Abstract
Objective
The study investigated the role of the quorum sensing (QS) regulator BpsR4 in the physiology and virulence of Burkholderia pseudomallei strain K96243.
Methods
A B. pseudomallei bpsR4 knockout mutant (ΔbpsR4) was constructed, and its gene expression profile was compared with that of the wild-type (WT) K96243 strain using RNA sequencing. In vitro assays were conducted to assess growth rates, stress responses, antibiotic sensitivity, motility, and biofilm formation. Ex vivo pathogenicity was evaluated using a human skin fibroblast infection model, and in vivo virulence was assessed in the Galleria mellonella model.
Results
bpsR4 deletion led to significant transcriptional reprogramming, including the downregulation of genes involved in iron acquisition, sulfur metabolism, oxidative stress response, and redox homeostasis and upregulation of genes linked to motility, chemotaxis, and membrane transport. ΔbpsR4 exhibited reduced tolerance to oxidative and heat stress and impaired biofilm formation but no significant change in motility. Additionally, ΔbpsR4 displayed decreased susceptibility to meropenem. In both in vitro and invertebrate infection models, the mutant demonstrated lower virulence than the WT strain.
Conclusion
This study highlighted the involvement of BpsR4 in stress response, antimicrobial susceptibility, and virulence in B. pseudomallei. Targeting QS pathways, particularly BpsR4 signaling, might represent a promising strategy to develop anti-virulence therapies that enhance antibiotic efficacy and improve clinical outcomes in melioidosis.
Introduction
Burkholderia pseudomallei is a gram-negative, rod-shaped, motile bacterium responsible for melioidosis, a potentially fatal infectious disease endemic to tropical regions, particularly Southeast Asia and northern Australia.1–3 Infection occurs through inhalation, ingestion, or direct skin inoculation,4,5 and its clinical manifestations range from asymptomatic infection to severe conditions such as pneumonia, septicemia, internal abscesses, and encephalomyelitis. 6 The global burden of B. pseudomallei infection is substantial, with an estimated 165,000 cases and 89,000 deaths annually, and its mortality rates range 9%–70%. 7 The disease is often underdiagnosed because of its non-specific symptoms and a lack of clinical awareness, leading to delayed treatment and poor outcomes. 8 The intrinsic resistance of the bacterium to multiple antibiotics further complicates treatment.9,10
The genome of B. pseudomallei contains two chromosomes with distinct functional roles. Specifically, one genome encodes essential metabolic functions, and the other encodes adaptive traits such as virulence and environmental survival.11,12 The bacterium's pathogenicity arises from its extensive arsenal of virulence factors, including type III and VI secretion systems (T3SS and T6SS), biofilm formation, flagella, pili, surface polysaccharides, and regulatory mechanisms such as quorum sensing (QS).13–16 These elements enable intracellular survival, immune evasion, and colonization of both phagocytic and non-phagocytic host cells.17–19
Central to the coordination of these virulence mechanisms is the QS system, which regulates gene expression in response to cell density via signaling molecules such as N-acyl-homoserine lactones (AHLs).20,21 In B. pseudomallei, the QS circuitry includes at least three canonical luxIR homolog pairs (bpsI1–bpsR1, bpsI2–bpsR2, bpsI3–bpsR3), together with two orphan LuxR homologs (bpsR4 and bpsR5).22–24 The canonical regulatory pair BpsI–BpsR facilitates C8-HSL synthesis and detection, enabling coordinated regulation of genes associated with biofilm maturation, motility, exopolysaccharide production, and antibiotic resistance.25,26
The transcriptional regulator BpsR has traditionally been considered part of a positive feedback loop with BpsI.23,24,27 However, emerging evidence from related bacterial systems suggests that LuxR-type proteins can both activate and repress gene expression depending on environmental and cellular contexts. This regulatory flexibility implies that BpsR's influence extends beyond bpsI activation and includes global modulation of virulence and stress adaptation genes.
Notably, most prior studies focused on bpsI mutants, leaving the regulatory scope of bpsR, particularly orphan variants such as BpsR4, underexplored. BpsR4 can function as a global regulator capable of modulating physiological pathways independent of AHL synthesis, potentially responding to alternative signals. Investigating bpsR4 mutants thus provides a unique opportunity to uncover QS-independent regulatory networks.
In this study, we hypothesized that BpsR4 serves as a master regulator of key phenotypes related to the pathogenicity and environmental adaptation of B. pseudomallei. To investigate this, we constructed a bpsR4 deletion mutant and performed transcriptomic analyses using RNA sequencing (RNA-Seq) to identify genes under BpsR4 control. Additionally, we conducted phenotypic assays to examine changes in stress resistance, antibiotic susceptibility, motility, and biofilm formation. Our findings highlighted the critical role of BpsR4 in controlling gene expression related to virulence, antibiotic resistance, and environmental resilience. These insights deepen our understanding of QS in B. pseudomallei, and they could potentially inform the development of QS-disrupting therapeutics aimed at mitigating melioidosis severity and improving clinical outcomes.
Methods
Bacterial strains, cell lines, and growth conditions
B. pseudomallei K96243 served as the wild-type (WT) strain in this study. B. pseudomallei were cultured in Luria–Bertani (LB) medium at 37°C.
Primary normal human dermal fibroblasts (NHDFs; C-12302, PromoCell, Heidelberg, Germany) were maintained in growth medium (PromoCell). The cells were cultured at 37°C in a 90% humidity-controlled incubator with 5% CO2, and the culture medium was replaced every 2 days. When the cells reached approximately 90% confluency, they were detached from the flasks using trypsin-EDTA solution (PromoCell).
Construction of B. pseudomallei ΔbpsR4 mutant and complementary strain
The B. pseudomallei ΔbpsR4 mutant and complementary strain were constructed using the markerless allele replacement method with some modifications. 28 To delete bpsR4, a DNA fragment consisting of upstream and downstream regions of B. pseudomallei, derived from GenBank (locus_tag = Bpss0312), was synthesized and cloned into the pUC57 vector (GenScript, Piscataway, NJ, USA). Subsequently, the fragment was cut via digestion with SmaI and EcoRI (New England Biolabs, Ipswich, MA, USA), followed by ligation to the pEXKm5 vector using the same restriction enzymes. The pEXKm5 plasmid containing the fragment was transformed into Escherichia coli RHO3 and mobilized into B. pseudomallei K96243 using the conjugation method. The obtained conjugants were selected on LB agar containing 1000 µg/mL kanamycin and incubated for 24–48 h at 37°C. To resolve the merodiploid, the bacteria were grown in yeast extract–tryptone agar containing 15% (w/v) sucrose at room temperature. The isolated colonies were screened for kanamycin-sensitive clones. bpsR4 deletion was validated by polymerase chain reaction (PCR) using primers flanking the deleted alleles (F-BpsR4 and R-BpsR4; Table 1). Additionally, the absence of the pEXKm5 plasmid was confirmed using oriT primers (Table 1).
To facilitate gene complementation, a similar pEXKm5-based allele exchange method was utilized. The full-length sequence of B. pseudomallei K96243 bpsR4 was amplified from genomic DNA using the primers F-BpsR4 and R-BpsR4 (Table 1). Then, the DNA product was subjected to SmaI and EcoRI digestion and subsequently ligated with the SmaI- and EcoRI-digested pEXKm5 vector. Similarly to that performed for ΔbpsR4 mutant, the pEXKm5 containing the entire sequence of bpsR4 was transformed into E. coli RHO3 for conjugation into the B. pseudomallei ΔbpsR4 mutant. The complementation of bpsR4 was confirmed by PCR using primers flanking the deleted alleles (F-BpsR4 and R-BpsR4; Table 1) and verified by DNA sequencing.
Primers used in this study.

Transcriptomic study
Sample preparation
A single colony each of the WT and mutant strains (three biological replicates each) was subcultured 1:10 in 5 mL of LB broth and incubated at 37°C with shaking at 200 rpm for 18 h. Total RNA was extracted from B. pseudomallei strains using an easy-BLUE™ Total RNA Extraction Kit (iNtRON Biotechnology, Gyeonggi-do, South Korea) and treated with RNase-free DNase I (New England Biolabs). Then, the quality of the RNA samples was determined by measuring the optical density (OD) at 260 and 280 nm using a NanoDrop Lite spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA).
RNA-Seq
Libraries were prepared for 100-bp paired-end sequencing using a TruSeq RNA Sample Preparation Kit (Illumina, San Diego, CA, USA). Namely, mRNA molecules were purified and fragmented from 2 µg of total RNA using oligo (dT) magnetic beads. The fragmented mRNAs were synthesized as single-stranded cDNAs through random hexamer priming. Double-stranded cDNA was prepared using the single-stranded cDNAs as the template. After sequential end repair, A-tailing, and adapter ligation, cDNA libraries were amplified by PCR. The quality of the cDNA libraries was evaluated using the Agilent 2100 BioAnalyzer (Agilent, Santa Clara, CA, USA). The libraries were quantified using the KAPA library quantification kit (Kapa Biosystems, Wilmington, MA, USA) according to the manufacturer's protocol. Following cluster amplification of denatured templates, sequencing was performed as paired-end (2 × 100 bp) using the Illumina HiSeq 2500 system.
Transcriptome data analysis
Raw sequencing reads were filtered using in-house scripts to remove low-quality reads based on the following criteria: > 10% undetermined bases, > 40% of bases with quality scores <20, and average quality score <20. High-quality reads were aligned to the B. pseudomallei reference genome using STAR v2.4.0b. Gene expression was quantified by Cufflinks v2.1.1 using species-specific annotation, excluding non-coding regions and applying multi-read and fragment bias corrections. Differentially expressed genes (DEGs) were identified using the TCC package in R (The R Foundation for Statistical Computing, Vienna, Austria), which applies DEGES/edgeR-based normalization. Read counts were generated with HTSeq-count v0.6.1p1 (https://htseq.readthedocs.io/en/master/), and DEGs were selected on the basis of q-value < 0.05 (adjusted using the Benjamini–Hochberg method). Gene Ontology (GO) enrichment analysis was performed using Fisher's exact test (p < .001) to identify significantly enriched biological processes, cellular components, and molecular functions. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis was also conducted to identify significantly enriched metabolic and signaling pathways associated with DEGs. Enrichment significance was assessed using adjusted p-values, and gene counts were used to represent the number of B. pseudomallei BpsR4-associated genes within each KEGG category.
Bacterial growth assay
A single isolated colony of B. pseudomallei was inoculated in LB broth and incubated at 37°C with shaking at 200 rpm for 24 h. Then, the overnight culture of bacteria was washed with phosphate-buffered saline (PBS) and adjusted to OD600 of 0.5. To examine growth kinetics, the prepared bacteria were added to fresh LB medium at a ratio of 1:500 and incubated at 37°C with shaking at 200 rpm. OD600 was measured at predetermined time points.
Heat resistance assay
A heat stress resistance assay was performed as described previously with some modifications.
29
Briefly, B. pseudomallei strains cultured in LB medium at 37°C for 18 h were washed with PBS and resuspended in PBS to OD600 of 0.15. One milliliter of the bacterial suspension was then added into a prewarmed tube and incubated at 50°C for 15 min. Before and after heat challenge, bacterial survival was determined on LB agar plates after incubation at 37°C for 24 h. The number of surviving bacteria was expressed as a percentage of the number of viable cells as follows: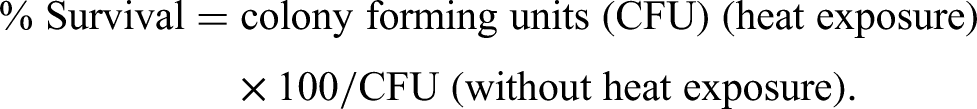
Oxidative stress assay
The survival of B. pseudomallei under oxidative conditions was investigated as described previously with some modifications.
29
Briefly, B. pseudomallei strains cultured in LB medium at 37°C for 18 h were washed with 1× PBS. The bacterial concentration was adjusted to OD600 of 0.15. Then, 100 µL of the bacterial suspension were treated with H2O2 (at a final concentration of 1 µmol/L) or left untreated at room temperature for 15 min. Treated and untreated bacteria were diluted 10-fold and plated on LB agar. After incubation at 37°C for 24 h, colonies were counted, and the number of colonies was compared between treated and untreated bacteria (without oxidant) and presented as percent survival as follows: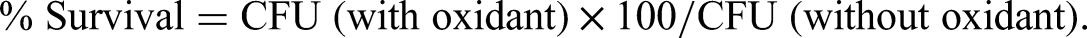
Motility assays
Motility assays were performed as previously described. 29 In brief, approximately 1 × 106 CFU of B. pseudomallei were cultured overnight on nutrient agar plates. The motility of the bacteria was analyzed on 0.5% agar plates supplemented with 5 g/L tryptone and 2.5 g/L NaCl. To reduce variation between plates, 30 mL of medium were poured into each plate in a laminar flow hood and left for 1 h with the lids off, after which the plates were used immediately. A 1-µL drop of freshly grown culture (OD600 of approximately 0.6) was then placed at the center of each plate to evaluate swarming motility, which was assessed by measuring the diameter of the swarming zones. For each bacterial strain, assays were performed at least three times.
Biofilm production and cell morphology
To assess biofilm formation, B. pseudomallei colonies were dissolved in PBS (pH 7.4), and the turbidity of the solutions was adjusted to 0.5 McFarland standard. Twenty microliters of the prepared cells were added to a 96-well microtiter plate containing 180 µL of LB broth and incubated at 37°C for 24 h, after which the fluids were drained. The wells were washed with 200 µL of PBS (pH = 7.4), and the cells were stained with 1% crystal violet and incubated at 37°C for 30 min. The wells were washed with water and allowed to air-dry at room temperature. The amount of biofilm produced by the bacteria was determined via destaining by adding 200 µL of 33% acetic acid. The absorbance of the resulting solution was measured at 595 nm using a microplate spectrophotometer. The test was repeated five times.
Antimicrobial sensitivity assays
The minimum inhibitory concentrations (MICs) of antibiotics (tetracycline, imipenem, meropenem, ceftazidime, trimethoprim/sulfamethoxazole, amoxicillin/clavulanate, and chloramphenicol) against B. pseudomallei were determined using the Epsilometer test (E-test; Liofilchem, Roseto Degli Abruzzi, Italy). Susceptibility testing and interpretation were conducted in accordance with the guidelines provided by the European Committee on Antimicrobial Susceptibility Testing.
Invasion and intracellular replication assay
NHDFs were seeded at a density of 5 × 104 cells per well in a 24-well plate. The next day, the medium was removed and replaced with 200 µL of fresh antibiotic-free growth medium. Overnight cultures of B. pseudomallei strains were adjusted to 1 × 106 cells/mL by measuring OD600 and used to infect the cells at a multiplicity of infection (MOI) of 20–100. After a 2 h co-culture, the infected cells were washed twice with PBS, and 500 µL of fresh growth medium containing 250 µg/mL kanamycin (Sigma-Aldrich, St Louis, MO, USA) were added, followed by incubation at 37°C for 1 h to eliminate any extracellular bacteria. To recover the invading bacteria, B. pseudomallei-infected NHDFs were then washed thrice with PBS before cell lysis with 100 µL of 0.1% (w/v) Triton X-100. The number of viable bacteria was determined as the CFU by performing a serial dilution. Ten microliters of each dilution were dropped onto LB agar plates and incubated at 37°C for 24–48 h.
At 4, 6, 8, and 10 h post-infection, the intracellular bacteria were recovered as previously described to assess the intracellular replication of B. pseudomallei strains in NHDFs.
Determination of multinucleated giant cell (MNGC) formation
At 10 h post-infection, NHDFs cells infected with B. pseudomallei strains were stained with Giemsa (Merck, Darmstadt, Germany) as previously described. 30 MNGC formation, as defined by the presence of at least three nuclei in a cell, was evaluated in 100 fields of view using an Olympus BX41 microscope (Olympus, Tokyo, Japan). The rate of MNGC formation was determined using the following formula: (number of nuclei in a multinucleated giant cell/total number of nuclei counted) × 100. A minimum of 1000 nuclei were counted per experiment.
Galleria mellonella killing assay
G. mellonella killing assays were performed as previously described. 31 Forty larvae (length, 2–2.5 cm; weight, 250–300 mg; melanization-free) were used in this experiment. After 18 h of growth in broth with shaking, the concentration of B. pseudomallei was adjusted to 100 CFU in PBS. A Hamilton syringe was used to inject an aliquot of the bacterial suspension into the body cavity of G. mellonella larvae via the proleg. Each control larva was injected with PBS. Following injection, larvae were incubated in the dark at 37°C. At 24–48 h post-injection, larvae were individually investigated for pigmentation and mobility. Larvae were considered dead when they displayed no movement in response to gentle prodding with a pipette tip. The number of dead larvae and the times of death were recorded, and the survival graph was plotted. This study followed the appropriate EQUATOR Network guideline, the ARRIVE (Animal Research: Reporting of In Vivo Experiments) guidelines for reporting animal research. 32
Statistical analysis
All assays were conducted in biological triplicate. Analysis of variance were used to test statistical differences between the control and test groups. For the G. mellonella killing assay, the log-rank (Mantel–Cox) test was used to compare survival curves. Statistical analyses were conducted using GraphPad Prism version 10.6.1 (GraphPad, Boston, MA, USA). Significance was denoted by p < .05.
Results
Generation of stable ΔbpsR4 and complemented strains in B. pseudomallei K96243
To investigate the role of QS in B. pseudomallei, a stable, markerless bpsR4 knockout mutant was constructed using a pEXKm5-based allelic replacement method. Successful deletion of bpsR4 was confirmed by PCR and DNA sequencing (Figure S1 in the supplementary materials). A complemented strain (ΔbpsR4::bpsR4) was generated by reintroducing bpsR4 into the ΔbpsR4 mutant using the same vector (pEXKm5), following previously established protocols. Complementation was similarly confirmed by PCR and DNA sequencing (Figure S1 in the supplementary materials).
Growth analysis in nutrient-rich LB medium revealed that the ΔbpsR4 mutant and complemented strains had no significant differences in growth rates versus the WT strain (Figure 1). These results indicate that bpsR4 deletion does not impair overall bacterial growth under these conditions. Subsequently, WT B. pseudomallei K96243 and the ΔbpsR4 mutant were subjected to transcriptomic analysis.

Growth kinetic of B. pseudomallei. The growth of B. pseudomallei K96243, ΔbpsR4 mutant, and the complementary strain in LB broth was determined by OD600 measurement. The data points and error bars represent mean OD600 and the standard deviations (SD) from triplicate experiments.
Transcriptomic profiling reveals differential gene expression in ΔbpsR4
To investigate the global transcriptional impact of BpsR4 in B. pseudomallei, we performed RNA-Seq to compare the ΔbpsR4 and WT strains. RNA was isolated from cultures at 18 h for transcriptome analysis because this time point corresponds to the late exponential/early stationary phase, when QS activity and associated regulatory pathways are likely active in B. pseudomallei and coordinated with broader stress response networks, including regulation by RpoS. 33 Our transcriptomic analysis revealed that BpsR4 regulates a diverse regulon comprising 545 genes. Hierarchical clustering of the top DEGs was visualized using a heatmap (Figure 2). The heatmap suggested a clear and consistent separation between the ΔbpsR4 and WT transcriptomes, with all replicates clustering tightly within their respective groups. This indicates high intragroup consistency and highlights substantial transcriptional divergence between the mutant and WT strains.

Transcriptomic heatmap showing DEGs of B. pseudomallei. Hierarchical clustering heatmap of the top DEGs obtained from RNA-Seq analysis of B. pseudomallei WT K96243 strain (K9) and ΔbpsR4 mutant (dBpsR4), each represented by three biological replicates. Gene expression values are presented as row-wise Z-scores of normalized counts. Red and blue colors indicate upregulation and downregulation, respectively, relative to the mean expression across all samples. The heatmap reveals distinct clustering between the mutant and WT groups, highlighting extensive transcriptional reprogramming. Gene annotations are shown on the right, with corresponding Ensembl gene IDs. Dendrograms indicate hierarchical relationships based on gene expression similarity.
The expression profiles revealed two major gene clusters in the ΔbpsR4 mutant relative to the WT strain, including one significantly downregulated and one markedly upregulated cluster. These patterns suggest that BpsR4 functions as a global transcriptional regulator capable of both activating and repressing gene expression. Notably, several genes involved in iron acquisition (e.g., fhuA, pvdA), sulfur assimilation (e.g., cysI, cysL), and oxidative stress response (e.g., catC, katG) were downregulated in the mutant, indicating that BpsR4 positively regulates pathways associated with stress adaptation and nutrient metabolism. Conversely, genes related to motility, outer membrane transport, and several uncharacterized hypothetical proteins were upregulated, suggesting potential derepression of these pathways in the absence of BpsR4. These findings demonstrate that BpsR4 plays a critical regulatory role in orchestrating multiple functional networks in B. pseudomallei, including those associated with virulence, environmental adaptation, and metabolic flexibility, through pathway-specific mechanisms.
To further explore the biological significance of these DEGs, we performed GO and KEGG enrichment analyses. Figure 3 summarizes the enriched GO and KEGG terms derived from the transcriptomic data. The bar graph depicts significantly upregulated and downregulated biological processes in the ΔbpsR4 mutant compared with those in the WT strain, with enrichment scores calculated as –log10 (p-value). Among the downregulated genes in the ΔbpsR4 mutant, GO analysis revealed significant enrichment in biological processes such as iron ion transport (GO: 0006826), sulfur compound metabolic process (GO: 0006790), response to oxidative stress (GO: 0006979), and cell redox homeostasis (GO: 0045454). Consistent with these results, KEGG pathway analysis highlighted significant downregulation of sulfur metabolism (bpe00920), ABC transporters (bpe02010), and glutathione metabolism (bpe00480). These findings suggest that BpsR4 positively regulates essential pathways related to nutrient acquisition, redox balance, detoxification, and stress response. Conversely, upregulated genes in ΔbpsR4 were enriched in GO terms such as flagellum-dependent cell motility (GO: 0071973), chemotaxis (GO: 0006935), and membrane transport (GO: 0055085). KEGG pathway enrichment further revealed significant involvement of the upregulated genes in bacterial chemotaxis (bpe02030) and flagellar assembly (bpe02040). These observations suggest that motility and environmental sensing pathways, particularly those responsive to chemical stimuli, are derepressed in the absence of BpsR4, potentially as a compensatory mechanism to altered regulatory dynamics. Collectively, the transcriptomic and enrichment analyses revealed that BpsR4 functions as a global regulator modulating diverse physiological processes, including bacterial growth and survival, stress resistance, metabolic adaptation, and virulence-associated traits, in B. pseudomallei.

Potential functional association of BpsR4 based on GO and KEGG enrichment analyses. Bar graph showing selected GO biological processes and KEGG pathways significantly enriched among DEGs in the ΔbpsR4 mutant compared to the B. pseudomallei K96243 strain. Enrichment scores are expressed as –log12(adjusted p-value). Blue bars indicate downregulated categories, while red bars represent upregulated ones. The analysis highlights BpsR4-dependent regulation of oxidative stress response, sulfur metabolism, nutrient transport, and motility-related functions.
BpsR4 contributes to oxidative stress resistance and heat tolerance
To evaluate the role of BpsR4 in oxidative stress responses, equal numbers of bacterial cells were exposed to 1 µmol/L H2O2 for 15 min, followed by plating on LB agar to assess survival (Figure 4A). In the presence of H₂O₂, the ΔbpsR4 mutant had a significantly lower survival rate than the WT strain (1.45% ± 0.22% vs. 3.97% ± 1.26%, p = .0379). These findings indicate that BpsR4 is involved in mediating resistance to oxidative stress in B. pseudomallei.
The role of BpsR4 in heat resistance was also investigated. As presented in Figure 4B, the percentage of surviving bacteria was significantly lower in ΔbpsR4 than in the WT strain (13.71% ± 2.79% vs. 19.31% ± 1.25%, p = .0473). Notably, the complemented strain displayed comparable survival as the WT strain. These results suggest that BpsR4 contributes to B. pseudomallei's resistance to heat stress.

Oxidative and heat stress response of B. pseudomallei. (A) Susceptibility to oxidative stress of B. pseudomallei K96243, ΔbpsR4 mutant, and the complementary strain to killing by 1 µmol/L H2O2. The treatment was determined at 15 min. Surviving bacteria were enumerated on LB agar plates after incubation at 37°C for 24 h and were expressed as the % survival. The data were obtained from three experiments. (B) Percent survival of B. pseudomallei K96243, ΔbpsR4 mutant, and the complementary strain after heat treatment at 50°C for 15 min. 100% viability corresponds to the colony forming unit count of unexposed bacteria. Data represent the mean ± SD from three independent experiments. Individual data points are shown as dots to illustrate the distribution of experimental replicates. *p < .05.
BpsR4 modulates antibiotic responses in B. pseudomallei
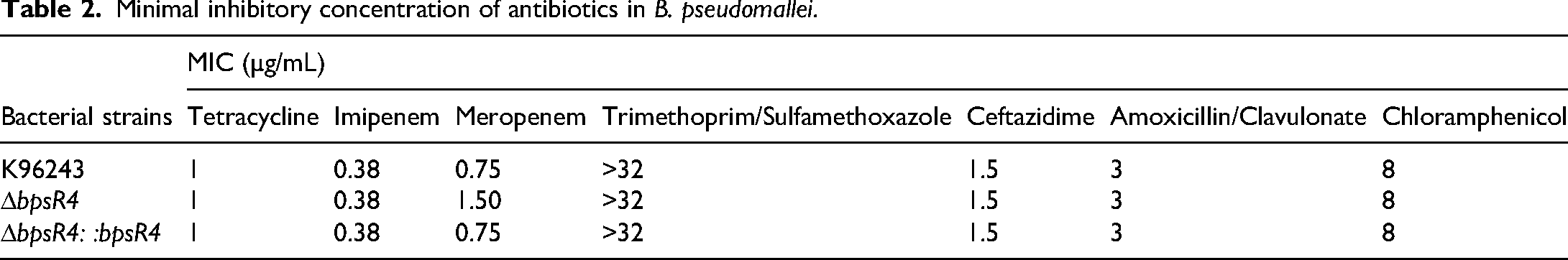
The impact of bpsR4 deletion on the antibiotic resistance profile of B. pseudomallei was assessed via comparisons among the ΔbpsR4 mutant, WT K96243, and ΔbpsR4::bpsR4 strains. Overall, the MICs for most antibiotics tested were comparable between the ΔbpsR4 mutant and WT strain, indicating that resistance to these antibiotics is largely bpsR4-independent. However, the ΔbpsR4 mutant exhibited an increased MIC for meropenem (Table 2), and this alteration was reversed in the complemented strain. These findings suggest that bpsR4 deletion can influence susceptibility to meropenem, indicating a potential regulatory role for bpsR4 in meropenem resistance in B. pseudomallei.
BpsR4 influences biofilm formation in B. pseudomallei
Biofilm formation was assessed in the WT, ΔbpsR4 mutant, and ΔbpsR4::bpsR4 strains by crystal violet staining (Figure 5A). Statistical analysis revealed significantly lower biofilm formation in ΔbpsR4 than in the WT (p = .0053). By contrast, biofilm formation did not significantly differ between the ΔbpsR4::bpsR4 and WT strains (p > .05). These findings indicate that bpsR4 deletion impairs biofilm formation, whereas complementation restores this phenotype, emphasizing the essential role of bpsR4 in promoting robust biofilm development in B. pseudomallei.
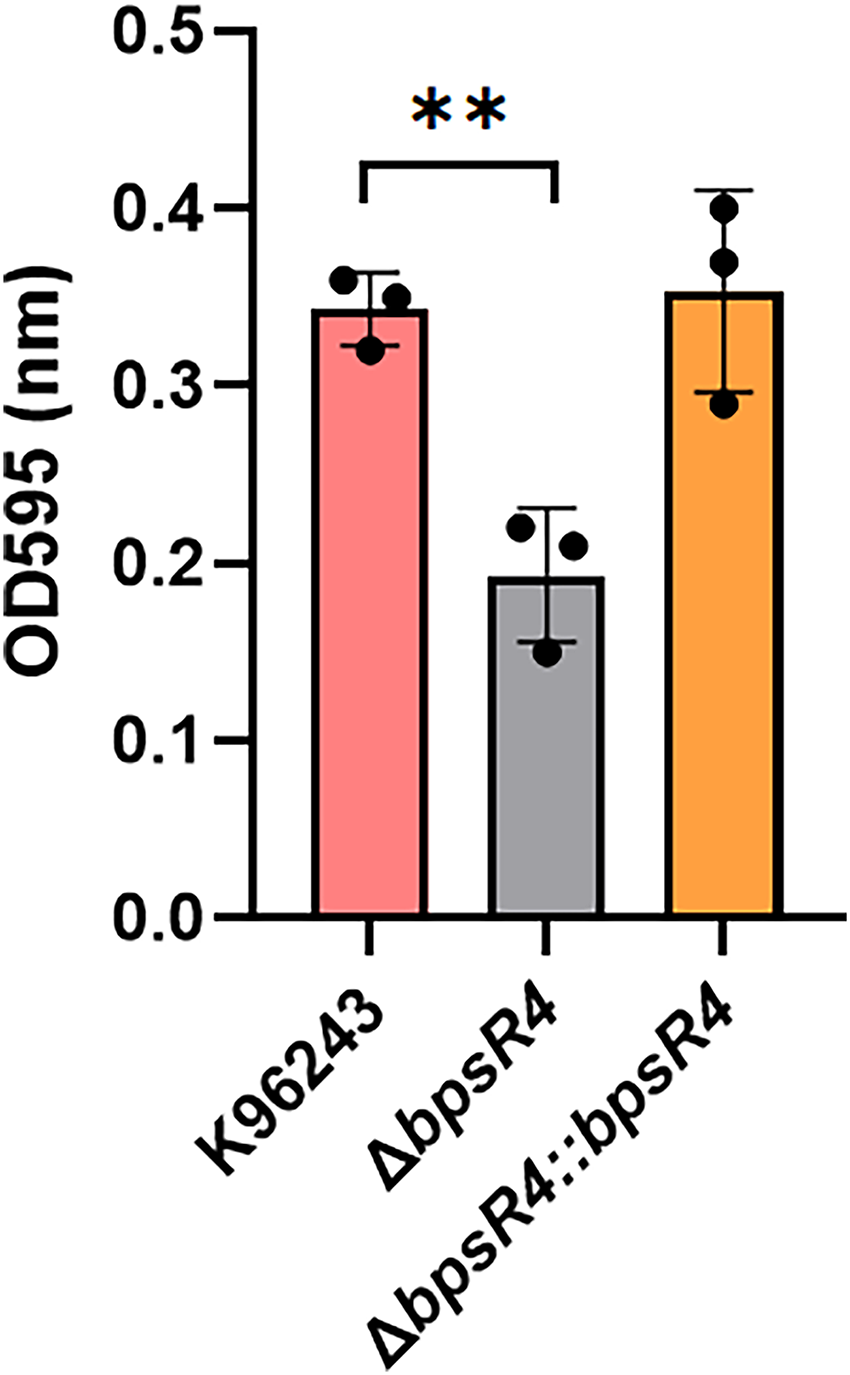
Biofilm formation of B. pseudomallei. Biofilm formation of B. pseudomallei K96243, ΔbpsR4 mutant, and the complementary strain was determined by a microtiter plate assay. Biofilm formation of B. pseudomallei was induced in trypticase soy broth at 37°C for 24 h. After incubation, the adherent bacteria were stained for 15 min with 1% crystal violet. The quantity of biofilm was measured at 630 nm. Data represent the mean ± SD from three independent experiments. Individual data points are displayed as dots to illustrate the variability among replicates. **p < .001.
Minimal inhibitory concentration of antibiotics in B. pseudomallei.
BpsR4 is not required for swarming motility
We investigated whether bpsR4 deletion affects swarming motility in B. pseudomallei. As presented in Table S1 in the supplementary materials, the size of the swarming zone did not differ between the ΔbpsR4 mutant and WT. Thus, bpsR4 deletion does not apparently affect the swarming motility of B. pseudomallei.
Loss of BpsR4 reduces intracellular survival in host cells
To understand the mechanism by which BpsR4 mediates cell pathogenesis, we performed in vitro infection assays using NHDFs infected with the WT B. pseudomallei K96243 or ΔbpsR4 mutant strain. The invasion efficiencies of B. pseudomallei strains in NHDFs were 0.42% ± 0.02%, 0.41% ± 0.10%, and 0.43% ± 0.01% at 3 h post-infection with the WT, ΔbpsR4, and ΔbpsR4::bpsR4 strains, respectively (p = 0.467, Figure 6A). Thus, the absence of bpsR4 did not affect the ability of B. pseudomallei to invade NHDFs. Meanwhile, the WT and ΔbpsR4 strains exhibited decreased intracellular replication rates in NHDFs for up to 10 h post-infection (p = 0.034, Figure 6B), suggesting that BpsR4 is required for intracellular replication in B. pseudomallei in NHDFs.
Intercellular spread during intracellular survival can be assessed according to the number of MNGCs. The percentage of MNGCs at 10 h post-infection with the ΔbpsR4 mutant was 11.57% ± 0.81%, compared with 18.42% ± 2.99% for the WT strain (p = .0441, Figure 7). Meanwhile, the percentage of MNGCs was comparable between the WT and complemented strains (Figure 7). Collectively, these findings demonstrate the positive regulatory role of BpsR4 in intracellular replication and intercellular spread of B. pseudomallei in NHDF cells.

Invading and surviving B. pseudomallei in NHDFs. NHDFs were infected with B. pseudomallei K96243, ΔbpsR4 mutants, and the complementary strains at an MOI of 20. (A) The invading bacteria into the NHDFs were recovered at 3 h post-infection. (B) The intracellular bacteria in the NHDFs were determined at 4, 6, 8, and 10 h post-infection. Values are presented as the mean ± SD from three independent experiments. Individual data points are shown as dots to illustrate replicate variation. *p < .05.

MNGC formation of NHDFs infected by B. pseudomallei. A) MNGC formation induced by B. pseudomallei K96243, ΔbpsR4 mutant, and the complementary strain. NHDFs were stained with Giemsa. Images were captured by standard light microscopy with a 40× objective lens. B) The percentage of MNGC formation induced by B. pseudomallei strains. Values are presented as the mean ± SD from three independent experiments. Individual data points are shown as dots to illustrate variability among replicates. *p < .05.
BpsR4 is essential for full virulence in the G. mellonella infection model
G. mellonella larvae are widely used as in vivo models to study various pathogens.34,35 In this study, we explored the effect of BpsR4 on bacterial virulence using an in vivo G. mellonella infection model. In comparison with the findings for WT K96243-infected G. mellonella, the number of dead worms was significantly lower for ΔbpsR4 strain-infected G. mellonella (p < .0001, Figure 8). B. pseudomallei WT-infected G. mellonella exhibited 100% mortality at 36 h post-infection, whereas ΔbpsR4-infected G. mellonella exhibited 100% mortality at 42 h post-infection (Figure 8). No mortality was observed in PBS-infected larvae.

Testing of B. pseudomallei virulence in an in vivo infection model. G. mellonella larvae were infected with 1 CFU of each strain of B. pseudomallei K96243, ΔbpsR4 mutant, and the complementary strain. The survival of larvae was monitored at 24 h post-infection until to 48 h post-infection. Each data set is representative of a single trial with the percent survival of infected larvae. Control larvae (PBS-injected larvae) did not die in any given trial.
Discussion
Our study demonstrated that BpsR4, a component of the QS regulatory system in B. pseudomallei, plays a central role in orchestrating gene expression programs essential for bacterial survival, environmental adaptation, and pathogenicity. Transcriptomic analysis of the ΔbpsR4 mutant and WT B. pseudomallei K96243 strains revealed that BpsR4 acts as a global regulator, modulating both activation and repression of key biological pathways encompassing a substantial regulon of 545 genes. Given the diversity of the pathways influenced, it is likely that BpsR4 also exerts indirect effects on gene expression by modulating additional transcription factors, thereby amplifying its impact on cellular physiology.
Notably, the loss of BpsR4 led to the significant downregulation of genes involved in iron acquisition (e.g., fhuA, pvdA), sulfur assimilation (e.g., cysI, cysL), and oxidative stress responses (e.g., catC, katG), indicating that BpsR4 positively regulates nutrient acquisition and stress resilience, which are crucial functions for survival in hostile environments, including during host infection. These transcriptomic shifts were further supported by the results of GO and KEGG enrichment analyses, which revealed marked enrichment in biological processes related to redox homeostasis, ABC transport, sulfur metabolism, and detoxification. Collectively, these findings position BpsR4 as a key activator of metabolic and stress-responsive systems that support the environmental persistence and intracellular fitness of B. pseudomallei.
Conversely, genes associated with motility, chemotaxis, and membrane transport were upregulated in the ΔbpsR4 mutant, suggesting that BpsR4 also represses locomotion and environmental sensing pathways under specific conditions. Enrichment analysis revealed increased representation of genes involved in flagellar assembly and bacterial chemotaxis. This might reflect a compensatory regulatory response in which the bacterium enhances motility and chemotaxis to offset stress or metabolic imbalance, potentially to relocate to more favorable niches.
These findings are consistent with the broader roles of LuxR-type QS regulators across gram-negative bacteria. In Aeromonas hydrophila, the LuxR-type regulator luxR08110 globally regulates gene networks related to sulfur metabolism, motility, and stress response, thereby promoting intracellular survival in host macrophages. 36 Knockdown of luxR08110 reduced motility, adhesion, and survival under oxidative and acidic stress, similar to the phenotypes of the ΔbpsR4 strain. Similarly, in Vibrio fischeri, the canonical LuxR regulator controls bioluminescence and a wide range of other genes in response to QS signals such as 3OC6-HSL. 37 In V. scophthalmi, LuxRV regulates biosynthesis, osmoprotection, and oxidative stress responses; and represses biofilm formation, and promotes PHA biosynthesis and stress tolerance. 38 These examples illustrate the conserved yet context-specific regulatory flexibility of LuxR-type proteins across species. Taken together, our findings identify BpsR4 as a central regulatory node within B. pseudomallei's QS network that contributes to the ecological versatility and virulence potential of the bacterium. Through dual regulatory control, BpsR4 enables precise modulation of gene expression to optimize bacterial fitness under fluctuating environmental or host-related conditions. Given its broad regulatory scope, targeting BpsR4 or its downstream effectors might represent a strategy to attenuate virulence and improve therapeutic outcomes in melioidosis.
The observation that the ΔbpsR4 mutant and WT strains exhibited comparable growth rates in LB medium suggests that BpsR4 is not essential for bacterial replication under nutrient-rich conditions. This aligns with findings in other gram-negative bacteria, in which LuxR-type QS regulators such as LasR (Pseudomonas aeruginosa) and AbaR (Acinetobacter baumannii) primarily regulate behaviors such as virulence, biofilm formation, and motility rather than core growth functions.39,40 The absence of a growth defect in ΔbpsR4 supports the idea that BpsR4 functions under specific environmental cues, particularly those involving host interactions or nutrient limitations. This also reinforces the hypothesis that QS in B. pseudomallei operates in a modular fashion, with distinct regulators driving different phenotypic traits independently of proliferation.
Our results also highlighted BpsR4's role in stress adaptation in B. pseudomallei, as the ΔbpsR4 strain exhibited markedly reduced survival under oxidative and heat stress, both of which were rescued by complementation. These findings suggest that BpsR4 exerts regulatory effects beyond canonical QS roles to include protective stress responses. In P. aeruginosa, LasR and RhlR regulate oxidative stress by modulating catalase and superoxide dismutase expression. 41 Similarly, in Staphylococcus aureus, the Agr system activates redox-sensitive networks that confer resistance to oxidative damage. 42 QS systems also contribute to thermal tolerance in other species; for example, luxS disruption in Glaesserella parasuis increases heat sensitivity, 43 and LuxR regulators have been found to interact with temperature-sensitive sigma factors such as σ^E. 44 Some LuxR regulators even function as thermosensors, adjusting QS output in response to temperature changes. The dual role of BpsR4 in mediating both oxidative and heat tolerance suggests that it regulates a broader stress response network critical for B. pseudomallei's survival in harsh environments, such as tropical soils or within host cells.
In the context of antibiotic resistance, BpsR4 appears to influence susceptibility to meropenem in B. pseudomallei. Although the susceptibility to most antibiotics was not altered by bpsR4 deletion, the ΔbpsR4 mutant exhibited increased resistance to meropenem, which was reversed by complementation. This indicates a specific regulatory role for BpsR4 in modulating carbapenem susceptibility. In P. aeruginosa, QS regulators such as LasR and RhlR modulate β-lactam susceptibility by regulating efflux pumps, membrane permeability, and biofilm pathways. 45 The selective impact of BpsR4 on meropenem, as opposed to other β-lactams, suggests it can regulate specific carbapenem resistance determinants. The finding that complementation restored susceptibility further supports the specificity and functional relevance of this regulation. Therefore, future proteomic studies could help identify downstream effectors of BpsR4 that mediate meropenem resistance, building upon the transcriptomic insights presented in this research. More broadly, in antimicrobial research, disrupting QS regulators such as BpsR4 could enhance antibiotic efficacy and reduce the risk of resistance development.
Furthermore, our findings confirmed that BpsR4 is a key regulator of biofilm formation. The ΔbpsR4 mutant displayed significantly impaired biofilm development, reinforcing the role of QS regulators in biofilm regulation of B. pseudomallei, consistent with previous reports.22,23 Similar roles have been described for LasR and RhlR in P. aeruginosa and HapR in V. cholerae, with these proteins modulating biofilm formation based on signal concentrations and population density.46–48 Although B. pseudomallei possesses a distinct QS system from those of other species, the impaired biofilm formation observed in the ΔbpsR4 mutant suggests that BpsR4 performs a similar regulatory role as well-known QS regulators in other bacteria. This underscores the potential of BpsR4 as a target for anti-biofilm strategies, particularly given the role of biofilms in orchestrating multicellular behaviors that contribute to chronic infection and antimicrobial resistance.
Regarding motility, transcriptomic data indicated upregulation of motility-related genes in the ΔbpsR4 mutant. However, swarming assays revealed no significant differences among the strains, suggesting that gene expression changes do not necessarily translate into altered motility under the conditions tested. Alternative assays or environmental stimuli might be required to fully evaluate BpsR4's role in flagellar function.
To further investigate BpsR4's role in pathogenesis, we employed an in vitro model using NHDFs. Despite its broad regulatory influence, BpsR4 did not affect cellular invasion efficiency. This indicates that BpsR4 is not involved in regulating the genes required for host cell entry. Importantly, this challenges the common assumption that QS broadly governs all virulence-associated traits and implies that the mechanisms underlying invasion by B. pseudomallei might function independently of BpsR4-mediated regulation. Contrarily, bpsR4 deletion led to a significant reduction in MNGC formation, an essential feature of intracellular spread, and this effect was reversed upon complementation. The reduced MNGC formation observed in the ΔbpsR4 mutant was likely influenced by the decreased intracellular bacterial burden at 10 h, resulting in fewer bacteria available to deploy virulence factors such as T3SS to promote cell fusion. 30 This finding highlights the interconnection between intracellular bacterial survival and MNGC formation, suggesting that the phenotype could represent an indirect consequence of impaired bacterial persistence rather than a direct defect in host cell fusion pathways. These results contrast with earlier findings in RAW264.7 cells, in which deletion of QS genes enhanced MNGC formation. 49 Differences in bacterial strain, host cell type, or QS signaling molecules might explain this discrepancy, underscoring the context-dependent nature of QS regulation.
In the G. mellonella infection model, the ΔbpsR4 mutant exhibited significantly reduced virulence, as evidenced by delayed larval mortality and improved overall survival compared with the findings in WT strain-infected larvae. The absence of mortality in PBS-injected controls confirmed the specificity of the observed pathogenic effects. BpsR4 has not been previously characterized; however, our findings suggest it functions as a key virulence regulator. This role is further supported by earlier findings implicating BpsR1, another member of the BpsR family, in QS-mediated virulence in both BALB/c mouse and C. elegans infection models.16,22 Although the Galleria model does not reflect the full complexity of mammalian immunity, it has been widely validated as a surrogate infection model in B. pseudomallei pathogenesis studies, with attenuation in larvae frequently correlating with reduced virulence in murine models of melioidosis.50,51 Given that related QS regulators, such as BpsR1, are required for virulence in mammalian systems, it is reasonable to hypothesize that BpsR4 also contributes to mammalian pathogenesis. 22 Together, these results expand our understanding of BpsR family regulators and highlight BpsR4 as a previously unrecognized contributor to pathogenicity.
Although our findings provide new insights into the roles of BpsR4 in B. pseudomallei physiology and pathogenesis, several limitations should be acknowledged. First, the transcriptomic analyses were performed under in vitro conditions, which might not fully capture the regulatory dynamics of BpsR4 during infection in vivo. Second, although the NHDF model and G. mellonella infection assays provided important functional insights, these models cannot fully recapitulate the complexity of mammalian systems. Additional studies in murine or other mammalian models will be required to validate the relevance of BpsR4 in host–pathogen interactions. Finally, our study was restricted to transcriptional profiling, and complementary proteomic or metabolomic approaches are needed to confirm downstream functional outcomes. Taken collectively, these limitations highlight the need for future work to corroborate and expand our observations.
Conclusions
BpsR4 functions as a dual regulator within the QS network of B. pseudomallei, governing critical pathways involved in stress adaptation, metabolism, biofilm formation, and virulence. The regulatory flexibility of BpsR4 enables the bacterium to respond efficiently to environmental fluctuations and host-imposed stresses. Targeting BpsR4 or its downstream effectors might represent a promising strategy for therapeutic intervention against melioidosis that disrupts pathogenic behaviors without compromising bacterial viability.
Supplemental Material
sj-pdf-1-sci-10.1177_00368504251394544 - Supplemental material for Transcriptomic profile of BpsR4 and its roles in stress response, antibiotic susceptibility, biofilm formation, and pathogenesis in Burkholderia pseudomallei
Supplemental material, sj-pdf-1-sci-10.1177_00368504251394544 for Transcriptomic profile of BpsR4 and its roles in stress response, antibiotic susceptibility, biofilm formation, and pathogenesis in Burkholderia pseudomallei by Amporn Rungruengkitkun, Phan Khoi Nguyen, Witawat Tunyong, Thida Kong-Ngoen, Sumate Ampawong, Thaniya Sricharunrat, Nitaya Indrawattana, Narisara Chantratita and Pornpan Pumirat in Science Progress
Supplemental Material
sj-pdf-2-sci-10.1177_00368504251394544 - Supplemental material for Transcriptomic profile of BpsR4 and its roles in stress response, antibiotic susceptibility, biofilm formation, and pathogenesis in Burkholderia pseudomallei
Supplemental material, sj-pdf-2-sci-10.1177_00368504251394544 for Transcriptomic profile of BpsR4 and its roles in stress response, antibiotic susceptibility, biofilm formation, and pathogenesis in Burkholderia pseudomallei by Amporn Rungruengkitkun, Phan Khoi Nguyen, Witawat Tunyong, Thida Kong-Ngoen, Sumate Ampawong, Thaniya Sricharunrat, Nitaya Indrawattana, Narisara Chantratita and Pornpan Pumirat in Science Progress
Footnotes
Acknowledgements
Authors acknowledge Prof. Dr Herbert Schweizer, Northern Arizona University for providing the pEXKm5 plasmid. We thank the support from the Department of Microbiology and Immunology, Central laboratory and the Office of Research Services, Faculty of Tropical Medicine for supporting the publication of this article.
ORCID iDs
Ethical considerations
All experiments and methods were performed in accordance with relevant guidelines and regulations. This project was exempted from ethics approval by the Ethics Committee of the Faculty of Tropical Medicine, Mahidol University (Bangkok, Thailand; reference No: TMEC 23-079). The B. pseudomallei infection protocol using G. mellonella was approved by the Animal Ethics Committee of the Faculty of Tropical Medicine, Mahidol University (certificate no: FTM-ACUC 031/2023). As G. mellonella is an invertebrate species that is not covered under vertebrate animal welfare legislation, such as the Guide for the Care and Use of Laboratory Animals, all experiments were conducted in compliance with the institutional biosafety approval and the ethics of animal experimentation guidelines of the National Research Council of Thailand.
In terms of biosecurity, both animal and bacterial laboratory work was carried out under the strict security and safety regulations of Mahidol University. All procedures involving infectious agents complied with national standards for handling pathogens. Institutional Biosafety Committee approval was obtained for this project (approval number: MU 2023-044; approval date: December 18, 2023).
Authors’ contributions
AR and PP contributed to the conception and design of the study. AR, PKN, WT, TK, SA, TS, NI, NC, and PP performed the experiments. AR, PKN, and PP analyzed the data. AR, PKN, and PP wrote the manuscript. PP edited the manuscript. All authors read and approved the final version of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Mahidol University (Fundamental Fund by National Science Research and Innovation Fund [SRF]), (fiscal year 2024).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
The raw data of transcriptomic analysis have been deposited in the NCBI Sequence Read Archive (SRA) database under accession number: PRJNA1313809.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
