Abstract
Introduction:
Virulent footrot of sheep caused by Dichelobacter nodosus is associated with tremendous economic losses due to recurrent treatment costs and increased culling rates. This organism being a fastidious anaerobe is difficult to isolate on ordinary media that does not support its growth. The D. nodosus serogroup B isolate described in the present study has been used in the preparation of the whole-cell killed vaccine against footrot in India. D. nodosus serogroup B is the predominant serogroup involved in virulent footrot (lesion score 4) in India as well as in many sheep-rearing countries of the globe.
Methods:
Genomic DNA was extracted using wizard Genomic DNA purification kit. The whole genome of the D. nodosus strain B was sequenced using an Illumina HiSeq 2500 platform and annotated according to functional gene categories. Annotations were performed using in-house developed Perl scripts using Nr/Nt database, uniprot, Pfam, KEGG, Panther DB, and GO database.
Result:
The assembled genome size is 1.311,533 Mb and GC content is 44.38. A total of 1215 protein-coding genes, 44tRNA and 7 rRNA were identified. The genome shows 98.63% sequence homology with the reference genome. However, 21 new genes have been identified in this genome. The information will provide insights into the various genes and regulators necessary for D. nodosus growth and survival.
Discussion:
The genome information of this serogroup B of D. nodosus isolate involved in 85–90% cases of virulent footrot of sheep in India provides further insights for improvement of the killed vaccine (B serogroup) developed recently in India. For the development of an efficacious vaccine against virulent footrot, it is essential to know the serological diversity as well as the virulent status of the strains of the D. nodosus. This serogroup isolate is a potential vaccine candidate to mitigate ovine footrot in India as the majority of virulent footrot cases belong to serogroup B of D. nodosus.
Introduction
Ovine footrot caused by a gram negative bacterium Dichelobacter nodosus, is a highly contagious disease of feet of ruminants, especially in sheep and goats that causes lameness, significant production, and economic losses worldwide. 1 The disease process begins as an inflammation of the interdigital skin of sheep and may progress to the severe separation of the claw capsule from the underlying soft tissues. The severity of footrot is dependent on the virulence of D. nodosus the infecting strain primarily, and on other factors like farm management, 2 sheep breed, 3 environmental/ climatic conditions, 4 and the presence of co-infecting bacteria at the site of infection. In some soils this fastidious organism has been reported to survive for two weeks. 5
The chronicity and endemicity of the disease leads to substantial economic loss by way of loss of body weight, condition and wool growth, decreased lambing percentage, and reduced value at the sale of the affected sheep. 6 The D. nodosus organism harbors fimbriae which are highly immunogenic for sheep and are considered to be the major protective immunogens for the host 7 Based on fimbrial antigens D. nodosus isolates are classified into at least ten distinguishable serogroups designated from A-I and M. 8 These serogroups have been further subdivided into 21 serotypes A1, A2, B1, B2, B3, B4, B5, B6, C1, C2, D, E1, E2, F1, F2, G1, G2, H1, H2, I, and M. 9 Serogroup B has been detected in 92.46% of footrot affected sheep with serotype B5 (83.0%) a new subtype of serogroup B as the predominant serotype of D. nodosus associated with severe footrot (lesion score 4) in sheep across the state of Jammu & Kashmir (J&K), India. 10 Serogrouping of D. nodosus isolates provides valuable epidemiological data on strain diversity which is a prerequisite for effective vaccination programs. 11
The isolation of D. nodosus from a footrot lesion is an extremely cumbersome procedure as the organism is extremely fastidious and ordinary enrichment media are unable to support the growth under anaerobic conditions. Hoof agar media prepared from sterilized sheep hoof capsules supplement with arginine and serine (TAS agar) is required for primary culture, isolation, and purification which could take 3–4 weeks.
Several important virulence factors of D. nodosus like extracellular subtilisin-like serine proteases (or subtilases), type IV fimbriae, the vrl, and vap genomic islands 12 have been incriminated in its pathogenesis. D. nodosus strains exist globally in two distinct forms the virulent and the benign. 13 Virulent strains of D. nodosus produce three homologous extracellular subtilases, namely AprV5, AprV2, and BprV. The AprV2 protease is thermostable and is responsible for the elastase activity of the organism. 14 A single amino acid substitution (Tyr92Arg) makes AprV2 more virulent than its benign counterpart AprB2. The difference at the gene level, between the protease gene variants aprV2 (accession no. L38395) and AprB2, is due to a 2-bp change from TA to CG at position 661–662. 15 The other two proteases are acidic protease 5 (AprV5 and AprB5) and basic protease (BprV and BprB), respectively. Stauble 16 developed a real-time PCR assay using the protease genes AprV2 and AprB2, to distinguish the virulent and benign strains of D. nodosus. The whole genome shotgun sequencing of a large number of D. nodosus isolates from different countries has revealed that some of the isolates contain a single nucleotide polymorphism (SNP) in the region of the 3′ end of the forward primer. 13 Previously, the intA gene (integrase) has also been suggested to be associated with virulence. 17
Methodology
The sample had been previously collected from sheep foot (lesion score 4) at the apex of the cleft between the horn of the hoof and the sensitive underlying tissues after scoring the animals for footrot lesion as per the protocol of Stewart and Claxton. 18
Isolation of D. nodosus
The organism is extremely fastidious and difficult to grow in the laboratory. The swab sample of the isolate used in the present study had been previously streaked on trypticase arginine serine (TAS) agar containing 4% hoof powder under anaerobic conditions (Anaerobic jar, Oxoid, UK) with gas packs (Becton Dickinson, MD, USA) followed by incubation at 37°C. Suspected colonies, post 5 days of incubation, were subcultured on 4% TAS agar again until pure colonies of D. nodosus were obtained. The suspected colonies of D. nodosus were confirmed by demonstration of typical cellular morphology in Gram-stained smears as well as by species-specific 16S rRNA gene by PCR. The culture was lyophilized for future use.
Extraction of bacterial DNA and 16s rRNA PCR for D. nodosus detection
The pure colony was suspended into150 µl of sterile distilled water. Snap chill method of DNA extraction was followed by boiling the suspension for 10 min, cooling on ice for 5 min, and centrifugation at 10,000 × g for 10 min. 2 µl of the supernatant was used as the template for PCR reaction. The extracted DNA was subjected to the amplification of the 16S rRNA gene for the detection of D. nodosus. PCR conditions and primer details have been previously described. 10
Serogrouping of D. nodosus
For serogrouping of the D. nodosus isolates, the multiplex PCR was carried out by using nine (A–I) serogroup-specific primers as described by Dhungyel et al. 19 was followed. Serogroup specific DNA controls were previously provided by Dr O.P. Dhungyel. The PCR products were analyzed in 1–2% agarose gels, stained with ethidium bromide, visualized under ultraviolet illumination, and photographed by Gel Documentation System (UVP, UK).
Virulence characterization
Gelatin gel test 20 to detect the thermostable proteases, as well as PCR for detection of intA gene (integrase) 17 in virulent strains, was carried out with slight modification as described previously. 10
Genomic DNA isolation and sequencing
The pure lyophilized culture was revived on TAS agar for extraction of DNA. DNA was extracted using the Wizard Genomic DNA purification kit (Promega Corp; Madison, WI, USA). Whole genome sequence of D nodusus was carried out with paired-end 150 bp multiplex on an Illumina HiSeq 2500 platform (Illumina Inc., San Diego, CA). After the filtering step, clean reads were assembled using SPAdes, ABySS, and Velvet. The raw reads were quality checked using in-house developed Perl scripts and processed by the cutadapt program 21 for removing adapter sequences and low-quality bases trimming towards 3′- end. Reads having quality above Q30 were taken for further analysis. Raw reads were transformed to clean reads using in-house developed Perl scripts, by removing the adapters sequences, low-quality reads and those containing undermined bases. After the filtering step, clean reads were assembled using SPAdes, ABySS, and Velvet. All clean reads (Phred quality scores >30 and length >25 bp) were mapped to the D. nodosus reference genome assembly using Bowtie-v2.2.6 29. We utilized genome assembly from the Velvet program for downstream analysis and better statistical parameters than other assemblers. The final draft assembly contained seven contigs.
Genome annotation
Genes were identified using RAST (Rapid Annotation using Subsystem Technology) 22 and the overview of the annotated genome was completed with the SEED viewer. 23 This was followed by manual curations by using the NCBI-Blast program against NCBI nr/nt database. UniProt, 24 Pfam, 25 KEGG, 26 Panther DB, 27 and GO 28 database. The tRNA ScanSE tool 29 was used to find tRNA genes.
Results & discussion
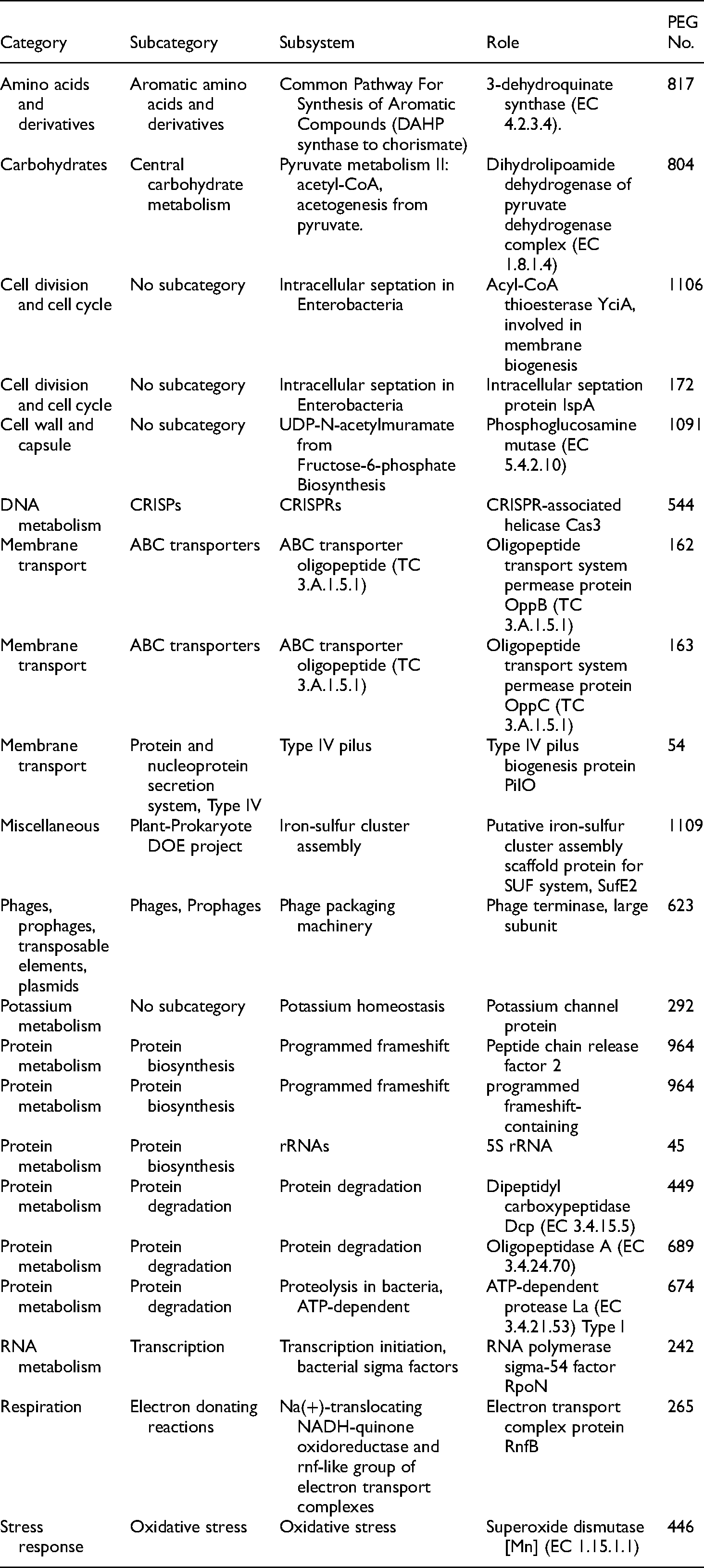
We report here the whole genome sequence of D nodosus JKS-07B serogroup B isolate. This strain was isolated from virulent footrot of sheep and produced thermostable protease in gelatin gel. The G + C content of the genome was 44.38%. Protein-coding gene prediction was performed using the Glimmer program. A total of 1215 protein-coding genes were identified and annotated using homology-based search through the National Center for Biotechnology Information (NCBI) Ref Seq non-redundant database using BLASTX program with an E-value cutoff of 10−5. A total of 44 tRNA genes, four non–coding RNAs, nine pseudogenes and seven rRNAs genes were identified (Table 1). A graphical map showing the entire genome sequence with sequence feature information is depicted in Figure 1. The functional parts of the metabolic reconstruction of this organism were compared to that of reference D. nodosus VCS1703A. We found a total of 21 genes that were unique to this isolate of D nodosus, (serogroup B) and have not been reported in the reference strain (Table 2). The genes comprise mainly of oligopeptide transport system permease protein OppB (TC 3.A.1.5.1), Oligopeptide transport system permease protein OppC (TC 3.A.1.5.1), Type IV pilus biogenesis protein PilO, intracellular septation protein IspA etc.

Draft genome of Dichelobacter nodosus serogroup B. Dark pink circle represent is the open reading frames with purple as the start and pink as the stop site, inner circles are GC-skew plot with black as GC content (magenta portion-GC-skew negative, olive green portion-GC-skew positive).
Genome characteristics of Dichelobacter nodosus JKS-07 serogroup B.

Genes associated with a subsystem in Dichelobacter nodusus JKS-07B serogroup B isolate from J&K compared with reference genes.
Nucleotide sequence accession No: This whole genome project has been deposited in Genbank as
The first D. nodosus genome to be sequenced is strain VCS1703A (GenBank Accession No: NC_009446.1) and has been used in several virulence studies. The genome of D. nodosus is 1,311,533 bp, of which one-fifth is believed to have been acquired by lateral gene transfer –an incorporated Mu-like bacteriophage. 30 The D. nodosus PubMLST database contains 171 isolates with 115 STs (release date 11 January 2021). The database suggests a high level of diversity with a low level of recombination which is reflected in the grouping of isolates and branch lengths shown in the cgMLST and wgMLST analysis. 10
Oligopeptide transport system permease protein OppB and OppC are involved in the binding-protein-dependent transport system for the translocation of the substrate across the bacterial membrane. Type IV pilus biogenesis protein PilO has a role in bacterial motility, and its DNA uptake from the surroundings while intracellular septation protein IspA has been shown to have a part in cell division and intracellular septation. This may account for the severity of lesion score (3 or 4) in footrot cases from sheep farms across J&K. D. nodosus serogroup B has 1215 coding sequences divided into 190 subsystems for connecting the genes in the new genome to functional roles (Figure 2). Among the various subsystems, protein metabolism (164), cofactors, vitamins, prosthetic groups, pigments (70), DNA metabolism (45), stress response (25) virulence, disease, and defense (17) genes are important.

Number of assigned functional subsystems.
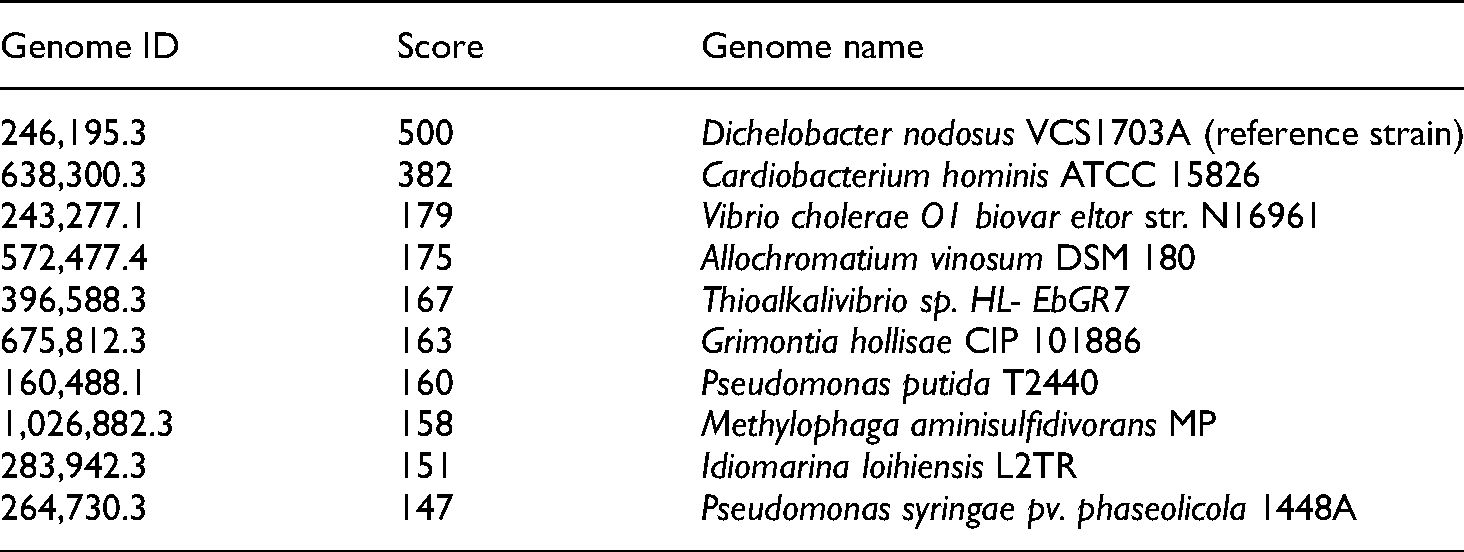
Among the closest neighbour serogroup B is highly related to D. nodosus VCS1703A with a score of 500 followed by Cardiobacterium hominis ATCC 15826 with a score of 382 (Table 3). A set of universal proteins plus up to 200 ‘unduplicated’ proteins were used to compare abinitio GLIMMER3 gene candidates to generate the closest phylogenetic neighbors and score them. These gene candidates were also used to help ‘bootstrap’ iterative retraining of GILMMER 3 and are not retained in the final annotation.
Closed neighbor identity of Dichelobacter nodosus JKS-07B serogroup B from isolate from J&K.
A total of 2140706 QC passed reads were mapped with the D. nodosus reference genome assembly VCS1703A (GenBank accession No: NC_009446.1) using BWA-MEM version 0.7.17.1. The genome shows 98.63% sequence homology with the reference genome. We found a total of SNP's at 2933 positions using VarScan 2 31 and at 92 positions an InDel compared to the reference genome. The pathogenicity of the genome was determined using Pathogen Finder version 1.1. 32 The data identifies 40 genes that correlate with the pathogenicity of this bacteria compared with the reference genome.
This study presents the genomic information of a D. nodosus JKS-07B serogroup B isolate from India. The genome information of this strain provides insights for further work on a universal vaccine in Asia as this serogroup has been isolated from >90% cases of virulent footrot in India. The whole genome information pertaining to circulating strains is important in knowing the relationship between virulent and benign strains as well as the relationship of isolates from different geographical regions. The infection due to virulent strains is often difficult to treat and occasionally results in therapeutic failures and complicated outcomes of the disease. The sequenced strain belongs to predominant serogroup B in India which has already been used in the preparation of a killed vaccine with promising results in Jammu & Kashmir (J&K), India.
Footnotes
Acknowledgments
The authors acknowledge the infrastructural facilities and funds utilized under the Indian Council of Agricultural Sciences, New Delhi under the Niche Area of Excellence in Anaerobic bacteriology.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
