Abstract
Objective
Breast cancer remains a leading cause of cancer-related mortality worldwide, yet the molecular mechanisms underlying its progression are not fully understood. This study aimed to identify novel hypoxia-responsive genes that promote tumor aggressiveness and could serve as prognostic markers or therapeutic targets.
Methods
The integrative bioinformatics analyses were performed using the Gene Expression Profiling Interactive Analysis (GEPIA) and User-Friendly Analysis of Cancer Gene Expression Data (UALCAN) databases to screen for genes associated with survival and tumor stage in breast cancer. Functional validation was conducted in 4th tumor, 1st clone (4T1) and M.D. Anderson-Metastatic Breast-231 (MDA-MB-231) cells under hypoxia. Gene expression was analyzed by quantitative polymerase chain reaction and western blotting. Stemness and tumorigenic potential were evaluated using sphere formation, viability, and proliferation assays.
Results
TBC1 domain family member 24 (TBC1D24) was identified as a hypoxia-responsive gene significantly associated with poor prognosis. Mechanistically, hypoxia-inducible factor-1α upregulated TBC1D24, which in turn increased adenosine diphosphate-ribosylation factor 6 (ARF6) expression and activated the phospholipase D (PLD) signaling pathway. These changes promoted cell viability, proliferation, and stem-like characteristics.
Conclusions
TBC1D24 mediates hypoxia-induced tumor progression by activating the ARF6/PLD signaling axis and promoting stemness in breast cancer cells, supporting its potential as a prognostic biomarker and therapeutic target.
Introduction
Breast cancer continues to be one of the most prevalent malignancies worldwide,1,2 with a significant portion of associated mortality attributed to metastatic dissemination and treatment resistance. 3 Despite substantial advancements in our understanding of the disease, the molecular underpinnings of breast cancer progression remain inadequately elucidated.4,5
Hypoxia is closely associated with tumor progression and metastasis, 6 and the expression and activation of hypoxia-inducible factor-1α (HIF-1α) play key roles in this process.7,8 HIF-1α contributes to multiple steps of the metastatic cascade, 9 including angiogenesis, extracellular-matrix remodeling, cell motility, and immune evasion. 10 These effects may be mediated through mechanisms such as DNA methylation, histone modifications, and chromatin reorganization, which collectively contribute to malignant transformation.11–13 Collectively, clarifying the molecular links between hypoxia and transcriptional or epigenetic reprogramming may help identify novel targets in breast cancer.
TBC1 domain family member 24 (TBC1D24) encodes a protein that contains the Tre2/Bub2/Cdc16 (TBC) domain, which is also found in Rab GTPase-activating proteins. 14 Proteins with TBC domains regulate vesicular transport and sorting primarily by modulating Rab-GTPase activity. 15 Previous studies have reported that TBC1D24 gene plays a role in regulating synaptic vesicle transport as well as the development of brain and somatic cells. 16 Pathogenic mutations in TBC1D24 lead to significant impairment of neuronal development.17,18 Previous research has reported that TBC1D24 is overexpressed in breast cancer tissues and correlates with poor prognosis and immune cell infiltration, suggesting its relevance to tumor progression. 19 Nonetheless, the molecular function and upstream regulation of TBC1D24 in breast cancer remain largely unexplored, especially under physiologically relevant conditions such as hypoxia.
Prior non-oncologic studies have shown that TBC1D24 can interact with and modulate adenosine diphosphate-ribosylation factor 6 (ARF6) activity. However, whether this interaction contributes to breast cancer malignancy, particularly in response to hypoxic stress, has not been investigated. In this study, we hypothesize that HIF-1α transcriptionally activates TBC1D24, which in turn enhances ARF6/phospholipase D (PLD) signaling to promote tumor cell stemness and progression. By elucidating this pathway, we aim to provide new insights into the hypoxia-driven molecular mechanisms of breast cancer and establish TBC1D24 as a potential prognostic marker and therapeutic target.
ARF6, a small GTPase-regulating membrane dynamics and actin remodeling,20,21 plays a critical role in cancer invasion and metastasis by activating the PLD signaling pathway.22,23 This process influences the generation of phosphatidic acid and the subsequent activation of downstream signaling cascades, which in turn modulate various cellular processes such as proliferation, differentiation, and migration. 24 Previous investigations have demonstrated that TBC1D24 can interact with and modulate ARF6 activity, 25 thereby playing a crucial role in diseases such as epilepsy and deafness. 26 However, whether this interaction contributes to breast cancer malignancy, particularly in response to hypoxic stress, has not been investigated. In this study, we hypothesize that HIF-1α transcriptionally activates TBC1D24, which in turn enhances ARF6/PLD signaling to promote tumor cell stemness and progression. By elucidating this pathway, we aim to provide new insights into the hypoxia-driven molecular mechanisms of breast cancer and establish TBC1D24 as a potential prognostic marker and therapeutic target.
Materials and methods
Bioinformatics analysis
To explore the prognostic value and regulatory mechanisms of TBC1D24 in breast cancer, multiple bioinformatics platforms were employed. The Gene Expression Profiling Interactive Analysis (GEPIA, v2) and User-Friendly Analysis of Cancer Gene Expression Data (UALCAN) databases were used to assess differential gene expression and correlations with overall survival (OS), disease-free survival (DFS), and TP53 mutation status. For GEPIA-based differential gene expression analyses, the significance threshold was set as |log2 fold change (log2 FC)| > 1 and p-value <0.01. A Kaplan–Meier plotter was utilized for independent survival validation. Survival curves were generated using median-split TBC1D24 expression (p < 0.05). Protein–protein interaction networks were predicted using the STRING database (v11.5), while signaling pathway associations were analyzed via cBioPortal. To investigate potential transcriptional regulation, the JASPAR (2022) and hTFtarget databases were applied to predict HIF-1α-binding motifs in the TBC1D24 promoter region. These analyses guided subsequent experimental validation of TBC1D24 as a hypoxia-responsive gene and its functional role in breast cancer progression.
Cell culture
This study employed human (M.D. Anderson-Metastatic Breast-231, MDA-MB-231) and murine (4th tumor, 1st clone, 4T1) triple-negative breast cancer (TNBC) cell lines in vitro experiments, which are widely used preclinical models for investigating tumor aggressiveness, therapy resistance, and hypoxia-associated signaling in TNBC, with broad applicability in both in vitro mechanistic exploration and in vivo modeling of TNBC progression. These cell lines were obtained from Zqxzbio (Shanghai, China). The cells were maintained in Roswell Park Memorial Institute (RPMI)-1640 medium, which was supplemented with 10% fetal bovine serum (FBS) (Thermo Fisher Scientific, Waltham, MA, USA), 1% penicillin/streptomycin (Sigma-Aldrich, St Louis, MO, USA), and incubated at 37°C and 5% CO2 humidified atmosphere. For hypoxic culture, cells were incubated in a hypoxia chamber (1% O2, 5% CO2, 94% N2) for 24 h. The culture medium was refreshed every other day to ensure optimal cell growth conditions.
Cell transfection and viability assay
Cell viability was assessed by 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide colorimetric assay (MTT) assay (Solarbio, Hefei, China). The cells were inoculated into 96-well plates at a concentration of 3 × 103 cells per well, suspended in 100 µL of culture medium. Following this, the plasmids encoding TBC1D24 and HIF-1α (Ribobio, Guangzhou, China) were transfected into the cells using a Transfection kit (Ribobio) according to manufacturer's protocol. Cells were transfected for 24 h before subsequent assays. To assess cell viability, 10 µL of 5 mg/mL MTT solution was introduced to each well at predetermined intervals on days 0, 1, 2, 3, 4, and 5 of the growth period. The plates were subsequently maintained at 37°C for an extra hour. Post-incubation, absorbance measurements were conducted using a spectrophotometer (Thermo Fisher Scientific) to evaluate the metabolic activity of the cells, providing an indication of cellular viability.
Cell treatment
4T1 and MDA-MB-231 cancer cells were cultured at a density of 2 × 106 cells/mL in six-well plates. The cell lines were maintained in RPMI-1640 culture medium (Sigma-Aldrich) supplemented with 10% FBS and 1% penicillin/streptomycin (Sigma-Aldrich). The cells were incubated at 37°C under a humidified atmosphere containing 5% CO2, with the culture medium being replaced every other day. For the CoCl2 (Sigma-Aldrich) treatment, 4T1 and MDA-MB-231 cells were cultured in RPMI-1640 medium (Sigma-Aldrich), either with or without 200 μM CoCl2, for 48 h. Cells were collected for quantitative polymerase chain reaction (qPCR) and western blot analysis.
Cell scratch test
Cells were seeded into a six-well plate, after achieving confluence. To specifically evaluate cell migration, cells were pretreated with mitomycin C (10 μg/mL) for 2 h prior to scratching to inhibit proliferation. A linear wound was then generated across the cell monolayer using a 10 μL sterile pipette tip. Subsequently, the cells were incubated in serum-free medium to exclude any effects of FBS on cell migration. Scratch images were captured at regular intervals over a period of 48 h using an Olympus inverted microscope. The width of the scratch was quantified using Olympus CellSens Dimension software.
Quantitative real-time PCR
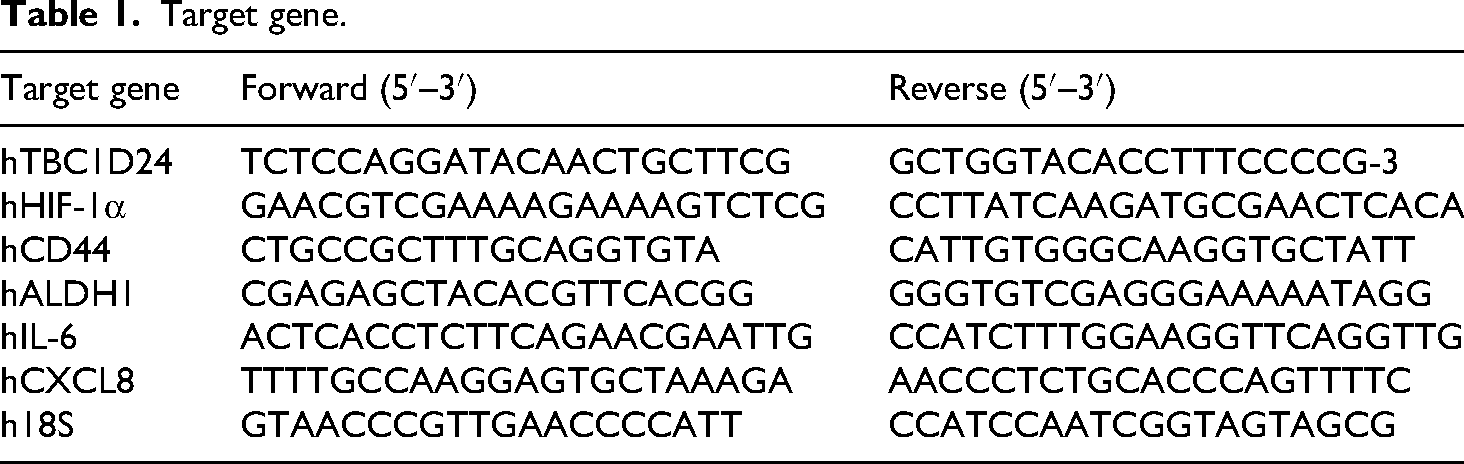
Total RNA was extracted with TRIzol (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's protocol. To eliminate potential genomic DNA contamination, RNA purification was performed. Subsequently, reverse transcription into first-strand complementary DNA (cDNA) was carried out using a First-Strand cDNA Synthesis Kit (Cwbio, Beijing, China). This involved combining 1 μg of total RNA with the reverse transcription mix as per the kit instructions. Quantitative real-time PCR analysis was then performed using a ABI 7500 real-time PCR system (Applied Biosystems, Foster City, CA, USA). Target-specific primers were added to the cDNA, and SYBR qPCR Master Mix (Vazyme, Nanjing, China) was used for the amplification reaction. The primers used were specifically designed for species-specific primers (see Table 1 for primer sequences). The relative quantification of gene expression was analyzed using the 2−ΔΔCt method, with 18S serving as the internal control to normalize the data. This approach provides a normalized and calibrated measure of gene expression levels between samples.
Target gene.
Sphere formation assay
The sphere formation assay was performed as previously described.
27
In brief, 100 cancer cells were seeded in each well of an ultralow attachment 96-well plate (Corning, New York, USA) containing 100 μL of FBS-free RPMI-1640 medium (Sigma-Aldrich) supplemented with
Luciferase assay
The TBC1D24 promoter fragment upstream of the transcription start site was cloned into pGL3-Basic to generate the wild-type (WT) promoter reporter; the hypoxia-response element (HRE) motif (5′-RCGTG-3′) was mutated to obtain the HRE-mutant construct. In addition, the pGL3-Basic empty vector was included as a negative control. Sequence-verified plasmids were obtained from Genechem (Shanghai, China). 4T1 cells were co-transfected with the WT, mutant, or empty vector reporter constructs, pRL-TK Renilla (10:1), and either HIF-1α expression plasmid or control vector. Dual-luciferase activity was measured after 48 h and normalized to Renilla.
Western blotting
Proteins were extracted using a Protein Extraction Kit (Biosharp, Hefei, China). The protein concentrations were quantified using the Bradford assay (Beyotime, Shanghai, China). A total of 30 μg of protein from each sample was loaded onto a sodium dodecyl sulfate-polyacrylamide gel electrophoresis system for separation, followed by transfer onto polyvinylidene difluoride membranes (Merck-Millipore, Billerica, MA, USA). The membranes were subsequently blocked with 5% bovine serum albumin in Tris-buffered saline containing 0.1% Tween 20 (TBST) for 2 h at room temperature to minimize non-specific binding. Following blocking, the membranes were incubated overnight at 4°C with primary antibodies targeting TBC1D24 (1:1000; Abcam, Cambridge, UK), ARF6 (1:1000; CST, Massachusetts, USA), PLD2 (1:1000; Abcam), HIF-1α (1:1000; Abcam), and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) (1:1000; Abcam). After a thorough washing with TBST to remove unbound primary antibodies, the membranes were incubated with horseradish peroxidase-conjugated anti-rabbit secondary antibodies (1:10 000; Proteintech, Wuhan, China) at room temperature for 1 h. Protein detection was performed using an enhanced chemiluminescence detection system (Merck-Millipore), with GAPDH serving as a control to normalize the expression levels of the target proteins. To ensure reproducibility and reliability, all experiments involving transfected cells were independently replicated at least three times.
Statistical analysis
All data are expressed as the mean ± standard deviation (SD). Statistical analyses were performed using GraphPad Prism version 8.0 (GraphPad Software, CA, USA). For comparisons between multiple groups, one-way analysis of variance was conducted, followed by post-hoc testing to identify specific differences. In cases involving pairwise comparisons, Student's t-test was employed. Prior to parametric testing, data were assessed for normality using the Shapiro–Wilk test and for homogeneity of variances using Levene's test. Statistical significance was defined as a p value less than 0.05. For extremely small p values, GraphPad Prism reports results in threshold format (e.g. p < 0.0001). Accordingly, statistical significance in figures is consistently indicated using the following symbols: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Results
TBC1D24 associated with survival outcomes in breast cancer
We initially analyzed the most differentially expressed survival genes (MDSGs) in datasets of breast cancer patients using the GEPIA database28,29 (http://gepia.cancer-pku.cn/). The top 500 MDSGs were identified from both the OS and DFS groups, using the median as the cutoff threshold (Supplemental Tables S1 and S2). The findings indicate that TBC1D24 is one of the MDSGs shared among the top 500 genes in both the OS and DFS cohorts, as highlighted in Supplemental Table S3. This overlap underscores the gene's pivotal role in differentiating the two groups, a conclusion further supported by the significant statistical disparities observed in Figure 1(a). To investigate the differences in TBC1D24 expression between the OS and DFS groups, we conducted an analysis using the GEPIA database. The results (Figure 1(b) and (c)) demonstrated that TBC1D24 is significantly correlated with both the OS and DFS rates in breast cancer. Next, we conducted a detailed analysis of TBC1D24 gene expression in breast cancer and normal tissues using the GEPIA database, which revealed significantly higher expression levels in tumor tissues (Figure 1(d)).

TBC1D24 associated with survival outcomes in breast cancer. (a) The Venn plot shows the intersection set of most differential survival genes in breast cancer patients in the GEPIA dataset. (b) Kaplan–Meier analysis of overall survival of breast cancer patients based on different TBC1D24 levels in the GEPIA dataset. (c) Kaplan–Meier analysis of recurrence-free survival of breast cancer patients based on different TBC1D24 levels in the GEPIA dataset. (d) The expression level of TBC1D24 mRNA in tumor tissues (1065 cases from TCGA) and normal tissues (291 cases from GTEx) was compared. (e) Expression of TBC1D24 in breast cancer patients with different TP53 status in the TCGA dataset. (f) Kaplan–Meier plots of overall survival of breast cancer patients stratified at the TBC1D24 level in the TCGA dataset. Data are expressed as mean ± SD. *p < 0.05, **p < 0.01. TBC1D24: TBC1 domain family member 24.
TP53 is one of the most frequently mutated genes in breast cancer and is closely linked to poor prognosis and genomic instability. To explore its relationship with TBC1D24, we analyzed TCGA data via the UALCAN platform. 30 The results revealed that patients with TP53 mutations exhibited significantly higher TBC1D24 expression compared to those without mutations (Figure 1(e)), suggesting a potential interaction between TP53 status and TBC1D24 expression. Additionally, we assessed patient prognosis using the Kaplan–Meier plotter database confirmed that patients with elevated TBC1D24 expression had significantly worse OS outcomes (Figure 1(f)). Collectively, these findings indicate that TBC1D24 is highly expressed in breast cancer tissues and is closely associated with poor prognosis, especially in patients with TP53 mutations, suggesting its potential value as a prognostic biomarker.
TBC1D24 promotes breast cancer cell proliferation
To elucidate the functional role of TBC1D24 in cellular processes, we initially overexpressed TBC1D24 in mouse TNBC 4T1 cells and human breast cancer MDA-MB-231 cells, both of which are widely used models for studying tumor proliferation and metastasis. qPCR and western blot analysis results confirmed the efficiency of TBC1D24 overexpression (Figure 2(a) and (b)). MTT assays revealed that TBC1D24 overexpression significantly increased the viability of both 4T1 and MDA-MB-231 cells compared to vector controls (Figure 2(c) and (d)). Additionally, scratch assays demonstrated that TBC1D24-overexpressing cells exhibited enhanced migratory capacity, further supporting a pro-tumorigenic role of this gene (Figure 2(e) to (h)). Furthermore, we performed RNA interference-mediated knockdown in both cell lines, with silencing efficiency confirmed by qPCR (Figure 2(i)). The knockdown of TBC1D24 resulted in a significant inhibition of cell viability in both 4T1 and MDA-MB-231 cell lines, as anticipated (Figure 2(j) and (k)). Collectively, these findings indicate that TBC1D24 promotes breast cancer cell viability and migration, suggesting its involvement in tumor progression.

TBC1D24 promotes breast cancer cell proliferation. (a, b) The expression levels of TBC1D24 mRNA and protein in 4T1 and MDA-MB-231 cells transfected with TBC1D24 expression. (c) The cell viability of 4T1 cells transfected with TBC1D24 expression by MTT assay. (d) The cell viability of MDA-MB-231 cells transfected with TBC1D24 expression by MTT assay. (e) Cell migration was detected by scratch assay in 4T1 cells transfected with a TBC1D24-expressing or a control. Scale bar, 100 μm. (f) Quantification of the cell migration rate in 4T1 cells. (g) Cell migration ability of MDA-MB-231 cells transfected with TBC1D24 expression or control was detected by scratch method. (h) Quantification of the cell migration rate in MDA-MB-231 cells. (i) The expression level of TBC1D24 mRNA in 4T1 and MDA-MB-231 cells transfected with TBC1D24 shRNA was detected by qPCR. (j, k) The cell viability of 4T1 and MDA-MB-231 cells transfected with TBC1D24 shRNA by MTT assay. Data are expressed as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. TBC1D24: TBC1 domain family member 24; qPCR: quantitative polymerase chain reaction.
TBC1D24 promotes cell stemness in breast cancer cells
To investigate the role of TBC1D24 in regulating breast cancer stemness, we first employed the sphere formation assay, which revealed that overexpression of TBC1D24 significantly enhanced the sphere formation ability of both 4T1 and MDA-MB-231 cells (Figure 3(a) to (d)). This result suggests that TBC1D24 may play a role in promoting the stem cell-like properties of these cancer cells. Figure 3(e) demonstrated that TBC1D24 expression in the sphere cells was higher compared to that in the normal adherent cells. This finding indicates that TBC1D24 is upregulated in breast cancer stem cells (CSCs). To explore the role of TBC1D24 in breast cancer stemness further, we analyzed the expression of stem cell markers in 4T1 cells using qPCR. The results showed that overexpression of TBC1D24 led to a significant upregulation breast CSC markers,31–33 including cluster of differentiation 44 (CD44), aldehyde dehydrogenase 1 (ALDH1), interleukin-6 (IL-6), and C-X-C motif chemokine ligand 8 (CXCL8) (Figure 3(f)). These markers are known to be associated with stemness and the tumorigenic potential of breast cancer cells. Collectively, these findings indicate that TBC1D24 promotes breast cancer stemness by enhancing the expression of CSC markers and supporting tumorsphere formation. These results highlight TBC1D24 as a potential regulator of CSC phenotypes and a promising target for preventing tumor recurrence.

TBC1D24 promotes cell stemness in breast cancer. (a) Tumor ball formation assay in 4T1 cells transfected with the TBC1D24-expressing plasmid. Scale bar, 300 μm. (b) Analysis of tumor ball formation results in 4T1 cells. (c) Experimental image of tumor ball formation in MDA-MB-231 cells after transfection with the TBC1D24 expression plasmid. Scale bar, 300 μm. (d) Analysis of tumor ball formation results of MDA-MB-231 cells. (e) Expression level of TBC1D24 mRNA in 4T1 or MDA-MB-231 parent cells and tumor balls. (f) Results of marker mRNA in 4T1 cells transfected with TBC1D24 expression plasmid. Data are expressed as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. TBC1D24: TBC1 domain family member 24.
Hypoxia induces TBC1D24 expression in breast cancer cells
To investigate whether hypoxia could induce the expression of TBC1D24, we cultured 4T1 and MDA-MB-231 cells under either normoxic or hypoxic conditions. As shown in Figure 4(a) to (c), both the mRNA and protein levels of TBC1D24, as well as HIF-1α, were significantly elevated under hypoxic conditions in both cell lines, indicating that hypoxia upregulates TBC1D24 expression. To further validate this finding, we treated cells with cobalt chloride (CoCl2), a hypoxia mimetic. Consistently, CoCl2 treatment led to a marked increase in TBC1D24 mRNA levels (Figure 4(d)), confirming that hypoxia-induced TBC1D24 expression is a robust and reproducible phenomenon. Next, we sought to determine whether HIF-1α, a key transcription factor activated under hypoxic conditions, could directly regulate TBC1D24 expression. To test this, we overexpressed HIF-1α in 4T1 cells (Figure 4(e)). The results from qPCR showed that overexpression of HIF-1α significantly increased the mRNA expression of TBC1D24 (Figure 4(f)).

Hypoxia induces TBC1D24 expression. The expression of TBC1D24 (a) or HIF-1α and (b) in 4T1 or MDA-MB-231 cells cultured in normal or hypoxia culture was detected by qPCR. (c) Western blot analysis of the expression of TBC1D24 and HIF-1α in 4T1 or MDA-MB-231 cells cultured under normal oxygen and hypoxia conditions. (d) The expression of TBC1D24 in CoCl2-treated 4T1 or MDA-MB-231 cells was detected by qPCR. (e) The expression of HIF-1α in 4T1 cells transfected with HIF-1α-expressing plasmid was detected by qPCR. (f) The expression of TBC1D24 in 4T1 cells transfected with HIF-1α-expressing plasmid was detected by qPCR. (g) Schematic diagram of the HIF-1α binding site in the TBC1D24 promoter region. (h) Detection of TBC1D24 promoter reporter luciferase reporter gene in 4T1 cells transfected with HIF-1α-expressing plasmid. Data are expressed as mean ± SD. ns was not significant, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. TBC1D24: TBC1 domain family member 24; HIF-1α: hypoxia-inducible factor-1α; qPCR: quantitative polymerase chain reaction.
To further investigate the transcriptional regulation of TBC1D24 by HIF-1α, we used bioinformatics tools such as the hTFtarget Database and JASPAR to predict transcription factor binding sites in the promoter region of TBC1D24. Our analysis identified a binding site for HIF-1α in the TBC1D24 promoter (Figure 4(g)). To functionally verify this regulatory interaction of TBC1D24 by HIF-1α, we conducted a luciferase reporter driven by the WT TBC1D24 promoter, as well as a mutant version with a disrupted HRE motif. As shown in Figure 4(h), HIF-1α overexpression significantly increased the luciferase activity of the WT promoter compared to vector, whereas this effect was abolished in the HRE-mutated construct. Notably, the baseline activity of the mutant promoter remained comparable to that of the pGL3-Basic empty vector. These results confirm that HIF-1α enhances TBC1D24 transcription via a functional HRE motif in its promoter. This finding suggests a mechanistic link between hypoxia and the stemness of breast cancer cells, possibly contributing to the aggressive behavior and therapeutic resistance of breast cancer.
TBC1D24 activates ARF6/PLD signaling in breast cancer cells under hypoxia
To investigate potential protein interactions, we interrogated the STRING database (https://string-db.org/), which predicted a strong association between TBC1D24 and ARF6 (interaction score: 0.953; Figure 5(a) to (c)). Although STRING reflects predicted protein–protein interactions rather than expression regulation, this finding prompted us to investigate whether TBC1D24 may modulate ARF6 expression in breast cancer. Subsequently, pathway enrichment analysis using the cBioPortal (http://cbioportal.org) to analyze signaling pathways involving TBC1D24 in the TCGA breast cancer patient cohort, we observed a notable correlation between TBC1D24 and the PLD signaling pathway (Figure 5(d)). Consistent with prior literature, ARF6 has been implicated in modulating cell proliferation, differentiation, and invasion through its regulation of the PLD signaling pathway. Based on these findings, we hypothesized that TBC1D24 influences breast cancer cell function via the ARF6/PLD signaling axis. Next, we overexpressed TBC1D24 in 4T1 and MDA-MB-231 cells, and found a marked increase in ARF6 protein levels and the activation of the PLD subunit PLD2 (Figure 5(e) to (g)). Additionally, we examined the impact of hypoxia on ARF6 expression. Our results demonstrated that 4T1 cells cultured under hypoxic conditions or treated with CoCl2 to mimic hypoxia exhibited significant upregulation of ARF6 mRNA (Figure 5(h) and (i)). These findings are consistent with previous studies reporting that hypoxia can activate ARF6 and its downstream signaling pathways to facilitate tumor progression.34,35 Together, these results suggest that TBC1D24, upregulated under hypoxia, may enhance ARF6 expression and activate the downstream PLD signaling cascade, thereby contributing to the malignant progression of breast cancer.

TBC1D24 activates ARF6/PLD signaling in breast cancer under hypoxia. (a–c) Using the STRING database to analyze breast cancer data, ARF6 was identified as a potential protein. (d) Analysis of TCGA breast cancer data using cBioPortal indicates that TBC1D24 may be involved in the PLD signaling pathway. (e–g) The expression analysis of ARF6 and its downstream effector, PLD2, were conducted in 4T1 and MDA-MB-231 cells transfected with TBC1D24. (h) The expression of ARF6 in cells cultured under normoxia or hypoxia conditions was determined by qPCR. (i) The expression of ARF6 in 4T1 cells treated with CoCl2 was determined by qPCR. Data are expressed as mean ± SD. *p < 0.05, ***p < 0.001. TBC1D24: TBC1 domain family member 24; ARF6: adenosine diphosphate-ribosylation factor 6; PLD: phospholipase D; qPCR: quantitative polymerase chain reaction.
Discussion
Breast cancer represents a highly lethal malignancy, the pathogenic mechanisms of which remain incompletely elucidated.36,37 Hypoxia has been recognized as a key physiological feature in the deterioration of breast cancer cells.38,39 However, the downstream mediators through which hypoxic tumor cells regulate their own proliferation and invasion remain largely unclear. Thus, identifying novel biomarkers and therapeutic targets is of paramount importance.
In this study, we systematically screened for hypoxia-associated genes linked to survival and tumor progression using bioinformatics tools such as GEPIA and UALCAN, and identified TBC1D24 as a promising candidate. Mechanistic experiments confirmed that HIF-1α directly upregulates TBC1D24, which in turn enhances the expression of ARF6 and activates its downstream effector PLD2. This activation of the ARF6/PLD signaling pathway promotes proliferation, invasion, and stem-like properties in breast cancer cells. These findings delineate a hypoxia-driven pathway in which TBC1D24 regulates ARF6-dependent PLD signaling. Moreover, the identification of TBC1D24 as a transcriptional target of HIF-1α provides new insight into how the hypoxic tumor microenvironment promotes tumor progression and lays the foundation for future therapeutic strategies targeting hypoxia-related pathways.
We observed that TBC1D24 plays a key role in the proliferation, migration, and invasion of breast cancer cells, and there is a clear correlation between hypoxia and tumor cell TBC1D24 expression, indicating a clear link between the tumor microenvironment and TBC1D24 activation. Functional assays further showed that TBC1D24 enhances colony formation and increases stemness-related gene expression, while its silencing leads to marked inhibition of malignant behaviors. Our study also found that TBC1D24 regulates the downstream PLD pathway through ARF6, which is consistent with previous reports. 17 Importantly, we further observed that this regulatory relationship is enhanced under hypoxic conditions in breast cancer cells. These findings suggest that TBC1D24 may serve as a biomarker and therapeutic target in hypoxia-adapted, stemness-enriched breast cancer. Given the association of TBC1D24 with TP53 mutation and stemness features, and based on our observations of its hypoxia-induced pro-malignant effects in both MDA-MB-231 and 4T1 TNBC models, TBC1D24 may serve as a promising therapeutic target, particularly in aggressive breast cancer subtypes driven by hypoxia and characterized by poor prognosis, such as TNBC.
There are some limitations to this study. First, although CoCl2 stabilizes HIF-1α it does not reproduce the spatial–temporal oxygen gradients of tumors; accordingly, CoCl2 data should be interpreted alongside results from controlled-oxygen hypoxia. Mechanistically, although ARF6/PLD lies downstream of TBC1D24, the route by which TBC1D24 regulates ARF6 remains undefined, and other hypoxia-responsive pathways may contribute. Our stemness readouts quantified sphere number and marker expression but not viability or clonal heterogeneity, motivating the use of fluorescein diacetate/propidium iodide staining, flow cytometry, and CD44+/CD24− immunofluorescence in future work. Additionally, while HIF-1α has been shown to transcriptionally activate TBC1D24, it remains to be determined whether upstream epigenetic mechanisms (chromatin accessibility, DNA methylation, histone modifications) contribute to this regulation. These insights may inform novel therapeutic strategies targeting hypoxia-induced reprogramming in breast cancer.
Conclusion
This study provides evidence that hypoxia induces HIF-1α-dependent upregulation of TBC1D24, which activates the ARF6/PLD axis and promotes breast cancer progression. These findings highlight the potential of targeting the HIF-1α–TBC1D24–ARF6/PLD pathway may offer a therapeutic strategy for aggressive breast cancer.
Supplemental Material
sj-docx-1-sci-10.1177_00368504251367649 - Supplemental material for TBC1D24 promotes the progression of the breast cancer cells via ARF6/PLD axis under hypoxia
Supplemental material, sj-docx-1-sci-10.1177_00368504251367649 for TBC1D24 promotes the progression of the breast cancer cells via ARF6/PLD axis under hypoxia by Qingyang Li, Kangkang Xie, Beibei Lin and Yudiao Gong in Science Progress
Supplemental Material
sj-docx-2-sci-10.1177_00368504251367649 - Supplemental material for TBC1D24 promotes the progression of the breast cancer cells via ARF6/PLD axis under hypoxia
Supplemental material, sj-docx-2-sci-10.1177_00368504251367649 for TBC1D24 promotes the progression of the breast cancer cells via ARF6/PLD axis under hypoxia by Qingyang Li, Kangkang Xie, Beibei Lin and Yudiao Gong in Science Progress
Supplemental Material
sj-docx-3-sci-10.1177_00368504251367649 - Supplemental material for TBC1D24 promotes the progression of the breast cancer cells via ARF6/PLD axis under hypoxia
Supplemental material, sj-docx-3-sci-10.1177_00368504251367649 for TBC1D24 promotes the progression of the breast cancer cells via ARF6/PLD axis under hypoxia by Qingyang Li, Kangkang Xie, Beibei Lin and Yudiao Gong in Science Progress
Footnotes
Acknowledgments
The experiments in this study were conducted at the Affiliated Cangnan Hospital of Wenzhou Medical University. We are particularly grateful to the developers and maintainers of GEPIA, UALCAN, JASPAR, and STRING—all open-access resources for the research community—whose online analysis pages provided the database used in this study.
Ethic statement
This in vitro study complied with institutional biosafety and ethics guidelines.
Author contributions
Qingyang Li: project administration and data analysis. Kangkang Xie: writing—original draft and investigation. Beibei Lin: formal analysis and methodology software. Yudiao Gong: supervision, writing—review and editing, and funding acquisition.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by The Science and Technology Project of Cangnan County (grant number 2021S06).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
All the data can be obtained by contacting the authors.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
