Abstract
Objective
Type 1 myocardial infarction (T1MI) is primarily caused by the formation of coronary thrombi, which leads to acute myocardial ischemia and hypoxia and is associated with high morbidity and mortality rates. However, the effects of thrombus-derived exosomes (TEs) on endothelial cell function remain unclear. The aim of this study was to investigate the interaction between lncRNA LOC101928697, which is enriched in TEs, and FUS proteins, as well as their impact on endothelial cell function.
Methods
Thrombus-derived exosomes were extracted from thrombi collected from patients with T1MI, and lncRNAs with differential expression were identified through microarray analysis. Bioinformatic analyses were used to predict the interaction between LOC101928697 and FUS protein. The effects of TEs and LOC101928697 on human umbilical vein endothelial cells (HUVECs) were evaluated using CCK8 assays, scratch assays, cell cycle analysis, fluorescence in situ hybridization, and RT-PCR. The role of LOC101928697 was further confirmed by knockdown experiments.
Results
Our study indicates that TEs exert a significant inhibitory effect on the proliferation and migration of HUVECs, and that this effect is mediated by LOC101928697. LOC101928697 is highly enriched in TEs and has been shown to regulate the expression levels of CyclinD1 and PCNA by binding to FUS proteins, thereby inhibiting the proliferative capacity of endothelial cells. Furthermore, the inhibition of HUVEC proliferation and migration induced by TEs was significantly ameliorated by knockdown of LOC101928697.
Conclusion
This study provides new insights into the mechanism by which TEs suppress endothelial cell function via the LOC101928697-FUS axis. These findings highlight a molecular target that may contribute to the progression of T1MI and provide a foundation for future research into diagnostic and therapeutic strategies for cardiovascular diseases.
Keywords
Introduction
Type 1 myocardial infarction (T1MI), one of the most fatal cardiovascular diseases worldwide, is characterized by the formation of intracoronary thrombi as its primary pathological mechanism. 1 This obstruction of myocardial blood flow leads to ischemia and necrosis, frequently resulting in severe complications such as heart failure and arrhythmias. 2 Although significant progress has been made in treatments such as thrombolysis, interventional procedures, and coronary artery bypass surgery, the high morbidity and mortality rates associated with T1MI remain a global public health concern. 3 Currently, there is a lack of effective early intervention strategies for intracoronary thrombosis, particularly with regard to improving vascular injury and remodeling after T1MI. More importantly, enhancing endothelial cell function remains a central challenge in treatment, necessitating urgent research breakthroughs.
Exosomes, as critical mediators of intercellular communication, have emerged as a promising area of research in cardiovascular disease. 4 These small vesicles, released by cells, contain a diverse array of biological molecules, including proteins, lipids, and nucleic acids, which enable them to modulate the activity of recipient cells through direct interaction. 5 Previous studies from our group have shown that thrombi contain a significant number of thrombus-derived exosomes (TEs), which can profoundly influence the functional regulation of cells adjacent to thrombi, thereby playing a key role in vascular health and the progression of cardiovascular diseases. 6 However, the precise mechanisms by which TEs contribute to cardiovascular pathology remain poorly understood, especially regarding their role in modulating endothelial cell function, which is still an area of ongoing exploration.
Non-coding RNAs, particularly long non-coding RNAs (lncRNAs), have been recognized as critical players in the progression of cardiovascular diseases. 7 Although they lack protein-coding capacity, lncRNAs play essential roles in processes such as gene regulation, cell proliferation, migration, and apoptosis. 8 Numerous studies have demonstrated that specific lncRNAs are closely associated with the pathogenesis of conditions such as myocardial infarction, 9 cardiomyopathy, 7 and heart failure, 10 thereby positioning them as potential therapeutic targets. 11 In the context of thrombosis, lncRNAs are believed to modulate thrombus stability by regulating the expression of associated proteins. 12 Furthermore, a growing body of research suggests that lncRNAs from specific sources are actively involved in the biological functions of endothelial cells. 13 The hypothesis that particular lncRNAs may be enriched in TEs provides an intriguing avenue for further investigation and opens up new research directions.
Previous research has revealed that TEs may participate in both thrombus development and consolidation through their inhibitory effects on endothelial cell proliferation and migration. These extracellular vesicles may also be involved in the processes of vascular endothelial damage that occur after T1MI. 6 Using gene chip analysis, we found that lncRNA LOC101928697 is significantly enriched in TEs. Our hypothesis is that this lncRNA influences endothelial cell behavior through its interaction with FUS proteins within these cells. Specifically, we propose that TEs affect endothelial cell growth and migration by means of the binding of their abundant lncRNA LOC101928697 to FUS proteins, thereby serving as a critical factor in the pathogenesis of T1MI. This research aims to clarify the relationship between lncRNA LOC101928697 and FUS proteins within TEs and to explore its functional significance in vascular endothelial cell damage induced by acute oxygen deprivation.
Materials and methods
Cell culture and TEs extraction
All procedures involving human participants were conducted in accordance with the ethical standards of the institutional and/or national research committee and with the 1975 Declaration of Helsinki as revised in 2024. All clinical samples in this study were obtained from Guizhou Provincial People's Hospital. All patients were recruited and enrolled with informed consent as described above. The recruitment period was from August 2023 to August 2024.
Coronary thrombi were obtained by thrombus aspiration from patients with T1MI (n = 4) at Guizhou Provincial People's Hospital who underwent emergency PCI. The thrombi were lysed using our previously established protocol (trypsin 20 μg/ml plus collagenase 50 U/ml), and exosomes were isolated from the thrombi by ultracentrifugation. 6 For the control group, exosomes were extracted from the serum of healthy volunteers (n = 3) and purified in the same manner. Human umbilical vein endothelial cells (HUVECs; ATCC, Manassas, VA, USA; RRID:CVCL_2959) were obtained from ATCC. The cells were cultured in RPMI-1640 medium (Gibco, Thermo Fisher Scientific, USA) supplemented with 10% fetal bovine serum (FBS, Gibco) under standard conditions (37°C, 5% CO₂).
Identification of TEs
In this study, exosomes were characterized based on three features: morphology, size, and specific markers. The morphology and size of exosomes were assessed by fixing the extracted exosomes onto a 200-mesh copper grid using 1% osmium tetroxide, followed by electron microscopy. Exosome particle size and concentration were measured using nanoparticle tracking analysis (NTA) at VivaCell Bioscience with the ZetaView PMX 110 system (Particle Metrix, Meerbusch, Germany) and the corresponding ZetaView 8.04.02 software. The exosomal markers CD63 and TSG101 were identified using western blot analysis. Cellular debris generated during exosome extraction was used as a negative control.
Co-culture of TEs with HUVEC
Based on published data, an exosome concentration of 128 ng/μl was selected for cell co-culture experiments. 6 Thrombus-derived exosomes were co-cultured with HUVECs for 72 h. As a positive control, acute hypoxia in HUVECs was induced using the liquid paraffin seal method (hypoxia for 2 h at 37°C). 6 The experimental design included three groups: a TEs-treated group (exosome group), a hypoxia-treated group (hypoxia group), and a normal untreated group (control group). Cell viability was assessed using the Cell Counting Kit-8 (CCK-8) assay. Additional assays, including the scratch assay (wound healing assay), flow cytometric analysis of the cell cycle, western blotting (WB), and reverse transcription quantitative polymerase chain reaction (RT-qPCR), were performed to evaluate cell proliferation, migration, and the expression levels of relevant proteins.
RT-qPCR analysis
Total RNA was extracted from cells using Trizol reagent (Takara, Japan) according to the manufacturer's protocol. cDNA was synthesized using the PrimeScript™ RT Reagent Kit (Takara, Japan). RT-qPCR was performed with the SYBR® Premix Ex Taq™ II kit (Takara, Japan) on a CFX96 Real-Time PCR Detection System (Bio-Rad, USA). All primers were synthesized by Sangon Biotech (Shanghai, China). The primer sequences used were as follows:
LOC101928697: Forward: 5′-GGGAAGGCAGTGGGGTCAGA-3′; Reverse: 5′-GACCGGTTCCTCTGGAGGTG-3′.
FUS: Forward: 5′-GGGAGTGGAGAGGAGGAGAA-3′; Reverse: 5′-TGAGAGCAGCAGAGCAGGAT-3′.
CyclinD1: Forward: 5′-GCTGCGAAGTGGAAACCATC-3′; Reverse: 5′-CCTCCTTCTGCACACATTTGAA-3′.
PCNA: Forward: 5′-CTGAAGGGACCTTAGACTGGAC-3′; Reverse: 5′-CAGTCTAGCTGCACCAAAGG-3′.
GAPDH: Forward: 5′-GAAGGTGAAGGTCGGAGTCA-3′; Reverse: 5′-GAAGATGGTGATGGGATTTC-3′.
The qPCR cycling conditions were: 95°C for 30 s, followed by 40 cycles of 95°C for 5 s and 60°C for 30 s. Each sample was analyzed in triplicate. Relative gene expression was calculated using the 2–ΔΔCt method and normalized to GAPDH.
Wound healing assay
Human umbilical vein endothelial cells were seeded in six-well plates and cultured until reaching 90–100% confluence. A sterile 200-μl pipette tip was used to create a scratch in the cell monolayer. After washing with PBS to remove detached cells, the cells were treated under the designated experimental conditions. Images of the wound area were captured at 0 h and 24 h using an inverted microscope. The wound healing rate (%) was calculated using ImageJ software based on the reduction in scratch width over time.
Overexpression and knockdown of lncRNA LOC101928697
The expression of lncRNA LOC101928697 was silenced using small interfering RNA (siRNA) technology, and an overexpression model was established using an overexpression plasmid in HUVECs. The effects of LOC101928697 on endothelial cell function were evaluated using the wound healing assay, cell cycle assay, western blotting (WB), and RT-qPCR.
Western blot (WB)
A total of 20 µg of exosomal protein was separated on 6–12% SDS-PAGE gels and transferred to PVDF membranes. After blocking with Protein Free Rapid Block buffer, the membranes were incubated with primary antibodies (FUS: Proteintech, Cat No. 11570-1-AP, 1:5000; PCNA: Proteintech, Cat No. 10205-2-AP, 1:5000; CyclinD1: Proteintech, Cat No. 60186-1-Ig, 1:5000; GAPDH: Proteintech, Cat No. 60004-1-Ig, 1:30000) overnight at 4°C. The following day, after three washes with TBST, the membranes were incubated with HRP-linked anti-rabbit IgG secondary antibodies for 2 h at room temperature. Protein bands were visualized using an enhanced chemiluminescence system (P36599; Millipore, Boston, MA, USA), and band intensity was quantified using ImageJ software (Bethesda).
Exosome electron microscopy
Isolated exosomes were reconstituted in 10–20 μl PBS, and a droplet was applied to glow-discharged 300-mesh copper grids coated with heavy-duty carbon-coated formvar (ProSciTech, Kirwan, QLD, Australia) for 5 min. Excess liquid was removed with filter paper (Whatman, Maidstone, UK). The grids were rinsed twice with Milli-Q water and then negatively stained with 2.5% uranyl acetate solution. All prepared samples were visualized using an H-7650 electron microscope operating at 100 kV.
Nanoparticle tracking analysis (NTA)
Exosomal particle size and concentration were determined using nanoparticle tracking analysis (NTA) performed at VivaCell Bioscience with the ZetaView PMX 110 system (Particle Metrix, Meerbusch, Germany) and ZetaView 8.04.02 software. The purified exosome preparations were appropriately diluted in 1× PBS buffer (Biological Industries, Israel) before analysis. Measurements were collected and evaluated at 11 different positions. System calibration was performed using 110 nm polystyrene reference particles, and the ambient temperature was maintained between 23°C and 30°C throughout the analysis.
Flow cytometric analysis
Cells (2 × 10^5) were stained with DCHF-DA antibodies during a 1-h incubation at 4°C under light-protected conditions. Following two PBS washing steps, the labeled cell populations underwent analysis via flow cytometry (Beckman Coulter, Fullerton, CA, USA).
RNA pull-down assay
To investigate the interaction between LOC101928697 and FUS protein, an RNA pull-down assay was performed. The full-length sequence of LOC101928697 (NC_000013.11:19182033-19187465) was amplified by PCR with a T7 promoter sequence added to the 5′ end of the template. In vitro transcription was carried out using the MEGAscript™ T7 Transcription Kit (Thermo Fisher Scientific, Cat# AM1334). Biotin-labeled RNA was synthesized using the Biotin RNA Labeling Mix (Roche, Cat# 11685597910). The biotinylated RNA was incubated with Dynabeads™ M-280 Streptavidin magnetic beads (Thermo Fisher Scientific, Cat# 11205D) at room temperature. Human cell lysates were prepared using IP lysis buffer (Beyotime, Cat# P0013), pre-cleared, and incubated with the RNA-coated beads at 4°C for 2–4 h with rotation. After extensive washing, the bound proteins were eluted, separated by SDS-PAGE, and analyzed by Western blot using anti-FUS antibody (Proteintech, Cat No. 11570-1-AP, dilution 1:5000) to detect the enrichment of FUS protein. Antisense RNA pull-down and input controls were included in parallel to confirm specificity and efficiency.
Primer sequences used for in vitro transcription of full-length LOC101928697 RNA:
Forward (with T7 promoter): 5′-TAATACGACTCACTATAGGGGGGAAGGCAGTGGGGTCAGA-3′
Reverse: 5′-AGGCCTTTGAGGCCGCATGT-3′
Bioinformatics analysis
Differentially expressed lncRNAs were identified using the limma package (version 3.52.2, Bioconductor, https://bioconductor.org/packages/limma/) in R software (version 4.2.2, R Foundation for Statistical Computing, Vienna, Austria; https://www.r-project.org/). Genes were considered differentially expressed if they had an adjusted p-value (FDR) < 0.05 and |log2 fold change| > 1. Annotation and interaction predictions were performed using AnnoLnc2 database (http://annolnc.gao-lab.org/), 14 the RBPDB database (http://rbpdb.ccbr.utoronto.ca/), 15 RPISeq database (http://pridb.gdcb.iastate.edu/RPISeq/), 16 and catRAPID website (http://service.tartaglialab.com/page/catrapid_group) 17 to predict the interaction between lncRNA LOC101928697 and FUS proteins. In addition, the public dataset GSE163772 was used to examine the expression of FUS genes in endothelial cells from a myocardial infarction model. This dataset was derived from endothelial cells isolated from a mouse model of myocardial infarction (n = 6). All data were subjected to FDR correction, with p-FDR < 0.05 considered statistically significant.
Fluorescence in situ hybridization experiment
A gene probe labeled with a red fluorescent marker was designed based on the promoter NR_126054.1 of lncRNA LOC101928697, with the sequence: GCTCACAGAGAGCTCCAAGAAG. The FUS protein was labeled with a green fluorescent marker, and the nuclei were stained blue using DAPI. The subcellular localization of lncRNA LOC101928697 and FUS protein was analyzed in HUVECs using fluorescence in situ hybridization (FISH) and immunofluorescence experiments.
Statistical analysis
All experimental data in this study were analyzed, and graphical representations were generated using SPSS 20.0 and GraphPad Prism 7. Independent t-tests were used for comparisons between two groups, while one-way analysis of variance was performed for comparisons among multiple groups. A p-value < 0.05 was considered statistically significant.
Results
Extraction and characterization of TEs
Building on our group's previous methodology, we successfully extracted vesicular material from thrombi. As shown in Figure 1(a) to (c), these vesicles exhibited the characteristic disc-shaped structure with a central depression (Figure 1(a)) and had particle sizes of approximately 100 nm (Figure 1(b)). Furthermore, exosomal markers CD63 and TSG101 were identified using western blot analysis. Our results showed positive expression of both CD63 and TSG101 in the extracted material (Figure 1(c)). These findings are consistent with the characteristic features of exosomes.4,18 Therefore, we conclude that we have successfully isolated TEs.

Extraction and characterization of thrombus-derived exosomes. (a) Electron microscopic observation of thrombogenic exosomes (at a magnification of approximately 80,000–100,000 times). The results show that the exosomes are within 200 nm in diameter and are disc-like structures with a concave middle. (b) Nanoparticle tracking analysis (NTA) was performed to observe the size distribution of exosomes. The NTA results showed that the exosome particle size was between 30 and 150 nm. (c) Western blot detection of exosome marker CD63 and TSG101 expression. Cell debris was used as a negative control group, and the results showed positive expression of CD63 and TSG101 in exosomes.
Effect of TEs on functional proliferation and migration of HUVEC
To evaluate the effects of TEs on endothelial cell function, TEs were co-cultured with HUVECs (exosome group) for 24, 48, and 72 h. An endothelial cell model of acute hypoxia, constructed using the liquid paraffin sealing method (37°C, 2 h), served as a positive control (hypoxia group). The results of the CCK8 assay (Figure 2(a)) showed that cell viability in the exosome group was significantly reduced compared with the control group at 72 h (p < 0.01), with no significant difference between the exosome group and the hypoxia group (p > 0.05). In the subsequent wound healing assay, as shown in Figure 2(b) and (c), TEs significantly inhibited HUVEC migration at 24 h of co-culture compared with the control group (p < 0.05). Similarly, cell migration in the hypoxia group was significantly lower than in the control group at 72 h of co-culture (p < 0.05).

Effect of thrombus-derived exosomes (TEs) on functional proliferation and migration of human umbilical vein endothelial cell (HUVEC). (a) CCK8 assay to detect the activity of each group of cells. (b) Wound healing assay to detect cell migration ability. (c) Statistical graph of the wound healing assay (4×). (d) Cell cycle test by flow cytometry. (e) Statistical graph of cell cycle assay, where proliferation index (PI) = (Number of cells in S phase + Number of cells in G2/M phase)/Total number of cells × 100%. (f) Western blot detection of cyclinD1 and PCNA, key proteins in the cell cycle. (g) Statistical graph of the ratio of cyclinD1 to GAPDH in the western blot experiment. (f) Statistical graph of PCNA to GAPDH ratio in western blot experiment. (i) Statistical graph of mRNA level expression of cyclinD1 in each group of cells detected by RT-qPCR. (j) Statistical graph of mRNA level expression of PCNA detected by RT-qPCR in each group of cells. ****p < 0.0001 compared to control group.
Consistent results were observed in cell cycle experiments (Figure 2(d) and (e)), where the proliferation index in the exosome group was significantly reduced compared with the control group (p < 0.01). To further investigate the effects of TEs, we examined the expression of proliferation-related proteins CyclinD1 and PCNA at both the protein and mRNA levels using western blotting (Figure 2(f) to (h)) and RT-qPCR (Figure 2(i) and (j)). Thrombus-derived exosomes consistently downregulated CyclinD1 and PCNA expression at both the protein and mRNA levels. Compared with the control group, the exosome group exhibited significantly decreased levels of CyclinD1 and PCNA (p < 0.01).
lncRNA microarray analysis of TEs
We performed lncRNA microarray analysis on TEs extracted from T1MI patients (n = 4) and serum-derived exosomes from healthy individuals (n = 3) to identify key effector genes present in thrombus exosomes. The results of the microarray analysis are presented in Figure 3(a) and (b). The heatmap is shown in Figure 3(a), and the principal component analysis is displayed in Figure 3(b). Figure 3(c) and (d) is volcano and hyperbolic volcano plots, respectively. While using identical thresholds, the hyperbolic (curved) volcano plot applies a more stringent, composite criterion for significance, requiring both a high absolute fold change and a low p-value. In contrast, the classic volcano plot marks as significant any genes meeting either threshold, resulting in many more hits. As a result, only our target gene (LOC101928697) falls outside the hyperbolic boundary, highlighting its outstanding change relative to all others. To validate the bioinformatics findings based on the lncRNA microarray, total RNA was extracted from the TEs group and serum-derived exosomes from healthy individuals (healthy serum exosome group), and the expression of lncRNA LOC101928697 was assessed using RT-PCR. The RT-PCR results (Figure 3(e)) demonstrated that LOC101928697 expression was significantly upregulated in the thrombus exosome group compared with the control group (p < 0.001). This result further confirms the reliability of our lncRNA microarray analysis.
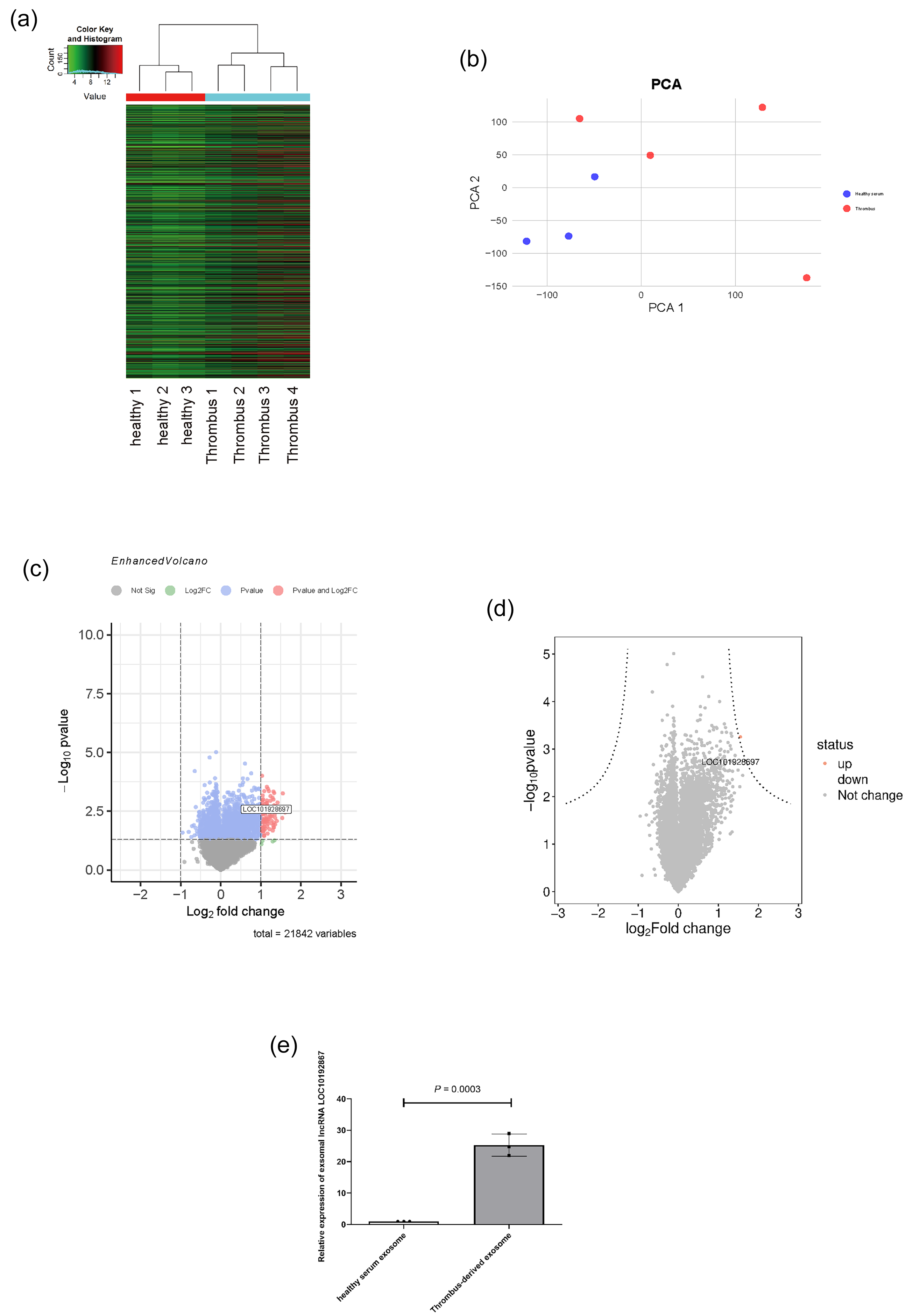
lncRNA microarray analysis of thrombus-derived exosomes (TEs). (a) Heatmap of lncRNA microarray analysis with colors indicating high or low levels of gene expression, red for high expression and green for low expression. (b) Principal Component Analysis plot. (c) Volcano plot. (d) Hyperbolic volcano plot. (e) RT-qPCR detection of lncRNA LOC101928697 expression in two group exosomes.
Effect of exosomal lncRNA LOC101928697 on HUVEC proliferation and migration
Thrombus-derived exosomes were co-cultured with HUVECs, and the expression of LOC101928697 in HUVECs was assessed by RT-qPCR. The RT-qPCR results (Figure 4(a)) showed that both the hypoxia group and the exosome group exhibited significant upregulation of LOC101928697 expression compared with the control group (p < 0.05). Notably, the exosome group showed significantly higher expression of LOC101928697 compared with the hypoxia group (p < 0.05).

Effect of exosomal lncRNA LOC101928697 on human umbilical vein endothelial cell (HUVEC) proliferation and migration. (a) RT-qPCR to detect the expression of lncRNA LOC101928697 in HUVEC cells of each group. (b) Cell models with inhibited and overexpressed lncRNA LOC101928697 were constructed, and the expression of lncRNA LOC101928697 in various cells was detected using RT-qPCR. (c) Wound healing assay to detect cell migration ability(4×).(d) Statistical graph of the wound healing assay. (e-g) Separate western blot experiments to detect cyclinD1 (f) and PCNA (g) protein-level expression. (h-i) are RT-qPCR experiments to detect the mRNA-level expression of cyclinD1 (h) and PCNA (i), respectively. (j) A cell cycle experiment, and (k) A statistical graph of PI values in a cell cycle experiment. ap < 0.05 compared to control group. bp < 0.05 compared to exosome group. cp < 0.05 compared to siRNA + exosome group.
Subsequently, LOC101928697 expression was inhibited using siRNA or enhanced through plasmid-based overexpression in HUVECs (Figure 4(b)). In the siRNA group, thrombus exosomes and siRNA-LOC101928697 were added to the cells simultaneously for co-culture. This experiment was divided into four groups: control group, exosome group, siRNA + exosome group (co-cultured with thrombus exosomes and treated with siRNA-LOC101928697), and overexpression group (overexpression of LOC101928697 only). The wound healing assay results (Figure 4(c) and (d)) demonstrated that inhibition of LOC101928697 expression significantly enhanced HUVEC migration compared with the exosome group (p < 0.05). Conversely, the overexpression of LOC101928697 significantly reduced HUVEC migration compared with the control group (p < 0.05). Western blot (Figure 4(e) to (g)) and RT-qPCR analyses (Figure 4(h) and (i)) showed that inhibition of LOC101928697 expression upregulated PCNA and CyclinD1 protein and mRNA levels compared with the exosome group (p < 0.05). Consistent with these findings, cell cycle analysis (Figure 4(j) and (k)) showed that inhibition of LOC101928697 expression significantly mitigated the exosome-induced reduction in cellular proliferation indices (p < 0.05). These results suggest that lncRNA LOC101928697 plays a major role in the inhibition of HUVEC proliferation and migration mediated by TEs.
Analysis of the correlation between FUS protein and myocardial infarction
Analysis of the lncRNA microarray data in this study revealed significant enrichment of lncRNA-associated genes in the RNA transport pathway (Figure 5(a)). Using the AnnoLnc2 database (http://annolnc.gao-lab.org/), 14 LOC101928697 was found to have only one CLIP-Seq-validated RNA-binding protein, FUS, which is widely recognized and studied as an RNA transport protein (Figure 5(b)). Further predictions using the RBPDP database (http://rbpdb.ccbr.utoronto.ca/) 15 confirmed a 100% correlation between LOC101928697 and FUS (Figure 5(c)).

Analysis of the correlation between FUS protein and myocardial infarction. (a) Enrichment Analysis Bar Plot based on differential gene expression profiles in lncRNA microarray analysis.(b) Detection information about lncRNA LOC101928697 binding to FUS proteins in AnnoLnc2 database. (c) Detection information about lncRNA LOC101928697 binding to FUS protein in RBPDP database. (d) Scores in the RPISeq database on the model of lncRNA LOC101928697 binding to FUS protein. (e-g) Prediction information about lncRNA LOC101928697 binding to FUS protein in catRAPID website, (e) Statistical map information about protein and RNA binding sites, (f) Total scoring information, and (g) Interaction map showing the interaction region between protein and RNA. (h-i) Analyses about bioinformatics techniques based on GSE163772 in the GEO database, where (h) is a statistical map of FUS gene expression in endothelial cells of a mouse model of myocardial infarction, and (i) A scatter plot about the correlation between the level of FUS gene expression and the disease state (control vs. myocardial infarction).
To validate the interaction between LOC101928697 and FUS, the RPISeq database (http://pridb.gdcb.iastate.edu/RPISeq/) was used, 16 showing that LOC101928697 directly binds to FUS protein, with prediction scores exceeding 0.5 in both RF classifier and SVM classifier models, indicating a positive interaction (Figure 5(d)). Similarly, predictions from the catRAPID website (http://service.tartaglialab.com/page/catrapid_group) 17 identified multiple potential interaction sites between LOC101928697 and FUS, with Z-scores >0 and Interaction Propensity scores >50 (Figure 5(e) to (g), Table 1, and Table 2).
Information on the interaction site of lncRNA LOC101928697 with FUS protein.
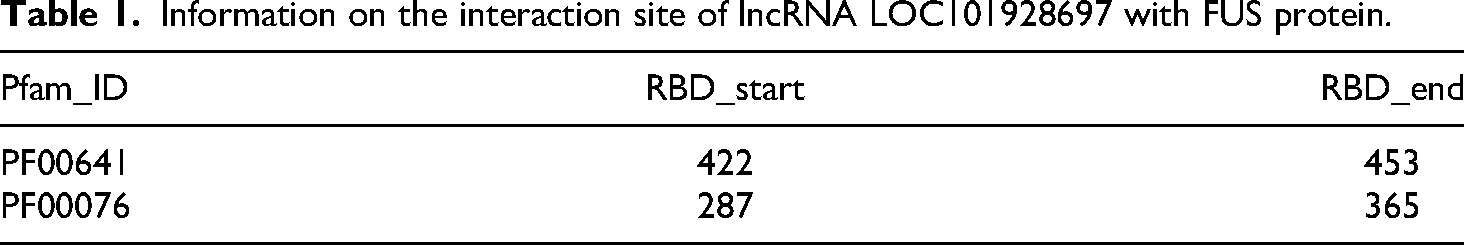

Information on the interaction site of FUS protein with lncRNA LOC101928697.

To further investigate the relationship between FUS and myocardial infarction, we performed a bioinformatics analysis using endothelial cell microarray data from a myocardial infarction mouse model in the Gene Expression Omnibus (GEO) database (GSE163772). The results (Figure 5(h) and (i)) revealed that FUS was highly expressed in endothelial cells from myocardial infarction samples compared with the control group (healthy individuals), with a strong correlation coefficient (R = 0.95).
Interaction of exosomal lncRNA LOC101928697 with FUS proteins
To further investigate the association between LOC101928697 and FUS protein in endothelial cells under acute hypoxia, western blot (Figure 6(a) and (b)) and RT-qPCR (Figure 6(c)) were used to measure FUS expression at the protein and mRNA levels. The results indicated that acute hypoxia significantly upregulated FUS mRNA expression in HUVECs compared with the control group (p < 0.05), while the protein level showed no significant change (p > 0.05). Additionally, thrombus exosomes significantly promoted both the mRNA and protein expression of FUS in HUVECs compared with the control group (p < 0.05). To explore the spatial relationship between LOC101928697 and FUS, we designed red fluorescence-labeled probes based on the promoter NR_126054.1 of LOC101928697 and performed FISH. As shown in Figure 6(d), the green fluorescence-labeled FUS protein and the red fluorescence-labeled LOC101928697 probe colocalized predominantly in the nuclear region, suggesting a direct interaction between the two. To verify the direct interaction between lncRNA LOC101928697 and FUS protein, an RNA pull-down assay combined with Western blot analysis was performed. As shown in Figure 6(e) and (f), the sense RNA pull-down group effectively captured and enriched FUS protein compared with the antisense RNA pull-down control. Densitometric analysis further demonstrated that the enrichment of FUS protein in the sense RNA pull-down group was significantly higher than that observed in the antisense group (p < 0.05). These findings indicate that lncRNA LOC101928697 is capable of specifically binding to FUS protein.

Interaction of exosomal lncRNA LOC101928697 with FUS proteins. (a and b) The western blot detection of FUS protein expression in each group of cells and the statistical graph. (c) Statistical graph of RT-qPCR to detect the expression of FUS at the mRNA level in each group of cells. (d) The fluorescence graph of fluorescence in situ hybridization (FISH) experiment. In which FUS was labeled with green fluorescence, lncRNA LOC101928697 was labeled with red fluorescence, and the nucleus was labeled with blue fluorescence (20×). (e) Western blot detection of FUS protein following RNA pull-down using sense or antisense LOC101928697 transcripts. (f) Quantification of FUS protein enrichment in sense RNA pull-down versus antisense control, based on densitometric analysis. (g-h) Western blot detection of FUS protein expression in each group of cells after knockdown or overexpression of lncRNA LOC101928697 and the statistical graphs. (i) Statistical graph of mRNA level expression of FUS in each group of cells after knockdown or overexpression of lncRNA LOC101928697 by RT-qPCR assay. ap < 0.05 compared to control group. bp < 0.05 compared to exosome group. c p < 0.05 compared to siRNA + exosome group.
Furthermore, we used western blot and RT-qPCR to analyze FUS expression in HUVECs with either overexpression or silencing of LOC101928697. The results (Figure 6(g) and (i)) showed that thrombus exosomes significantly upregulated FUS protein and mRNA expression compared with the control group (p < 0.05), an effect that was mitigated by siRNA-mediated suppression of LOC101928697 (p < 0.05). Conversely, the overexpression of LOC101928697 alone significantly increased FUS protein and mRNA levels compared with the control group (p < 0.05). These findings further support the regulatory role of LOC101928697 in FUS expression within HUVECs.
Discussion
The primary objective of this study was to investigate how TEs, lncRNA LOC101928697, and FUS proteins influence endothelial cell proliferation and migration. Through a series of in vitro experiments, we found that TEs significantly inhibited the proliferation and migration of endothelial cells by interacting with FUS proteins through their abundant lncRNA LOC101928697. Further analysis suggested that this effect may be related to changes in the expression of key cell cycle proteins, such as Cyclin D1 and PCNA. These findings not only reveal the critical role of TEs in regulating endothelial cell function but also provide new perspectives for understanding the relevance of cardiovascular disease, especially coronary thrombosis, to vascular physiology.
Exosomes have increasingly been recognized as essential mediators of intercellular communication, particularly in the context of cardiovascular diseases. 19 These vesicles are known to transport a variety of biomolecules, including proteins, RNAs, and lipids, enabling them to interact with recipient cells and influence their functions. 20 In our earlier studies, we were the first to identify TEs and to preliminarily validate their functional roles. 6 In the present study, we found that TEs significantly inhibited the migration and proliferation of HUVECs, suggesting that TEs act as potent negative regulators of endothelial cell function. This finding is consistent with previous reports on the inhibitory effects of TEs on vascular endothelial cells and further supports the notion that TEs contribute to endothelial cell damage during the pathological progression of T1MI under acute hypoxic conditions. Endothelial dysfunction is well known to play a key role in maintaining vascular wall stability, with the proliferation and migration capacities of endothelial cells being critical for the prognosis of patients with myocardial infarction.21,22 Our results suggest that TEs may exacerbate this dysfunction, potentially accelerating thrombus formation and stabilization while promoting localized vessel wall damage. These insights provide a deeper understanding of the role of TEs in the pathology of cardiovascular diseases.
Our data also revealed that TEs were significantly enriched in lncRNA LOC101928697. The binding of this lncRNA to FUS proteins in endothelial cells appears to be a key mechanism underlying the inhibitory effects of TEs on endothelial cell function. In recent years, FUS has emerged as a key RNA-binding protein critically involved in the regulation of cardiovascular and endothelial cell functions. Notably, the recent study by Hou et al. 23 demonstrated that FUS modulates the alternative splicing of cell proliferation-related genes in HUVECs, thereby influencing a variety of pathways including cell cycle progression, apoptosis, and signal transduction. Furthermore, Xiao et al. 24 revealed that the lncRNA THRIL exerts its regulatory effects on endothelial progenitor cell autophagy and proliferation via direct interaction with FUS, thus underscoring the importance of FUS–non-coding RNA complexes in maintaining endothelial homeostasis. Additionally, Garikipati et al. 25 reported that circFndc3b promotes post-infarction angiogenesis through its interaction with FUS, further highlighting the essential role of FUS-mediated non-coding RNA regulatory networks in endothelial cell function and vascular repair. Taken together, these high-impact studies provide strong theoretical support and literature evidence for the mechanism proposed in the present work, namely that exosomal LOC101928697 regulates endothelial cell function via its interaction with FUS.
Research on LOC101928697 is limited, with only one prior study suggesting a possible association between this lncRNA and vascular injury in Kawasaki disease, although the precise mechanisms remain unclear. 26 However, extensive studies on lncRNAs have established their regulatory roles in diverse biological processes, often mediated through interactions with target proteins.27,28 These roles are particularly evident in the pathogenesis of various diseases, including cardiovascular conditions. For instance, lncRNA MALAT1 has been shown to bind MT-ND1 protein, influencing cardiac energy metabolism. 29 Similarly, lncRNA PVT1 has been demonstrated to regulate cardiomyocyte apoptosis during myocardial infarction via the microRNA-216/Ccnd3 axis. 30 In this study, we combined bioinformatics analysis and experimental validation to investigate the potential role of lncRNA LOC101928697 in regulating endothelial cell proliferation and migration through binding to the RNA-binding protein FUS,31,32 also known as FUS/TLS. FUS is a well-characterized DNA/RNA-binding protein that plays a significant role in gene transport, nuclear translocation, cell cycle regulation, and intracellular RNA metabolism.33,34 Previous studies have shown that lncRNA HOTAIR regulates cardiomyocyte pyroptosis in diabetes mellitus by binding to FUS and modulating SIRT3 expression. 35 Moreover, the HOTAIR/FUS/SIRT3 axis has been implicated in cardiomyocyte damage during myocardial ischemia-reperfusion injury. 36 In our experiments, we observed a marked upregulation of FUS protein in endothelial cells treated with thrombus exosomes. Furthermore, lncRNA LOC101928697 was shown to regulate endothelial cell proliferation and migration via its interaction with FUS. These findings indicate that the LOC101928697/FUS axis plays a critical role in mediating the inhibitory effects of thrombus exosomes on endothelial cell function. This highlights the potential importance of FUS in the pathophysiology of cardiovascular diseases and further supports the emerging significance of lncRNA-protein interactions in regulating endothelial cell behavior.
In addition, Cyclin D1 and PCNA are key regulatory proteins essential for cell cycle progression. Both play critical roles in facilitating the transition from the G1 phase to the S phase, and changes in their expression levels often reflect the proliferative capacity of cells. 37 In this study, we observed a significant downregulation of Cyclin D1 and PCNA expression in HUVECs treated with TEs, suggesting that these exosomes may inhibit endothelial cell proliferation by suppressing the expression of these cell cycle proteins. Considering the observed interaction between lncRNA LOC101928697 and FUS proteins, it is plausible that lncRNA LOC101928697 indirectly regulates the expression of Cyclin D1 and PCNA through its interaction with FUS, thereby leading to the inhibition of endothelial cell proliferation. The proliferation and migration of endothelial cells are critical for vascular repair, and their inhibition by TEs may contribute significantly to thrombus formation and stabilization, as well as to the progression of cardiovascular diseases such as atherosclerosis.18,38 Therefore, the regulation of Cyclin D1 and PCNA expression via the lncRNA LOC101928697/FUS axis may represent a crucial mechanism by which TEs impair endothelial cell function, providing new insights into their role in cardiovascular disease pathology.
Importantly, the clinical relevance of the LOC101928697/FUS molecular axis extends beyond its mechanistic role in endothelial cell dysfunction. Endothelial repair is a critical determinant of vascular homeostasis following myocardial infarction, 39 influencing both the restoration of the vascular barrier and the subsequent process of cardiac remodeling. 40 Impaired endothelial proliferation and migration, as demonstrated in our study, can delay or compromise re-endothelialization, thereby exacerbating ischemia-induced injury and promoting adverse ventricular remodeling. Recent clinical and preclinical studies have underscored that rapid and effective endothelial regeneration after infarction not only reduces the risk of recurrent thrombosis but also attenuates the development of heart failure by preserving microvascular integrity and limiting myocardial fibrosis.41,42 In this context, our findings suggest that upregulation of LOC101928697 in TEs, via its interaction with FUS, may serve as a negative regulator of these reparative processes. Targeting the LOC101928697/FUS axis could, therefore, represent a novel therapeutic strategy to enhance endothelial repair, mitigate pathological remodeling, and ultimately improve clinical outcomes in patients suffering from acute myocardial infarction. Future studies, including in vivo validation and clinical correlation analyses, are warranted to explore the translational potential of this axis in cardiovascular regenerative medicine.
The present study sought to elucidate the function of exosomal lncRNA LOC101928697 in the context of coronary thrombosis. To this end, TEs were isolated from T1MI patients, and preliminary evidence was provided that LOC101928697 plays a significant role in endothelial cell injury by binding to the nuclear translocator protein FUS.
Limitation
However, there are certain limitations to this study. One major limitation is the relatively small sample size of the included clinical studies, which may have led to potential statistical false positives. In addition, it should be acknowledged that GAPDH was used as a reference gene for our hypoxia model. Although some studies have indicated that short-term hypoxic treatment does not lead to significant changes in GAPDH expression in HUVECs, 43 we nevertheless urge researchers to interpret the corresponding results with caution. Furthermore, this study did not confirm the specific binding site between lncRNA LOC101928697 and FUS, which will be a key focus for further research by our team. Moreover, our discussion regarding the downstream effector molecules of FUS in relation to LOC101928697 remains relatively superficial. In future studies, we aim to investigate the mechanisms underlying TE-induced endothelial cell injury in greater depth, providing a more comprehensive understanding of these molecular pathways.
Conclusion
In summary, the findings of this study confirm that TEs inhibit the proliferation and migration of endothelial cells through their enriched lncRNA LOC101928697, which binds to FUS proteins. This mechanism may play a significant role in the onset and progression of T1MI. Additionally, FUS proteins appear to act as key regulators of lncRNA LOC101928697, influencing the cell cycle progression of endothelial cells. Changes in the expression levels of Cyclin D1 and PCNA are likely important indicators of this process. The insights into the effects of TEs on endothelial cells presented in this study not only reveal their potential impact on vascular pathology but also identify promising molecular targets for cardiovascular diseases, including myocardial infarction. In future research, we aim to further investigate the interactions between TEs and other lncRNAs, proteins, and RNA-binding proteins. These efforts will deepen our understanding of their roles in cardiovascular diseases and pave the way for innovative approaches to early diagnosis and targeted therapies.
Footnotes
Acknowledgements
The authors sincerely thank the Department of Cardiology of Guizhou Provincial People's Hospital (Guizhou Cardiovascular Hospital) for providing the thrombus.
Ethical considerations
The study was conducted in accordance with the Declaration of Helsinki, and was approved by the Ethics Committee of Guizhou Provincial People's Hospital (Ethical Approval Number: Lun Audit (Research) 2023-071). Written informed consent was obtained from all participants prior to their inclusion in the study. Patient recruitment was conducted from 2023/08 to 2024/08 and included four patients with a confirmed diagnosis of T1MI and three healthy volunteers as controls. Four patients diagnosed with T1MI by emergency coronary angiography from the Department of Cardiology of Guizhou People's Hospital were recruited in the preliminary phase of this study. Written informed consent was obtained from the patients during the preoperative interview to facilitate the extraction of coronary thrombus and peripheral blood. All patients were provided with written formal consent prior to the acquisition of samples.
Consent to participate
All patients were provided with written formal consent prior to the acquisition of samples.
Author contributions
Youfu He, Wei Liu, Yu Zhou, Liqiong Ai, Xiaoxue Huang, and Yu Qian contributed to Data curation. Liqiong Ai, Xiaoxue Huang, Yu Qian, and Lei Peng contributed to Formal analysis. Youfu He and Wei Liu contributed to Funding acquisition. Yu Zhou, Liqiong Ai, and Lei Peng contributed to Investigation. Youfu He, Yu Zhou, Yu Qian, and Lei Peng contributed to Resources. Liqiong Ai contributed to Software. Youfu He, Xiaoxue Huang, Yu Qian, and Lei Peng contributed to Validation. Youfu He contributed to Writing—original draft. Yu Zhou and Lei Peng contributed to Writing—review & editing. All authors will be updated at each stage of manuscript processing, including submission, revision, and revision reminder, via emails from our system or the assigned Assistant Editor.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Guizhou Provincial Science and Technology Agency Project (Guizhou Provincial high-level innovative talents, Qian Ke He Foundation ZK [2023] General 217, Qian Ke He Foundation ZK [2023] General 216), Linping District Science and Technology Plan General Project (LPWJ2024-02-64), Guizhou Provincial Health and Wellness Commission (gzwkj 2021-102, the Key Advantageous Discipline Construction Project of Guizhou Provincial Health Commission in 2023, the Provincial Key Medical Discipline Construction Project of the Health Commission of Guizhou Province from 2025 to 2026) and The National Natural Science Foundation of China (No. 82260084).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The thrombus-derived exosome-related gene expression dataset was obtained from the Gene Expression Omnibus database (![]() ) under accession number (GSE213115). Additionally, the public dataset GSE163772 was utilized to examine the expression of FUS genes in endothelial cells from a myocardial infarction model.
) under accession number (GSE213115). Additionally, the public dataset GSE163772 was utilized to examine the expression of FUS genes in endothelial cells from a myocardial infarction model.
