials and methods
Patients
The research involved 196 patients with a verified diagnosis of non-valvular AF along with chronic kidney disease (CKD) at different stages. Among them, 104 patients were administered apixaban, while 92 patients were given rivaroxaban.
The study sample included 97 patients with hemorrhagic complications and 99 patients without such complications. The participants were recruited from the clinical facilities of State Budgetary Institution of Health Care “Hospital of Veterans No. 2 of DZM” and State Budgetary Institution of Health Care “S.S. Yudin Municipal Clinical Hospital of DZM.”
This work represents a multi-center observational case–control study based on secondary analysis of biological samples and clinical data obtained from four prospective pharmacogenetic cohorts conducted between 2020 and 2023. Each study was approved by a local ethics committee, and all participants provided written informed consent. More detailed methodological information on each contributing cohort, including recruitment settings and patient characteristics, can be found in the corresponding original studies.15–18 For the present work, we selected patients with documented bleeding events (cases) and patients without any bleeding history during DOAC therapy (as controls).
The reporting of this study conforms to the general principles of the STROBE guidelines. 19
All included patients had non-valvular AF and met the standard indication for oral anticoagulant therapy, defined as a CHA2DS2-VASc score ≥1 for men and ≥2 for women.
DOAC dosing (apixaban 2.5–5 mg twice daily; rivaroxaban 10–20 mg once daily) was standardized, and bleeding severity was uniformly classified according to the International Society on Thrombosis and Haemostasis (ISTH) criteria: major, clinically relevant non-major, and minor. Events were identified from hospital records. In brief, events leading to a reduction in hemoglobin levels by at least 2 g/dl, fatal bleeding, or events occurring in critical body locations were considered major bleeding events. Events that required hospitalization, medical or surgical intervention, or a change in antithrombotic treatment were considered clinically relevant non-major. Any acute bleeding event not meeting the criteria for major or clinically relevant non-major bleeding was classified as a minor bleeding episode.
Because the present analysis integrated patients from four previously conducted pharmacogenetic studies with partially different designs and follow-up durations, the duration of DOAC therapy and the time between treatment initiation and bleeding onset varied across cohorts. For consistency, information on DOAC type and dosing regimen was harmonized. When available, data on treatment duration and time-to-bleeding were used for internal verification of event classification but were not pooled for statistical analysis because of methodological heterogeneity between source studies.
All plasma and DNA samples were re-analyzed under a unified protocol for exome sequencing and pharmacokinetic assessment. Clinical and laboratory data (age, sex, body mass index (BMI), renal function, hemoglobin, platelet count, comorbidities, HAS-BLED score [hypertension, abnormal renal/liver function, stroke, bleeding history or predisposition, labile international normalized ratio, elderly, drugs/alcohol concomitantly], and concomitant therapy) were harmonized across the cohorts. Inclusion/exclusion criteria were consistent with the original studies (non-valvular AF diagnosis, age ≥ 18 years, absence of mechanical heart valves, pregnancy, or severe hepatic impairment). No cases of clinically significant DOAC intolerance requiring treatment discontinuation were reported in the analyzed cohorts. Liver function tests (alanine aminotransferase, aspartate aminotransferase, total bilirubin) were available for two of the four contributing cohorts and showed no clinically significant abnormalities; these parameters were not included in the pooled analysis due to incomplete data coverage.
The sample size was calculated to ensure adequate statistical power and reliability. Using a confidence level of 95% (α = 0.05), a margin of error of 7%, and an assumed population proportion of 0.5, the minimum required sample size was 196 individuals. Additionally, power analysis for group comparisons with an effect size of 0.5, α = 0.05, and power = 90% determined a required sample size of 85 participants per group.
A chi-square test comparing proportions of bleeding outcomes (apixaban: 56%, rivaroxaban: 44%) and non-bleeding outcomes (apixaban: 51%, rivaroxaban: 49%) showed no significant association (χ2 = 0.3379, p = 0.561). This supports the random nature of the sample and its adequacy for detecting medium-sized effects.
Measurement of plasma concentrations of apixaban and rivaroxaban
To determine the drug concentrations, venous blood specimens were collected using vacuum tubes containing EDTA-K3 (tripotassium ethylenediamine tetraacetic acid) anticoagulant. Plasma samples were collected after at least 5 days of continuous DOAC therapy, corresponding to steady-state conditions (approximately 4–5 T1/2). The blood samples were then centrifuged at 3000 r/min for 15 min to isolate the plasma. The separated plasma was divided into Eppendorf tubes and stored at −70 °C until further analysis.
Quantification of apixaban and rivaroxaban levels in the blood plasma samples was conducted through high-performance liquid chromatography (HPLC) with an Agilent 1200 chromatograph equipped with a four-channel pump, mobile phase degasser, and chromatographic column thermostat. Detection was carried out with an Agilent TripleQuad LC/MS 6410 mass spectrometer (triple quadrupole type).
Sample preparation involved the precipitation of blood plasma proteins. First, plasma samples were thawed at room temperature. Next, 100 μl of plasma was dispensed into Eppendorf plastic tubes, followed by the addition of 250 μl of a 9:1 mixture of methanol and 0.1% hydrochloric acid (HCl). The mixture was vortexed, left to stand for 10 min, and vortexed again. The samples were then centrifuged at 10,000 r/min for 10 min. The supernatant was transferred to chromatographic vials and loaded onto the autosampler of the chromatograph.
An Agilent Polaris 3 C18-A column (length: 50 mm; inner diameter: 3.0 mm; particle size: 3.0 μm) was utilized for the separation process. The column temperature was maintained at 40 °C. The mobile phase consisted of two constituents: solution “A” (1 ml of concentrated formic acid diluted with deionized water to a total volume of 1 L) and solution “B” (1 ml of concentrated formic acid diluted with acetonitrile to a total volume of 1 L). Chromatographic separation was conducted using a gradient elution method. The blending of the mobile phase components during the analysis followed the program outlined in Table S1 in the Supplemental materials.
The volume of the injected sample was 5 μl, and the analysis was completed within a 10-min timeframe. The spectra of apixaban and rivaroxaban were obtained using multiple reactions monitoring with electrospray ionization in the positive ionization mode. The atomizer gas pressure was set at 35 lbf/in2, with a drying gas flow rate of 11 L/min at a temperature of 350 °C. The fragmentation voltage was maintained at 135 V, and the collision cell voltage was set to 25 V.
CYP3A4 phenotyping
Urine samples for CYP3A4 phenotyping were obtained during routine clinical testing without strict timing constraints. A CYP3A4 activity was determined by estimating the ratio of 6-β-hydroxycortisol to cortisol concentrations in patient urine collected in the morning. Cortisol is a specific CYP3A4 substrate. By calculating the metabolic ratio of the concentrations of cortisol and its metabolite, 6-β-hydroxycortisol, the activity of CYP3A4 was assessed: high ratio values indicate high isoenzyme activity, while low values indicate low activity. The methodology for determining a CYP3A4 activity is also generally accepted. 20
Cortisol and its metabolite were determined by HPLC with mass spectrometric detection using an Agilent 1200 liquid chromatograph (Agilent Technologies Inc., USA, 2008) and an AgilentTripleQuad LC/MS 6410 mass spectrometer. Data were processed with Agilent MassHunter Workstation Software LC/MS Data Acquisition for 6400 Series Triple Quadrupole (version B.08.02).
For chromatographic determination, the sample preparation technique and conditions for chromatographic analysis described in the work by Smirnov et al. were applied. 20
Exome sequencing
Venous blood samples were obtained from patients using vacuum tubes containing a minimum of 4 ml of EDTA-K3 anticoagulant. Genomic DNA was extracted from venous blood samples utilizing a QIAamp DNA Blood Mini Kit (Qiagen, Hilden, Germany). Libraries were prepared from 500 ng of genomic DNA using an MGI Easy Universal DNA Library Prep Set (MGI Tech, Shenzhen, China) following the manufacturer's instructions. The DNA fragmentation process was carried out through ultrasonication on a Covaris S-220 instrument (Covaris, Inc., Woburn, MA, USA), resulting in an average fragment size of 250 bp. Enrichment of DNA libraries was achieved through pre-pooling as described in the work by Belova et al., 21 employing SureSelect Human All Exon v8 probes (Agilent Technologies, Santa Clara, CA, USA) designed to cover the entire human exome. Quantification of DNA and libraries was conducted using a Qubit Flex instrument with a dsDNA HS Assay Kit in accordance with the manufacturer's guidelines (Invitrogen, Waltham, MA, USA). Subsequent quality assessment of the libraries was performed on a Bioanalyzer 2100 utilizing a High Sensitivity DNA kit (Agilent Technologies, Santa Clara, CA, USA) following the manufacturer's protocol.
The libraries were subsequently circularized and subjected to paired-end sequencing on the DNBSEQ G-400 platform utilizing the DNBSEQ-G400RS High-throughput Sequencing Set PE100 following the protocol provided by the manufacturer (MGI Tech, Shenzhen, China), achieving an average coverage of 100×. FastQ format files were produced through the application of the manufacturer's basecallLite software (ver. 1.0.7.84) (MGI Tech, Shenzhen, China).
Bioinformatic quality control and correction of sequencing data were performed utilizing FastQC v0.11.9 22 and bbduk v38.96 software. 23 Subsequent bioinformatic processing of sequencing data for each sample included alignment to the GRCh38 human reference genome using bwa-mem2 v2.2.1 24 and SAMtools v1.9. 25 Quality metrics for exome enrichment were obtained using Picard v2.22.4. 26 Variants were called with bcftools v1.9 27 and deepvariant v1.5.0, 28 and annotating using AnnoVar 29 and Intervar v2.2.2. 30 Several in-house scripts were employed to enhance the quality of the final variant annotation files. As a final step, MultiQC v1.16 31 software was executed for quality control following the bioinformatic pipeline.
Exome-wide association study
The study cohort comprised of 97 patients with bleeding and 99 without bleeding, totaling 461,938 unique polymorphic markers with a genotyping quality of 99.47%. Genetic variants were filtered using PLINK v1.90b6.24. 32 Multi-allelic variants were treated as false heterozygotes (–vcf-half-call r). Markers genotyped in fewer than 98% of samples were filtered out, removing 16,542 variants. Samples with genotyping quality below 98% were also discarded. Variants on the X and Y chromosomes were removed, and variants with a minor allele frequency below 0.2 were filtered out, leaving 66,720 variants. A Hardy–Weinberg equilibrium test was performed on 60,142 markers (p < 0.00001), yielding a final genotyping quality as 99.99%. Association analysis was conducted using logistic regression with additive, dominant, and recessive models, incorporating covariates such as the first principal components or the drug administered. The resulting p-values were adjusted for multiple testing by Bonferroni corrections (padj < 0.05).
Polygenic risk scores (PRSs) were generated using PRSice-2 33 with parameters –stat OR and –binary-target F to predict the occurrence of bleeding events, without providing a file for –pheno. Additional quality control measures were applied, including the elimination of duplicates, resulting in a final set of 11,938 variants.
To exclude hereditary forms of bleeding, the HP:0001892 and HP:0000790 panels were employed. 34 Diplotyping of pharmacogenes was performed using the Panno resource, with a focus on the European population. 35
Statistical analysis
Statistical analysis and data visualization were carried out using the R programming language (RStudio Pro 2023.09.1) and GraphPad Prism 8.0.1 software (GraphPad, La Jolla, CA, USA). Missing values were imputed using the amelia package. 36 Feature selection was carried out using the Boruta algorithm, 37 which identifies relevant predictors by comparing their importance with that of randomly permuted “shadow” variables. Predictors classified as confirmed (“green”) or tentative (“red”) by Boruta were retained for regression modeling. In addition, PRS was included in all models as a variable of primary interest.
Results
Clinical features of the individuals
The baseline demographic, clinical, and laboratory characteristics of the patient cohorts are summarized in Table 1. Of all 97 bleeding events, 5 (5.2%) were classified as major, 23 (23.7%) as clinically relevant non-major, and 69 (71.1%) as minor according to ISTH criteria. Detailed individual patient data, including specific bleeding events and comorbidities, are provided in Table S2 in the Supplemental materials.
Clinical and demographic characteristics of the study samples.

BMI: body mass index; AH: arterial hypertension; CHF: chronic heart failure; CHD: coronary heart disease; DM 2: type 2 diabetes mellitus; CKD: chronic kidney disease.
Data are presented as the median [25th percentile; 75th percentile] unless stated otherwise.
Boruta's algorithm 37 was utilized to evaluate the significance of predictors determining Css min/D (ng/ml/mg). Due to the replacement of two missing values with negative values (−5.173 and −1.724) by the amelia package, the natural logarithm of each Css min/D value was calculated, with 1 added (ln Css min/D + 1). BMI, the medications themselves, the presence of a history of coronary heart disease (CHD) and CKD, and the ratio of 6-β-hydroxycortisol to urinary cortisol concentrations were identified as the most crucial factors influencing medication levels. Notably, differences were observed between the drug groups. For apixaban, the concentration of urinary 6-β-hydroxycortisol (ng/ml) and its ratio to cortisol were significant predictors. For rivaroxaban, these parameters, along with age, the presence of CHD, and group membership (bleeding and non-bleeding) were also found to be important (Figure S1 in the Supplemental materials).
Interestingly, the concentration of urinary 6-β-hydroxycortisol and urinary cortisol (ng/ml) emerged as predictors of bleeding development, with the stage of CKD emerging as an additional predictor (Figure S2 in the Supplemental materials). In this analysis, the unadjusted minimum concentration at the steady-state (Css min/D) value was utilized.
To assess the predictive capability Css min and Css min/D, we used the R package “pROC” 38 (Figure 1). The results revealed statistical insignificance, with low sensitivity and specificity, emphasizing the need to incorporate additional variables to better estimate bleeding risk.
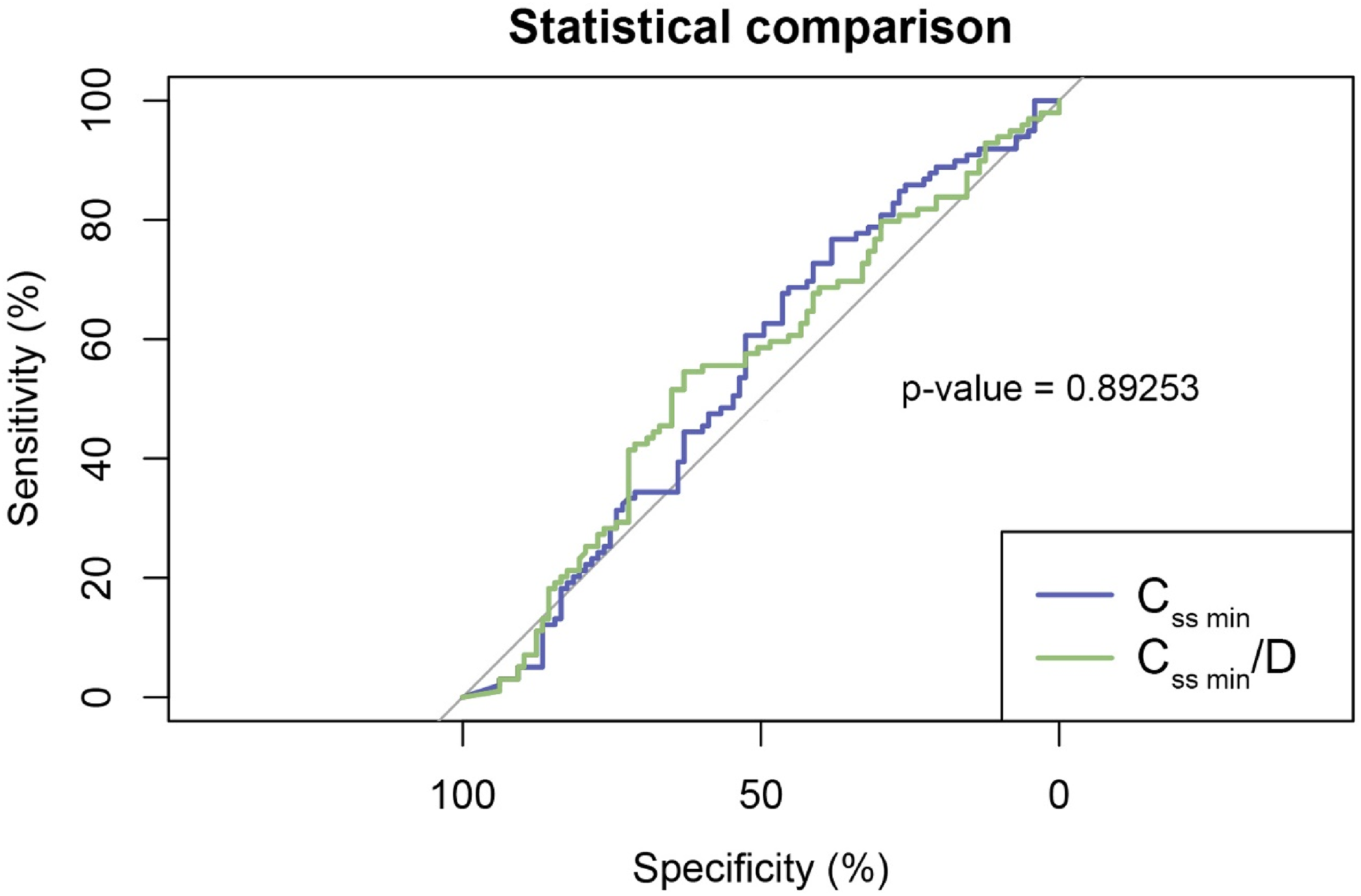
Assessment of binary classification (“with bleeding” and “without bleeding”) demonstrated that neither indicator alone provided adequate predictive accuracy.
The residual equilibrium concentration adjusted for daily dose (Css min/D) did not follow a normal distribution, as shown in Figure 2. A similar analysis was conducted for Css min and blood platelet counts, as depicted in Figure S3 in the Supplemental materials.

Histogram illustrating the distribution of the residual equilibrium concentration adjusted for daily dose (Css min/D, ng/mL/mg) in the two sample groups (a); Q–Q plot (Shapiro–Wilk test revealed significant deviation from normality for Css min/D: W = 0.8493, p < 0.0001) (b).
A permutation factorial analysis of variance was performed to evaluate differences in Css min/D between patients with and without bleeding. The results indicated no statistically significant differences (F(1,185) = 0.6927, p = 0.4001). However, when accounting for drug type within groups, Css min/D levels were significantly higher in patients using apixaban (p < 0.0001 in both the groups) (Figure 3). Among the patients using apixaban, the difference in Css min/D between the subgroups with and without bleeding was not significant (p = 0.8098), but in patients using rivaroxaban, the significance level was p = 0.0309.

Residual equilibrium concentration adjusted for daily dose (Css min/D, ng/mL/mg) was compared between two groups of subjects based on the specific drug administered (apixaban or rivaroxaban) using permutation factorial analysis of variance and post-hoc analysis (estimated marginal means).
The study revealed that patients with bleeding disorders had a higher BMI compared to those without bleeding disorders (F(1,187) = 4.6603, p = 0.0306), with no significant differences based on drug intake. Furthermore, patients using apixaban exhibited a more advanced stage of CKD (F(1,192) = 9.5407, p = 0.0023), although no differences were observed between the groups with and without bleeding. Post-hoc analysis suggested a potential statistical significance within the bleeding group (p = 0.0809). Permutation factorial analysis of variance demonstrated a significant impact of the drug factor on HAS-BLED scores (F(1,183) = 25.47, p = 0.0001). Specifically, bleeding patients taking apixaban had higher HAS-BLED scores compared to those taking rivaroxaban (p < 0.0001), with similar differences observed in the non-bleeding group (p = 0.0048). Notably, age, blood hemoglobin concentration, and platelet count did not exhibit significant differences across the groups and subgroups.
The exome-wide association study did not reveal any statistically significant signals
The study participants were screened for hereditary diseases associated with abnormal bleeding, such as hematuria. The analysis identified 14 carriers of pathogenic variants among the patients with bleeding disorders. These variants were located on autosomes and affected the following genes: F5 (rs6025-T), F11 (rs121965064-C), VWF (rs41276738-T, rs900907976-A), ATP7B (rs76151636-T, rs28942076-T, rs780558532-TG, rs780558532-T), F7 (rs36209567-T), ABCC6 (rs745900279-delAG, rs72653706-A, rs1423674851-C), FANCA (rs397507553-delAGA), GP6 (rs368858591-A, rs199588110-A), FYB1 (rs769310295-insA), TNXB (rs200718357-T), SIK3 (NM_001366686.3):c.741+1G>A, and VPS33B (NM_018668.5):c.1105+2T>C. Based on full-exome sequencing data, none of the participants were found to be at risk of hemorrhagic events with no apparent clinical cause or late onset.
The exome-wide association study (EWAS) conducted on the samples from the two groups revealed an average sequencing coverage depth of 86.88×, with an average coverage completeness of 97.15% at a depth of 10×. Figure 4 illustrates the population affiliations of the samples, where it was not possible to definitively identify two or more clusters.

A scatter plot generated with two principal components indicated a low level of genetic heterogeneity within the samples.
The variant rs7299963-C demonstrated the highest level of statistical significance in the recessive model (p = 2.282 × 10−4, C allele frequency 0.3454, T allele frequency 0.5707; odds ratio (OR) 0.3968 [95% confidence interval (CI), 0.2638–0.597]). In the additive and dominant models, rs539002185-G yielded significant results (p = 9.345 × 10−6 in both instances, G allele frequency 0.1237, A allele frequency 0.2828; OR 0.358 [95% CI, 0.2112–0.6067]). The inclusion of covariates, such as the first six principal components and medication, did not alter the statistical significance. Both variants were located in intronic regions and exhibited a protective effect. The genomic factor prior to adjusting for different inheritance patterns was 1.03156.
The variant most strongly associated with bleeding across all the three inheritance models, including the drug covariate, was rs4298115-T (p = 0.0007236 in the additive model, p = 0.06448 in the recessive model, and p = 0.0001805 in the dominant model, before correction; T allele frequency 0.5876, C allele frequency 0.404: OR 2.102 [95% CI, 1.405–3.145]). However, Bonferroni correction increased the p above 0.05. The protective variant with the lowest p before correction for multiple testing was rs343122-C (C allele frequency, 0.1443, A allele frequency, 0.3485; OR 0.3153 [95% CI, 0.1921–0.5177]).
Filtering the .vcf file based on regions containing the specified genes and rerunning the EWAS with the previously described algorithm did not reveal statistically significant differences, even with a reduction in the burden of multiple testing. To establish the PRSs for predicting bleeding, further quality control measures were implemented, resulting in the removal of duplicates and leaving a total of 11,938 variants. We developed PRS focusing solely on the presence or absence of bleeding (Figure 5). In one male participant, the specific type of bleeding remained undetermined. The PRS showed a strong correlation with the investigated trait. An intriguing observation was the absence of overlap in PRS values between the two groups.
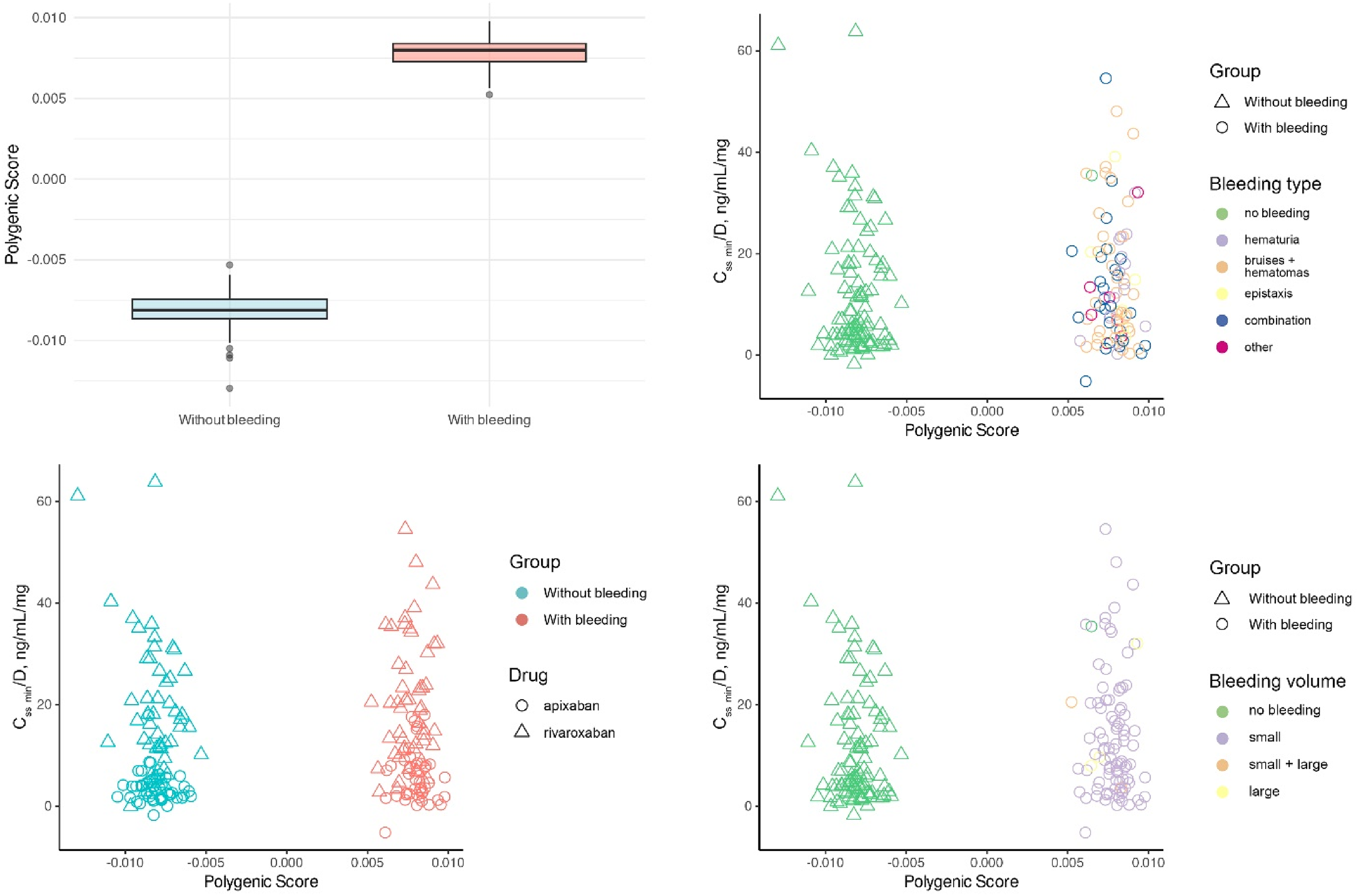
Polygenic bleeding risk scores (PRS) based on residual equilibrium concentration adjusted for daily dose (Css min/D, ng/mL/mg). Scatter plots of best-fit PRS by drug, bleeding type, and bleeding volume. Note: “Other” in bleeding type included gynecologic bleeding, gastrointestinal bleeding, and gingival bleeding.
We hypothesize that these results may have been due to the small sample sizes and the minor effects of each variant. Additionally, we determined PRS, based only on the assumption of additive variant effects. Therefore, we decided to use PRS as one of the independent variables and predict the risk of bleeding by incorporating clinical parameters.
Frequencies of variants in the target genes
Genotype frequencies of genes encoding enzymes metabolizing apixaban and rivaroxaban, including CYP3A4, CYP1A2, CYP2C8, CYP2C9, CYP2C19, CYP2J2, CYP3A5, as well as genes for the DOAC transporters ABCB1 and ABCG2, 39 were analyzed using PLINK (Table S3 in the Supplemental materials). No initial quality control measures were conducted on the .vcf file in this instance. A total of 216 variants were examined, and linkage disequilibrium (LD) was calculated for them (Table S4 in the Supplemental materials). The rs2472304-G variant (CYP1A2) exhibited the most significant association with the risk of hemorrhagic events (prior to adjustment for multiple testing). In the additive model, the p was 0.008736, while in the dominant model, the p was 0.007099 before adjustment. The G allele frequency was 0.4794, the A allele frequency was 0.3434, with OR 1.76 [95% CI, 1.172–2.644]. The variant rs2472304 is not part of the known star allele and is in high LD with rs2470890 (r2 = 0.99013).
Upon constructing logistic regression models including only patients administered a single anticoagulant, rs2472304-G demonstrated the strongest association in the rivaroxaban group across all three inheritance models: additive (p = 0.00529), recessive (p = 0.06489), and dominant (p = 0.006274), along with rs2470890 (CYP1A2), prior to correction for multiple testing. In the apixaban group, rs6977165-C (CYP3A5) showed significance in the additive and dominant models (p = 0.02277), while rs2235046 (ABCB1) and rs10276036 (ABCB1) were nominally associated under the recessive model (p = 0.14), with no evidence of LD between them (r2 < 0.2).
No significant differences were found in the prevalence of CYP2B6, CYP2C19, CYP2C8, CYP2C9, CYP2D6, CYP3A4, CYP3A5, CYP4F2, DPYD, NUDT15, SLCO1B1, TPMT, and UGT1A1 haplotypes between the two sample groups (Table S5 in the Supplemental materials).
The study also examined the diplotypes of the CYP3A4 gene, responsible for metabolizing rivaroxaban and apixaban, as presented in Table 2. Contingency tables were constructed, and statistical analysis was conducted using Fisher's exact test, except for CYP3A4*1 and CYP3A4*36 (rs2242480) where a chi-square test with Yates correction was applied. The results indicated no significant difference in the distribution of diplotypes. Notably, a trend toward statistical significance was observed in patients with the CYP3A4*1 and CYP3A4*36 diplotypes (χ2 = 2.0471, df = 1, p = 0.1525). Conversely, a less pronounced difference was noted for the CYP3A4*36 haplotype compared to other CYP3A4 haplotypes across the four groups (χ2 = 6.8, df = 3, p = 0.079). Furthermore, a slightly higher incidence of bleeding was observed in patients receiving apixaban, whereas an opposite pattern was noted for those on rivaroxaban.
Diplotypes of pharmacogenes in two groups of patients.

Linear regression models
We observed a statistically significant correlation between PRS and urinary 6-β-hydroxycortisol and cortisol concentrations in patients using both medications: r = −0.27, p = 0.0053 and r = −0.21, p = 0.0298 for apixaban, and r = −0.35, p = 0.0006 and r = −0.26, p = 0.0119 for rivaroxaban; but their ratio was significant only for rivaroxaban (r = −0.28, p = 0.007) (Figure 6). Additionally, the random forest algorithm indicated that the PRS for apixaban was the least influential factor in determining ln Css min/D + 1 compared to other predictors.
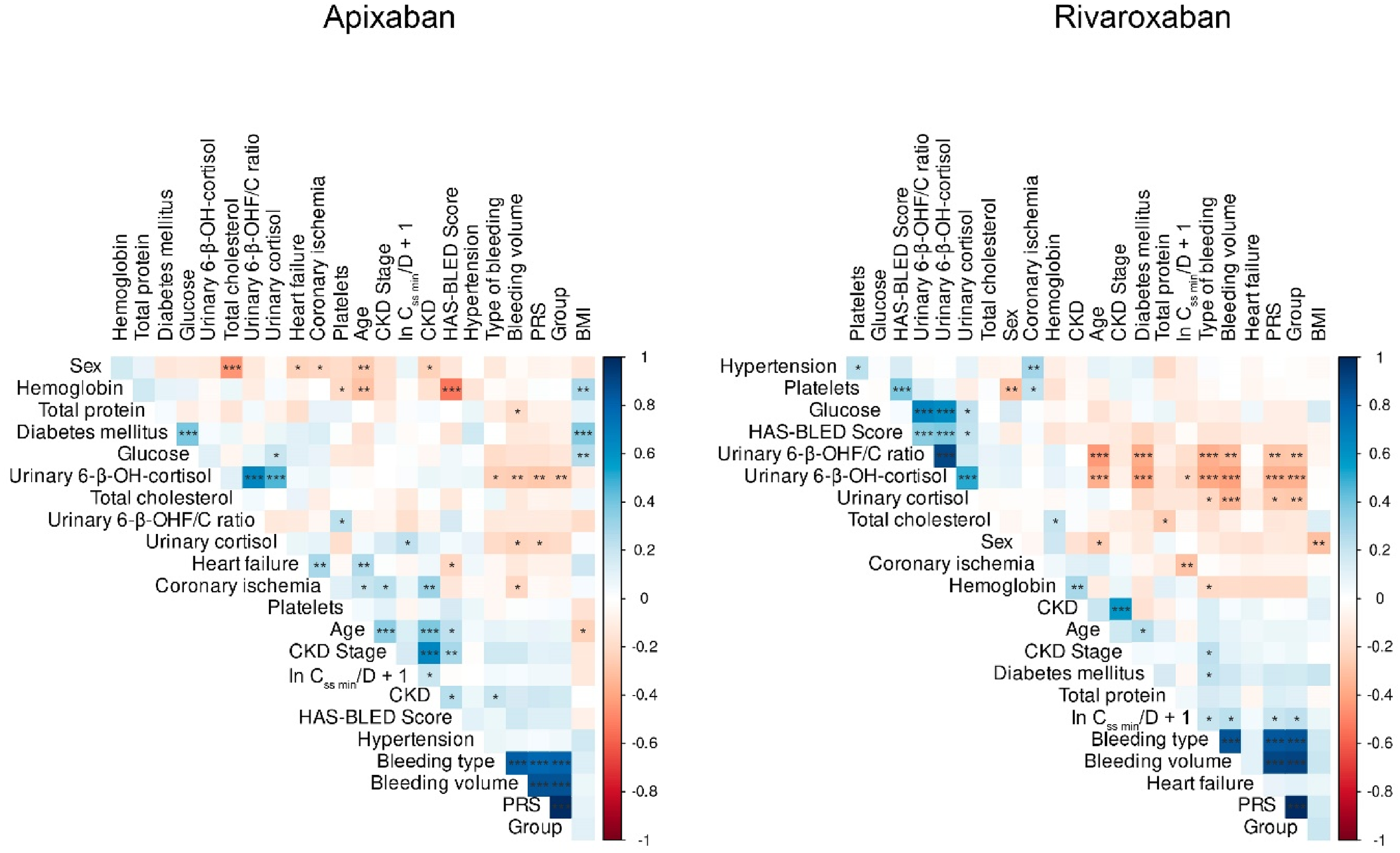
Correlation matrices were generated for apixaban and rivaroxaban, with asterisks denoting the confidence level of statistical significance (the matrices were constructed using the corrplot library in R, employing Pearson's coefficient).
A linear regression model (the lm() function in R) was developed to analyze patients using apixaban, incorporating predictors such as PRS, urinary 6-β-hydroxycortisol concentration, the ratio of 6-β-hydroxycortisol to cortisol, and the presence of CKD (refer to Figure S4 in the Supplemental materials). The model demonstrated homoscedasticity (Breusch-Pagan test, BP = 3.472, df = 4, p = 0.4822), meaning that the predictor values had no effect on the residuals’ variance, which remained constant. However, as indicated by the low multiple R2 (0.08972) and F-statistic value (2.439, p = 0.05187), the model's explanatory power was limited. Notably, the ratio of urine 6-β-hydroxycortisol to cortisol concentrations was the only variable to exhibit a significant coefficient (p = 0.0216), with an increase of one unit in the ratio resulting in a 0.1038 decrease in ln Css min/D + 1. These results suggest a poor predictive capacity of PRS, implying that genetics was not a major factor in apixaban-using patients.
The development of a linear regression model for patients receiving rivaroxaban revealed significant heteroscedasticity of the residuals, as indicated by the results of the Breusch–Pagan test (BP = 14.021, df = 5, p = 0.01548). Predictors such as age, PRS, CHD, urine 6-β-hydroxycortisol concentration, and its ratio to cortisol were all included in the model. A generalized least-squares (GLS) method was used, applying different standard deviations for each age value to address the heteroscedasticity issue. Significant improvements were observed after applying GLS: all predictor coefficients attained significance (p < 0.05), and there were no longer any indicators of heteroscedasticity in the residuals (p = 0.8567). The accuracy of the model estimates increased as the residual standard error dropped below 0.0138. Notably, ln Css min/D + 1 increased by 2.835 when PRS increased by one unit (t = 20.4307, p < 0.0001). On the other hand, age had a significant positive effect; an increase of 1 year led to a rise of 0.00066 in ln Css min/D + 1 (t = 3.2296, p = 0.0018). The presence of coronary ischemia was associated with a 0.867 decrease in ln Css min/D + 1 (t = −458.8183, p < 0.0001). Furthermore, a unit increase in urinary 6-β-hydroxycortisol concentration resulted in a 0.00384 decrease in ln Css min/D + 1 (t = −57.8738, p < 0.0001), while its ratio to cortisol was linked to a 0.157 increase in ln Css min/D + 1 per unit (t = 49.0953, p < 0.0001). It means that for patients taking rivaroxaban, genetics appears to be a major determinant.
Discussion
The existing scientific literature contains a considerable number of studies investigating the associations between specific single-nucleotide variants (SNVs) in genes related to transporter proteins and enzymes involved in the primary metabolic pathways of apixaban (ABCB1, CYP3A4) and rivaroxaban (ABCB1, CYP3A4, CYP3A5). The authors emphasize the impact of SNV markers of CYP3A4 (rs35599367), CYP3A5 (rs776746), and ABCB1 (rs1128503, rs2032582, rs1045642, rs4148738) genes on the pharmacokinetic parameters of DOAC.12,1340–45 The relationship between genetic variations influencing DOAC pharmacokinetics and the risk of bleeding remains unclear. Research has indicated a higher incidence of bleeding in patients with AF who have elevated peak drug concentrations during DOAC administration, reaching 156% and 266% of the baseline for apixaban and rivaroxaban, respectively. 46 Pharmacogenetic investigations have suggested that specific genotypes and allele variants of ABCB1 (rs4148732) and ABCG2 (rs2231142) may lead to increased peak plasma concentrations of apixaban and rivaroxaban, although no direct associations with elevated bleeding risk have been established.13,46,47
The search results from the PharmGKB database 48 for apixaban and rivaroxaban reveal the presence of pharmacogenetic markers with a level of evidence 3 (low). This level of evidence does not support the consideration of these markers for the integration of pharmacogenetic testing into clinical practice. The limitation is attributed to the lack of reproducibility of results across different studies. 47 Currently, there are no recommendations with a high level of evidence concerning the utilization of SNV of genes encoding metabolizing enzymes (such as CYP3A4, CYP3A5, etc.) and transporter genes (such as ABCB1, ABCG2, etc.) for optimizing the therapy of patients with indications for prevention of AF-related stroke. Findings from association studies suggest that the carriage of pharmacogenetic markers does not significantly impact the exposure to DOACs, which would otherwise manifest in an increased risk of hemorrhagic complications.
No statistically significant difference was found between the genotypes of rs1045642 (3 GG, 29 AG, 17 AA in the healthy group and 8 GG, 22 AG, 13 AA in the bleeding group), despite claims that patients with the GG genotype may have decreased clearance of rivaroxaban compared to those with the AA or AG genotypes. As for apixaban, the only variant with level of evidence 3 was rs2231142, where increased concentrations were observed in individuals with the TT genotype. We did not detect this genotype in the healthy group (14 TG, 36 GG), however, there was no significant difference compared to those with bleeding (1 TT, 11 TG, 42 GG).
Several reasons for the absence of these associations can be contemplated. Initially, while the examined variations impact the functional activity of apixaban and rivaroxaban biotransformation catalysts, additional enzymes (such as CYP1A2, CYP2C8, CYP2C9, CYP2C19, and CYP2J2) might also contribute to their metabolism, 45 thereby offsetting the potential impact of SNVs in the primary pathway enzymes and maintaining the overall drug metabolism.
Second, within our context, the occurrence of bleeding as a clinical outcome cannot solely be attributed to genetically determined variations in the functional activity of DOAC metabolizing enzymes. The risk of hemorrhagic complications in individuals undergoing anticoagulant treatment is impacted by various clinical and demographic factors, with genetics representing just one aspect. Conversely, blood coagulation involves an intricate network of interactions among cascades of factors and cofactors, making it challenging to pinpoint the singular most critical component within this system.
Finally, studies using the candidate gene approach are guided by specific hypotheses and offer distinct advantages in terms of the robustness of statistical inference when identifying associations between genes and drugs. In the context of apixaban and rivaroxaban metabolism, the candidate gene approach suggests that variations in the ABCB1 P-glycoprotein gene and the CYP enzymes responsible for DOAC metabolism (CYP3A4/5, CYP1A2, CYP2C8, CYP2C9, CYP2C19, and CYP2J2) may be linked to drug concentration levels in the bloodstream and, consequently, to variations in clinical outcomes. Nonetheless, such analyses may overlook numerous other genes and millions of SNVs that could potentially impact the variability in the pharmacokinetics and pharmacodynamics of these medications. Another limitation of candidate gene studies is their inability to fully consider the combined effects of multiple genetic variants simultaneously, known as polygenic effects. Research indicates that the variability in drug response among individuals may be attributed to polygenic inheritance.49,50
To date, there has been a paucity of GWASs and EWASs investigating genetic risk factors linked to drug response or pharmacokinetics of DOACs in patients.13,51
In our study, we conducted an EWAS analysis to identify novel markers of bleeding risk. Our results revealed that none of the genetic variations in our sample exhibited a statistically significant correlation with the risk of hemorrhagic episodes at the exome level in both the apixaban and rivaroxaban treatment groups. Notably, the most noteworthy genetic variations in both the groups were located within the CIT and ADPRHL1 genes, which have not been previously linked to coagulation disorders. Upon adding medication as a covariate, robust correlations were observed in the ATP10D and UTP15 genes, which also have not been discussed in the existing literature in the context of bleeding risk. In a separate analysis of SNVs of candidate enzyme metabolizer genes of the studied DOACs, only the rs2472304-G variant of the cytochrome CYP1A2 gene exhibited a significant association with bleeding risk under the additive and dominant inheritance models (prior to adjustment for multiple comparisons). Furthermore, a tendency toward a notable correlation with hemorrhagic incidents was observed in patients with CYP3A4*1 and CYP3A4*36 diplotypes who were prescribed apixaban.
The data obtained suggest that clinical and demographic parameters, such as BMI, age, and concomitant CKD and CHD, were predictors of Css min/D values. Patients taking apixaban, regardless of bleeding status, exhibited higher average Css min/D values (p < 0.0001). In contrast, the association between Css min/D values and bleeding risk was observed only in the rivaroxaban group.
An additional discovery indicated that urinary levels of cortisol, its metabolite, and their ratio played a significant role in predicting Css min/D values for both apixaban and rivaroxaban. Cortisol is a specific substrate of CYP3A4, the enzyme responsible for metabolizing these drugs. The determination of CYP3A4 activity can be achieved by calculating the metabolic ratio between cortisol and its metabolite 6-beta-hydroxycortisol: higher ratios reflect increased isoenzyme activity, while lower ratios suggest reduced activity. However, the reliability of this approach for CYP3A4 phenotyping remains a subject of debate among researchers, with some questioning its practical utility in clinical decision-making.52–54
PRS demonstrated an inverse correlation with the concentrations of 6-β-hydroxycortisol and cortisol in urine, showing a more significant impact in patients undergoing rivaroxaban treatment. Notably, the ratio of 6-β-hydroxycortisol to cortisol had varying effects on the ln Css min/D + 1 for the two medications: an inverse relationship was observed for apixaban, while a direct relationship was noted for rivaroxaban. Additionally, in prognostic models, it was found that in patients treated with rivaroxaban, the ln Css min/D + 1 index increased with PRS, age, the ratio of 6-β-hydroxycortisol to cortisol concentration in urine, and decreased with higher levels of 6-β-hydroxycortisol in urine, as well as the presence of CHD as a concomitant disease. Detailed explanations for these findings cannot yet be provided.
Limitation of our study
The study is subject to several limitations. The samples used were from patients with AF receiving DOACs, treated in various institutions within a single city, Moscow. These genetic samples were obtained as part of distinct observational pharmacogenetic studies focusing on apixaban and rivaroxaban with different designs and follow-up durations, which limited the ability to uniformly assess treatment duration and time-to-bleeding across all participants. The observational design of the studies conducted across different clinics may have led to variations in routine clinical practices, potentially introducing biases in the collected data.
Patient data were extracted from medical records, which limits the scope of the study to specific datasets (real-world data). The data collection and analysis focused mainly on standard clinical parameters typically obtained from patients diagnosed with AF. Nevertheless, it is important to acknowledge that discrepancies in clinical protocols among various healthcare institutions existed.
The clinical sample size was constrained by limited resources for patient recruitment and genetic analysis. Future studies with larger samples and improved control over inclusion criteria homogeneity may provide further insights into the relationship between DOACs and genetic factors.
Conclusions
Thus, no new polymorphic variants associated with an increased risk of bleeding following the use of apixaban and rivaroxaban were identified in this study using EWAS analysis. The findings suggest that the residual equilibrium concentration of anticoagulants, including dose-adjusted values, may not independently predict hemorrhagic complications in patients with non-valvular AF. Although no statistically significant differences were found for the PharmGKB Level 3 variants rs1045642 (rivaroxaban) and rs2231142 (apixaban), these markers—along with suggestive trends observed for rs2472304, rs6977165, and the CYP3A4*1/CYP3A4*36 diplotype—may contribute to bleeding susceptibility and warrant further investigation in larger, well-powered cohorts.
Supplemental Material
sj-png-1-sci-10.1177_00368504251398881 - Supplemental material for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants
Supplemental material, sj-png-1-sci-10.1177_00368504251398881 for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants by Dmitry A Sychev, Anastasiia A Buianova, Sherzod P Abdullaev, Karin B Mirzaev, Vera A Belova, Anna O Shmitko, Valery V Cheranev and Oleg N Suchalko, Lyudmila V Fedina, Svetlana V Batyukina, Natalia A Shatalova, Pavel O Bochkov, Sergey V Glagolev, Denis V Rebrikov, Dmitriy O Korostin in Science Progress
Supplemental Material
sj-png-2-sci-10.1177_00368504251398881 - Supplemental material for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants
Supplemental material, sj-png-2-sci-10.1177_00368504251398881 for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants by Dmitry A Sychev, Anastasiia A Buianova, Sherzod P Abdullaev, Karin B Mirzaev, Vera A Belova, Anna O Shmitko, Valery V Cheranev and Oleg N Suchalko, Lyudmila V Fedina, Svetlana V Batyukina, Natalia A Shatalova, Pavel O Bochkov, Sergey V Glagolev, Denis V Rebrikov, Dmitriy O Korostin in Science Progress
Supplemental Material
sj-png-3-sci-10.1177_00368504251398881 - Supplemental material for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants
Supplemental material, sj-png-3-sci-10.1177_00368504251398881 for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants by Dmitry A Sychev, Anastasiia A Buianova, Sherzod P Abdullaev, Karin B Mirzaev, Vera A Belova, Anna O Shmitko, Valery V Cheranev and Oleg N Suchalko, Lyudmila V Fedina, Svetlana V Batyukina, Natalia A Shatalova, Pavel O Bochkov, Sergey V Glagolev, Denis V Rebrikov, Dmitriy O Korostin in Science Progress
Supplemental Material
sj-png-4-sci-10.1177_00368504251398881 - Supplemental material for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants
Supplemental material, sj-png-4-sci-10.1177_00368504251398881 for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants by Dmitry A Sychev, Anastasiia A Buianova, Sherzod P Abdullaev, Karin B Mirzaev, Vera A Belova, Anna O Shmitko, Valery V Cheranev and Oleg N Suchalko, Lyudmila V Fedina, Svetlana V Batyukina, Natalia A Shatalova, Pavel O Bochkov, Sergey V Glagolev, Denis V Rebrikov, Dmitriy O Korostin in Science Progress
Supplemental Material
sj-xlsx-5-sci-10.1177_00368504251398881 - Supplemental material for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants
Supplemental material, sj-xlsx-5-sci-10.1177_00368504251398881 for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants by Dmitry A Sychev, Anastasiia A Buianova, Sherzod P Abdullaev, Karin B Mirzaev, Vera A Belova, Anna O Shmitko, Valery V Cheranev and Oleg N Suchalko, Lyudmila V Fedina, Svetlana V Batyukina, Natalia A Shatalova, Pavel O Bochkov, Sergey V Glagolev, Denis V Rebrikov, Dmitriy O Korostin in Science Progress
Supplemental Material
sj-xlsx-6-sci-10.1177_00368504251398881 - Supplemental material for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants
Supplemental material, sj-xlsx-6-sci-10.1177_00368504251398881 for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants by Dmitry A Sychev, Anastasiia A Buianova, Sherzod P Abdullaev, Karin B Mirzaev, Vera A Belova, Anna O Shmitko, Valery V Cheranev and Oleg N Suchalko, Lyudmila V Fedina, Svetlana V Batyukina, Natalia A Shatalova, Pavel O Bochkov, Sergey V Glagolev, Denis V Rebrikov, Dmitriy O Korostin in Science Progress
Supplemental Material
sj-xlsx-7-sci-10.1177_00368504251398881 - Supplemental material for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants
Supplemental material, sj-xlsx-7-sci-10.1177_00368504251398881 for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants by Dmitry A Sychev, Anastasiia A Buianova, Sherzod P Abdullaev, Karin B Mirzaev, Vera A Belova, Anna O Shmitko, Valery V Cheranev and Oleg N Suchalko, Lyudmila V Fedina, Svetlana V Batyukina, Natalia A Shatalova, Pavel O Bochkov, Sergey V Glagolev, Denis V Rebrikov, Dmitriy O Korostin in Science Progress
Supplemental Material
sj-xlsx-8-sci-10.1177_00368504251398881 - Supplemental material for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants
Supplemental material, sj-xlsx-8-sci-10.1177_00368504251398881 for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants by Dmitry A Sychev, Anastasiia A Buianova, Sherzod P Abdullaev, Karin B Mirzaev, Vera A Belova, Anna O Shmitko, Valery V Cheranev and Oleg N Suchalko, Lyudmila V Fedina, Svetlana V Batyukina, Natalia A Shatalova, Pavel O Bochkov, Sergey V Glagolev, Denis V Rebrikov, Dmitriy O Korostin in Science Progress
Supplemental Material
sj-xlsx-9-sci-10.1177_00368504251398881 - Supplemental material for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants
Supplemental material, sj-xlsx-9-sci-10.1177_00368504251398881 for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants by Dmitry A Sychev, Anastasiia A Buianova, Sherzod P Abdullaev, Karin B Mirzaev, Vera A Belova, Anna O Shmitko, Valery V Cheranev and Oleg N Suchalko, Lyudmila V Fedina, Svetlana V Batyukina, Natalia A Shatalova, Pavel O Bochkov, Sergey V Glagolev, Denis V Rebrikov, Dmitriy O Korostin in Science Progress
Supplemental Material
sj-docx-10-sci-10.1177_00368504251398881 - Supplemental material for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants
Supplemental material, sj-docx-10-sci-10.1177_00368504251398881 for Exome-wide association study of bleeding events in patients receiving direct oral anticoagulants by Dmitry A Sychev, Anastasiia A Buianova, Sherzod P Abdullaev, Karin B Mirzaev, Vera A Belova, Anna O Shmitko, Valery V Cheranev and Oleg N Suchalko, Lyudmila V Fedina, Svetlana V Batyukina, Natalia A Shatalova, Pavel O Bochkov, Sergey V Glagolev, Denis V Rebrikov, Dmitriy O Korostin in Science Progress
Footnotes
ORCID iDs
Ethics approval
This study was carried out in accordance with the guidelines of the Declaration of Helsinki and approved by the local ethics committee of the Russian Medical Academy of Continuous Professional Education, Moscow, Russia (protocol No. 16 and date of approval November 25, 2020; protocol No. 12 and date of approval October 20, 2021; protocol No. 15 and date of approval October 25, 2021). This study was conducted prospectively. The recruitment period began on November 27, 2020, and ended on May 21, 2023. All patient data were anonymized prior to analysis to ensure complete de-identification.
Consent to participate
Written informed consent was obtained from all participants prior to their inclusion in the study.
Consent for publication
Written informed consent was obtained from all participants for the publication of any potentially identifiable data included in this article.
Author contributions
Methodology, AAB; software, VVC and ONS; formal analysis, AAB; investigation, ShPA, VAB, AOSh, POB, VVC and ONS; resources, LVF, SVB, and NASh; project administration, KBM, SVG, and DVR; supervision, DAS and DOK. All authors have read and agreed to the published version of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Russian Science Foundation, grant No. 22-15-00251 “Personalized use of Direct Oral Anticoagulants Based on Pharmacogenomic Approach,” and by the Ministry of Science and Higher Education of the Russian Federation allocated to the Center for Precision Genome Editing and Genetic Technologies for Biomedicine, grant No. 075-15-2019-1789. The authors have no other relevant affiliations or financial involvement with any organization or enterprise that has a financial interest or financial conflict with the subject matter or materials discussed in the manuscript, other than those disclosed.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and material
The datasets generated and analyzed during this study are not publicly available due to ethical restrictions and patient confidentiality protections under Russian Federation laws on personal data protection (Federal Law No. 152-FZ). However, anonymized data supporting the findings may be made available upon reasonable request from qualified researchers to the corresponding author. Requests should include a detailed research proposal and data protection plan.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
