Abstract
Recent advances in sports genomics and epigenetics have significantly enhanced our understanding of the biological underpinnings of sprint, strength, and power performance. While genetic predisposition influences muscle fiber composition, contractile speed, and neuromuscular efficiency, epigenetic mechanisms are increasingly recognized for their role in modulating gene expression in response to training and environmental stimuli. In this narrative review, we synthesize current findings on key genetic variants, including polymorphisms in ACTN3 (R577X), ACE, and AMPD1, which have been associated with muscle hypertrophy, anaerobic capacity, and explosive strength. Genome-wide association studies (GWAS) and candidate gene approaches have identified over 150 polymorphisms relevant to power-oriented sports, with ACTN3 R577X emerging as a strong predictor of sprinting ability. However, interindividual variability in performance cannot be explained by genetics alone. Epigenetic modifications—such as DNA methylation, histone acetylation, and microRNA (miRNA) activity—regulate skeletal muscle plasticity, recovery, and adaptation. Notably, miRNAs act as key post-transcriptional regulators, influencing pathways involved in oxidative metabolism and neuromuscular remodeling. Furthermore, telomere biology has emerged as a complementary factor, with telomere length serving as a biomarker of biological aging and regenerative capacity, particularly relevant to high-intensity, power-based sports. By integrating genomics, epigenomics, and transcriptomics, this narrative review provides a comprehensive framework for understanding the complex biology of elite athletic performance and highlights future directions for personalized training and talent identification in sport science.
Introduction
Sports performance in disciplines such as sprinting, strength, and power exhibits significant inter-individual variability, even among athletes of similar age, morphology, and training history. 1 This variability can be partially attributed to genetic inheritance. Numerous studies have identified associations between genetic variants and athletic performance, particularly in elite athletes competing at the highest international levels, such as the Olympic Games. 1 Due to their genetic homogeneity and access to specialized resources (e.g. expert coaching, tailored nutrition, and advanced training facilities), these athletes serve as an optimal model for investigating the influence of genetic factors on exercise-related traits, including maximal oxygen consumption (VO₂ max), muscle strength, and peak power output.1,2
Early family and twin studies have estimated the heritability of sports performance to be approximately 66%, even after accounting for environmental influences. 3 Furthermore, findings from the HERITAGE study demonstrated that individual physiological responses to training, particularly improvements in VO₂ max, are also genetically influenced, with heritability reaching 47% in sedentary individuals. 4 These findings suggest that certain individuals possess a favorable “genetic blueprint” that predisposes them to excel in specific athletic disciplines. The heritability of muscle-related traits, such as muscle mass and strength, is particularly relevant to sprint and power performance, with genetic contributions estimated to range from 31% to 78% in family-based studies. 5 Additionally, the heritability of maximal power output and total explosive power has been reported at 74% and 84%, respectively. 6 However, research on power performance remains less developed than that on endurance, largely due to the complexity of isolating genetic effects in heterogeneous populations and the lack of consistent replication across independent studies.
While genetic factors provide the foundation for athletic potential, their influence is dynamically modulated by epigenetic mechanisms and environmental stimuli. Epigenetics refers to heritable changes in gene expression that occur without alterations to the DNA sequence itself, allowing for dynamic regulation of how genes are activated or silenced in response to external factors. In the context of athletic performance, epigenetic regulation is strongly influenced by training load, recovery protocols, nutritional status, and other lifestyle factors.7–9 Three major epigenetic mechanisms—DNA methylation, histone modifications, and microRNA (miRNA) activity, play central roles in shaping skeletal muscle adaptation, neuromuscular remodeling, and metabolic efficiency.8–10 Recent advances in exercise epigenetics have demonstrated that these mechanisms are responsive to both acute and chronic training stimuli, thereby contributing to individual variability in strength, sprint, and power performance.8–12
Beyond gene expression regulation, another emerging layer of biological complexity relevant to athletic performance lies in telomere biology. Telomeres are repetitive nucleotide sequences (TTAGGG)n located at the ends of linear chromosomes, serving as protective caps that maintain genomic integrity and regulate cellular replicative lifespan. 13 Each time a cell divides, telomeres shorten gradually, a process that can eventually lead to cellular senescence or apoptosis. As such, telomere length is widely recognized as a biomarker of biological aging. In skeletal muscle and other performance-related tissues, telomere dynamics are increasingly recognized as being linked to regenerative capacity, recovery efficiency, and tissue resilience, all essential physiological traits in elite athletes involved in high-intensity, short-duration sports such as sprinting, powerlifting, and strength-based disciplines. 13
Consequently, a deeper understanding of the interplay between genetic predisposition, epigenetic modulation, and telomere maintenance is essential to fully elucidate the molecular underpinnings of individual variability in elite athletic performance.
The present work is a narrative review that aims to synthesize the current knowledge on the genetic and epigenetic determinants of sprint, strength, and power performance. Focusing on studies involving elite athletes, it explores key genetic polymorphisms, examines the role of epigenetic modifications, and discusses the emerging evidence on telomere biology. Finally, it proposes future research directions to advance the integrated understanding of how genetic and environmental factors converge to shape athletic excellence.
To provide a conceptual framework for this review, we propose an integrative model summarizing the main molecular contributors to sprint, strength, and power performance. This model outlines the roles of genetic predisposition, epigenetic regulation, transcriptomic responses, and telomere dynamics in shaping individual variability in performance traits (Figure 1).

Integrated molecular contributors to sprint, strength, and power performance.
The model illustrates the interplay between genetic variants (e.g. ACTN3, PPARA), epigenetic mechanisms (DNA methylation, histone modifications, miRNAs), transcriptomic modulation by training, and telomere biology in supporting muscle adaptation, recovery, and long-term performance capacity.
Genetic basis of sprint, power, and strength performance
Sprint, power, and strength performance are complex phenotypes shaped by an array of genetic factors that influence skeletal muscle contractility, fiber type composition, neuromuscular coordination, and anaerobic energy pathways. These characteristics are fundamental to success in high-intensity, short-duration sports such as sprinting, jumping, throwing, and weightlifting.14,15 A key biological determinant of such performance is the relative abundance and functional efficiency of type II (fast-twitch) muscle fibers. These fibers are specialized for rapid, forceful contractions and are highly dependent on anaerobic metabolism. Their prevalence and performance capacity are, to a large extent, genetically determined. Athletes engaged in sprint and power disciplines consistently show a predominance of fast-twitch muscle fibers and reliance on anaerobic energy pathways, which contributes to superior speed, explosiveness, and strength. 14
While the genetic determinants of endurance performance have been more extensively studied, recent advances in sports genomics have enabled the identification of several variants associated with explosive athletic capabilities. Research based on both candidate gene approaches and genome-wide association studies (GWAS) has highlighted the polygenic nature of these traits. Variants in genes such as ACTN3, ACE, IGF1, CKM, PPARA, and BDNF have been associated with differences in muscle power, hypertrophy potential, and anaerobic metabolism. These associations often reflect functional impacts on fast-twitch muscle fiber development, ATP turnover rate, and neuromuscular efficiency.16–22
Power- and strength-oriented athletes tend to exhibit distinct genotypic profiles that favor rapid force production and high intramuscular energy availability.18,22 Heritability estimates for muscle strength range from 30% to 85%, while those for muscle mass have been reported to range from 50% to 95%, underscoring the substantial genetic contribution to variability in these performance-related traits.19–21
In addition to static genotypic associations, several studies have incorporated phenotypic performance tests and direct physiological assessments to better define genotype–phenotype relationships. Tests such as the vertical jump, countermovement jump (CMJ), sprint time over 10–30 m, and one-repetition maximum (1RM) strength assessments have been commonly used to evaluate the functional relevance of genetic variants in elite athlete cohorts.23,24 Moreover, muscle biopsies have provided direct evidence linking specific genotypes to muscle fiber type composition, metabolic enzyme activity, and contractile properties, offering critical insights into the molecular basis of performance differences.25,26
Although many of these associations remain to be fully validated across different populations and sport disciplines, the accumulation of evidence supports the existence of a genetically favorable profile for power and sprint performance. The functional relevance of these polymorphisms is further explored in the sections that follow, which detail their roles in cardiovascular function, muscle structure, energy metabolism, and adaptive plasticity.
Cardiovascular and oxygen transport mechanisms
The proper functioning of the cardiovascular system is critical for power and strength athletes, as it determines their ability to achieve peak performance during high-intensity efforts. 27 Efficient cardiovascular function ensures optimal oxygen and nutrient delivery to muscles, enabling sustained strength and power output during short, explosive activities. 28 Moreover, good cardiovascular health facilitates rapid recovery, allowing athletes to train more frequently and intensively. 27 Power and sprint athletes often exhibit genetic predispositions that enhance their cardiovascular function, contributing to their ability to generate force quickly and sustain high-intensity performance over short durations.27,29,30 Table 1 summarizes the genetic variants linked to cardiovascular and vascular health in power and sprint athletes.
Key genetic variants associated with cardiovascular and vascular health in power and strength athletes.

NA: not applicable.
Renin–angiotensin system and blood pressure regulation
Several genes play a key role in blood pressure regulation, including the ACE gene. The D allele of the rs1799752 polymorphism in this gene has been associated with greater muscle power, increased muscle volume, and a higher percentage of fast-twitch muscle fibers.27,29,30 Some studies have confirmed that the prevalence of the D allele is significantly higher in power and sprint athletes,27,29,30,44 while other studies have not found this association.24,45 Furthermore, this allele has been linked to strength status in athletes competing at international and national levels, as well as in weightlifters.27,31,32 Additionally, the C allele of the rs699 polymorphism in the AGT gene has been associated with elevated levels of angiotensin-II, conferring a performance advantage in sprinting by promoting muscle hypertrophy. 28 Association studies have reported an over-representation of the CC genotype in elite power athletes, facilitating superior sprint performance. 33 Similar findings have been observed in Polish athletes, 28 as well as in academy footballers, where the CC genotype correlated with faster sprint times over 5- and 20-m distances. 34 Furthermore, this genotype has been significantly associated with strength status in weightlifters and has been shown to enhance explosive strength following training.35,36
Research by Mustafina et al. (2014) found that the A allele of the rs11091046 polymorphism in the AGTR2 gene was more prevalent among female power athletes than in non-athletes. 37 Guilherme et al. (2018) confirmed these findings in Brazilian power athletes, particularly those competing at the international level. Their study reported allele frequencies of 11.2% in the control group versus 17.4% in international male athletes, while the A/A genotype was 11.3% more common among international female athletes compared to non-athletes. 38 Additionally, the -786T/C (rs2070744) polymorphism in the NOS3 gene has been linked to athletic performance, with the TT genotype correlating with significant improvements in sprinting and jumping ability, key attributes in strength sports.39,40,46 For instance, young basketball players carrying the TT genotype demonstrated superior sprinting and jumping performance. 39 Furthermore, this genotype was more prevalent among elite sprinters and jumpers compared to endurance athletes and non-athletes.40,41
Angiogenesis and tissue oxygenation
The HIF1A gene is influenced by the Pro582Ser polymorphism (rs11549465), where the T allele enhances gene stability and activity, leading to improved responses to hypoxia and enhanced carbohydrate metabolism.43,47 In the context of sports performance, the Ser allele has been significantly associated with power-oriented athletes, including Russian weightlifters and wrestlers,43,47 as well as power specialists among Ukrainian athletes. 41 These findings were further validated in a cohort of Polish athletes. 42 However, studies on Israeli sprinters and Iranian athletes did not replicate the same associations.48,49 Additionally, a case–control study conducted on Russian professional strength athletes revealed that the A allele (rs890022) of the ANGPT2 gene was significantly overrepresented in strength athletes (15%), including powerlifters, compared to controls (6.3%) (P = 0.03). 22
Muscle fiber composition, function, and structural development
In power and strength sports, performance depends on muscle fiber composition and function. 50 These fibers support efficient ATP mobilization, muscle hypertrophy, and sustained force production under high loads. 50 Their structural and functional adaptations are influenced by specific genetic factors, summarized in Table 2.
Key genetic variants linked to muscle fiber composition, function, and structural development in power/strength athletes.

NA: not applicable.
Muscle fiber composition and function
The ACTN3 gene, particularly its R577X polymorphism, is one of the most extensively studied genetic factors associated with muscle power and sprint performance in athletes. This association is strongly supported by mechanistic evidence from mouse models lacking this gene. 14 The RR genotype enables the normal production of alpha-actinin-3, a protein essential for fast and powerful muscle contractions. 75 Athletes with this genotype benefit from greater explosive strength and speed. In contrast, individuals with the XX genotype, characterized by the absence of this protein, may be at a disadvantage in these types of performance. 75 Most studies investigating the association between this polymorphism and power performance in athletes have confirmed a significant relationship, showing notable differences compared to the non-athletic population and endurance athletes.23,27,30,51–53 However, some studies have not found this association.24,76–78 Furthermore, the RR genotype has been reported at significantly higher frequencies among strength athletes, such as bodybuilders and national-level weightlifters, compared to control groups in multiple studies.27,32,54,55
Additionally, carriers of the G allele (rs8111989) in the CKM gene exhibit greater muscle power and strength,66,67 whereas the A allele is more strongly associated with endurance capacity.66,79 Another notable polymorphism, rs17602729 in the AMPD1 gene, significantly impacts enzymatic activity. Individuals with the CC genotype exhibit high AMPD activity, optimizing energy production, whereas those with the TT genotype display reduced activity, leading to increased susceptibility to early fatigue and muscle cramps. The frequency of this polymorphism varies across populations, being more common in Europeans (11%), rare in Africans (1%), and absent in Asians. 61 A meta-analysis by Ipekoglu et al. (2023) found that the C allele was more prevalent among strength athletes, suggesting that rs17602729 could serve as a genetic marker for power performance. 60
The IGF2 gene, which plays a crucial role in regulating muscle growth and function, is characterized by the rs680 polymorphism. The G allele is associated with higher IGF2 mRNA expression than the A allele, indicating a major effect on gene transcription. 80 This allele has been overrepresented in strength athletes, including sprinters and jumpers, 72 as well as judokas 73 and football players. 34 Furthermore, among decathletes, the prevalence of the IGF2 rs680 GG genotype was significantly higher than in other athlete groups. Decathletes also exhibited a higher prevalence of the IGF1 rs35767 TT genotype, which has been linked to superior athletic performance. 70 Similarly, the TT genotype of IGF1 (rs35767) was significantly associated with strength status in weightlifters.22,71
The ACTG1 gene encodes a key protein involved in the cytoskeleton of cells, particularly in striated muscles. 56 The A allele (rs6565586) has been linked to strength status in athletes and was significantly more frequent among powerlifters than in control groups. 22 This finding was further supported by a GWAS, which identified this allele as being associated with hand grip strength. 56
Furthermore, a study conducted on 60 Turkish athletes (31 power, 29 endurance) and 20 controls investigated the role of the ARK2N gene in muscle physiology. The results indicated that its expression was significantly higher in power athletes (p = 0.042) and was associated with a greater proportion of oxidative fast-twitch muscle fibers (p = 0.017). Additionally, the T allele of the rs6507691 polymorphism, which enhances ARK2N expression (p = 3.8 × 10−¹²), was linked to larger fast-twitch muscle fibers (p = 0.015). This allele was also overrepresented among elite power athletes (38.6%; OR = 2.2, p = 0.023) and wrestlers (33.8%; OR = 1.8, p = 0.044). 63
Muscle development and resilience
The TGF-β superfamily regulates muscle growth, with myostatin acting as a key inhibitor. 81 The rs2854464 polymorphism in ACVR1B may alter miR-24 binding, potentially enhancing muscle signaling and improving sprint and power performance.58,82,83 Studies indicate that this genetic variant influences strength and muscle mass, with the A allele being more prevalent among Caucasian athletes specializing in sprinting and power events, suggesting a possible ethnic influence.57,58 Similarly, the rs10501089 polymorphism in the BDNF gene, particularly the A allele, has been associated with higher circulating BDNF levels and a greater proportion of fast-twitch muscle fibers. Carriers of this allele exhibit superior grip strength and enhanced performance in anaerobic tests.64,65 The BDNF gene's plasticity also contributes to muscular adaptation and hypertrophic potential in individuals carrying this polymorphism. 84
Additionally, a recent study found that the C allele of the AGRN rs4074992 variant is significantly more frequent in Turkish sprint and power athletes compared to endurance athletes. This allele has been linked to an increased cross-sectional area of fast-twitch muscle fibers in Russian athletes, a key determinant of muscle power and strength. 59 Moreover, the G allele of the rs3213537 polymorphism in the CPNE5 gene has been associated with sprint performance in British footballers (5–20 m sprint), young Polish women (10 m sprint), and the overrepresentation of Russian sprinters compared to endurance athletes. 34
The UBR5 gene, encoding an E3 ubiquitin ligase, is essential for muscle hypertrophy and recovery. The A alleles of the rs10505025 and rs4734621 polymorphisms have been linked to larger fast-twitch muscle fibers, favoring strength and power in athletic populations. 74 The rs41274853 polymorphism in CNTFR (particularly the T allele) has been linked to strength-based activities like weightlifting and sprinting.68,69 Additionally, miR-675-5p has been proposed to regulate CNTFR expression by targeting the 3′-UTR region of the gene, which contains this polymorphism. 69 This interaction may influence CNTFR gene expression, potentially explaining the superior strength and speed performance observed in individuals homozygous for the T allele (TT genotype) in strength-based disciplines. 69 The TT genotype has also been associated with weightlifting performance in strength athletes.68,69
Metabolic regulation and energy availability
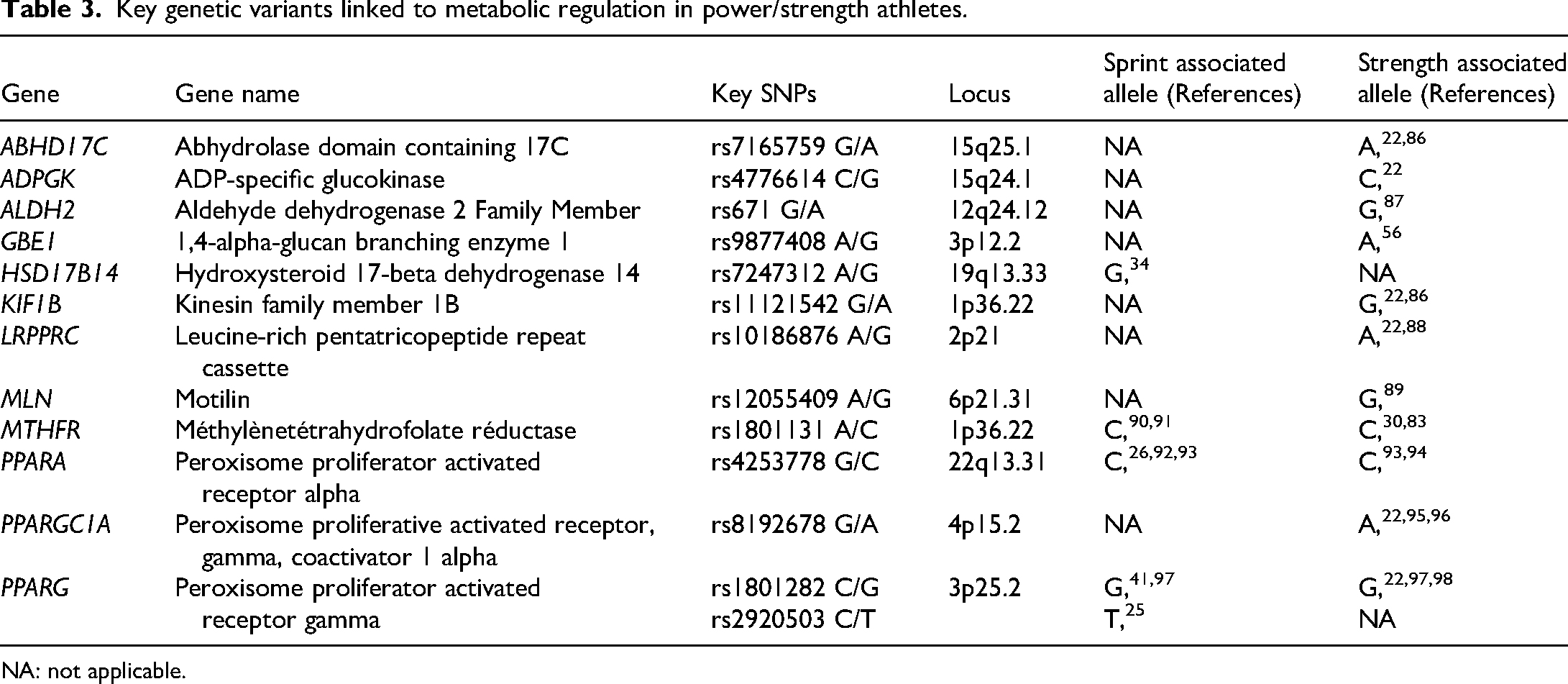
Metabolic pathways and hypoxia response are crucial for power athletes, ensuring efficient energy production under high-intensity conditions. 85 Rapid ATP generation, muscle glycogen mobilization, and anaerobic glycolysis provide immediate energy, while hypoxia-inducible factors regulate oxygen homeostasis and metabolic adaptations.42,43,47 Genetic variants influencing hypoxia signaling, muscle fiber composition, and energy metabolism impact power performance, endurance, and recovery.25,33 Table 3 summarizes key polymorphisms linked to metabolic regulation in power and sprint athletes.
Key genetic variants linked to metabolic regulation in power/strength athletes.
NA: not applicable.
Energy metabolism
Among the genes involved in energy metabolism, the PPARA gene is characterized by the rs4253778 G/C polymorphism, where the C allele has been associated with increased speed and power in multiple studies. 26 A study on Russian athletes reported a higher frequency of this allele in individuals excelling in short-term anaerobic events (p = 0.029). 26 Similarly, Ginevičienė et al. (2010) found a greater prevalence of the C allele in Lithuanian athletes compared to the general population. 92 Subsequent research demonstrated that individuals with the CC genotype exhibit superior performance in explosive muscular efforts, favoring muscular hypertrophy and anaerobic capacity, particularly in sprinters and weightlifters. 93 In a study by Maciejewska et al., the CC genotype was significantly more frequent in power/strength athletes, including elite Polish, Italian, and Lithuanian gymnasts, compared to controls. 99 Similar findings were observed in footballers, where the CC genotype correlated with higher strength capacity, as measured by the one-repetition maximum (1RM) test, compared to other genotypic groups (GG, GC). 78 Moreover, carriers of the C allele demonstrated better strength and speed performance in the 30-s Wingate test (WT30). 94 In addition, the A allele (rs8192678) of the PPARGC1A gene has been linked to strength status in powerlifters and weightlifters.22,95,96 Other significant genetic markers include rs7165759 (ABHD17C) and rs4776614 (ADPGK), where the A and C alleles, respectively, are associated with increased strength in elite athletes.22,86 The A allele (rs2273555) of the GBF1 gene and the G allele (rs12055409) of the MLN gene were more prevalent in weightlifters than in non-athletic controls. 89 A dominant effect of the A allele of GBF1 was linked to increased grip strength (p = 2.3 × 10 ⁻ ³). 56 Moreover, carriers of the Ala allele (rs1801282 C/G) in the PPARG gene exhibit a better glycemic response to exercise 100 and improved glucose uptake in skeletal muscle. 101 This allele was also found at higher frequencies in power athletes, with 23.8% in Russian athletes versus 15.1% in controls (p < 0.0001), a result replicated in a study on Ukrainian athletes.41,97 The same association has also been found in strength athletes such as weightlifters and highly elite strength athletes.22,97,98 Additionally, the G allele (rs7247312) of the HSD17B14 gene, involved in steroid metabolism, has been linked to improved athletic performance. 34 Regarding MTHFR polymorphisms, the rs1801131 (1298A > C, E429A) variant reduces enzymatic activity, affecting DNA methylation and gene expression. 102 The CC genotype was overrepresented in sprint/power and strength athletes, potentially conferring an advantage in these sports.22,88,90,91,103 The C allele may contribute to faster acclimatization to high-intensity exercise by regulating genes involved in energy metabolism and muscular hypertrophy. 90 However, this association remains debated, as the CA genotype could neutralize both the beneficial and adverse effects of homocysteine regulation. 90
Bioenergy regulation
A GWAS on 223,315 individuals identified 101 loci associated with grip strength (p < 5 × 10⁻⁸), including the G allele (rs11121542) of the KIF1B gene. 86 This association was confirmed in another study, where the G allele was significantly more frequent in powerlifters compared to controls. 22 Similarly, the A allele (rs10186876) of the LRPPRC gene was linked to handgrip strength, with a higher prevalence among elite strength athletes, particularly weightlifters.22,88
Detoxification and metabolism of toxic compounds
A study by Saito et al. investigated the ALDH2 rs671 polymorphism in weightlifters, finding no significant differences in genotype and allele frequencies concerning overall performance. However, athletes with the GA + AA genotype had lower strength performance relative to body weight than those with the GG genotype (7.1 ± 1.2 vs. 7.8 ± 1.0; p = 0.010, partial η² = 0.08), suggesting that this polymorphism may influence strength performance in weightlifters. 87
Structural integrity and glycosylation in power athletes
Glycosylation plays a vital role in power athletes by enhancing cellular stability, communication, and mechanical strength under intense physical stress.21,104 This process, which involves adding sugars to proteins and lipids, improves cell signaling, protein stability, and receptor function, contributing to more efficient muscle contractions and faster recovery.21,104,105
Genetic studies have identified polymorphisms associated with sprint and power performance. A GWAS found that the G allele (rs10196189) of the GALNT13 gene was significantly more prevalent in elite Jamaican, African-American, and Japanese sprinters. 21 A meta-analysis confirmed this association, 21 and further validation by McAuley et al. (2024) linked the G allele to superior sprint performance and counter-movement jumping ability in young footballers. 104
Additionally, Rankinen et al. reported that the T allele of GALNTL6 (rs558129) was associated with a 5–7% increase in power output in the Wingate test and was more frequent among Russian strength athletes. 106 However, its prevalence did not differ significantly between endurance athletes and controls. 107 Zwijewski et al. (2021) later confirmed that T allele carriers were 1.56 times more likely to excel in sprint and strength events, particularly among Polish swimmers. 105
Stress response and recovery mechanisms
The body's ability to manage stress and recover efficiently is essential for power athletes. 108 High-intensity sports, such as sprinting and weightlifting, impose acute physiological stress, demanding efficient energy production and muscle repair under extreme conditions.22,86,88 Power athletes depend on a robust stress response and rapid recovery, including efficient ATP replenishment and muscle fiber repair. 109
This section explores genetic factors contributing to stress resistance and recovery pathways. Table 4 summarizes the key genetic variants influencing stress response and recovery in power and sprint athletes.
Key genetic variants linked to stress response and recovery in power/strength athletes.

NA: not applicable.
Oxidative stress response
The T allele of the Ala16Val polymorphism (rs4880) is associated with a less efficient MnSOD enzyme, encoded by the SOD2 gene, which reduces the body's ability to manage oxidative stress. 115 Several association studies have reported that the TT genotype and T allele are rare among power and sprint athletes, suggesting that these genotypes may not be advantageous in high-power sports.113,114 Researchers have observed that carriers of the T allele exhibit increased oxidative stress and muscle damage, 115 as evidenced by elevated levels of creatine kinase following intense exercise. 116 Consequently, the T allele appears to restrict exercise capacity in activities where oxidative stress management is crucial.
Furthermore, IL-6 expression in muscle increases during intense exercise in proportion to exercise intensity and duration. 109 The -174 C/G (rs1800795) polymorphism in the IL6 gene influences its transcriptional response, with the C allele associated with lower levels of plasma-derived interleukin-6. Ruiz et al. (2010) reported a higher frequency of the GG genotype and G allele in Spanish strength athletes, correlating these variants with improved performance in strength-based disciplines. 109 This finding was corroborated by Eider et al. (2013). 117 Additionally, McAuley et al. (2024) identified a significant association between the IL6 GG genotype and sprint and jump performance in young footballers, noting a higher prevalence of the G allele among professional players training in elite sports academies.34,104,110 However, Eynon et al. (2011) found no significant differences in IL6 allele frequencies between endurance and power athletes in an Israeli cohort compared to non-athletic controls, suggesting potential interpopulation variability. 118 Nevertheless, in weightlifters and high-level strength athletes, the G allele was significantly more frequent than in control populations. 22
Cellular repair mechanisms
A recent study published in 2024 found that the D allele of the rs35493922 polymorphism in the MDM4 gene was overrepresented in strength athletes, including sprinters, compared to controls and endurance athletes. 111 This allele, along with the DD genotype, was associated with increased MDM4 gene expression, greater lean mass, taller stature, and larger fast-twitch muscle fibers, all of which are essential for sprinting and jumping performance. 111 MDM4 functions as an inhibitor of the p53 protein, which plays a role in promoting muscle hypertrophy and limiting muscle atrophy, particularly in fast-twitch muscle fibers that are crucial for power-based sports. 111 Moreover, a study by Nakamichi et al. demonstrated that the gain-of-function variant (rs572934641) in the PIEZO1 gene enhances instantaneous power in mice, leading to improved jumping and running speed through modifications in tendon structure. Additionally, a high frequency of the E756del mutation was observed in Jamaican sprinters, suggesting that these PIEZO1 variants may be key genetic factors contributing to sprint performance. 112 Other genetic variants linked to strength-related attributes include the C allele (rs303760) of the RMC1 gene, the A allele (rs12599952) of the DHODH gene 86 and the T allele (rs9320823) of the MMS22L gene, all of which have been associated with enhanced athletic strength.22,88
Hormonal and cellular signaling in strength and power athletes
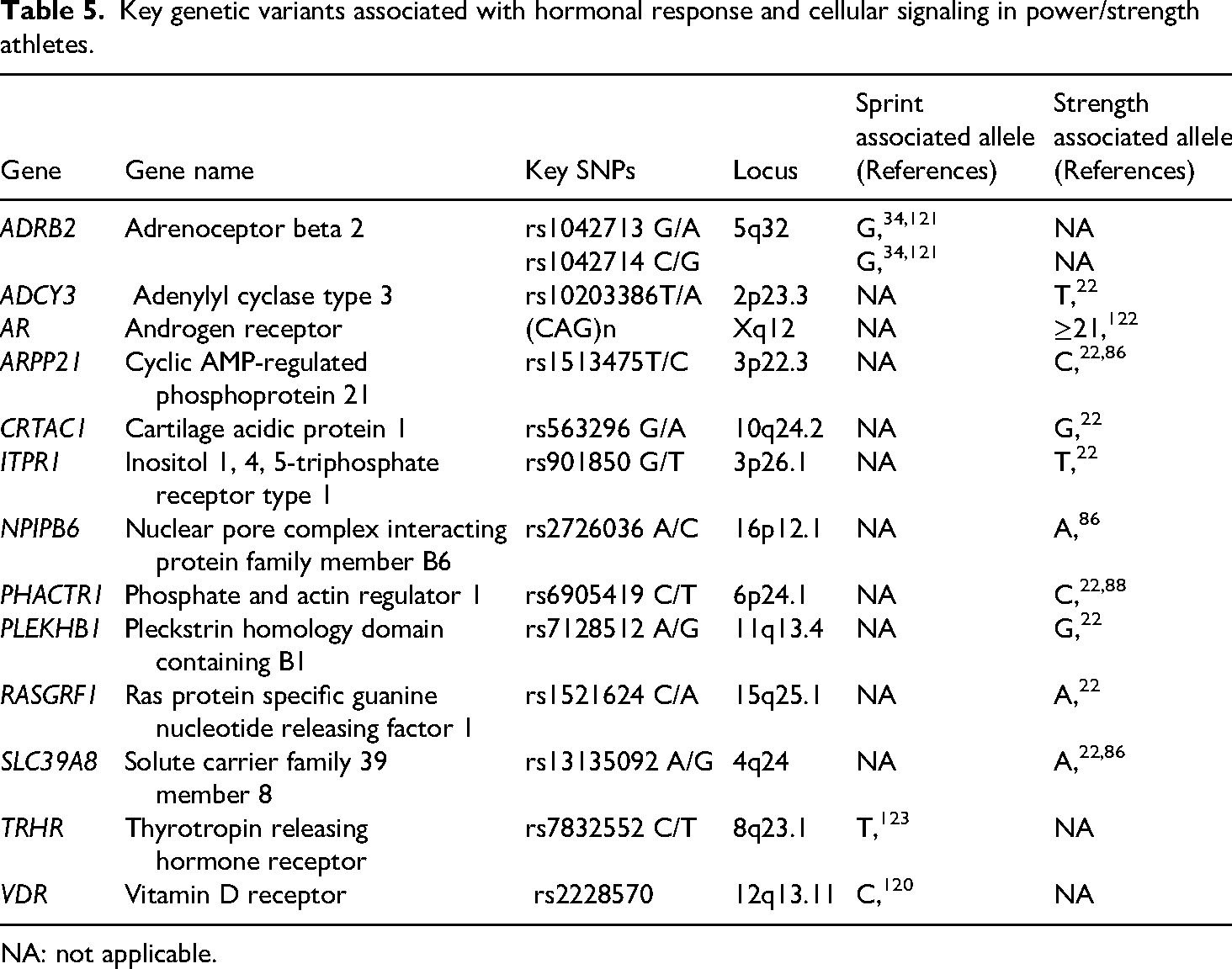
Hormonal regulation plays a crucial role in the performance of power and strength athletes, influencing muscle development, strength, and post-exercise recovery.119,120 Specific genetic variants modulate these hormonal responses and affect intracellular signal transduction, thereby impacting performance and adaptation to high-intensity training. Table 5 summarizes the key genetic variants associated with hormonal response and cellular signaling in power and sprint athletes.
Key genetic variants associated with hormonal response and cellular signaling in power/strength athletes.
NA: not applicable.
Receptors and hormone regulation
Two polymorphisms (Gly16Arg and Glu27Gln) in the ADRB2 gene have been associated with power-oriented athletic status. Sawczuk et al. reported a correlation between these variants and a predisposition for success in strength sports among Polish athletes. 121 These findings were later confirmed by Pickering et al., who observed an overrepresentation of the G allele in both polymorphisms among soccer players compared to British and European controls. 34 Additionally, thyroxine, a hormone essential for muscle development and the prevention of tissue decline with aging, is secreted through activation of the hypothalamic–pituitary–thyroid axis by the TRHR gene. 119 Low thyroxine levels are associated with muscle weakness. 119 A genome-wide association study identified the TT genotype of the TRHR rs7832552 polymorphism as being associated with greater lean body mass in Caucasian American subjects, 124 and potentially linked to sprint or power performance. 123 Similarly, the VDR gene, which regulates calcium metabolism and muscle development, shows that the CC genotype of the rs2228570 polymorphism is more frequent in elite wrestlers. This genotype is believed to provide an advantage in explosive power and neuromuscular function. 120
Guilherme et al. also investigated the influence of AR gene polymorphism, characterized by the number of CAG repeats, in athletes such as bodybuilders, sprinters, and elite weightlifters. Their results indicated that bodybuilders (both male and female) carrying long alleles (≥21 CAG repeats) exhibited higher body mass indices, greater absolute muscle mass, larger arm and thigh circumferences, and significantly greater upper and lower limb strength (p < 0.05) compared to individuals with short alleles (<21 CAG repeats). Moreover, a higher prevalence of long alleles was observed in power athletes compared to controls (p = 0.0076). 122 For the NPIPB6 and ADCY3 genes, the frequency of the A allele (rs2726036) and T allele (rs10203386), respectively, was significantly higher among elite weightlifters and strength athletes compared to controls.22,86
Transduction of intracellular signals
Several polymorphisms have been identified as overrepresented in strength athletes, particularly elite weightlifters, compared to non-athletic controls. These include the T (rs901850) allele of the ITPR1 gene, 22 the A (rs1521624) allele of the RASGRF1 gene, 22 the A (rs13135092) allele of the SLC39A8 gene,22,86 the G (rs563296) allele of the CRTAC1 gene, 22 the C (rs1513475) allele of the ARPP21 gene22,86 the C (rs6905419) allele of the PHACTR1 gene22,88 and the G (rs7128512) allele of the PLEKHB1 gene. 22 All were found at significantly higher frequencies in strength athletes than in the general population. 22
Regulation of gene expression and transcription factors
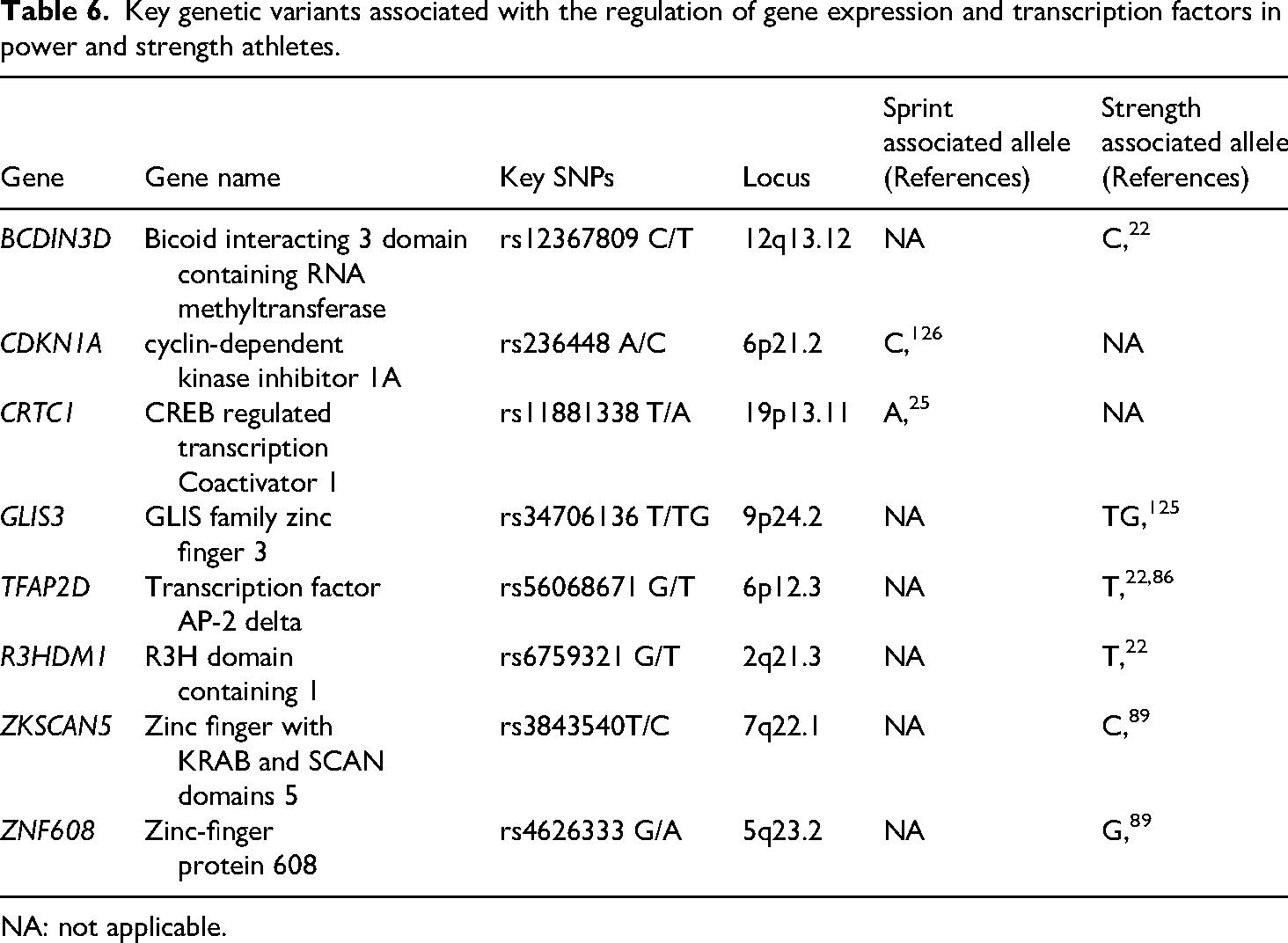
For strength athletes, the regulation of gene expression and transcription factors is essential for muscle growth, exercise adaptation, and post-exercise recovery.86,89 Certain genetic variants modulate the expression of transcription factors, activating pathways that govern muscle hypertrophy, strength, and training resilience (Table 6).65,86,89 These genetic factors influence not only protein synthesis and muscle fiber composition but also metabolic efficiency, contributing, alongside technical and psychological factors, to success in strength and power sports.65,86,89 Notably, the C allele (rs12367809) of the BCDIN3D gene, the T allele (rs6759321) of the R3HDM1 gene, and the C allele (rs3843540) of the ZKSCAN5 gene were significantly overrepresented in weightlifters, powerlifters, and other strength athletes compared to controls. 22 The TG genotype of the rs34706136 polymorphism in the GLIS3 gene was also more frequent among weightlifters. 125 Furthermore, the T allele (rs56068671) of the TFAP2D gene22,86 and the G allele (rs4626333) of the ZNF608 gene 89 were significantly more prevalent in weightlifters compared to non-athletes.
Key genetic variants associated with the regulation of gene expression and transcription factors in power and strength athletes.
NA: not applicable.
Genomic predictors of rapid walking performance and elite sprinter status
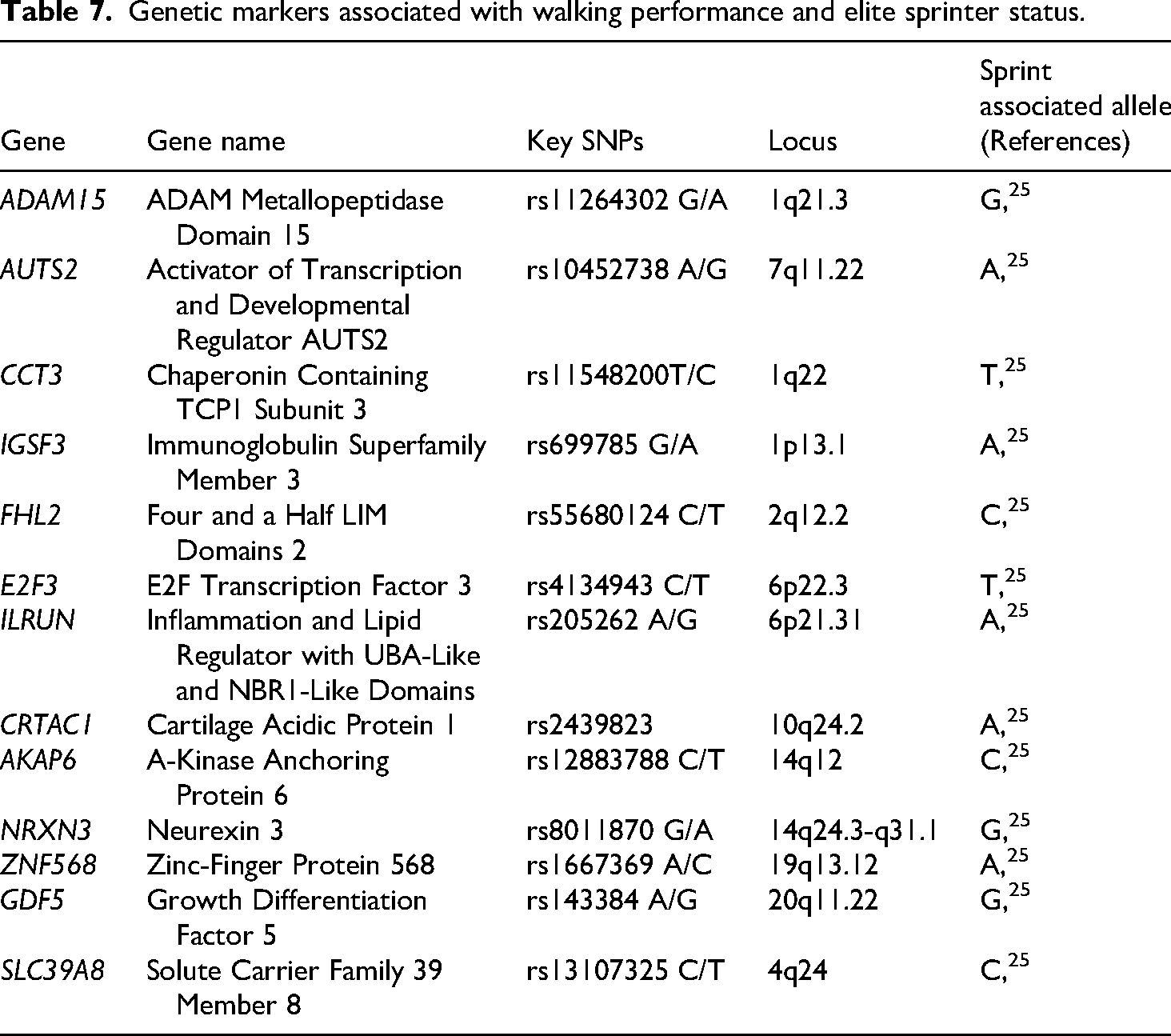
Guilherme et al. explored genetic associations with self-reported walking pace and elite sprinter status in a cohort of 137 elite sprinters and ethnically matched controls. They examined 70 candidate SNPs to identify genetic variants linked to traits favorable for sprinting, such as a higher proportion of fast-twitch muscle fibers, increased height, and lower BMI. The goal was to pinpoint polymorphisms optimizing speed and power while enhancing understanding of the genetic factors influencing sprinting performance. 25 They identified 20 SNPs more prevalent in elite sprinters than in controls, 15 of which were associated with both rapid walking and elite sprinter status. Among these, rs699785 in the IGSF3 gene was linked to a higher proportion of fast-twitch fibers, increased height, and reduced BMI, traits advantageous for sprint performance. Other associated SNPs are presented in Table 7. These genes are associated with favorable traits for sports performance, including increased lean mass, muscle strength, height, and reduced BMI. A total genetic score (TGS) derived from these polymorphisms showed better accuracy in identifying elite sprinters. However, the study's limitations include a small sample size and a lack of replication in an independent cohort. 25
Genetic markers associated with walking performance and elite sprinter status.
Role of epigenetics in sprint, power, and strength performance
Beyond the genetic foundation of athletic traits, epigenetic mechanisms play a key role in modulating gene expression in response to external stimuli relevant to sports performance. 7 In sprint, strength, and power disciplines, epigenetic regulation contributes to muscle plasticity, neuromuscular adaptation, and recovery efficiency.7,8 This section focuses on the specific epigenetic pathways that influence short-duration, high-intensity athletic performance.
Impact of DNA methylation on athletic performance
DNA methylation, an essential epigenetic modification, involves the addition of a methyl group to cytosine residues within CpG sites, mediated by DNA methyltransferases (DNMT1, DNMT3A, DNMT3B). 8 This process regulates gene expression depending on the genomic context. Hypermethylation, particularly in promoter regions, is generally associated with gene silencing, whereas hypomethylation may lead to gene activation. 8
Exercise has been shown to influence DNA methylation patterns in skeletal muscle, modulating the expression of genes involved in muscle function, energy metabolism, and recovery.127–130 One study demonstrated that intense exercise induces global DNA hypomethylation in skeletal muscle biopsies from sedentary but healthy individuals. This epigenetic modification was associated with a dose-dependent increase in the expression of PGC-1α, PDK4, and PPARD, alongside hypomethylation of their respective promoters. 130 Similar findings were observed in mouse soleus muscle and cultured myotubes exposed to caffeine, suggesting a conserved mechanism linking muscle contraction to DNA methylation changes. However, 48 h after a three-week training program, DNA methylation levels remained unchanged despite increased mRNA expression of PGC-1α and TFAM, indicating that DNA hypomethylation is a transient event associated with mRNA synthesis. 130 These findings suggest that while DNA methylation contributes to exercise-induced gene activation, other regulatory mechanisms, such as transcription factor recruitment and chromatin remodeling, likely drive long-term adaptations. A supervised eight-week physical training program further demonstrated significant DNA methylation changes in skeletal muscles, influencing genes and pathways related to muscle function and metabolism. 129 Notably, FSTL3 and RP11-624M8.1 exhibited increased methylation, whereas CNGA1, FCGR2A, KIF21A, MEIS1, SLC26A7, and ZNF280C showed decreased methylation. 129 Pathway analysis revealed enrichment in myogenesis regulation and actin cytoskeleton pathways, highlighting the role of DNA methylation in muscle growth and repair. 129 These findings underscore the impact of physical training on the epigenetic landscape, suggesting a direct influence on strength and power performance in athletes.
In a related study, post-exercise changes in DNA methylation and mRNA expression profiles in skeletal muscle were largely similar between low-load and high-load resistance exercise bouts, despite differences in the number of repetitions performed. Both bouts resulted in significant changes in DNA methylation, particularly 3 h after exercise, and these changes were associated with the regulation of genes involved in focal adhesion, MAPK signaling, and PI3K-Akt signaling. 127 These molecular alterations suggest that the volume of exercise (repetitions) rather than load intensity may play a key role in influencing the skeletal muscle response at the molecular level.
microRNAs as regulators of muscle adaptation and training response in athletes
microRNAs are small, non-coding RNAs that regulate gene expression at the post-transcriptional level by binding to untranslated regions of target mRNAs, leading to translational repression or mRNA degradation. While their primary function is to downregulate gene expression, some miRNAs have also been reported to enhance mRNA translation in specific contexts. 8 In the field of sports science, accumulating evidence suggests that miRNAs play a pivotal role in muscle adaptation, recovery, and inflammation in response to training (Table 8).
Exercise-induced microRNA expression profiles and their functional implications.
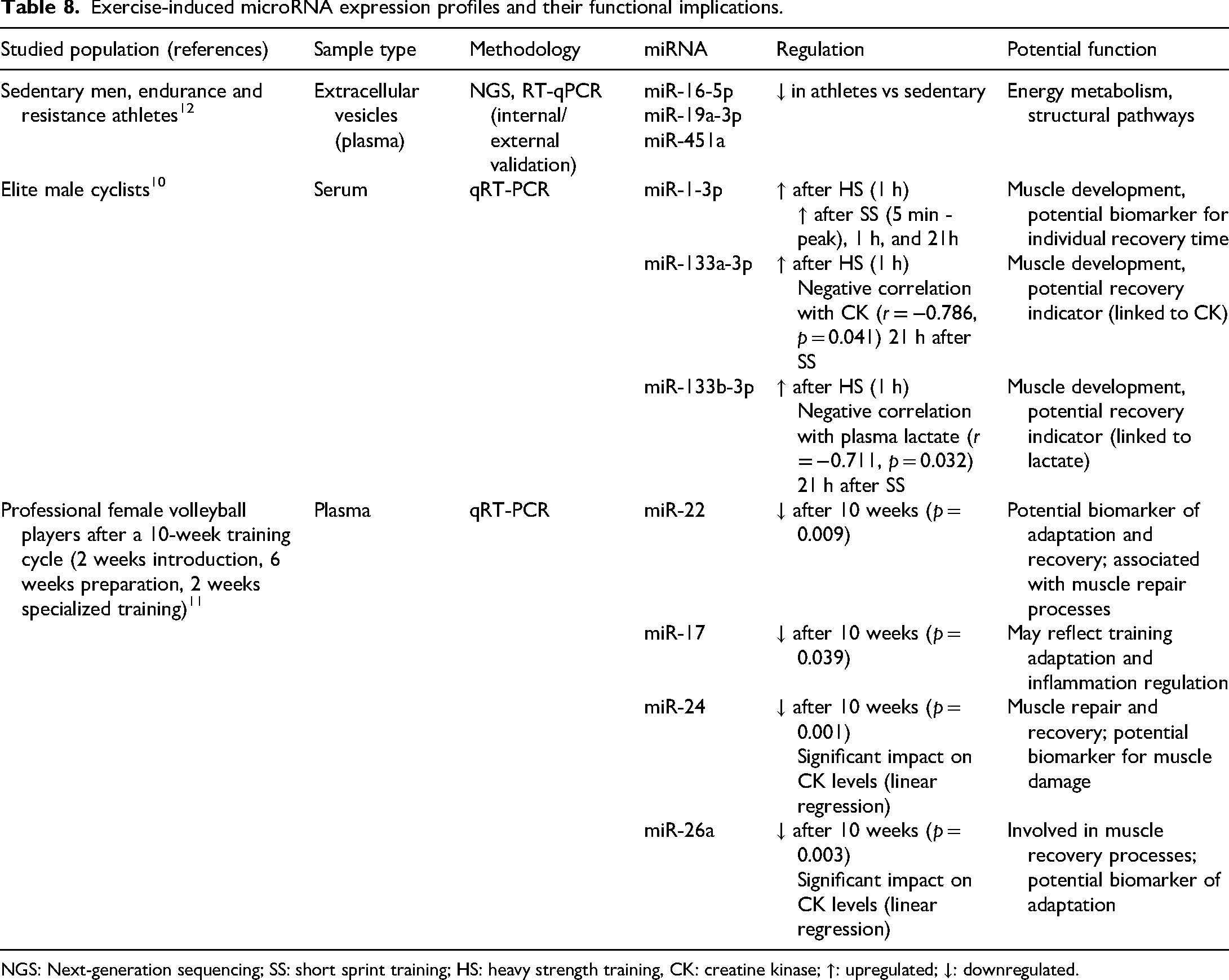
NGS: Next-generation sequencing; SS: short sprint training; HS: heavy strength training, CK: creatine kinase; ↑: upregulated; ↓: downregulated.
For instance, a study investigating the expression of miR-223, miR-320a, and miR-486 in professional female volleyball players over a 10-week training cycle demonstrated their association with IGF1R signaling, a key pathway in muscle adaptation and recovery. Specifically, miR-223 and miR-320a were significantly correlated with IGF1R expression, while miR-486 was implicated in the regulation of insulin-like growth factor 1 (IGF-1) signaling, essential for muscle hypertrophy. Among these, miR-320a exhibited the strongest association with IGF1R expression, alongside creatine kinase levels, suggesting its potential as a biomarker for training adaptation. 131 Similarly, differences in circulating miRNA profiles have been observed between endurance- and strength-trained athletes. Notably, miR-222, miR-21, miR-146a, and miR-221 were found at significantly higher levels in endurance-trained athletes compared to their strength-trained counterparts, indicating their potential role in distinguishing training modalities. 132 Furthermore, high-intensity interval training (HIIT), particularly cycling sprints, has been shown to induce significant alterations in circulating miRNA expression related to muscle function and metabolism. Following HIIT, plasma levels of miR-1, miR-133a, miR-133b, miR-122, and miR-16 were significantly reduced, while miR-206 and miR-499 remained unchanged. Interestingly, miR-133b exhibited a strong correlation with peak power output (r = 0.712, p = 0.001), and miR-122 correlated with the peak power ratio (r = 0.665, p = 0.003), further reinforcing their potential as biomarkers of anaerobic performance. 133
These findings underscore the role of miRNAs as key regulators of skeletal muscle adaptation to different training stimuli. The dynamic alterations in miRNA expression profiles in response to exercise highlight their potential as biomarkers for monitoring athletic performance, recovery, and muscle remodeling.
Histone modifications and chromatin remodeling in response to training
Epigenetic modifications play a fundamental role in orchestrating skeletal muscle adaptations to exercise. Among these, histone post-translational modifications (PTMs), including acetylation, methylation, phosphorylation, and ubiquitination, dynamically regulate chromatin accessibility and gene transcription. 8 These modifications influence the structural organization of DNA around histone proteins, thereby modulating gene expression programs essential for muscular plasticity and performance optimization. 8
Histone acetylation, primarily catalyzed by histone acetyltransferases (HATs), is strongly associated with gene activation. Exercise has been shown to enhance histone acetylation at key loci involved in mitochondrial biogenesis and oxidative metabolism, such as PGC-1α, a master regulator of endurance adaptation. 134 Recent findings indicate that a single bout of resistance exercise (RE) induces profound histone modifications in human skeletal muscle. 135 Specifically, there is a significant increase in histone H3 acetylation (+235%), a modification that enhances gene transcription by loosening chromatin structure. Additionally, activating histone methylation marks, such as H3K4me1 (+290%) and H3K27me3 (+849%), are upregulated, indicating enhanced transcriptional activity at specific gene loci. 135 Notably, after 10 weeks of RE training, a sustained increase in H3 acetylation (+40%, p < 0.05) was observed, suggesting that exercise induces long-term epigenetic memory, potentially facilitating lasting adaptations in muscle function and metabolism. 135
Conversely, histone deacetylation, primarily mediated by histone deacetylases (HDACs), serves as a repressive mechanism that can constrain skeletal muscle plasticity. Class II HDACs are highly expressed in human skeletal muscle and are functionally modulated by exercise.134,136 For instance, HDAC5 interacts with GLUT4 and MEF2, leading to the deacetylation of histones at their promoter regions and subsequent transcriptional repression. However, following exercise, HDAC5 is actively exported from the nucleus, relieving transcriptional repression and enhancing the expression of GLUT4 and MEF2, thereby promoting glucose uptake and muscle remodeling. Furthermore, HDACs regulate PGC-1α expression in response to acute exercise. Notably, after just 3 h of intense exercise in sedentary individuals, PGC-1α expression increases by up to 10.8-fold, underscoring the rapid and profound influence of exercise-induced epigenetic reprogramming. 9
Collectively, these findings illustrate that histone modifications represent a central epigenetic mechanism underpinning skeletal muscle adaptation to training. The interplay between histone acetylation, deacetylation, and methylation orchestrates transcriptional programs that optimize muscle function, recovery, and metabolic efficiency. This dynamic epigenetic landscape not only facilitates immediate exercise responses but may also contribute to long-term muscle memory, enhancing athletic performance and adaptation across diverse training modalities.
Telomere biology and its relevance to sprint, strength, and power performance
Building on the pivotal role of telomeres in genomic stability and cellular aging, recent studies have begun to highlight their specific relevance in the context of elite athletic performance. In particular, the regenerative demands of high-intensity, short-duration sports such as sprinting and strength-based events place considerable stress on musculoskeletal and neuromuscular systems, tissues in which telomere dynamics are especially important. 13
Emerging evidence suggests that regular physical activity, particularly resistance and high-intensity training, may exert telomere-protective effects. A large-scale population study involving 4814 adults reported that individuals engaging in ≥90 min of weekly strength training exhibited significantly longer leukocyte telomere length, equivalent to approximately 3.9 years of biological age deceleration compared to inactive individuals. 137 Similarly, elite athletes demonstrate more favorable telomeric profiles. In a comparative cross-sectional study, elite power athletes exhibited significantly longer leukocyte telomeres than sedentary age-matched controls (0.89 ± 0.14 vs. 0.78 ± 0.13; p = 0.013), suggesting a training-induced delay in cellular aging and enhanced regenerative potential. 138 Likewise, in a pivotal study involving elite master sprinters aged approximately 50 years, leukocyte telomere length was not only significantly greater than in age-matched non-athletes, but also strongly correlated with individual performance metrics. Specifically, telomere length positively correlated with relative sprint performance (r = 0.641, p < 0.01) and inversely correlated with age-related decline in performance (r = –0.624, p < 0.01), suggesting that telomere integrity may underpin the capacity to preserve muscle power across the lifespan. 139
Adding further support, a recent systematic review and meta-analysis evaluated whether high-level athletes exhibit longer telomeres than age-matched non-athletes and explored the underlying biological mechanisms. Across 11 studies including 240 master athletes and 209 non-athletes, pooled analyses revealed significantly longer telomeres in athletes (SMD = 0.89; 95% CI: 0.45–1.33; p < 0.001). These athletes also demonstrated lower levels of oxidative damage (SMD = 0.59) and higher antioxidant capacity (SMD = –0.46), supporting a physiological environment conducive to telomere preservation. Furthermore, enhanced telomerase activity and increased expression of shelterin complex proteins were proposed as molecular mechanisms sustaining telomere integrity in long-term trained individuals. 140
Limitations of genetic association studies and the need for integrative approaches
GWAS and candidate gene approaches have identified several genetic variants linked to athletic performance; however, these studies come with inherent limitations that must be acknowledged. Most genetic polymorphisms associated with strength, power, or sprint phenotypes have small individual effect sizes and contribute only modestly to the overall phenotype. Athletic ability is highly polygenic and influenced by complex interactions between multiple genes and environmental factors, including training, nutrition, and recovery strategies. Moreover, the majority of genetic studies in exercise physiology have been conducted in European populations, raising questions about the transferability of findings to more diverse ethnic groups. Allele frequencies, linkage disequilibrium patterns, and gene–environment interactions may differ substantially across populations, limiting the generalizability and reproducibility of results. Additionally, a significant proportion of reported associations lack replication in independent cohorts, which further undermines their reliability.
Importantly, these genetic studies often fail to account for epigenetic regulation and its dynamic responsiveness to environmental stimuli. For instance, two athletes with identical genetic profiles may exhibit divergent performance outcomes due to differences in DNA methylation, microRNA expression, or histone modifications, all of which influence gene expression patterns and muscular adaptation. These mechanisms are highly plastic and may mediate the phenotypic response to training or recovery. Beyond this, emerging research suggests that telomere length, a biomarker of cellular aging and regenerative capacity, may also be relevant to power-oriented sports, particularly in terms of muscular recovery and tissue resilience. However, the interaction between genetic background, epigenetic modifications, and telomere biology remains largely unexplored in elite athletic populations.
To overcome these limitations and move toward a more comprehensive understanding of athletic performance, future research should embrace multi-omics approaches, integrating genomics, epigenomics, transcriptomics, proteomics, and telomere biology. Such integrative strategies would allow researchers to capture the layered complexity of gene–environment interactions and biological aging processes that influence performance traits. Longitudinal studies that track molecular adaptations across time, combined with advanced modeling techniques, could provide critical insights into how genetic predisposition, molecular plasticity, and cellular longevity converge to shape elite athletic potential. This systems-level approach is essential for building predictive models, refining talent identification protocols, and developing personalized training interventions grounded in biological evidence.
Conclusion and future perspectives
The relationship between genetics and strength or power performance is a rapidly advancing field with significant implications for sports science and athlete development. This review identified 49 genetic markers associated with power performance and 42 markers linked to strength, providing valuable insights into the biological mechanisms underlying muscle function, force production, and neuromuscular efficiency. For power-oriented athletes, the most strongly associated genetic markers include the R allele of the ACTN3 gene (rs1815739), the T allele of GALNTL6 (rs558129), the C allele of PPARA (rs4253778), the C allele of AMPD1 (rs17602729), and the D allele of MDM4 (rs35493922). Regarding strength performance, key genetic factors include the AR gene with ≥21 CAG repeats, the A allele of LRPPRC (rs10186876), the T allele of MMS22L (rs9320823), the C allele of PHACTR1 (rs6905419), and the G allele of PPARG (rs1801282).
Epigenetic mechanisms such as DNA methylation, miRNAs, and histone modifications play a crucial role not only in regulating gene expression in response to training, but also in modulating the expression of inherited genetic traits. These mechanisms can enhance or silence genetic predispositions, thereby shaping an athlete's phenotypic potential regardless of identical genotypes. In this way, epigenetics serves as a key interface between the genome and the environment, allowing for the dynamic regulation of gene expression across different stages of training, development, and recovery. Future research should focus on longitudinal epigenetic profiling and multi-omics approaches to uncover how these modifications influence the expression of performance-related genes over time, particularly in elite athletes.
Footnotes
Acknowledgments
This work was supported by the Mohammed VI Foundation for Science and Health.
Author's contributions
AEH designed and developed the study, conducted the literature review, wrote the manuscript, and contributed to the analysis and interpretation of the data; AM assisted with the literature review, reviewed and revised the manuscript; SK critically reviewed and revised the manuscript for its intellectual content, reviewed and approved the final version of the manuscript; CB and YC reviewed the paper based on their expertise in the field of sports medicine ; EEF, SB, and LB supervised, revised, and validated the last draft; RE and TD supervised, validated, and contributed to writing, reviewing, and editing.
All authors have read and approved the final version of the manuscript and agree with the order of presentation.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
