Abstract
Objective
To synthesize recent molecular advances that inform diagnosis, risk-stratification, and perioperative treatment in early-stage and locally advanced non-small cell lung carcinoma (NSCLC), with emphasis on comprehensive genomic profiling, minimal residual disease (MRD) detection by circulating tumor DNA (ctDNA), and the translation of biomarkers into targeted and immunotherapy strategies.
Methods
Systematic review registered in PROSPERO (CRD420251076423). Searches of PubMed, Scopus, Web of Science, and Embase (January 2015–April 2025) followed PRISMA 2020/PRISMA-S. From 4640 records, 890 duplicates were removed; 3750 titles/abstracts were screened; 150 full texts were assessed; 75 studies met inclusion criteria. Risk of bias used Newcastle–Ottawa Scale (NOS) for observational studies and Cochrane RoB 2 tool for randomized controlled trials; certainty was summarized with GRADE where applicable.
Results
Actionable alterations (e.g. EGFR, ALK, KRAS, MET, RET, BRAF, NTRK) are prevalent in early-stage NSCLC and comparable to advanced disease, supporting routine comprehensive genomic profiling in curative-intent settings. Next-generation sequencing (NGS) and ctDNA enable the detection of MRD, earlier relapse prediction, and dynamic treatment monitoring. Perioperative strategies integrating targeted therapy and immunotherapy (e.g. adjuvant EGFR-TKI, neoadjuvant chemo-immunotherapy) improve pathological and disease-free outcomes in selected biomarker-defined populations. Evidence profiles generally show low-to-moderate risk of bias and moderate-to-high certainty for key outcomes related to profiling and MRD, with heterogeneity across platforms and endpoints.
Conclusions
Molecular advances—particularly broad NGS and ctDNA-based MRD—are reshaping the perioperative management of early and locally advanced NSCLC, enabling precision selection for targeted and immunotherapy approaches. Standardization of testing workflows and reporting, and cost-effective implementation are priorities for equitable adoption and for future trials that combine NGS, MRD, and multi-omic/AI-driven risk stratification.
Keywords
Introduction
Non-small cell lung cancer (NSCLC) accounts for approximately 85% of all lung cancer diagnoses and remains the leading cause of cancer-related mortality worldwide. 1 Although advances in targeted therapies and immune checkpoint inhibitors have significantly improved outcomes in metastatic disease, early-stage and locally advanced NSCLC continue to exhibit high relapse rates—up to 70% within five years post-resection—underscoring the need for more effective, personalized treatment strategies. 2
The advent of comprehensive genomic profiling has revolutionized the landscape of NSCLC, uncovering a diverse array of molecular subtypes driven by specific genomic alterations. Actionable mutations in genes such as EGFR, ALK, ROS1, KRAS, MET, RET, BRAF, and NTRK have enabled the implementation of precision medicine approaches across disease stages.3,4 Notably, studies have revealed that the prevalence of actionable genomic alterations in early-stage NSCLC is comparable to that in advanced disease, positioning genomic analysis as a cornerstone for risk stratification and therapeutic decision-making even in potentially curable settings.5,6
Beyond primary oncogenic drivers, genomic profiling facilitates the detection of co-mutations with significant prognostic and predictive implications. For instance, co-alterations in TP53, present in nearly half of NSCLC cases, are linked to poor survival in EGFR-mutant tumors.7,8 Similarly, mutations in KEAP1 and STK11 have been associated with resistance to immunotherapy and altered responses to immune checkpoint blockade, highlighting the complexity of tumor-immune interactions and the need for individualized therapeutic strategies. 9
Recent landmark trials have further underscored the importance of integrating molecular profiling into clinical decision-making for early-stage NSCLC. The NeoADAURA trial demonstrated that neoadjuvant osimertinib, combined with chemotherapy, significantly improved major pathological response rates compared to chemotherapy alone, setting a new benchmark for targeted neoadjuvant therapy. Similarly, KEYNOTE-671 and CheckMate-816 highlighted the clinical benefit of combining immunotherapy with chemotherapy in the perioperative setting, with substantial improvements in event-free survival and pathological complete response rates. These studies confirm that biomarker-driven treatment strategies are no longer confined to advanced disease but are reshaping the curative-intent management of early-stage NSCLC. In this context, next-generation sequencing (NGS) is not only a diagnostic tool but also a cornerstone of precision oncology, enabling individualized therapeutic strategies from the earliest stages of disease.10,11,12
In parallel, the emergence of liquid biopsy technologies—particularly circulating tumor DNA (ctDNA)—offers a non-invasive method to detect minimal residual disease (MRD), predict recurrence, and dynamically monitor treatment response. These innovations have opened new avenues for tailoring therapy intensity and duration in real time, thereby enhancing the precision and personalization of early-stage NSCLC management. 13
These advances in next-generation sequencing and liquid biopsy technologies are central to the realization of precision oncology. By enabling comprehensive genomic characterization, early identification of actionable drivers, and non-invasive monitoring of minimal residual disease (MRD), they facilitate truly individualized treatment selection and dynamic therapy adaptation. Collectively, these approaches bridge the gap between molecular insights and clinical decision-making, ensuring that therapeutic strategies are both biologically informed and patient-centered.10,13
This systematic review aims to comprehensively analyze the current state of molecular advances in early-stage and locally advanced NSCLC, examining the evidence supporting molecular profiling, targeted therapies, immunotherapy approaches, and liquid biopsy applications. By synthesizing data from recent clinical trials, molecular studies, and translational research, this review seeks to provide a comprehensive framework for understanding how precision oncology approaches are transforming treatment paradigms and improving patient outcomes in these disease stages. The ultimate goal is to inform clinical practice and guide future research directions in the rapidly evolving field of molecular oncology for early-stage and locally advanced NSCLC.
Methods
This systematic review was conducted in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA 2020) and PRISMA-S guidelines, ensuring methodological transparency and reproducibility. The review protocol was prospectively registered with PROSPERO (CRD420251076423. The aim was to evaluate recent molecular advances in early-stage and locally advanced non-small cell lung cancer (NSCLC), with a focus on genomic profiling, resistance mechanisms, and targeted therapy integration.
Study selection
A comprehensive literature search was conducted across four databases: PubMed, Scopus, Web of Science, and Embase, covering publications from January 2015 to April 2025. The search terms used included: “early-stage NSCLC,” “molecular profiling,” “genomic alterations,” “targeted therapy,” “ctDNA,” “liquid biopsy,” “resistance mechanisms,” and “precision oncology.” Boolean operators (AND, OR) were used to refine search results. Reference lists of relevant studies were manually screened for additional articles. Full search strategies for each database, including search terms and Boolean operators, are provided in Appendix A. The search yielded 1340 articles from PubMed, 970 from Scopus, 1100 from Web of Science, and 1250 from Embase. A total of 4640 articles were retrieved. After removing 890 duplicates, 3750 articles were screened by title and abstract, and 150 were assessed in full text. Finally, 75 articles were included. A PRISMA 2020 flow diagram (Figure 1) summarizes this process.
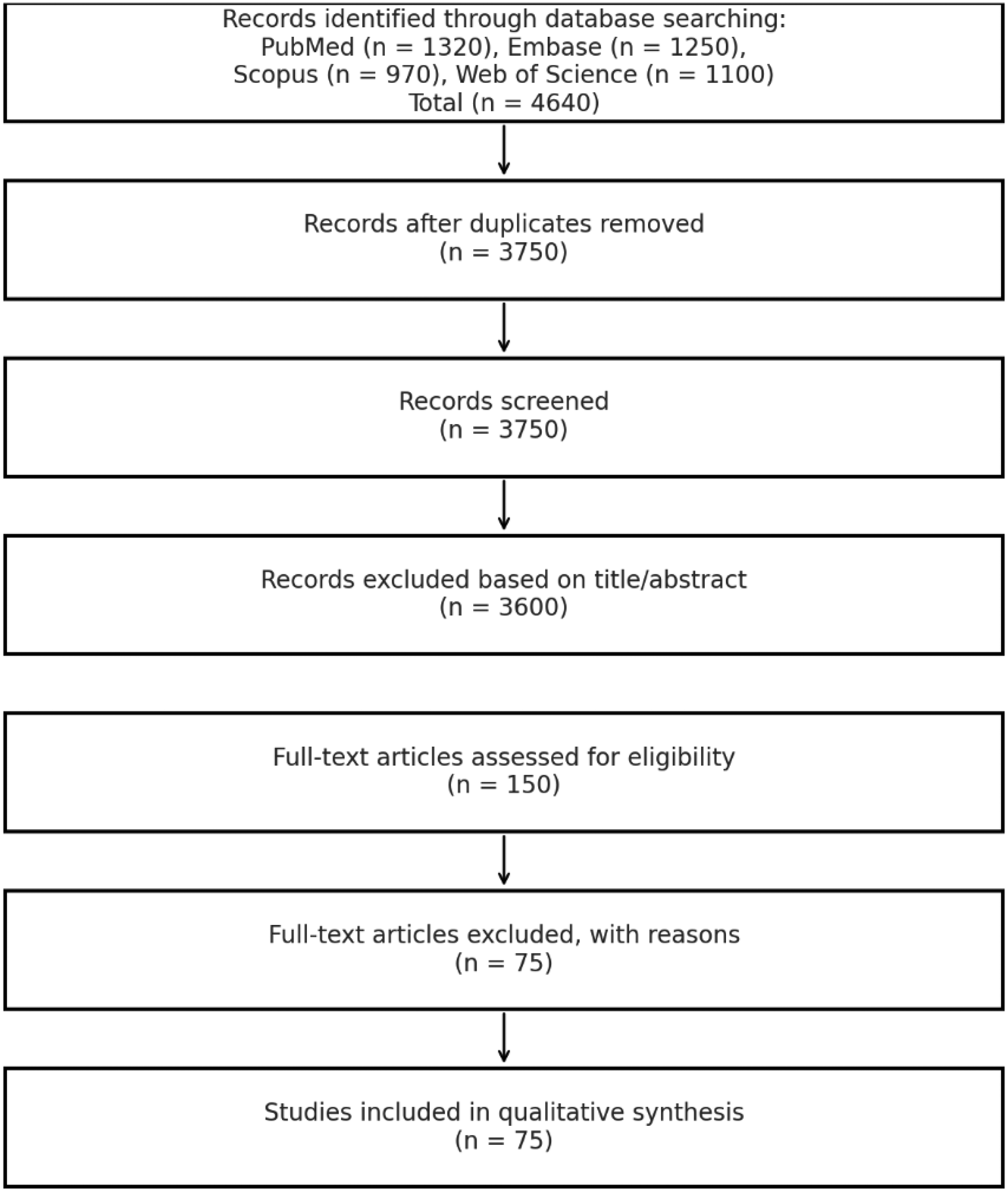
PRISMA 2020 flow diagram for study search.
Eligibility criteria
Inclusion criteria were: (1) original research articles or clinical trials published in English; (2) studies investigating molecular characterization or targeted therapies in early-stage or locally advanced NSCLC; (3) papers reporting outcomes of genomic profiling, ctDNA-based monitoring, or resistance to therapy. Exclusion criteria included: (1) reviews, editorials, and conference abstracts; (2) studies focused exclusively on metastatic NSCLC; (3) preclinical studies without clinical correlation.
Data extraction
Data extraction was conducted using a standardized form developed for this review. Key variables extracted include: study characteristics (authors, year, country, design), patient population (stage, histological subtype), molecular alterations (e.g. EGFR, ALK, KRAS, MET), type of molecular testing (tissue vs. liquid biopsy), intervention details (targeted therapy or immunotherapy), and reported clinical outcomes (e.g. disease-free survival, overall survival, recurrence rates, resistance mechanisms).
In addition to electronic database searches, the reference lists of all included articles were manually screened to identify additional eligible studies. Citation tracking (“snowballing”) was performed using tools such as Google Scholar and Scopus to identify newer studies that cite the included papers. Conference abstracts and proceedings from major oncology meetings (e.g. ASCO, ESMO, WCLC) were reviewed for relevant unpublished data. Where necessary, study authors were contacted for clarification or to obtain missing information.
For observational cohort studies, we used the Newcastle–Ottawa Scale (NOS), which is widely adopted for non-randomized designs. Studies scoring 7 or higher were considered high quality. For randomized controlled trials, we applied the Cochrane RoB 2 tool to better assess randomization, allocation concealment, and blinding. This dual-tool approach allowed tailored quality assessment according to study design.
All search results were imported into reference management software (EndNote X9, Clarivate Analytics; 2013. Philadelphia, PA, USA.) for duplicate removal. Screening of titles and abstracts, followed by full-text eligibility assessment, was conducted independently by two reviewers (WA and AA) as well as any risk of bias. Discrepancies were resolved through discussion or third-party adjudication (SA). In addition, the potential for selective outcome reporting and publication bias was evaluated by reviewing study protocols (if available), trial registries, and comparing registered outcomes with reported results. For randomized trials, discrepancies between prespecified and published outcomes were noted. Moreover, the overall certainty of the evidence was evaluated using the GRADE approach, where applicable. We assessed the body of evidence for each key outcome across studies based on risk of bias, consistency, directness, precision, and potential publication bias. The certainty was categorized as high, moderate, low, or very low.
Outcomes
The primary outcomes were the prevalence and prognostic implications of actionable molecular alterations in early-stage and locally advanced NSCLC. Secondary outcomes included the impact of genomic profiling and ctDNA analysis on treatment decisions, resistance mechanisms, and patient survival.
Review
Molecular profiling in early stage and locally advanced NSCLC
The implementation of comprehensive genomic profiling (CGP) in early-stage NSCLC has emerged as a critical tool for identifying actionable driver mutations and accelerating treatment decisions at disease recurrence. 3 Studies analyzing over 6697 patients with lung adenocarcinoma (LUAD) who underwent surgery between 2011 and 2021 demonstrated that broad genomic testing leads to faster rates of targeted treatment at recurrence with metastatic disease. 14 This approach proves particularly valuable given that up to 55% of patients with NSCLC face disease recurrence after resection, 15 yet currently only one in five patients with early disease receive the benefits of comprehensive molecular testing, 16 proving a critical gap in precision of molecular oncology implementation.
Insights from recent genomic studies
Recent advances in the genomic profiling of NSCLC have provided critical insights into the mutational landscape and its clinical implications. In a landmark study, 17 comprehensive analysis of 74 early-stage NSCLC patients revealed a diverse array of 285 mutations across both tissue DNA and circulating tumor DNA (ctDNA) samples. The most frequently mutated genes included EGFR, TP53, KMT2B, BRAF, PIK3CA, CDKN2A, and KRAS. Importantly, EGFR mutations were detected in 42.9% of tissue samples but only 12.8% of ctDNA, underscoring the limitations of plasma-based genotyping in early-stage disease. The clinical relevance of EGFR mutations was further supported by the ADAURA trial, 18 where adjuvant Osimertinib significantly improved disease-free survival (DFS) in patients with resected EGFR-mutated NSCLC. Moreover, a recent study 19 employing next-generation sequencing (NGS) of a 1021-gene panel in 36 Chinese NSCLC patients revealed distinct mutation profiles between tumor subtypes and metastatic status. CTNNB1 mutations were exclusively detected in non-metastatic lung adenocarcinoma (LUAD), while ARID1A mutations were significantly associated with non-metastatic lung squamous cell carcinoma (LUSC), highlighting subtype- and stage-specific genetic alterations. These findings highlight the critical need for integrating tissue and liquid biopsy-based genomic assessments to enhance precision oncology by guiding targeted therapies, monitoring disease evolution, and adapting treatment strategies in real time. This approach is particularly vital in early-stage NSCLC, where matching patients to therapies like osimertinib based on dynamic molecular profiles can significantly improve clinical outcomes.
Comparative analysis of early- vs. late-stage NSCLC
McGuire et al. (2022)
2
conducted a detailed comparative study to identify clinically actionable molecular alterations in early-stage versus late-stage NSCLC, utilizing an age- and sex-matched pair analysis approach. Their findings demonstrate that key actionable mutations—such as KRAS, EGFR, ALK, ROS1, RET, and MET exon 14 skipping—occur at comparable frequencies in both early and late-stage disease. While
In early-stage NSCLC, the distribution of actionable genetic alterations varies by molecular subtype. Muthusamy et al. (2024)
20
reported mutation frequencies of 16.1% for
Moreover, Yokochi et al. (2025)
22
conducted a retrospective database analysis assessing the prevalence and characteristics of druggable mutations detected by cancer gene panel (CGPa) testing among patients with NSCLC in Japan. The study included 1425 adult NSCLC patients without prior known actionable mutations. Results demonstrated that 44.6% of patients harbored druggable mutations, predominantly involving
Abbreviations: NSCLC; non-small cell lung carcinoma
Prognostic significance of actionable oncogenic drivers
The prognostic and predictive implications of actionable biomarkers differ substantially between early and advanced stages of NSCLC. For example,
Development of combination targeted therapies
The integration of comprehensive genomic profiling in EGFR-mutant NSCLC has revealed that co-occurring actionable mutations—such as de novo MET amplification and ERBB2 alterations—are associated with significantly shorter progression-free survival when treated with first-generation EGFR TKIs, underscoring their adverse prognostic impact. 27 These findings emphasize the need for thorough molecular characterization prior to initiating targeted therapy. Additionally, recent evidence from the ADAURA trial has confirmed the clinical benefit of adjuvant osimertinib in patients with resected EGFR-mutant NSCLC, particularly those harboring exon 19 deletions or L858R mutations, demonstrating improved outcomes and establishing adjuvant targeted therapy as a new standard of care in early-stage disease. 18 Moreover, a recent comprehensive synthesis demonstrated that molecularly selected patients with early-stage NSCLC derive substantial benefit from adjuvant and perioperative targeted therapies, showing significant and durable improvements in disease-free survival (DFS), particularly in EGFR- and ALK-altered cohorts, thereby reinforcing the integration of biomarker-driven strategies into curative treatment pathways. 27 Collectively, these insights highlight the critical role of molecular profiling in guiding both initial and adjuvant treatment strategies in EGFR-driven NSCLC.
Emerging evidence suggests that
Resistance mechanisms to single-agent and combination therapies
The introduction of targeted therapies against oncogenic drivers, such as EGFR, ALK, ROS1, RET, MET, BRAF, and KRAS, has revolutionized NSCLC management. However, resistance to these agents remains a major clinical challenge.
Resistance to EGFR tyrosine kinase inhibitors
Resistance to EGFR TKIs in NSCLC can be broadly categorized into EGFR-dependent and EGFR-independent molecular mechanisms, each posing distinct therapeutic challenges. Primary resistance, observed in approximately 20–30% of cases, is defined by a poor initial response to TKIs and is frequently attributed to pre-existing de novo mechanisms such as co-occurring genetic alterations or alternative pathway activations that bypass EGFR inhibition. 29 This fining underscore the necessity of extensive pre-treatment molecular profiling in advanced NSCLC to identify inherently TKI-resistant cases and to inform the development of combinatorial targeted therapies aimed at overcoming intrinsic resistance.
The
However, resistance to osimertinib emerges and involves multiple mechanisms, with
Additional resistance mutations include
Resistance to ALK inhibitors
For ALK-rearranged NSCLC, resistance often arises through secondary mutations in the ALK kinase domain (e.g. L1196 M and G1202R) or amplification of the ALK gene itself, which sustain downstream signaling despite TKI treatment. 35 Additionally, bypass signaling pathways—such as EGFR or KRAS activation—can compensate for ALK inhibition, enabling tumor survival. 36 These adaptive changes highlight the tumor's ability to exploit parallel signaling networks when primary oncogenic drivers are suppressed.
Resistance to ROS1 inhibitors
ROS1 rearrangements, while relatively rare (0.6% in early-stage disease), represent highly actionable alterations with approved targeted therapies.
37
The identification of these fusions through comprehensive genomic profiling has enabled access to effective treatments such as crizotinib and subsequent-generation ROS1 inhibitors.
37
Resistance in ROS1-rearranged NSCLC frequently involves
Resistance to MET inhibitors
In MET-driven NSCLC, particularly those with MET exon 14 skipping mutations or amplification, resistance to MET inhibitors (e.g. capmatinib, tepotinib) arises via several mechanisms. These include
A recent case report in Frontiers in Oncology demonstrated that even low-level MET gene copy gain in an EGFR-mutant, TKI-resistant metastatic NSCLC patient yielded a meaningful therapeutic response—specifically, a 7-month progression-free survival—when treated with combined osimertinib and savolitinib, underscoring the potential utility of targeted therapy against MET-mediated resistance even at sub-threshold amplification levels. 39
Resistance to BRAF inhibitors
Similarly, resistance to BRAF-targeted therapies in patients with BRAF V600E mutations, such as dabrafenib and trametinib, may occur through MAPK pathway reactivation via
Resistance to KRAS inhibitors
Recent advances in KRAS-targeted therapy, particularly with KRAS G12C inhibitors like sotorasib and adagrasib, have demonstrated promising initial responses in NSCLC patients. Nonetheless, resistance can develop through multiple routes, including
Other resistance mechanisms
The complexity of therapeutic resistance in NSCLC is further intensified by histological transformation and epigenetic reprogramming. A subset of tumors undergo
Impact of molecular advances on patient outcomes and survival improvements
Beyond targeted gene panels, emerging comprehensive sequencing technologies are increasingly applied in early-stage NSCLC. Whole-exome sequencing (WES) provides detailed information on mutational burden and novel driver events, while RNA sequencing enables the detection of gene fusions, alternative splicing, and expression-based signatures with therapeutic implications.19,20 Furthermore, liquid biopsy–based NGS platforms are now being investigated for minimal residual disease (MRD) detection, offering the ability to non-invasively monitor recurrence risk and treatment response. Collectively, these technologies expand the scope of precision oncology by integrating genomic, transcriptomic, and dynamic molecular data into therapeutic decision-making.
Integration of comprehensive genomic profiling (CGP)
Real-world evidence from comprehensive genomic profiling studies shows that patients who receive CGP have improved detection of actionable biomarkers (53% versus 14%) and greater use of matched therapies compared to single-gene testing approaches. 43 A 2023 multicenter cohort study investigating the timing of CGP in metastatic solid tumors demonstrated a pronounced survival benefit for patients with lung cancer who underwent earlier CGP. Specifically, patients who received CGP in the second and third tertiles after metastatic diagnosis experienced significantly higher mortality risks compared to those tested in the first tertile. 44 These findings underscore the critical importance of early integration of CGP in the management of metastatic lung cancer, as delayed testing may limit the clinical benefit of targeted therapies and adversely impact patient outcomes.
Moreover, trials such as LC-SCRUM (Lung Cancer Genomic Screening Project for Individualized Medicine) and BATTLE (Biomarker-integrated Approaches of Targeted Therapy for Lung Cancer Elimination) have demonstrated the feasibility and clinical utility of molecular profiling in real-time decision-making. The BATTLE trial, a pioneering study in adaptive randomization based on biomarker profiles, showed that tailoring therapy to individual tumor genomics improves disease control rates, particularly in previously treated NSCLC patients. 45 Similarly, the LC-SCRUM-Japan initiative established a nationwide genomic screening platform for lung cancer, facilitating early identification of rare but targetable alterations like ALK, ROS1, and RET fusions, which expanded therapeutic options and enrollment in matched clinical trials. 46 Moreover, the NCI-MATCH trial (Molecular Analysis for Therapy Choice) included NSCLC among various solid tumors and emphasized that assigning therapies based on specific molecular alterations, regardless of tumor origin, can lead to clinically meaningful responses. 47 These studies collectively underscore that early and comprehensive genomic analysis should be an integral part of NSCLC management to personalize treatment, avoid ineffective therapies, and enhance long-term survival.
The economic implications of routine CGP in early-stage NSCLC are increasingly recognized. In a recent U.S.-based cost-effectiveness analysis,
Table 2 summarizes the most recent clinical trials highlighting the importance of molecular diagnosis in early-stage and locally advanced NSCLC.
Summary for the recent key clinical trials (2019–2024) on molecular advances in early-stage and locally advanced NSCLC.
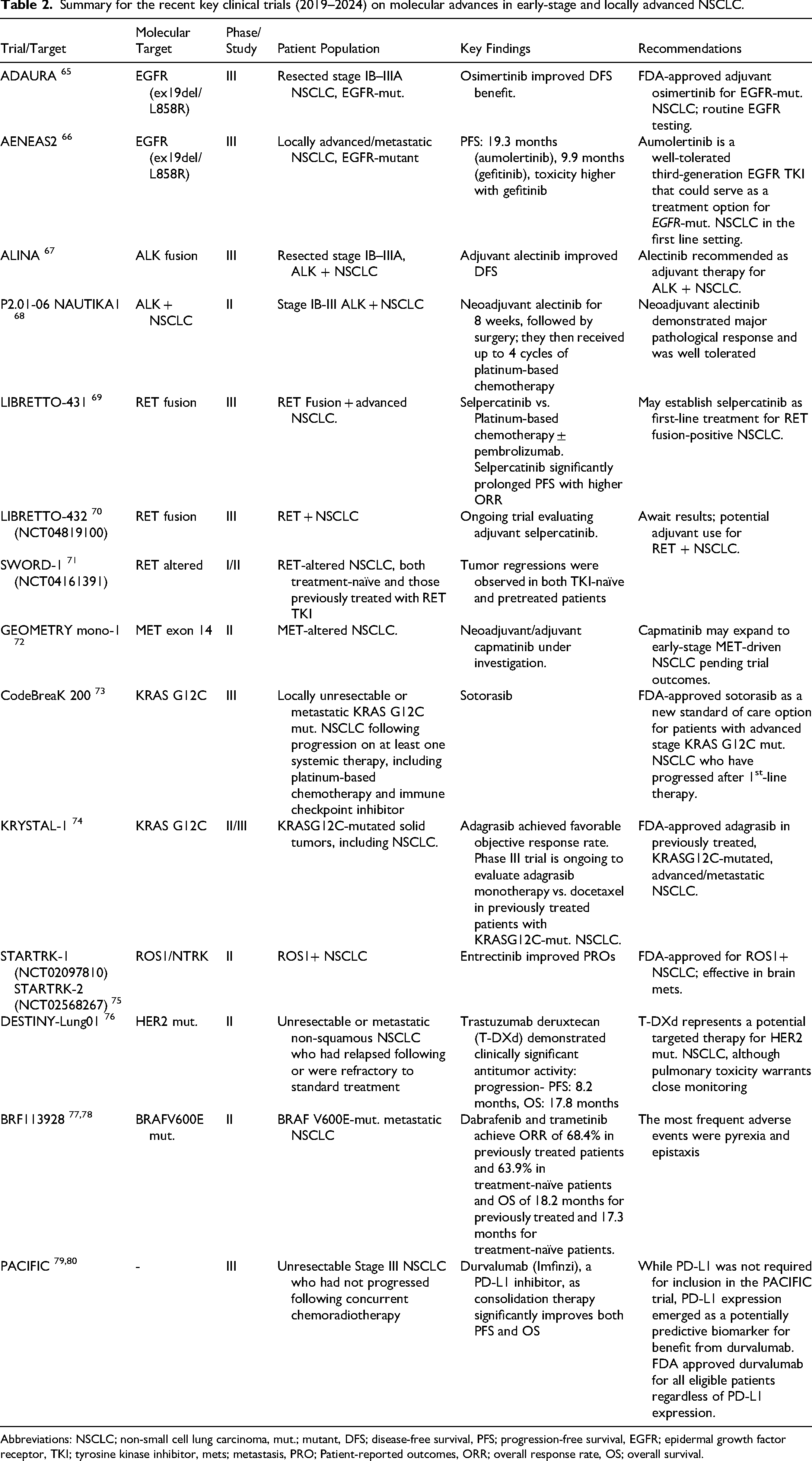
Abbreviations: NSCLC; non-small cell lung carcinoma, mut.; mutant, DFS; disease-free survival, PFS; progression-free survival, EGFR; epidermal growth factor receptor, TKI; tyrosine kinase inhibitor, mets; metastasis, PRO; Patient-reported outcomes, ORR; overall response rate, OS; overall survival.
Integration of circulating tumor DNA
The integration of ctDNA-based MRD monitoring has emerged as a transformative tool in NSCLC, enabling early detection of recurrence, stratification of relapse risk, and refinement of post-treatment surveillance strategies. It represents a promising approach for early-stage disease monitoring with significant clinical potential.
ctDNA-based MRD detection strategies can be broadly divided into tumor-informed and tumor-agnostic approaches. Tumor-informed assays, such as Signatera, require prior knowledge of patient-specific tumor mutations and enable highly sensitive detection of recurrence risk. Tumor-agnostic approaches, in contrast, use predefined gene panels without prior tumor sequencing, offering faster turnaround but generally lower sensitivity in early-stage disease. 48
In early-stage NSCLC, these approaches have been evaluated in landmark adjuvant and neoadjuvant immunotherapy trials. For example, ctDNA analyses in IMpower010 (atezolizumab), CheckMate-816 (nivolumab), AEGEAN (durvalumab), and NADIM (nivolumab + chemotherapy) have demonstrated prognostic value, with ctDNA clearance after surgery or neoadjuvant treatment correlating with improved disease-free survival.49,50,51 The integration of ctDNA into escalation/de-escalation strategies is an area of active investigation, aiming to tailor adjuvant therapy intensity based on MRD status rather than stage alone. Ongoing phase III trials, such as MERMAID-1 and MERMAID-2, are prospectively evaluating ctDNA-guided therapeutic strategies, highlighting the potential of this approach to optimize treatment selection in early-stage and locally advanced lung cancer. 50
In comparison with tissue-based next-generation sequencing (NGS), liquid biopsy has faster turnaround and facilitates longitudinal sampling, yet its sensitivity is reduced in early-stage disease due to low ctDNA burden. Comparative workflow analyses suggest that tissue NGS is more effective for baseline comprehensive profiling, whereas liquid biopsy is particularly valuable for serial monitoring, detecting MRD, and overcoming spatial heterogeneity. Thus, the two approaches are complementary rather than competitive, and their integration offers the most robust strategy for precision oncology in both early-stage and advanced NSCLC.42,48
Economic impact and the promise of targeted-immunotherapy synergy
The integration of comprehensive molecular profiling in NSCLC has demonstrated both clinical and economic value. By identifying patients with genomic alterations unlikely to benefit from immunotherapy, such profiling enables the avoidance of ineffective treatments and associated toxicities, leading to significant cost savings compared to limited approaches like EGFR-only testing. Furthermore, the combination of targeted therapies with immunotherapy is emerging as a promising strategy to enhance outcomes, particularly in subgroups with lower monotherapy efficacy—such as EGFR-mutant tumors. In adjuvant immunotherapy, the IMpower010 trial demonstrated that atezolizumab significantly improved DFS versus best supportive care after adjuvant chemotherapy in patients with stage II–IIIA NSCLC, particularly those with PD-L1 ≥ 1%, leading to its FDA approval. Similarly, PEARLS/KEYNOTE-091 evaluated pembrolizumab in the adjuvant setting and showed a DFS benefit irrespective of PD-L1 status, expanding applicability to a broader patient population.51,52
Emerging biomarkers and novel therapeutic targets
The landscape of biomarker discovery in NSCLC continues to expand beyond traditional driver mutations, with NGS revealing previously unrecognized therapeutic targets and prognostic markers.
Fanconi anemia pathway
Innovative projects implementing NGS in early-stage NSCLC have identified novel genetic alterations with potential roles as biomarkers of tumor evolution and patient monitoring. Analysis of 15 surgical samples using comprehensive cancer gene panels revealed that 39.36% of genes presented frequent mutations, with significant representation of Fanconi anemia pathway mutations in 80% of analyzed patients. 53
NOTCH signaling pathway
Notch signaling pathway alterations have emerged as a significant area of investigation, with NOTCH3 mutations identified among the genes with highest mutation frequencies in ctDNA from early-stage patients.54,55 The Notch pathway's role in lung cancer development and its potential as a therapeutic target continues to be explored through ongoing clinical investigations.
NTRK gene fusions
NTRK gene fusions, though rare, represent clinically actionable oncogenic drivers that enable targeted therapy with tropomyosin receptor kinase (TRK) inhibitors when identified through comprehensive molecular profiling. 56 These fusions result from chromosomal rearrangements involving NTRK1, or NTRK3 genes, leading to constitutively activated TRK fusion proteins that promote tumorigenesis in various NSCLC histologies, including LUAD, LUSC, and neuroendocrine carcinoma, without restriction to specific clinical features such as age, sex, or smoking history. 56 The identification of NTRK fusions in NSCLC has direct therapeutic implications, as first-generation tropomyosin receptor kinase (TRK) inhibitors—larotrectinib and entrectinib—have demonstrated high response rates and durable disease control, including in patients with central nervous system metastases. 57 These agents are now recommended as first-line treatment options for patients with advanced or metastatic NSCLC harboring NTRK fusions, offering a histology-agnostic approach to targeted therapy. Given the low frequency but high therapeutic relevance of NTRK fusions, broad molecular profiling—preferably with RNA-based NGS—is advocated for all NSCLC patients, particularly those lacking other actionable drivers. 58 This approach ensures that eligible patients are identified for precision therapy with TRK inhibitors, which can significantly improve outcomes in this otherwise challenging subset of lung cancer
Homologous recombination repair pathway
The characterization of homologous recombination repair pathway alterations has revealed new therapeutic opportunities in NSCLC. Comprehensive genomic profiling has identified alterations in the HRR pathway, including somatic BRCA1/2 mutations in advanced NSCLC, providing treatment options with platinum agents or PARP inhibitors. 59 This represents an expansion of precision medicine approaches beyond traditional lung cancer targets.
Microsatellite instability
Tumor mutational burden (TMB) and microsatellite instability (MSI) have emerged as pivotal tumor-agnostic biomarkers guiding immunotherapy decisions in NSCLC. High TMB and MSI-high (MSI-H) status are associated with increased neoantigen load and immune responsiveness, correlating with improved outcomes to immune checkpoint inhibitors. Studies indicate that up to 16% of NSCLC patients harbor such immunotherapy-predictive genomic alterations. 60 CGP enables simultaneous assessment of these biomarkers alongside actionable driver mutations, allowing for a more nuanced understanding of each tumor's molecular landscape. 60 This integrated approach enhances patient stratification and optimizes personalized immunotherapy strategies.
The certainty of evidence across key clinical and molecular outcomes was assessed using the GRADE approach, and the findings were summarized in Table 3.
GRADE evidence profile of molecular advances in NSCLC.
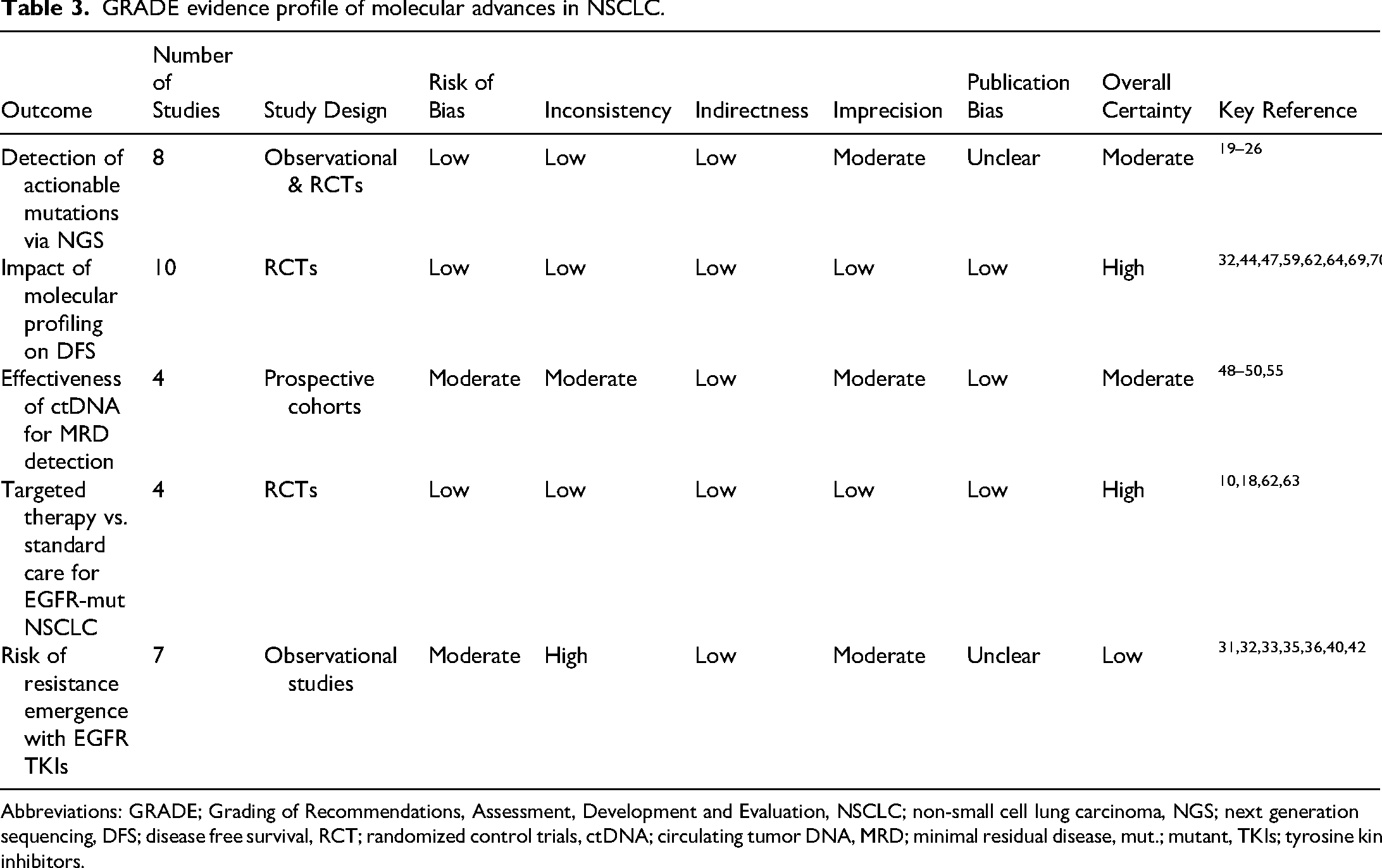
Abbreviations: GRADE; Grading of Recommendations, Assessment, Development and Evaluation, NSCLC; non-small cell lung carcinoma, NGS; next generation sequencing, DFS; disease free survival, RCT; randomized control trials, ctDNA; circulating tumor DNA, MRD; minimal residual disease, mut.; mutant, TKIs; tyrosine kinase inhibitors.
Bridging the gap-what are we still missing?
Stage-specific genomic differences
Despite recent advances in genomic profiling that revealed the molecular landscape of NSCLC, it shows that this cancer varies significantly between early-stage (operable) and stage IIIA (locally advanced) disease. There is a significant molecular difference between early-stage (I–II) and stage IIIA NSCLC. Early-stage tumors often harbor higher frequencies of KRAS and MET exon 14 skipping mutations, whereas EGFR, ALK rearrangements, and BRAF V600E mutations are more frequently observed in advanced stages.2,20,21 In non-metastatic subtypes, CTNNB1 mutations are enriched in LUAD, while ARID1A alterations are more common in LUSC. Stage IIIA tumors demonstrate increased TP53, KEAP1, and STK11 co-mutations, which are linked to immunotherapy resistance.7,8,26 These patterns highlight the need for stage-specific molecular profiling to guide therapeutic strategies.
Importantly, several of these alterations are already being explored in early-stage and locally advanced NSCLC clinical trials. For instance, ADAURA (NCT02511106) established the benefit of adjuvant osimertinib in resected EGFR-mutant NSCLC, while the ALINA trial (NCT03456076) demonstrated the efficacy of alectinib in ALK-positive resected tumors . Similarly, BRAF V600E-mutant NSCLC is under investigation in early-stage adjuvant settings using dabrafenib–trametinib combinations.27,28,31 These studies highlight that molecular profiling is not only descriptive but also directly informs trial-based therapeutic strategies in operable NSCLC.
Tumor microenvironment barriers
The tumor microenvironment (TME) significantly influences therapeutic efficacy in NSCLC, particularly in stage IIIA disease. Activated fibroblasts and elevated TGF-β levels impair the delivery and activity of EGFR TKIs. Additionally, immune exclusion, characterized by reduced CD8+ T-cell infiltration, diminishes response to checkpoint inhibitors . Hypoxia-induced resistance, mediated by HIF-1α, further compromises the effect of agents like Osimertinib.31,32,33 Targeting both tumor-intrinsic and TME-mediated resistance mechanisms is therefore essential for optimizing treatment outcomes.
Recent perioperative immunotherapy trials, including CheckMate 816 (NCT02998528), have shown improved pathological response rates when immune checkpoint inhibitors are used in resectable NSCL. However, resistance patterns associated with STK11, KEAP1, and TP53 co-mutations7,26 underscore the importance of integrating molecular context into trial design and patient stratification.
Liquid biopsy limitations
Liquid biopsy, particularly circulating tumor DNA (ctDNA) analysis, has emerged as a promising tool for monitoring minimal residual disease (MRD) and predicting recurrence in NSCLC. However, its sensitivity is substantially reduced in early-stage disease due to lower tumor burden.17,18 For instance, the detection rate of EGFR mutations in plasma is significantly lower than in tissue samples, with only a minority of stage I patients exhibiting detectable ctDNA. This limitation is primarily attributable to the low fraction of tumor-derived DNA in circulation at early stages. To overcome this challenge, combining ctDNA analysis with circulating tumor cell (CTC) enumeration has been shown to enhance sensitivity. Moreover, serial ctDNA monitoring can improve the prediction of recurrence, enabling more timely therapeutic interventions.48,49,50 Despite these advances, the clinical utility of ctDNA in early-stage NSCLC remains constrained by technical limitations, and further research is needed to optimize its application in this setting.
Prospective studies such as TRACERx and ongoing MERMAID-1 and −2 trials (NCT04642469, NCT04642492) are evaluating ctDNA-guided adjuvant strategies, demonstrating how molecular monitoring is increasingly embedded into early-stage trial frameworks.
Limitations of molecular profiling
Despite its transformative potential, molecular profiling in early-stage NSCLC faces several limitations. Tissue sampling in small tumors often results in inadequate DNA yield, and intratumoral heterogeneity may lead to sampling bias that underestimates genomic complexity.17,19 Also, liquid biopsy approaches have reduced sensitivity in early disease due to the low ctDNA fraction.48,49,50 NGS technologies present logistical challenges, including turnaround time, interpretation of variants of uncertain significance, and disparities in access across healthcare systems.19,20,22 Moreover, the cost-effectiveness of universal NGS in early-stage NSCLC remains debated, particularly in low- and middle-income countries with limited infrastructure . These limitations underscore the importance of combining molecular assays with clinical and pathological risk factors to ensure accurate patient stratification.
Ongoing MERMAID trials (NCT04385368) 61 are evaluating ctDNA-guided adjuvant strategies, demonstrating how molecular monitoring is increasingly embedded into early-stage trial frameworks.
Global implementation challenges
Comprehensive genomic profiling (CGP) demonstrates cost-effectiveness in high-income settings by improving quality-adjusted life years through personalized therapy. However, resource-limited regions face challenges including high cost, limited NGS infrastructure, and poor adherence to testing guidelines. These disparities necessitate strategic investment in diagnostics, capacity building, and policy reforms to achieve equitable implementation of precision oncology.
Recent advances in artificial intelligence (AI) and digital pathology are reshaping the landscape of precision oncology in NSCLC. AI-driven models trained on multi-omics data and radiogenomic features have demonstrated high accuracy in stratifying early-stage disease and predicting recurrence, complementing traditional clinicopathological risk factors. 62 Moreover, digital pathology platforms applying spatial transcriptomics and multiplex immunohistochemistry provide unprecedented resolution of tumor heterogeneity, allowing integration of molecular and morphological biomarkers. These innovations enhance patient stratification, refine minimal residual disease (MRD) monitoring, and inform perioperative therapeutic decisions, thereby addressing critical gaps highlighted in earlier targeted therapy and immunotherapy trials.
Moreover, a recent work demonstrates that radiogenomic machine learning algorithms, combining imaging features with molecular profiling, reliably predict key oncogenic mutations and recurrence risk in early-stage NSCLC with high accuracy—highlighting their potential as non-invasive, dynamic profiling tools that complement tissue and liquid biopsy approaches. 63
Future directions
Emerging innovations such as spatial transcriptomics and AI-powered digital pathology offer insights into intratumoral heterogeneity and molecular subtyping.53,54 These tools, combined with advanced MRD monitoring49,50 and novel targeted agents,56,57,59 are poised to enhance NSCLC personalization. Ongoing translational research is crucial to overcoming therapeutic resistance and realizing the full potential of molecular strategies in early and locally advanced NSCLC.
Emerging therapeutic approaches are poised to further transform the early-stage NSCLC landscape. mRNA-based cancer vaccines, designed to elicit robust antitumor immune responses against patient-specific neoantigens, have demonstrated promising results in early-phase clinical trials. 60 Similarly, recombination-based therapies, including engineered T-cell receptor (TCR) and CAR-T strategies, are being explored for their ability to target tumor-specific mutations and neoantigens. 64 These novel modalities complement ongoing efforts to integrate multi-omic profiling, spatial transcriptomics, and artificial intelligence into clinical workflows, offering unprecedented opportunities to refine patient selection and improve outcomes. Their incorporation into perioperative settings could represent the next frontier in curative-intent precision oncology.
Collectively, the integration of targeted therapies (ADAURA, ALINA), perioperative immunotherapy (CheckMate 816), and MRD-directed strategies (TRACERx, MERMAID) provides a clinically relevant roadmap. Future studies combining NGS-based detection, spatial transcriptomics, and AI-driven risk stratification will be critical for refining precision oncology in early and locally advanced NSCLC.53,65,66
In addition to molecular and technical considerations, the psychosocial dimensions of precision oncology warrant greater emphasis. Emerging survivorship studies demonstrate that genomic testing and precision-guided treatment may impose significant emotional, informational, and financial burdens on patients and families. 64 Anxiety surrounding genomic risk, expectations of targeted therapies, and uncertainty in test interpretation can exacerbate distress, particularly in resource-limited settings where access to novel agents remains inequitable. Incorporating psychosocial support, patient education, and culturally sensitive counseling into precision oncology pathways is therefore essential.
Summary and conclusion
In conclusion, the findings of this review highlight not only the expanding molecular understanding of NSCLC but also its direct translation into practical clinical algorithms for early and locally advanced disease. By linking genomic alterations and resistance mechanisms with perioperative trial evidence, the narrative underscores how molecular biology insights are increasingly shaping real-world treatment pathways for oncology practitioners. Advances in genomic and transcriptomic technologies, combined with liquid biopsy applications, have enabled the early detection of actionable alterations, the identification of resistance mechanisms, and the optimization of perioperative therapeutic strategies. Integration of biomarkers such as EGFR, ALK, KRAS, and PD-L1 into clinical practice has already reshaped standard care, while novel approaches—including mRNA vaccines, recombination therapies, and multi-omic platforms—are expanding the horizon of precision oncology. Nevertheless, challenges related to tissue sampling, heterogeneity, cost-effectiveness, and global accessibility remain. Continued translational research and equitable implementation are essential to realize the full potential of molecular advances in achieving durable cures for patients with early-stage and locally advanced NSCLC.
This review is limited by heterogeneity in included studies, differences in patient selection criteria for early-stage NSCLC, and underrepresentation of negative or unpublished studies. Variations in molecular testing methods and reporting standards may also introduce bias. As this is a narrative synthesis rather than a meta-analysis, effect estimates were not pooled.
Footnotes
Acknowledgements
The author thanks Dr Ahmed Ali (AA) and Dr Sara Ahmad (SA) for assistance with literature screening, study selection, and risk-of-bias assessment. AI-assisted tools were used only for language refinement/formatting; all scientific content was developed by the author.
Ethical considerations
This work is a systematic narrative review conducted in accordance with PRISMA guidelines and did not involve any new studies with human participants or animals performed by the authors. Hence, ethical approval was not required.
Author contributions
WA (Wael Abdo Hassan) conceived and designed the study; developed and registered the protocol; performed database searches, screening, eligibility assessment, and data extraction; conducted the qualitative synthesis and GRADE assessment; drafted and critically revised the manuscript; and approved the final version.
Declaration of conflicting interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author received no financial support for the research, authorship, and/or publication of this article.
Data availability statement
All data generated or analyzed in this study are included in the article and its supplementary materials.

