Abstract
Breast cancer recurrence remains a major cause of mortality, with up to 30% of early-stage patients relapsing as incurable metastatic disease. Conventional surveillance with imaging and serum markers (CA15–3, CEA) lacks the sensitivity and specificity to detect minimal residual disease. This narrative review examines non-invasive biomarkers such as circulating tumor DNA (ctDNA), circulating tumor cells (CTCs) and exosomes and the technologies enhancing their performance. Droplet digital PCR and next-generation sequencing detect ctDNA at allele frequencies below 0.1%, identifying molecular relapse a median of 10–12 months before radiologic progression. Microfluidic and affinity-based platforms isolate CTCs with over 75% sensitivity in metastatic settings. Nanoengineered sensors and standardized workflows improve exosome isolation, revealing miRNA and protein signatures predictive of recurrence. Proteomic and metabolomic profiling identify dysregulated metabolic pathways and protein networks, offering functional insights that complement molecular assays. Integrative multi-omics approaches merge genomic, transcriptomic, proteomic and metabolomic data; machine-learning frameworks detect subtle patterns and correlations, enabling dynamic, personalized surveillance. By detecting molecular and functional biomarkers early, clinicians can tailor therapy, monitor treatment response and intervene promptly. Challenges include low analyte abundance, assay variability, high costs and lack of standardized protocols, limiting clinical adoption. Prospective validation in large cohorts is critical. We highlight ongoing clinical trials such as ctDNA-guided adjuvant therapy and CTC-driven stratification studies that aim to establish clinical utility. Non-invasive biomarker platforms could shift breast cancer follow-up from reactive detection to proactive intervention, ultimately improving survival and quality of life through personalized, real-time monitoring.
Keywords
Introduction
Breast cancer (BC) is a heterogeneous disease, with varying molecular properties, multimodal therapeutic strategies and expected prognoses. 1 At present, BC recurrence is a major problem in oncology and is one of the most important causes of long-term mortality and morbidity in millions of women all over the world.2,3 Evolving treatments have improved the initial management. Nonetheless, approximately 25%–30% of women with early-stage breast cancer will experience recurrence, which may progress to metastatic disease. 4 In the disease recurrence, the outcome is dependent on several clinicopathological and immunohistochemical (IHC) characteristics including increased tumor burden, shorter metastatic interval as well as general patient performance factors. 5 Notably, the early identification of the recurrence is vital for the purpose of timely intervention leading to better prognosis. However, the methods that are currently used to monitor the recurrence including, imaging and serum tumor markers, are invasive, costly, less sensitive and non-specific.6,7 For instance, the detection of early-stage tumor or residual lesions is unsatisfactory, and its application in the evaluation of treatment efficacy and prognosis is also limited. 8 These limitations call for the need for non-invasive, reliable biomarkers that can give accurate and early diagnosis of BC recurrence.
Non-invasive biomarkers present a new approach in monitoring incidence and recurrence of cancer as they provide real time information about the tumor without the need for invasive procedures.9,10 They are different from the conventional methods of testing such as tissue biopsies or invasive imaging because they are able to detect the molecular markers of the recurrence of the cancer through the analysis of circulating tumor cells (CTCs), circulating tumor DNA (ctDNA), and exosomes, which are small particles called extracellular vesicles (EVs).11–14 These methods are not only less invasive than traditional methods but can also be used to monitor the dynamics of the disease and the treatment. 15 In an era of precision medicine, identifying and validating these biomarkers hold immense potential to transform the landscape of cancer care. 16
Technological advances have been vital in the development and use of non-invasive biomarkers. Development of new technologies such as ddPCR or optimized NGS has greatly improved the sensitivity, specificity and precision for the detection of ctDNA, thereby predicting cancer recurrence and minimal residual disease as well as early diagnosis of cancer patients.17,18 In addition, microfluidic technologies and other methods have advanced the isolation and characterization of CTCs and exosomes (

Overview of breast cancer biomarker workflow. This figure illustrates the workflow for breast cancer biomarker, spanning from sample collection to biomarker identification and analysis. The leftmost panel outlines various sources of biological samples, including breast tissue, body fluids (e.g. blood, urine, and saliva), cellular samples, and media supernatant from cell culture. These samples serve as inputs for downstream biomarker analysis. The central panel highlights key biomarker detection technologies such as ddPCR, next-generation sequencing (NGS), and microfluidic platforms for high-throughput genomic and transcriptomic analysis. Additionally, nuclear magnetic resonance (NMR) spectroscopy and mass spectrometry are utilized to investigate metabolic profiles and protein abundances. The breast cancer biomarkers identification through multi-omics pathway analysis, encompassing genomic, transcriptomic, proteomic, and metabolomic layers. Specific biomarkers, including circulating tumor DNA (ctDNA), circulating tumor cells (CTCs), exosomes, and extracellular vehicles (EVs), are identified using these technologies, providing valuable insights into the molecular mechanisms and pathways underlying breast cancer progression.
This narrative review is guided by the Scale for the Assessment of Narrative Review Articles (SANRA). 21 The objectives of the review were to (1) summarize recent advancements in non-invasive biomarkers for the detection and monitoring of breast cancer recurrence; (2) critically appraise their analytic performance and clinical utility; and (3) identify the challenges and future research directions. A targeted literature search was performed in different databases such as PubMed/MEDLINE, Embase, Scopus, and Web of Science for articles published until February 2025. The search strategy combined different keywords related to breast cancer, recurrence, and non-invasive biomarkers using Boolean operators to capture all relevant studies.
Current state of non-invasive monitoring of breast cancer recurrence
Monitoring BC recurrence is important for patient survival and as part of the extended cancer control strategy.22,23 Despite the advances in the initial treatment, the recurrence is still a problem of concern and occurs in up to 30% of the patients with early stage BC. 24 This relapse is normally associated with worse patient outcomes especially when the recurrence leads to metastatic disease. The conventional diagnostic approaches include imaging techniques such as mammography, ultrasound, magnetic resonance imaging (MRI), and positron emission tomography (PET) scans, and serum tumor markers including CA 15–3 and CEA. 25 Although these methods are useful in clinical practice, they have some drawbacks, which are particularly significant in the case of early detection of recurrence. 26 Although PET and MRI are sensitive imaging modalities, they are costly, not widely available, and cannot reliably distinguish between scar tissue and true malignancy. This limitation contributes to a higher rate of false-positive findings, often leading to unnecessary biopsies and interventions. Imaging techniques, which are commonly used and non-invasive, have low sensitivity for detecting small or residual disease. 27 Similarly, mammography has difficulties with detecting the recurrent disease in the dense breast in young women or those with postoperative changes. 28 In a similar way, while PET and MRI are sensitive, they are expensive and need special facilities, which are not readily available in resource-constrained settings. These imaging modalities cannot distinguish between scar tissue from surgery and real malignancy, which results in false positives and unnecessary interventions. 29 Other tumor markers, such as CA 15–3 and CEA, also have their limitations. These markers are not very specific and the levels of these markers do not always correlate with the disease recurrence, especially in the early stages.30,31 Abnormal levels may be due to non-malignant conditions and hence the diagnosis may be problematic. Furthermore, their sensitivity is too low to detect subclinical metastasis, thus leading to the detection of symptomatic metastases when it is already late. 32
New biomarker-based monitoring is an innovative strategy that replaces the conventional methods. Among them, liquid biopsy methods that include analysis of ctDNA, CTCs, and EVs are at the forefront of this revolution. 33 These techniques allow the monitoring of the tumor development and changes in the tumor load in real time, before they become apparent on imaging or clinically. 34 Moreover, the application of proteomic and metabolomic profiling in recurrence monitoring involves the analysis of proteins and metabolites in body fluids and identify changes that indicate tumor recurrence. 35 These areas have produced new biomarkers that complement the conventional approaches to the monitoring. Therefore, the current non-invasive monitoring techniques can be characterized as a shift from the use of conventional imaging and enzyme markers to biomarker-based approaches developed with the help of new technologies. Despite the fact that considerable advancement has been made, there are still some issues with the validation of these tools and their integration into clinical practice. However, the ability of recent diagnostic approaches to enhance the early detection and personalized medicine is an area of great interest in oncology. To conclude, while conventional diagnostic tools play a major role in clinical settings, the rise of biomarker-based tool helps in breakthrough in monitor breast cancer. These emerging technologies way more accurate, detect early and individualized reappearance detection, setting the stage for in-depth investigation of novel biomarkers and their clinical potential.
Emerging biomarkers for monitoring breast cancer recurrence
New biomarkers for the monitoring of BC recurrence are identified and applied, and these are of great importance in the field of oncology. Although current markers such as CA 15–3 have poor sensitivity and specificity, these new biomarkers contain more specific information that is more related to the molecular events that occur in the tumor. These are particularly of value for detecting the recurrence at an early stage when treatments are most effective. 36
ctDNA is one of the most promising biomarkers for the recurrence monitoring. 37 ctDNA is released into peripheral blood and other body fluids through apoptosis and necrosis of tumor cells, representing a small fraction of the cell-free DNA (cfDNA) in patients with cancer. 38 Emerging real-world data shows the utility of ctDNA in detecting molecular residual disease and in treatment-response monitoring, helping clinicians to optimize treatment and surveillance strategies. Several approaches to measure ctDNA have been developed, such as panel-based assays, next-generation sequencing and ddPCR. 38 Interestingly, recent studies correlated the presence of ctDNA in postoperatively patients with lung, 39 bladder, 40 gastrointestinal, 41 and colorectal cancers. 42 In BC, the ctDNA presence following resection has been proven to correlate with a poor prognosis and a higher rate of recurrence.43,44
CTCs are also becoming popular as biomarkers for the recurrence of cancer.
45
While ctDNA contains genetic information, CTCs are complete cancer cells that can be examined for their phenotypic and functional properties. The presence of CTCs in the blood is often linked to the metastatic potential, thus, they are useful in predicting the BC recurrence.46,47 The isolation and characterization of CTCs have been enhanced by advances in microfluidic technologies to the point where they can be used in clinical practice.
48
Similarly, exosomes and other EVs are emerging as another important category of biomarkers for BC recurrence.
49
These vesicles are released by tumor cells and contain a molecular payload of proteins, lipids and nucleic acids that describe the biological state of the tumor. Recent studies have identified exosomal RNA and protein signatures that are associated with the recurrence, providing another level of diagnostic accuracy.
50
Various studies have identified certain exosomal miRNAs to play a role in the progression and recurrence of BC. For instance, exosomal protein surviving was found in higher concentrations in BC patients than in healthy population.
51
Tumor-derived exosomal hsa-miR-21-5p, which is detected in peripheral blood, can efficiently discriminate between healthy people and cancer patients. It was highly unregulated in BC patients.
52
In BC patients, serum exosomal lncRNA DANCR levels are also elevated. The use of serum exosomal lncRNA DANCR in combination with A153 and CEA significantly improves diagnostic efficiency.
53
Also, exosomal miR-92b-5p offers a new management approach and serves as a non-invasive biomarker for the diagnosis of BC
54
(
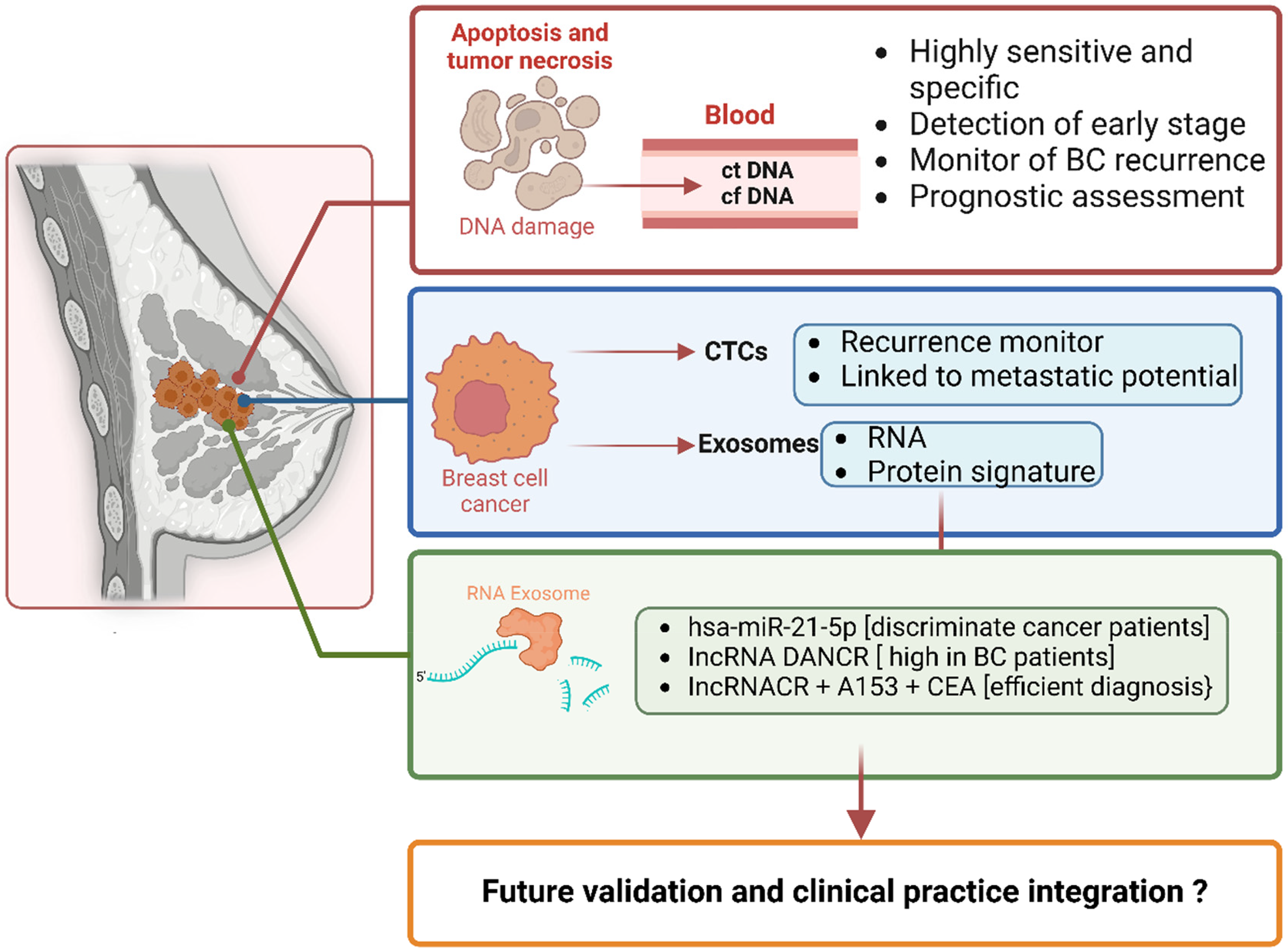
Biomarkers in breast cancer monitoring and prognosis. The figure highlights the role of circulating biomarkers in breast cancer detection, monitoring, and prognosis. The process begins with DNA damage resulting from apoptosis and tumor necrosis, leading to the release of circulating tumor DNA (ctDNA) and cell-free DNA (cfDNA) into the bloodstream. These markers are highly sensitive and specific, enabling early-stage detection, recurrence monitoring, and prognostic assessment. Circulating tumor cells (CTCs), derived from breast cancer cells, act as indicators of recurrence and metastatic potential. Exosomes, another key biomarker, carry RNA and protein signatures that provide valuable molecular insights. Specifically, RNA exosomes contain markers like hsa-miR-21-5p, which discriminates cancer patients, lncRNA DANCR, elevated in breast cancer patients, and a combination of lncRNACR, A153, and CEA, which support efficient diagnosis. This framework underscores the potential of these biomarkers, while also emphasizing the need for further validation and clinical integration for effective implementation.
Importantly, most of these liquid-biopsy markers can be applied across breast cancer subtypes, but certain analytes show subtype enrichment. For example, ctDNA assays targeting PIK3CA and ESR1 mutations are especially informative in hormone-receptor–positive/HER2–negative tumours, whereas ERBB2-specific ctDNA panels afford high specificity for HER2-amplified disease. CTC enumeration and characterization (EpCAM+/CK+/CD45− cells), by contrast, has been validated broadly in both early-stage and metastatic settings irrespective of molecular subtype, although phenotypic profiling (e.g. HER2 or PD-L1 on CTCs) can guide subtype-directed therapies. Exosomal biomarkers likewise span subtypes miR-21–5p and miR-1246 are elevated across luminal and triple-negative cohorts—but emerging panels (including lncRNA DANCR or protein cargo such as survivin) may yield enhanced accuracy in aggressive phenotypes such as triple-negative breast cancer. By delineating these subtype associations, clinicians can better tailor surveillance strategies to the patient's tumour biology.
Proteomic and metabolomic profiling are also increasing the number of biomarkers available for the diagnosis of recurrent BC. These technologies, including mass spectrometry and NMR spectroscopy, have been used to identify particular protein and metabolic patterns that are linked to recurrent BC.
55
However, there are some issues that come with the use of new biomarkers in the clinical setting. The biomarker expression levels may vary greatly between patients and tumors, and therefore, these biomarkers need to be properly validated in clinical studies to determine their effectiveness. Additionally, standardization of detection techniques, as well as incorporation of these biomarkers into clinical practice, is an important task.
56
Collectively, tumor biomarkers serve as critical tools across the continuum of clinical care, including early detection, diagnostic evaluation, prognostic assessment, and recurrence surveillance. Their integration into clinical practice holds significant potential to enhance patient outcomes by prolonging survival and improving quality of life. A comparative analysis of all these biomarkers is given in
A comparative analysis of non-invasive biomarkers for detection and monitoring of breast cancer.
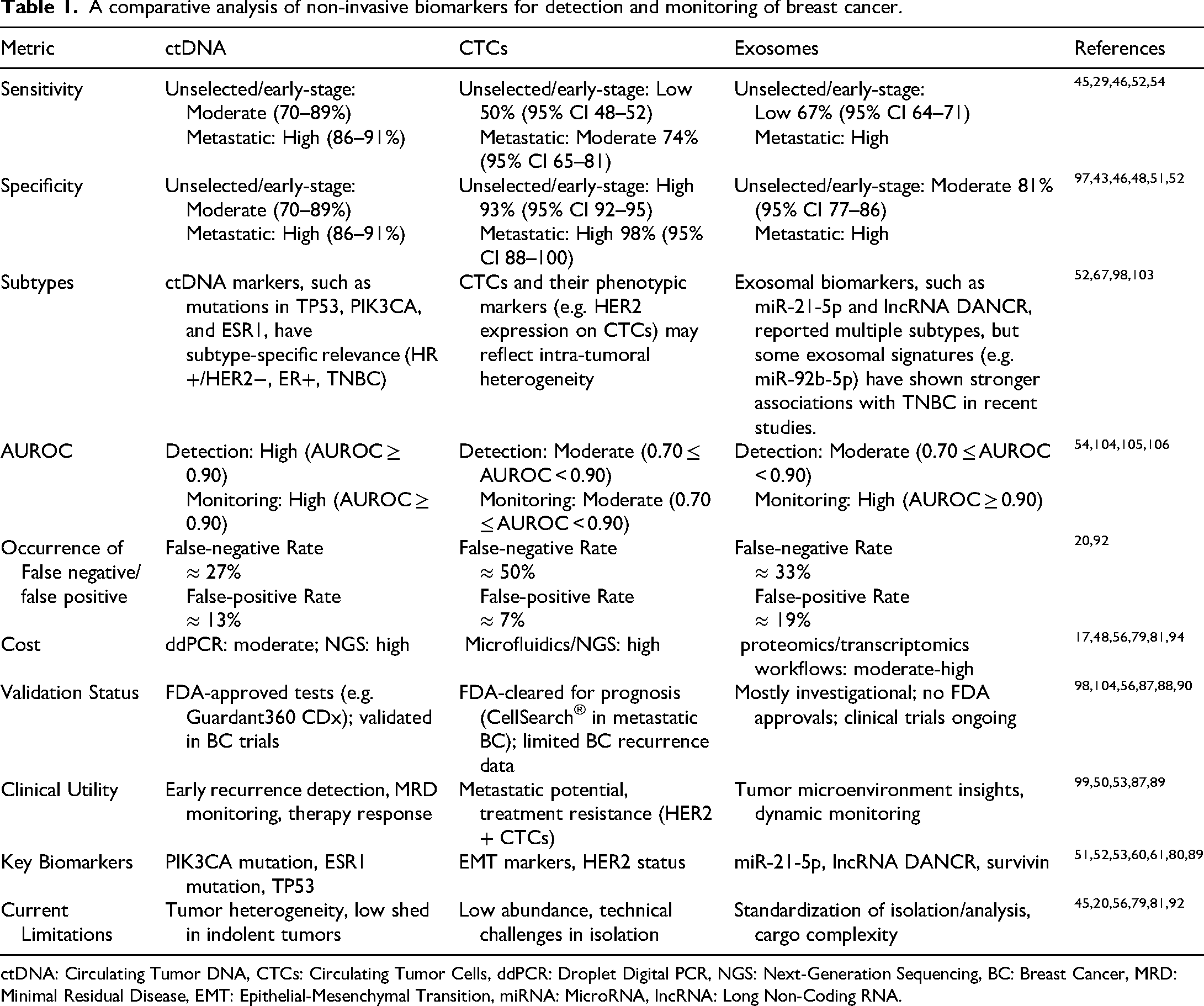
ctDNA: Circulating Tumor DNA, CTCs: Circulating Tumor Cells, ddPCR: Droplet Digital PCR, NGS: Next-Generation Sequencing, BC: Breast Cancer, MRD: Minimal Residual Disease, EMT: Epithelial-Mesenchymal Transition, miRNA: MicroRNA, lncRNA: Long Non-Coding RNA.
Role of multi-omics approaches in detecting breast cancer recurrence
Multi-omics approaches are a new concept of understanding and tracking the trend of BC recurrence, which offers a holistic understanding of the tumor behavior. 57 These approaches combine data from different omics platforms including genomics, transcriptomics, proteomics, and metabolomics to reflect the genetic and biochemical make up of the BC. 58 Such global profiling not only enhances the identification of recurrence but also opens new possibilities for the development of individualized monitoring strategies. 59
Genomics is the beginning of multi-omics methods that describe the alterations at the DNA level that are associated with the disease recurrence. For example, mutations in TP53, PIK3CA and BRCA1/2 are common in BC that has been used as indicators for early identification of the disease.60,61 NGS of DNA introduced rapid and cost-efficient way to identify genes involved in BC. 62 NGS enables simultaneous detection of numerous genetic alterations within a single assay, establishing the framework for “multigene sequencing” approaches. These methodologies have expanded the identification of genes implicated in BC susceptibility beyond previously characterized loci. 63 For instance, BARD1 (BRCA1-associated ring domain), which interacts directly with BRCA1, has emerged as a potential low-to-moderate penetrance gene linked to BC risk. 64 Studies report that loss-of-function (LoF) mutations in BARD1 occur in approximately 0.51% of BC patients. Furthermore, individuals with BARD1 mutations exhibit a notably younger average age at initial diagnosis (42.3 years; range 24–60 years) compared to the broader cohort (48.6 years; range 17–92 years). 65 Similarly, germline LoF mutations in BRIP1 (BRCA1-interacting protein C-terminal helicase 1), another gene with low penetrance, have been identified as contributors to BC risk, particularly in early-onset cases. 66
Transcriptomics is a follow up of the genomic data analysis and focuses on the level of gene expression, both the messenger and the non-coding RNAs. Certain RNA signatures, for example, miR-21 and miR-155, have been associated with BC progression and recurrence. Sinicropi et al. analyzed a cohort of 136 patients and identified over 2000 RNAs with significant associations to BC recurrence. Intriguingly, a substantial subset of these RNAs were intronic, with their corresponding exonic regions showing no discernible link to relapse. These RNA signatures shed light on molecular mechanisms underlying disease recurrence and could serve as potential targets for therapeutic strategies aimed at mitigating relapse.67,68
Proteomics and metabolomics are two layers of multi-omics that provide a functional context to the data and focusing on the changes in the tumor environment.
69
Through advances in mass spectrometry, specific protein markers associated with the recurrence of BC have been identified; these involve proteins regulating immune suppression and angiogenesis.70,71 The metabolome represents a dynamic biochemical profile, where shifts in metabolite concentrations or ratios can signal pathological states, revealing dysregulated metabolic pathways characteristic of complex diseases such as BC.
59
Metabolomic studies in BC commonly analyze biofluids (urine, serum, plasma, saliva) or tissue samples. Since metabolites serve as terminal products of cellular processes, their levels mirror the integrated response of biological networks, making them potential biomarkers for diagnostics and therapeutic monitoring.
72
Achieving comprehensive metabolic profiling typically requires integrating multiple analytical platforms. Current methodologies include GC-MS, LC-MS, and NMR spectroscopy.
73
While MS-based techniques often provide superior sensitivity, NMR spectroscopy enables non-destructive analysis of intact tissues and even in vivo metabolic observations, a capability termed magnetic resonance spectroscopy (MRS) in clinical settings. Metabolomic profiling highlights dysregulation in key metabolic pathways, such as upregulated glycolysis - a hallmark of aggressive tumor phenotypes. Additionally, metabolomics research has identified perturbations in protein and lipid metabolism as recurring themes in cancer biology, reflecting the systemic metabolic rewiring that supports malignant progression..
74
These functional understandings are very useful for the development of new methods of monitoring and treating the disease (

Multi-Omics approaches for detecting breast cancer recurrence. This figure outlines multi-omics strategies for identifying biomarkers associated with breast cancer recurrence. The genomics panel highlights DNA-level mutations, including BRCA1/2 (low to moderate risk), loss-of-function (LoF) mutations in BRAD1 (common in younger patients), and mutations in BRIP1, PIK3CA, and TP53, detected using next-generation sequencing (NGS). The transcriptomics section focuses on gene expression markers like miR-21 and miR-155, which are strongly linked to breast cancer progression and recurrence. Proteomics emphasizes the functional roles of protein markers in immune suppression and angiogenesis, with mass spectrometry serving as a key detection tool. Metabolomics investigates functional and tumor microenvironment changes through metabolite levels in urine, plasma, saliva, or tissue, employing sensitive techniques like GC-MS, LC-MS, and NMR spectroscopy. Together, these integrative approaches enable comprehensive detection and monitoring of breast cancer recurrence.
Overall, multi-omics approaches are valuable for the surveillance of BC recurrence. These approaches offer a holistic view of the tumor using genomic, transcriptomic, proteomic, and metabolomic data and thus enable early identification and more tailored approaches to management.
Comparative insights: metabolomic versus molecular markers
Metabolomic markers and molecular markers each offer distinct advantages for monitoring recurrence and treatment response. Molecular assays such as ctDNA and CTCs track tumor-specific genetic or cellular features, providing high specificity and reproducibility once a sufficient tumor burden sheds detectable material into the circulation. 45 However, these markers often require a threshold of tumor turnover before becoming positive, potentially delaying early detection or real-time treatment assessment.
By contrast, metabolomic profiling measures dynamic changes in small-molecule metabolites that reflect tumor physiology and microenvironmental shifts. Because metabolic adaptations occur rapidly in response to cellular stress or therapy, metabolomic markers can reveal emerging recurrence or treatment efficacy earlier than molecular assays.59,72 That said, metabolite levels are also influenced by non–tumor factors (diet, co-morbidities), and require stringent biofluid handling and analytical standardization to ensure reliable comparisons.
In practice, integrating both modalities can harness their complementary strengths: molecular assays confirm tumor-specific events with high specificity, while metabolomic readouts deliver sensitive, early warnings of biochemical relapse or response. Emerging workflows that combine serial metabolomic panels with molecular marker kinetics have demonstrated improvements in lead-time to detection and overall diagnostic accuracy, supporting a truly personalized, dynamic surveillance strategy. 75
Advances in biomarker detection technologies
Recent progress in the field of BC recurrence detection has enhanced the sensitivity of detection methods and technologies. These technological advancements enable the quantification and identification of biomarkers including exosomes, CTCs and ctDNA to the highest level of precision and address the limitations of traditional monitoring strategies and enable clinicians to detect recurrence in its early stages and improve patient care.
Among the most significant improvements are ddPCR and NGS for detecting ctDNA. Both technologies have high sensitivity and are able to detect small amounts of tumor-specific DNA fragments in the blood.17,76 ddPCR is especially useful because of its low cost and short time to result, making it suitable for use in a clinical setting. 77 On the other hand, NGS offers a broad view of the mutational spectrum of the tumor, and is thus able to detect many genetic alterations at once. 78 Thus, the two features of high sensitivity and comprehensive profiling make NGS the standard technique of precision oncology, especially for recurrence monitoring.
Another advancement involves the development of microfluidic devices that can sort and analyze CTCs, which are rare in the blood. 79 These platforms sort CTCs based on size, immune affinity or density. After being captured, single cell sequencing and phenotypic analysis of these cells can give important information on the metastatic potential and the molecular characteristics of these cells. For instance, changes in the level of EMT markers on CTCs have been associated with breast cancer recurrence and can be used as a prognostic factor as well as a detection method. 80
New materials and instruments based on nanotechnology are also expanding the frontiers of the field, particularly with regard to the detection and analysis of exosomes. These methods have enhanced sensitivity of exosome isolation and analysis using nanoparticle based sensors and bioengineered platforms. 81 Interestingly, recurrence monitoring using wearing devices and point of care technologies is also an exciting new area in cancer diagnosis.82,83 These devices seek to reflect tumor dynamics in real time through the integration of biomarker detection with sophisticated data analytics. For example, wearable sensors that can detect ctDNA or exosomal RNA in interstitial fluid are being developed, with the aim of bringing recurrence monitoring out of the hospital and into daily life.84,85 Although, the sensitivity, specificity and usefulness of detecting BC recurrence have been improved through the application of new technologies for detection, however, there are still issues that need to be worked on in order to adopt these technologies in the clinical practice. The cost and the complexity of the methods employed, such as NGS and microfluidics, limit their availability in resource constrained settings. Furthermore, the detection methods need to be standardized across different populations of patients and these tools need to be validated for their reliability and repeatability.
Clinical applications of non-invasive biomarkers for monitoring breast cancer recurrence
The application of biomarkers for the purpose of monitoring the recurrence of BC has already revealed its effectiveness in improving patient outcomes. Thus, they help in the early identification of the disease, better prognosis, and individualized therapy. Many of them have been tested in clinical practice through case studies and clinical trials. The use of liquid biopsy and other technologies such as ctDNA analysis has become popular in identifying recurrence of cancer even before it becomes clinically detectable. 86 In one of the studies, the authors found that the ctDNA was detectable for months before the recurrence was visible on the radiographic examination in patients who were undergoing the routine ctDNA monitoring. 87 Such findings are useful in predicting the behavior of ctDNA and its ability to predict the necessity for early therapeutic measures. Similarly, increased CTC counts during follow-up have been linked to an increased risk of metastatic BC recurrence and therefore more frequent surveillance and earlier treatment.2,88 In addition to detection, the phenotypic characterization of CTCs has revealed information about the mechanisms of treatment failure, including the development of HER2-positive CTCs in HER2-negative tumors. 89 This information has directly contributed to changes in treatment plans and has identified the role of CTCs in personalized medicine. Moreover, exosomal biomarkers have been explored for their ability to predict the recurrence of BC and assess the efficacy of treatments. At one point, exosomal RNA levels were monitored in a patient undergoing adjuvant therapy, and changes in the levels of certain miRNAs associated with recurrence led to further investigation and early detection and management of metastatic disease. 90 Therapeutic changes based on ctDNA have been shown to result in better progression-free survival than conventional approaches. 91 Likewise, CTC enumeration as a stratification helped in identifying the most appropriate treatments that patients should receive based on their risk profiles. Such examples are representative of how exosomes can offer real-time information on tumor behavior and therapy efficiency.
The clinical translation of non-invasive biomarkers is exemplified by many ongoing and completed clinical trials. The phase II DARE trial (NCT04567420) investigates ctDNA-guided adjuvant therapy escalation/de-escalation in 100 early-stage breast cancer patients. Similarly, the c-TRAK TN trial (NCT03145961) employs ctDNA testing to direct therapy in 208 post-surgical triple-negative BC patients, aiming to validate ctDNA's utility in high-risk cohorts. Notably, the TRAK-ER trial (NCT04985266) expands this approach to 1100 patients, using ctDNA assays to stratify relapse risk and personalize surveillance. For metastatic BC, the phase III NCT01710605 trial leverages CTC counts to guide therapy in 800 patients. NCT06016790 explores cfDNA multi-omics for early BC detection, while PORTRAIT (NCT05916755) integrates multi-omics profiling in 100 early triple-negative BC patients to identify predictive biomarkers of neoadjuvant chemotherapy and immunotherapy response. These efforts reflect a growing emphasis on large-scale validation, risk stratification, and multi-modal biomarker integration to address tumor heterogeneity and improve clinical utility (
Clinical trials highlighting translational progress in non-invasive biomarkers.

ctDNA: Circulating Tumor DNA, CTCs: Circulating Tumor Cells, cfDNA: Cell-free DNA, BC: Breast Cancer, NACT: Neoadjuvant Chemotherapy, ICI: Immune Checkpoint Inhibitor, eTNBC: early Triple-Negative Breast Cancer.
Currently, the FDA-approved non-invasive biomarkers focuses primarily on tests analyzing ctDNA and CTCs. Guardant360 CDx is a comprehensive liquid biopsy test that utilizes NGS to analyze ctDNA. It is approved for use in all solid tumors, including breast cancer, and helps to identify actionable mutations for guiding treatment decisions, particularly in targeted therapies. FoundationOne Liquid CDx also utilizes NGS technology to assess over 300 genes plus genomic signatures such as microsatellite instability (MSI) and tumor mutational burden (TMB) from ctDNA in the bloodstream. CellSearch is the first and only FDA-cleared test for CTCs in patients with metastatic breast, prostate, and colorectal cancer. It's a vital tool for prognostication and monitoring the effectiveness of treatments.
Despite their promise, non-invasive biomarkers face limitations that temper their current clinical utility. ctDNA detection remains challenging in tumors with low DNA shedding or indolent biology, leading to false negatives in early-stage or oligometastatic disease.20,92 Similarly, CTCs are exceedingly rare in circulation, particularly in non-metastatic settings, and their isolation relies on technically demanding platforms that are not yet standardized across institutions.45,79 Exosomal biomarkers, while rich in tumor-derived cargo, suffer from heterogeneity in isolation methods (e.g. ultracentrifugation vs. commercial kits) and a lack of consensus on diagnostic thresholds for recurrence.56,81 Additionally, cost and accessibility barriers persist, as technologies like NGS and microfluidics remain unaffordable in resource-limited settings, exacerbating disparities in recurrence monitoring. These limitations highlight the importance of ongoing research to refine detection protocols, validate biomarkers in diverse populations, and integrate multi-modal strategies to mitigate individual biomarker shortcomings.
Challenges and limitations of non-invasive biomarkers
Non-invasive biomarkers such as ctDNA, CTCs), and EVs hold great promise for early detection and monitoring of breast cancer recurrence. However, each modality faces distinct technical and biological hurdles that limit its current clinical utility.37,93
ctDNA is often present at very low allele fractions typically below 0.1% of total cell-free DNA in early-stage disease, complicating reliable detection. 94 In a comprehensive meta-analysis of operable breast cancer, sensitivity of ctDNA assays ranged widely from 31% to 100%, with specificity between 70% and 100%, and an average lead-time of about 10.8 months before clinical relapse; this heterogeneity reflects differences in assay design, patient cohorts, and sampling timepoints. 93 Ultra-sensitive personalized assays such as the Invitae PCM platform demonstrated 76.9% sensitivity (10/13 relapsed patients) and 100% specificity in a cohort of 61 high-risk patients, with a median lead-time of 11.7 months. 95 Serial postoperative ctDNA monitoring with the Signatera assay achieved 88.2% sensitivity (30/34 patients) for predicting relapse up to 38 months before radiologic detection, but also revealed false positives in 5/122 nonrelapsing patients underscoring challenges of clonal hematopoiesis and assay noise. 96
Circulating tumor cells offer a cellular complement to ctDNA but suffer from extreme rarity on the order of 1–10 cells per mL of blood and phenotypic heterogeneity that hampers enrichment and detection. 97 Meta-analysis of 16 studies found pooled sensitivity of 50.0% (95% CI 48.0–52.0%) and specificity of 93.0% (95% CI 92.0–95.0%) for CTCs in breast cancer diagnosis, with significant heterogeneity across detection platforms. 98 Use of a microfluidic CytoSorter system improved performance, achieving 76.56% sensitivity and 95.4% specificity in a recent cohort study. 99 However, epithelial-to-mesenchymal transition can downregulate EpCAM and cytokeratin markers used for CTC capture, leading to false negatives, and variations in enrichment methods contribute further to inconsistent results. 97
Exosomal biomarkers encapsulated in EVs combine lipid-protected stability with the potential to carry proteins, miRNAs, and other cargo reflective of tumor biology. Yet, EV isolation is hampered by co-isolation of non-vesicular particles, lack of standardization, and variable yield and purity across ultracentrifugation, precipitation, and affinity-based techniques. 100 A meta-analysis of 3423 patients identified 56 EV-based biomarkers associated with overall survival, disease-free survival, and metastatic progression, yielding a pooled AUC of 0.91 (95% CI 0.86–0.95) for predicting metastasis but with high inter-study heterogeneity (I²=76.8%) and occasional reports of AUC = 1.00 in small cohorts that require independent validation. 101 In triple-negative breast cancer, an EV-miRNA panel (miR-142–5p, miR-320a, miR-4433b-5p) achieved 93.33% sensitivity and 68.75% specificity in distinguishing patients from healthy controls, but reproducibility and clinical impact remain to be confirmed in larger, prospective trials. 102
Overall, while ctDNA, CTCs, and EVs each offer unique insights into tumor dynamics, their clinical implementation is constrained by low analyte abundance, assay variability, and lack of harmonized protocols. Future efforts should focus on multi-analyte integration, standardized pre-analytical workflows, rigorous validation in large cohorts, and comparative studies to define the optimal biomarker or combination thereof for guiding personalized surveillance and early intervention.
Conclusion
Biomarkers offer crucial information regarding the prognosis and treatment response of BC patients. Non-invasive biomarkers, in particular, present a promising alternative to conventional biomarkers, especially for monitoring disease recurrence or progression, assessing treatment response, and identifying targetable mutations to guide therapy. The integration of non-invasive biomarkers in the surveillance of BC recurrence is a new approach in oncology. These biomarkers are more sensitive and can give real-time information on tumor behavior compared to the conventional methods. Recent improvements in detection technologies, including ddPCR, NGS, and microfluidic platforms have also added value these to tools and make them useful for personalized and dynamic monitoring of recurrence. However, the use of non-invasive biomarkers has problems that need to be solved immediately. The problems include; biological variability, technical limitations, cost, and systemic barriers such as access disparities which need to be solved through research and development and professional collaborations. Additionally, strong validation studies and standardization are required to establish the credibility and applicability of these technologies in various populations and healthcare systems.
Critical opinion and future directions
While biomarkers hold significant promise for advancing breast cancer (BC) diagnosis, further research in biomarker discovery and development is necessary for their integration into routine clinical practice for BC. The use of new biomarkers for early detection is primarily hindered by the low signal levels emitted by early-stage BC, which directly impact the sensitivity of sequencing techniques. To improve sensitivity, ultra-deep sequencing methods need to be developed. Additionally, size selection of biomarker fragments and the use of single-strand DNA libraries for NGS represent effective strategies. To address false positives during the sequencing of multiple alterations with NGS, error-reduction approaches such as molecular barcodes and bioinformatic data analysis pipelines can be employed, reducing the risk of errors during library preparation and sequencing. In addition to refining analytical techniques, optimizing key pre-analytical processes such as collection, centrifugation, preservation, and extraction protocols for liquid biopsy biomarkers like ctDNA and CTCs will be crucial for ensuring biomarker quality. With the advancement of machine learning, it will be possible to harness more signals and integrate multiple parameters, further enhancing the discovery and detection of BC-specific molecular signatures.
Footnotes
Acknowledgements
To enhance the clarity, coherence, and technical precision of this manuscript, AI-assisted tools were employed during the drafting and revision process. These tools were utilized to support language refinement and writing flow. All scientific concepts, analyses, and interpretations remain the sole responsibility of the authors.
Author contributions
Conceptualization: MET,SAR, YET; Literature search: SAR, SMS; Visualization: RB; Writing - Original draft: MET, SAR, SMS, YET RB; Writing - Review & Editing: MET, SMS, YET; All authors read and approved the final manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/ or publication of this article.
Data availability statement
This review is based entirely on previously published studies and publicly available data. No new data were generated or analyzed in this study.
