Abstract
The frequency of drought is expected to rise in many parts of the world with increasing climate change. Despite being an economically valuable plant species, the molecular mechanisms regulating the responses of peach (Prunus davidiana) to drought stress and the functional genes conferring drought resistance are currently unknown. In this study, we investigated the phenotypic and physiological responses of peach seedlings to experimental conditions that included a control, a period of drought stress, and a rehydration period. We performed transcriptome sequencing and investigated differences in the transcriptome of peach seedlings exposed to different treatments. We also analyzed the functions and regulatory pathways of differentially expressed genes using GO and KEGG enrichment. The results showed that severe drought stress occurred in the peach seedlings on the sixth day of drought, and that the physiological responses of peach seedlings experiencing drought stress were significantly different from those in control conditions. We found 21348 differentially expressed genes in peach seedlings under drought stress, of which 10105 were up-regulated and 11243 were down-regulated in comparison with peach seedlings in control conditions. These differentially expressed genes were mainly involved in the biosynthesis of amino acids, metabolic pathways, antioxidant defense systems and the plant hormone signal transduction system. The results suggest that peach seedlings respond to severe drought stress by initiating antioxidant defense mechanisms to alleviate damages, activating different signal transduction pathways to transmit signals, regulating the synthesis of amino acids, and initiating metabolic mechanisms to enhance osmotic pressure. This study illuminates the mechanisms for drought resistance in peach seedlings at the molecular level. Overall, the findings provide a theoretical basis for the cloning and functional analysis of genes conferring drought resistance, and the cultivation of more drought resistant varieties of peach.
Keywords
Introduction
As a consequence of increasing climate change globally, drought has become the dominant driver of abiotic stress in plants. 1 Drought not only impacts a plant's water balance, but also causes oxidative stress, damages the cell membrane, thereby causing dehydration and even mortality. 2 These impacts of drought can severely limit the growth and yield of economically important plants. Recent studies highlight conserved molecular mechanisms across species, particularly in antioxidant defense and transcriptional regulation, which coordinate plant adaptation to drought. 3 Many plant species have evolved complex physiological and biochemical metabolic mechanisms as adaptive responses to stressful environmental conditions. First of all, in terms of phenotypic characteristics, plant water loss leads to slow plant growth, leaf wilting and drooping, leaf number and leaf area decreasing, and even plant death. 4 In addition to the phenotype, cell membrane permeability will also change due to water shortage, and changes in membrane permeability will affect relative conductivity. 5 In addition, the activity of superoxide dismutase (SOD) and the content of malondialdehyde (MDA) in plants are closely related to their drought tolerance. SOD scavenges O2-· radicals, while catalase (CAT) and peroxidase (POD) detoxify H2O2, collectively mitigating oxidative damage. 6 They can scavenge free radicals such as O2-·, H2O2 and ·OH in the body and prevent membrane lipid peroxidation, so as to protect the cell membrane. 7 Chlorophyll converts light energy into chemical energy in plant photosynthesis, and its content reflects the strength of plant photosynthesis. When plants encounter drought stress, chlorophyll synthesis is inhibited, which eventually leads to yellowing and wilting of plant leaves. Transcriptomic analyses in crops such as maize and wheat reveal that drought downregulates chlorophyll biosynthesis genes (e.g. POR, HEMA1), highlighting conserved molecular responses. 8 Carotenoids, as an auxiliary pigment, also have antioxidant effects. Carotenoids protect photosynthetic apparatus by quenching singlet oxygen and dissipating excess energy, a mechanism validated in both Arabidopsis and rice. 9 Therefore, the changes of chlorophyll a (Chl a), chlorophyll b (Chl b) and carotenoid (Car) contents are commonly used to reflect the degree of stress on plants. Carotenoids also have antioxidant effects. Therefore, the changes of chlorophyll a (Chl a), chlorophyll b (Chl b) and carotenoid (Car) contents are commonly used to reflect the degree of stress on plants. The results showed that the content of photosynthetic pigments in leaves decreased significantly under drought stress, 4 and the decrease of photosynthetic pigments caused the change of light reaction process, which led to the decrease of chlorophyll fluorescence parameters. 10 Some plant species can initiate a variety of drought response mechanisms in response to drought stress, for instance by altering their growth morphology, initiating drought resistance genes, activating protective enzymes, or regulating endogenous hormones and osmotic adjustment substances to alleviate any damages caused by drought stress. 11 For example, ABA-mediated stomatal closure and proline accumulation are universal strategies in both monocots and dicots.12–13 A variety of transcription factors and signaling molecules play key regulatory roles in these defense systems. 14 As a response to drought stress, various transcription factors in a plant can regulate the expression of downstream genes independently, or by binding to other proteins. 15 The NAC family transcription factor RD26 in Arabidopsis and its homolog SNAC1 in rice coordinate drought responses by regulating stomatal dynamics and root architecture, demonstrating functional conservation. 16 Therefore, the systematic study of gene transcription and gene regulation in plant cells based on transcriptomics is of great significance for understanding the molecular mechanism under drought stress and mining drought-resistant genes, thus providing a theoretical basis for cultivating stress-resistant varieties. 17 Previous studies have conducted in-depth analysis of transcriptomics of potato, 18 wheat,19–20 rape, 21 peanut, 22 maize, 23 mulberry 24 and other crops under drought stress, identified corresponding drought stress response genes, and confirmed that these differentially expressed genes are mainly enriched in oxidative phosphorylation, photosynthesis and plant metabolic pathways. However, there are few reports on transcriptomics in peach drought resistance. The peach (Prunus davidiana) is a perennial deciduous fruit tree that is native to China, where it has been cultivated for over 3000 years. The peach plant requires a large amount of water throughout its growth, and drought and water shortage can lead to a serious decline in yield and quality. Given the conserved roles of ABA signaling (PYR1 receptors) and antioxidant enzymes (SOD/CAT) in drought adaptation, targeting homologous pathways in peach could accelerate drought-resistant breeding. 25 Given the high economic value of the peach plant and the increasing frequency of drought events brought on by climate change, 26 it is imperative to elucidate the molecular mechanisms underlying the response of the peach plant to drought stress. It is hoped that such knowledge will help advance the cultivation of more drought resistant peach plants.
In this study, peach seedlings were subjected to drought stress, and the phenotypic changes of seedlings, physiological indexes and hormone contents in plants were observed. On this basis, transcriptome sequencing was conducted on the leaves of the peach seedlings, and a functional annotation as well as classification and metabolic pathway analyses were performed. The expression of key genes determining the responses of peach seedlings to drought stress was examined at the transcription level, and in particular, genes relating to drought resistance were identified. The findings provide a theoretical basis for the cloning and functional analysis of drought resistance-related genes in peach seedlings and the cultivation of more drought resistant peach varieties.
Materials and methods
Materials and samples
The experiment was carried out in the greenhouse of Henan Institute of Science and Technology (113°54′E, 35°18′N) in 2021. The daily temperature was 25-35 °C, the night temperature was 15-25 °C, and the air humidity was 70%. The test materials were three-month-old peach seedlings (Prunus davidiana), and the peach seeds are collected from Xinxiang City, Henan Province, China. Peach has a high degree of drought resistance, cold resistance, barren resistance, less pests and diseases. It is widely used as rootstock for peach, plum, apricot and other Rosaceae fruit trees. The tested soil was taken from the 0 ∼ 20 cm soil layer of the test site. After natural air drying, screening and impurity removal, it was mixed with the cultivation substrate in a capacity ratio of 1 ∶ 2 for reserve. The plastic basin with a diameter of 22 cm, a bottom diameter of 18 cm, and a height of 16 cm is filled with 3 kg of soil, and a tray is placed at the bottom to sink with water. One seedling was planted in each plastic basin, and its soil moisture content was maintained at field capacity to ensure the robust growth of seedlings.
Experimental design
Drought stress was simulated by pot water control method. There were two levels of soil moisture treatment. One was normal irrigation, and the soil relative water content was always maintained at (75 ± 5) %. The second treatment was drought stress. Drought stress adopts natural drying method. When the soil moisture content was 75%, the watering was stopped (0 days) until the plant wilted (6 days). On the 7th day, the seedlings under drought stress were rehydrated. The phenotypic changes of plants during treatment were observed. In order to study the response of plants to short-term drought stress, fresh leaves were taken on the 1st, 2nd, 3rd, 4th, 5th and 6th days of drought stress treatment to determine physiological indexes. Samples of fresh leaves were taken during the drought stress on Days 0, 3, 4, 5, 6, as well as during the rehydration period on Days 7, 8 and 9; these samples were recorded as CK, T3, T4, T5, T6, T7, T8 and T9, respectively. The fresh leaves were frozen in liquid nitrogen and stored in an ultra-low temperature refrigerator at −80 °C. The changes of hormones and gene expression in plants were determined. Each treatment was repeated three times, with four pots per replicate. Pot seedlings are arranged randomly.
Determination of physiological indices
For each sample, relative conductivity was determined using a Mettler FiveEasy plus conductivity meter. 27 Chlorophyll content was calculated using a formula modified from the Arnon method, as reported in the study by Lichtenthaler et al. 28 Superoxide dismutase (SOD) activity was determined using the nitrogen blue tetrazolium method. 29 Peroxidase (POD) activity was determined using the method by Zhang et al., 30 and the sample's absorbance at 470 nm was subsequently calculated. CAT activity was determined after Shah et al., 31 and the sample's absorbance at 240 nm was subsequently calculated. The contents of abscisic acid (ABA), indole acetic acid (IAA), gibberellin (GA3), isopentenyl adenosine (IPA), and methyl jasmonate (JA-ME) were detected using an enzyme-linked immunosorbent assay.
RNA library construction and transcriptome sequencing
A Takara MiniBEST Plant RNA Extraction Kit (Takara, Japan) was used to extract total RNA from the CK, T3, T4, T5, T6, T7, T8 and T9 samples. The quality of the total RNA extracted was determined using 1% agarose gel electrophoresis. For sequencing, mRNA with Poly (A) of all samples was enriched from total RNA using oligo (dT) magnetic beads.32–34 The DNA probe was digested using DNase I after hybridisation with rRNA to obtain the purified RNA. RNA was then converted into short fragments using a fragmentation buffer. First-strand cDNA was synthesised using random N6 primers, followed by second-strand cDNA synthesis. The ends of double cDNA were repaired; 5′ ends were phosphorylated, and 3′ ends formed cohesive ends with A-tailing. Then, cDNA was ligated to the sequencing adapters. The ligation products were amplified using PCR to build a cDNA library and sequenced on the DNBSEQ platform (BGI Genomics, Shenzhen, China). The integrity and purity of RNA and the quality of the library were determined using the DNBSEQ sequencing platform.
Marker reference genome and gene annotation
To obtain clean reads of the raw data using the DNBSEQ platform, the reads containing the adapter were deleted to obtain clean reads / poly-N tails and low-quality data. We used HISAT to align clean reads to the reference group sequence, and used Bowtie2 to align clean reads to the reference gene sequence. The gene reference group was formatted to the ini-index group format of HISAT (http://www.ccb.jhu.edu/software/hisat) using a HISAT Burrows Wheeler transformation as well as the format of the Ferragina-Manz (FM) index (which includes the whole genome). This facilitated comparative analysis of genes with HISAT comparison graphs. The gene annotation file (reference genome version: GCF_000346465.2_Prunus_persica_NCBIv2) was downloaded directly from the BGI geneomics systems website (https://biosys.bgi.com/#/report/login).
Quantitative analysis of gene expression
We filtered the raw data to remove all low-quality reads (i.e. reads with joint contamination and a high content of unknown base N). Using HISAT v0.1.6, we beta positioned effective high-quality reads into the Prunus persica genome (https://www.ncbi.nlm.nih.gov/assembly/GCF_000346465.2) to facilitate further analysis. We used RSEM to calculate gene expression levels, and obtained the number of reads mapped to exons per 10 million bases in each million mapping reads (FPKM). We then screened for differences between genes (DEGs), standard conditions for |log2FC| 1 or higher, q-value 0.05 or less. Analyses of genetic differences were performed using genetic databases (GO) (http://bioinfo.cau.edu.cn/agriGO/analysis.php), the Kyoto Encyclopedia Gene and Genome database (KEGG) (https://www.kegg.jp/kegg/pathway.html) and MapMan (Vvnifera_14, http://mapman.gabipd.org/web/guest/mapmanstore).
Real-time fluorescence quantitative PCR analysis
In order to verify the accuracy and reliability of the transcriptome sequencing, we randomly selected four differentially expressed genes (PpADC, PpPI4 K, PpHSP, PpC5B) and analyzed whether their qRT-PCR results were consistent with the transcriptome results. A Takara MiniBEST Plant RNA Extraction Kit (Takara, Japan) was used to extract total RNA from the CK, T3, T4, T5, T6, T7, T8 and T9 samples. The above time points were completely synchronized with the enzyme activity determination period to analyze the correlation between enzyme activity changes and gene expression at the same time point. Single-stranded cDNA was synthesized using PrimeScript™ RT reagent Kit with gDNA Eraser (Perfect Real Time) (Takara, Otsu, Japan). And then, the SYBR® Premix Ex Taq™ II (Tli RNaseH Plus) (Takara, Otsu, Japan) was used to amplify and quantify the prepared cDNA. Primer 6.0 was used to design primers, and Actb was used as the internal reference gene. 35 The qRT-PCR primer sequences are shown in Table 1. The qPCR amplification was performed as follows: initial denaturation at 94 °C 5 min, followed by 32 cycles of denaturation at 94 °C for 30 s, annealing at 58 °C for 30 s, and extension at 72 °C for 1 min; after the final cycle, terminalelongation was performed at 72 °C for 10 min, and the samples were stored at 4 °C.The relative expression levels of the selected genes were calculated using the 2−ΔΔCT method. 36 The experiments were performed with three biological replicates and three technical replicates in each treatment.
Primers for qRT-PCR validation of DEGs.
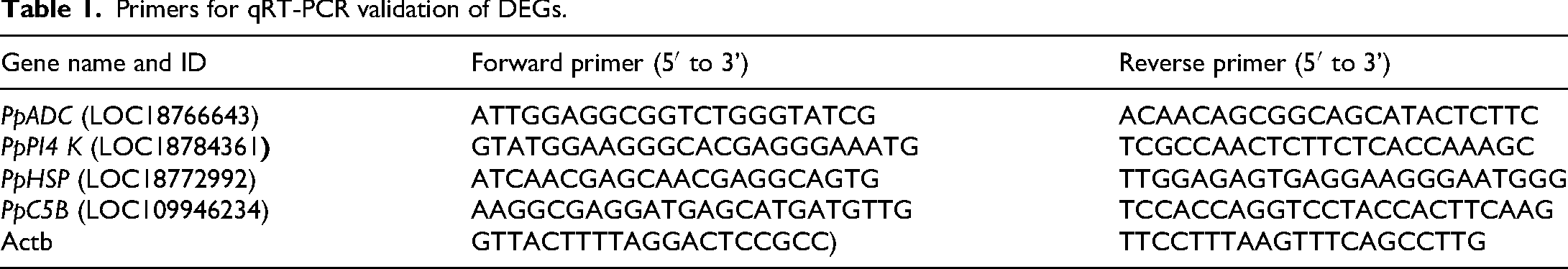
Statistical analyses
No a priori sample size calculation was performed. The number of biological replicates (n = 3 per group) was determined based on common practices in similar plant physiology studies (citation). While this sample size allows for preliminary statistical analysis, it should be noted that larger sample sizes may enhance the robustness of the results.
Experiments were conducted in a completely randomized design. Each analysis was conducted in triplicates. Results were presented as the means ± standard errors (SE) of three replicates. The significance of the differences among groups was determined according to Duncan's new complex range method.
Results
Phenotypic changes in peach seedlings experiencing drought stress
The peach seedlings started to wilt and gradually wilted until Day 6 (Figure 1). However, after being exposed to continuous rehydration in the three subsequent days, the plants essentially returned to normal growth. The time of severe drought symptoms of potted seedlings in this experiment was 6 days after drought stress.
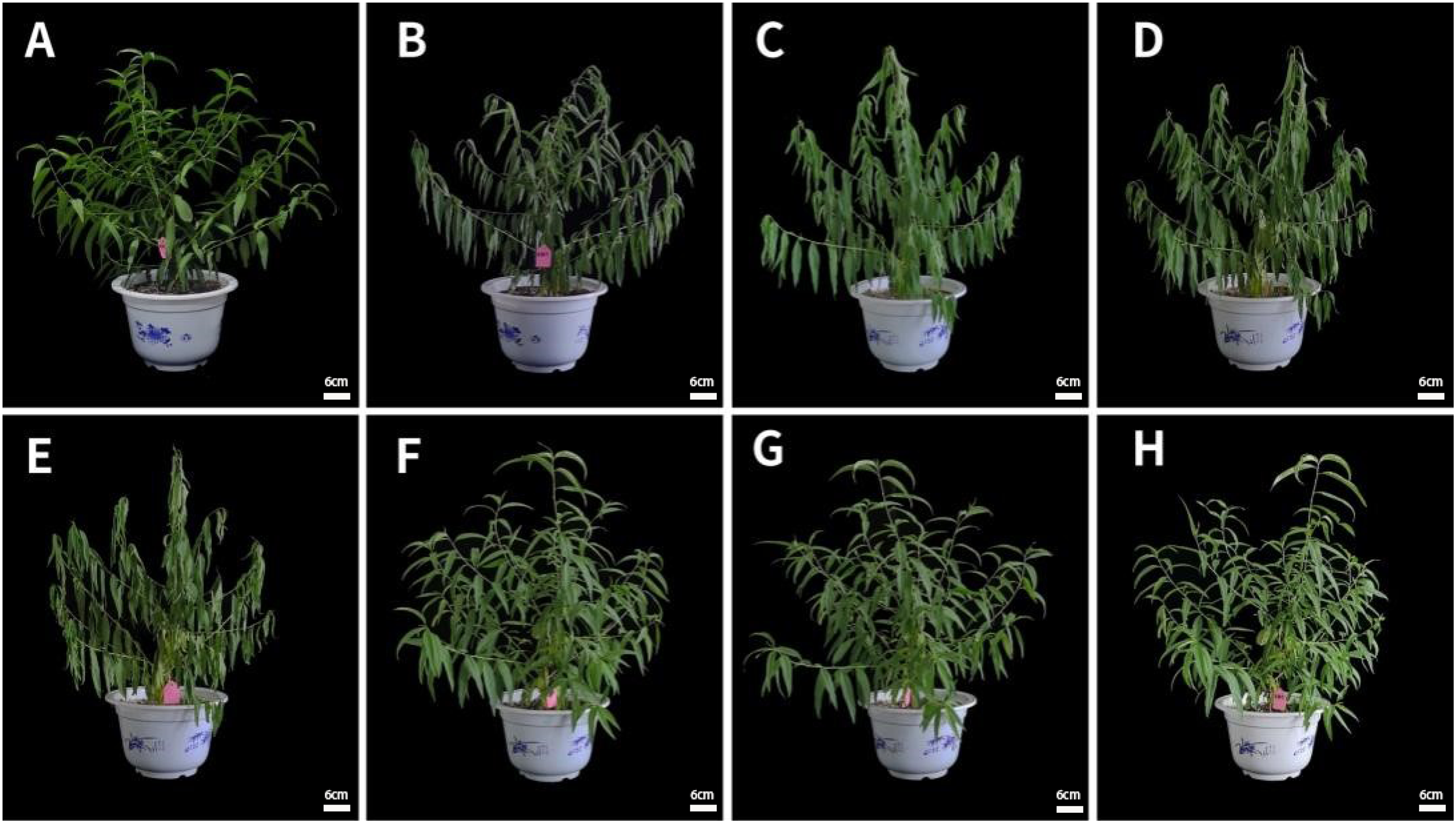

Phenotypic changes in a peach seedling throughout and after the drought stress. Note: (A) The plant under a normal watering regime (sample CK). (B)–(E): The plant on Days 3, 4, 5, and 6 of the drought stress (samples T3–T6). (F)–(H) The plant after one, two, and three days of rehydration (samples T7–T9).
Effect of drought stress on relative electrical conductivity of peach leaves
Stress can easily cause damage to a plant's cell membrane. This leads to exosmosis of the cell fluid and an increase in relative electrical conductivity. Hence, the relative electrical conductivity of plant can be measured to assess its membrane structure. We observed that the relative conductivity of the leaves of the peach plants peaked at 94.08% on Day 5 of the drought stress (Figure 2). Peach will have a short adaptation stage when suffering from water stress, and the relative electrical conductivity of peach leaves increases with the time of water stress. This may be related to the reactive oxygen species (ROS) produced by peach plants.

Changes in the relative electrical conductivity of peach leaves under drought stress.
Effects of drought stress on photosynthetic pigment content and enzyme activity in peach leaves
Drought stress causes plants to decreases their photosynthetic rate and photosynthetic intensity.37–39 We found that the contents of chlorophyll a, total chlorophyll and carotenoids in peach plants undergoing drought stress generally increased and subsequently decreased, while the contents of chlorophyll b generally increased over the experimental period (Figure 3). During the drought stress treatment, the chlorophyll a content of peach seedlings changed smoothly, and there was no significant difference. The content of chlorophyll b increased first and then decreased. By Day 4 of the drought stress, the carotenoid content of peach seedlings undergoing drought stress reached the highest value. Overall, these results indicate that the critical transition period was four days into the drought stress. By this time, the health of the peach seedlings had been critically impacted; the water content in leaf cells decreases, and the rate of chlorophyll synthesis slows down, thus affecting the formation of Ribosome. Protein synthesis is blocked, metabolism is slowed down, and chlorophyll biosynthesis is inhibited. Chlorophyll is decomposed, and chlorophyll content is reduced. 40
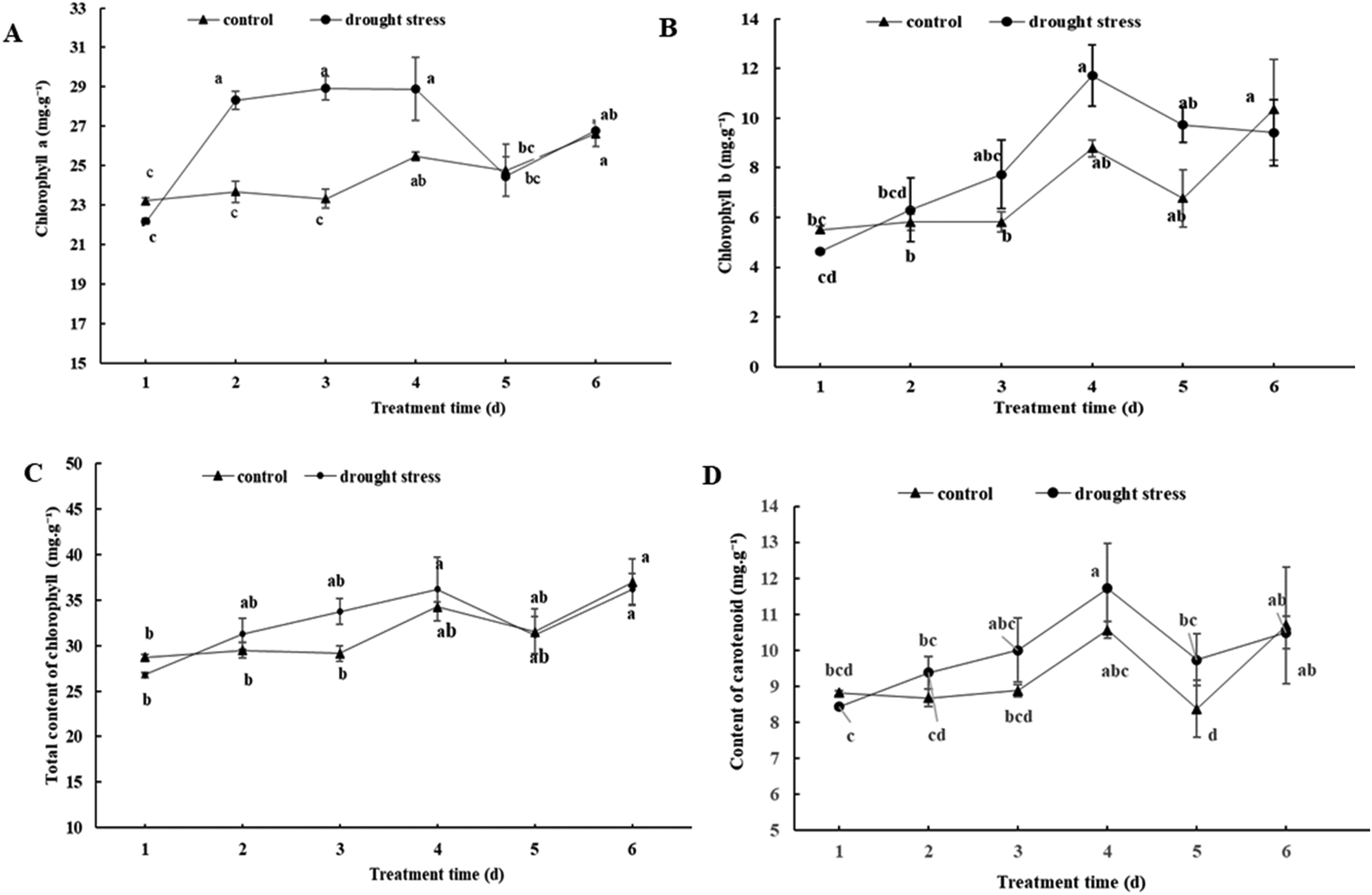
Changes in the chlorophyll a(A), chlorophyll b(B), total content of chlorophyll(C) and carotenoid contents(D) of peach leaves under drought stress. Mean values for each indicator marked with same lower case letter in each panel are not significantly different (P ≤ 0.05) according to Duncan's multiple range test.
The activity of SOD (Figure 4A) as well as POD (Figure 4B) in the peach leaves first decreased and subsequently increased over the period of drought stress, while the activity of CAT (Figure 4C) increased first and then decreased. The SOD activity of the peach leaves peaked at 384.18 U/g on Day 4 of the drought stress; this corresponded to a significant increase of 13.23% from the level of SOD activity in the peach leaves in the control. The SOD activity of the peach leaves had decreased sharply (by 38.32%) on Day 6 of the treatment. In general, SOD activity showed a substantial decrease from Day 4 to Day 6; it decreased by 34.48% between Day 4 and Day 5, and 91.22% between Day 5 and Day 6. The POD activity of the peach leaves also peaked on Day 4 of the drought stress at 13000 u/g; this corresponded to a 33.43% increase over the level of POD activity of peach leaves in the control. POD activity decreased by 34.48% between Day 4 and Day 5 of the drought stress.

Changes in the activity levels of SOD(A), POD(B) and CAT(C) in peach leaves experiencing drought stress.
The activity of CAT in the peach leaves peaked at 2503.11 u/g on Day 5 of the drought stress. Specifically, CAT activity significantly increased by 44.91% between Day 4 and Day 5, but significantly decreased by 61.68% between Day 5 and Day 6.
Effect of drought stress on the contents of endogenous hormones in peach leaves
The ABA content of the peach leaves increased continuously from Day 3 to Day 6 during the drought stress, the highest concentration was 110.318 ng/g·FW on the 6th day, before decreasing sharply over the next three days of rehydration(Figure 5A). The IAA content of the peach leaves fluctuated and increased over the course of the drought stress and peaked on Day 6(Figure 5B). Upon rehydration, the IAA content of the peach leaves decreased slightly. The GA3 content of the peach leaves peaked on Day 6 of the drought stress and decreased gradually after rehydration (Figure 5C). The IPA content of the peach leaves was generally constant over the course of the drought stress, and decreased gradually after rehydration (Figure 5D). The JA-ME content of the peach leaves increased at first, then decreased and then increased again under drought stress (Figure 5E). The JA-ME content increased continuously in the first two after rehydration before decreasing sharply on the third day. Trends in the contents of ABA, IAA, IPA, JA-ME and GA3 in the peach leaves under drought stress are shown in Figure 5. In particular, it is apparent that the content of ABA increased throughout the drought stress (Figure 5). We therefore suspect that changes in the levels of the endogenous hormone ABA may influence the expression of drought-related genes in peach plants experiencing drought stress.

Changes in the contents of ABA(A), IAA(B), GA3(C), IPA(D) and JA(E) in peach leaves under drought stress.
Identification of DEGs
A total of 21, 348 DEGs were screened from 20 intergroup comparisons. Of these DEGs, 10105 DEGs were up-regulated and 11243 DEGs were down-regulated. It was determined that 351 DEGs were up-regulated during the period of drought stress in comparison to the control (CK-T3, CK-T4, CK-T5, CK-T6). The Venn/UPSETR diagram in Figure 6 illustrates the numbers of DEGs that were common and unique to the various control-treatment comparison groups. A total of 2255 DEGs were common to the CK-T3, CK-T4, CK-T5, CK-T6, and CK-T7 groups, which had 252, 460, 426, 1306, and 764 unique DEGs, respectively. A total of 1225 DEGs were common among the CK-T3, CK-T6, CK-T9, T3-T6, and T3-T8 groups, which had 195, 703, 803, 133, and 1193 unique DEGs, respectively (Figure 6). As shown in Figure 6, in comparison with their state during the control, most DEGs tended to be upregulated as the duration of drought stress prolonged.
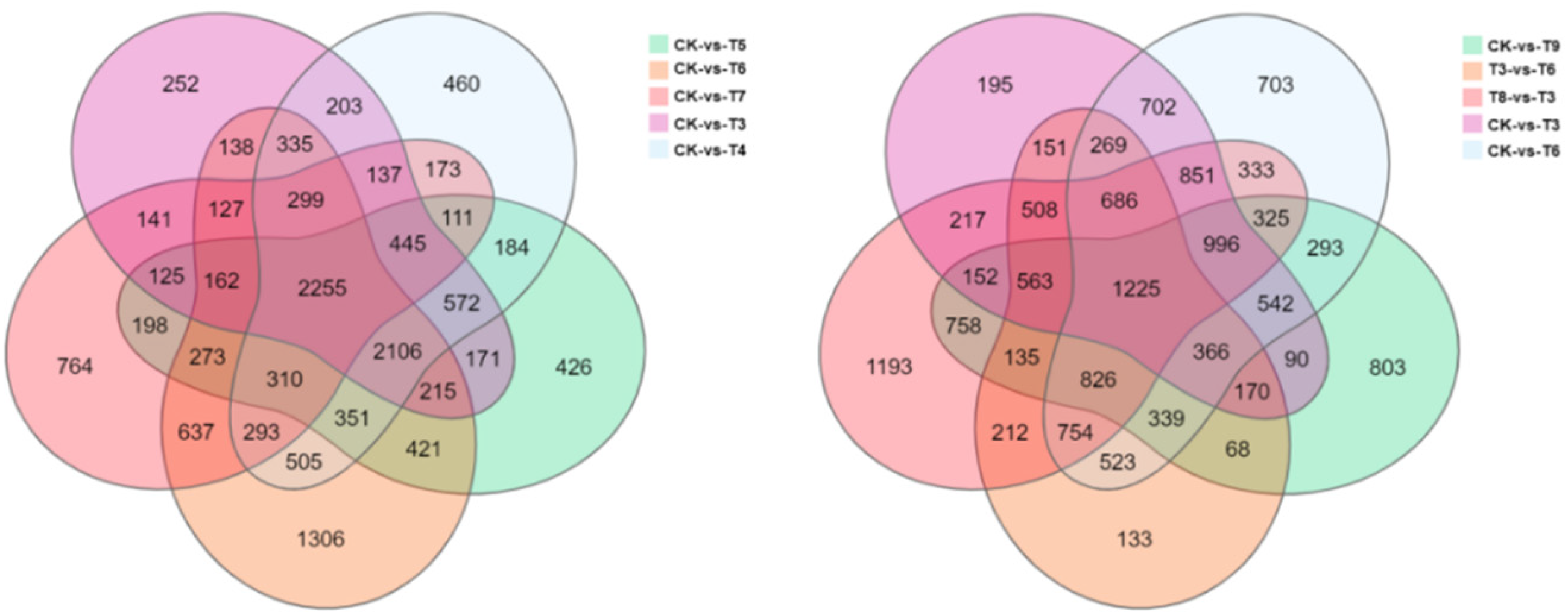
Numbers of common and unique DEGs among different control-treatment comparison groups. Note: Each polygon in the Venn diagram corresponds to a group of gene sets. Areas where different circles overlap indicate an intersection of gene sets, while non-overlapping areas indicate genes that are uniquely expressed. Numbers correspond to the number of genes.
Annotating the GO functions of DEGs
There are three different types of gene ontologies (GOs): molecular functions, cellular components, and biological processes. Under the domain of molecular functions, we found that the DEGs in peach plants in the drought stress (CK-T3, CK-T4, CK-T5, CK-T6) and in the rehydration process (CK-T7, CK-T8, and CK-T9) mainly served to enrich ATP binding (GO: 0005524), nucleotide binding (GO: 000166), transferase activity (GO: 0016740), hydrolase activity (GO: 0016787), catalytic activity (GO: 0003824), and GTP binding (GO: 0005525) (Figure 7). In terms of cell components, DEGs were mainly enriched in integral components of membrane (GO: 0016021), membrane (GO: 0016020), nucleus (GO: 0005634), cytoplasm (GO: 0005737) and chloroplast (GO: 0009507). In terms of biological processes, we found that the DEGs mainly enriched translation (GO: 0006412), phosphorylation (GO: 0016310), protein ubiquitination (GO: 0016567), RNA modification (GO: 0009451), responses to water deprivation (GO: 0009414) and other similar functions.
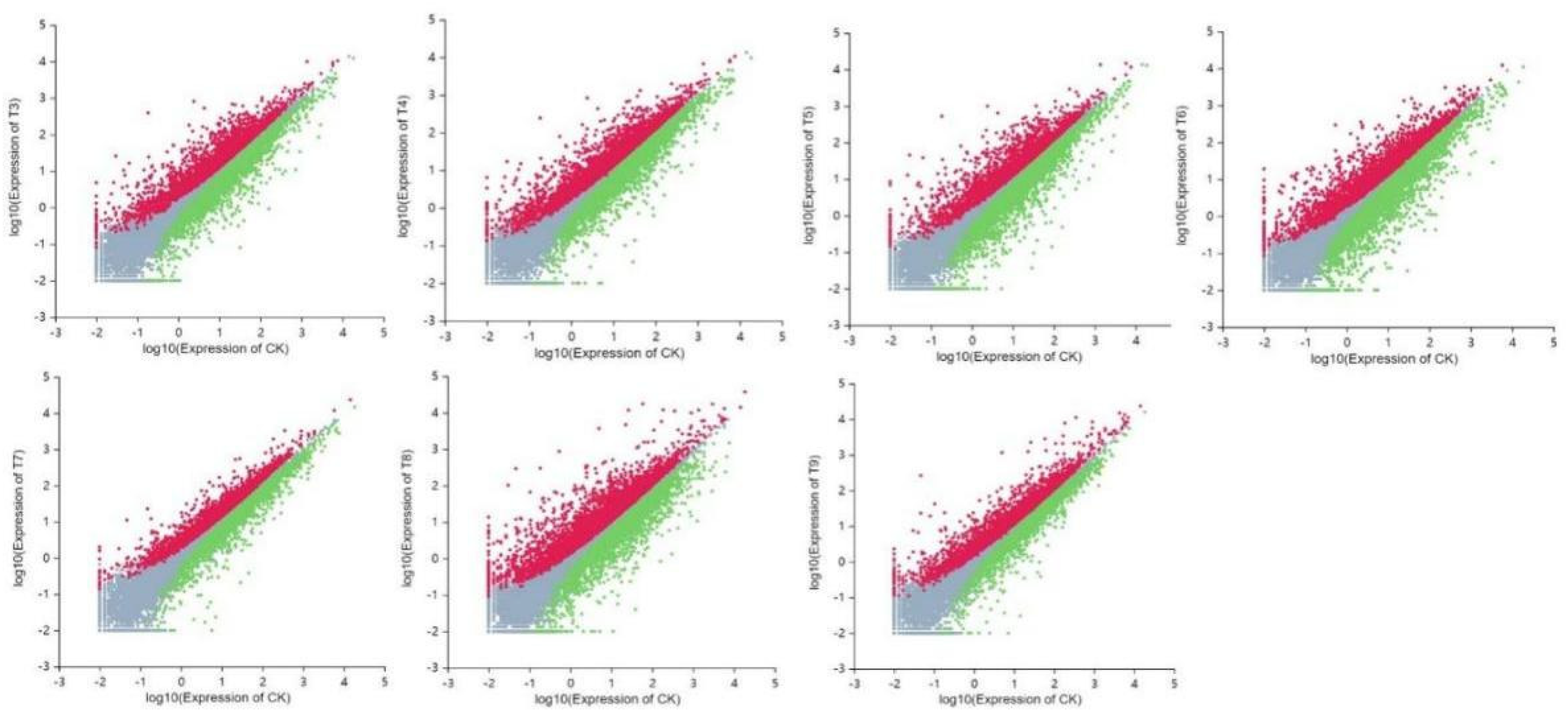
Scatterplots of differential genes between control-treatment comparison groups. Note: The x- and y- axes represent the scores of whole gene expression. Red dots indicate DEGs, blue dots indicate up-regulated DEGs, while green dots indicate non-DEGs.
Notes on the functions of DEGs from the KEGG
According to their notes in the KEGG, the DEGs of peach plants served to enrich the biosynthesis of amino acids and metabolism of carbon during the drought stress (CK-T3, CK-T4, CK-T5, CK-T6) and rehydration process (CK-T7, CK-T8, and CK-T9). Specifically, the EDGs enriched processes such as plant hormone signal transduction, ribosome, protein processing in endoplasmic reticulum, Spliceosome, plant-pathogen interaction, MAPK signaling pathway – pathogen interaction plant, Ubiquitin mediated proteolysis, peroxisome.
KEGG is enriched in amino acid biosynthesis and metabolism after drought stress and rehydration, and encodes genes related to the biosynthesis of arginine, as well as the metabolism of arginine and proline. The biosynthesis of arginine is mainly encoded by the L-Glutamine synthase gene (PpGS), the glyoxyl aminotransferase gene (PpGGAT), the δ-1-pyrrolidine-5-carboxylate synthase aminoacylase 1 gene (PpACY1), the aspartate transaminase gene (PpASP), the amino acid acetyltransferase gene (PpNAGS2), the acetylornithine deacetylase gene (PpNAO), the L-argininosuccinic acid synthase gene (PpASS), and the arginase 1 gene (PpARG1). The metabolism of arginine and proline is mainly encoded by the imidase gene (PpAS), the δ-1-pyrrolidine-5-carboxylate synthase gene (PpP5CS), the ornithine aminotransferase gene (PpOAT), the n-carbamylputamide acylhydrolase gene (PpCPA), the proline imidase peptidase gene (PpPLD), the guanidine imidase deimidase gene (PpAGDI), the huminyl 4-carboxylase 10 gene (PpP4H10), the huminyl 4-carboxylase 1 gene (PpP4H1), the spermidine decarboxylase gene (PpADC), the pyrrolidine-5-carboxylase reductase gene (PpP5CR), and the aminase gene (PpAS1).
The genes involved in arginine biosynthesis and the metabolism of arginine and proline were PpGS, PpGDH, PpASP, PpACY1, PpALT, PpASS, PpPAO2, PpP4H7, PpP5CR, PpP5CS, PpADC, and PpP4H4. These findings provide a candidate set of genes that are up-regulated in peach plants during drought stress, and which can be further screened to investigate the adaptive drought responses of this species.
We found that two CAT genes (gene IDs: 18776773, 1877304) and six SOD genes in peach plants may regulate their responses to drought stress. During the drought stress, the expression of both the above CAT genes and three SOD genes (gene IDs: 18786713, 18775015, 18789179, 18772363, 18774382, 18784827) were up-regulated. In particular, the expression of the SOD gene (gene ID: 18784827) was significantly up-regulated on Day 6 of the drought stress (T6, corresponding to severe stress), but became down-regulated after rehydration.
Plants undergoing abiotic stresses sense these environmental signals and transmit them to organelles to activate adaptive responses. We found that the activity of zeaxanthin cyclooxygenase (ZEP), a key enzyme involved in ABA synthesis, was up-regulated during drought stress and down-regulated following rehydration. The activity of ABA aldehyde oxidase (AAO) was up-regulated during drought stress, and showed an increasing-decreasing-increasing trend after rehydration. The activities of NCED and ZEP were up-regulated on Day 6 of the drought stress (i.e. a period of severe drought stress), but down-regulated on the first day after rehydration (T7). These results suggest that NCED (gene ID: 18792639), NCED (gene ID: 18780753), and ZEP (gene ID: 18770963) are key enzymes involved in the biosynthesis of ABA in peach plants responding to drought stress.
Real-time fluorescent quantitative PCR verification
To verify the results of the transcriptome sequencing, we randomly selected four genes—PpHSP (gene ID: 18773996), PpC5B (gene ID: 109946234), PpPI4 K (gene ID: 18784361), and PPADC (gene ID: 18766643)—for real-time qRT-PCR detection. We found that the trend of qRT-PCR expression for these genes was largely consistent with the RNA-seq data. This indicated that the results of transcriptome sequencing were accurate and reliable (Figure 8).

qRT-PCR identification based on transcriptomics. Note: a. b, c and d are transcriptome data, and a1, b1, c1 and d1 are validation data.
Discussion
The response of a plant to drought stress is essentially a complex biological process, which not only involves photosynthesis, osmotic regulation, the metabolism of antioxidants, plant hormones, and sugars, but also involves a variety of signal factors, transcription factors, and functional genes. 11 Elucidating these complex molecular mechanisms can advance understanding of the drought resistance of plant species. 41 The expression of DREB gene family is up-regulated when plants are subjected to drought stress. By regulating a series of functional genes downstream, plants can produce osmotic adjustment substances, change stomatal conductance, etc., and enhance drought resistance. 42 A previous study found that Pinus yunnanensis seedlings experiencing drought stress displayed significantly different physiological responses in comparison with those in a control. 43 Likewise, in the present study, we observed that the physiological indices of the leaves of peach seedlings under drought conditions differed from those in control conditions. We therefore sought to elucidate the molecular mechanisms determining these responses of peach seedlings to drought stress by transcriptome sequencing and a comparative analysis of the leaves of the seedlings under drought and control conditions. The study of differentially expressed genes (DEGs) in transcriptomics is to determine which genes play a major role in response to abiotic stress by comparing gene expression profiles under different tissues and conditions, 44 which provides a basis for further study of molecular responses of plants to environmental stress.45,46 Manohar et al. 47 took tall fescue plants and endophytes as research objects, and compared the transcriptome data of leaves, pseudostems, crowns and roots under drought stress. It was found that the main enrichment pathways of DEGs in other tissues except for the crown were photosynthesis, carbohydrate metabolism, plant hormone biosynthesis and signal transduction, cell tissue and transcriptional regulation. Maria 48 analysis showed that the expression of SOD and CAT genes increased under drought stress, indicating that the activation of antioxidant system was related to the response mechanism of species to drought stress. Willian et al. 49 studied the close relationship between amino acid metabolism and stress response. Our GO and KEGG enrichment analysis showed that there were 21348 differentially expressed genes in peach seedlings experiencing drought stress. This finding substantially enriches the available information on the genes of peach plants. Our annotation of gene functions showed that most differentially expressed genes were involved in the regulation of enzyme activity (e.g. SOD and CAT), the amino acid metabolism pathway, and plant hormone signal transduction.
The metabolism of amino acids plays an important role in the growth and development of plants. It involves many different metabolic pathways and acts as a precursor for the synthesis of numerous signal molecules involved in various intermediate pathways. Previous studies have shown that in tea plants, drought stress induces the expression of pyrroline-5-carboxylic acid synthase gene (P5CS), pyrroline-5-carboxylic acid reductase gene (P5CR), and γ-glutamyl phosphate reductase gene (GRR). 50 Similarly, in this study, the DEGs in peach plants experiencing drought stress were identified to be involved in the metabolism of amino acids. In particular, we observed that genes related to the biosynthesis of arginine as well as the metabolism of arginine and proline was significantly up-regulated on the sixth day of drought stress. The up-regulated expression of these genes likely strengthened the resistance of the peach plants to the stressful external environment.
Plants suffering stress typically produce a large number of reactive oxygen species that pose a threat to their cell structure and various biological processes. 41 Drought stress, in particular, can trigger an increase in the rate of ROS production. In addition to serving as a second messenger, ROS can induce oxidative damage during periods of drought. The ROS balance is related to photosynthesis, the metabolism of phenylalanine and the peroxidase pathway. Plant peroxisomes constitute a key component of the cellular antioxidant system, which acts to remove excessive ROS in a plant through a variety of metabolic pathways. 51 We observed dynamic changes in the activities of antioxidant enzymes in peach plants subjected to drought stress. Previous studies have found that the activities of CAT, SOD, and POD in the seedlings of fruit-bearing plants, 52 oat, 53 corn, 54 and potato 55 experiencing drought stress were higher than those in control conditions. Likewise, the activities of SOD, POD, CAT, APX, and GR in the leaves of Perilla frutescens, 56 Chinese seabuckthorn, 57 Dendrobium officinale, 58 lily, 59 and rice 60 first increased and subsequently decreased during periods of drought stress. In conclusion, our results together with these previous observations indicate that the activities of CAT, SOD, and POD constitute a coordinated response to drought stress. In particular, our results suggest that POD and SOD play a major role under mild and moderate levels of drought stress, while CAT plays a major role under severe levels of stress, where it acts to reduce membrane permeability and to maintain the integrity of the membrane structure. 61 Studies have shown that plants experiencing drought stress do not rely on singular enzyme systems to remove the ROS they produce under stress, but instead coordinate two or more enzymes in their enzyme systems to alleviate the damages caused by ROS, thereby improving their drought tolerances overall. 62 In the present study, we found that the expression of CAT (gene IDs: 18776773, 18777304) and SOD (gene IDs: 18772363, 18774382, 18784827) in peach seedlings was up-regulated during periods of drought stress. Similar responses in sea buckthorn were observed by Ye et al. 63 These findings indicate that peach seedlings experiencing drought stress remove ROS and alleviate drought-induced damages by enhancing the activity of CAT and SOD.
Although ABA has broad functions in plant growth and development, its main function is to regulate plant water balance and osmotic stress tolerance. 64 ABA can also control a plant responses to drought stress by influencing transcription, as well as through the post-transcriptional modification of downstream regulators in the signaling pathway. 65 ABA therefore enhances a plant's resistance to biotic and abiotic environmental stressors. In a previous study on apple plants responding to stress, ABA-related genes involved in signal transduction such as PYR / PYL and PP2C were found to be highly expressed. 66 The results of the present study are consistent with those of previous studies. Our analysis of the transcriptome data from peach plants showed that several genes encoding for enzymes involved in the ABA signaling pathway were stimulated during periods of drought stress. This included the up-regulated expression of ZEP, PP2C, and ABF genes, which may have occurred due to the rapid accumulation and activation of endogenous abscisic acids. Our results therefore indicate that the up-regulation of ABA signal transduction may improve the resistance of peach plants to drought stress. Overall, our results showed that key enzymes involved in ABA synthesis, namely zeaxanthin cyclooxygenase (ZEP), 9-cis-epoxycarotenoid dioxygenase (NCED), and ABA aldehyde oxidase (AAO), were up-regulated during periods of drought stress. These results were also consistent with the results of Li et al. 67
One limitation of this study is the absence of a priori sample size calculation. The sample size (n = 3 biological replicates) was selected based on methodological conventions rather than statistical power analysis. Although the results show significant trends (e.g. p < 0.05 for ABA accumulation), the limited sample size may increase the risk of Type II errors (false negatives), particularly for subtle phenotypic or gene expression changes. Future studies with larger sample sizes and power-based designs are recommended to validate these findings. In addition, this study simulated drought stress through water control in greenhouse pots, which may not fully replicate the natural conditions in the field. The controlled environment might have underestimated the complex stress responses that occur in natural ecosystems, potentially limiting the extrapolation of results to real-world drought scenarios. In later experiments, the results can be verified in field experiments under natural drought conditions or more complex stress models (e.g. shelters) can be used to simulate ecological stress.
Conclusion
The physiological responses of peach seedlings experiencing drought stress were significantly different from those in control conditions. A total of 21348 DEGs were identified based on transcriptome differences in peach seedlings experiencing drought stress and those in control conditions. Of these DEGs, 10105 DEGs were up-regulated and 11243 DEGs were down-regulated. A functional annotation showed that genes involved in the biosynthesis and metabolism of amino acids, peroxidase, and plant hormone signal transduction may be important in determining the responses of peach plants to drought stress. When nine genes relating to drought-stress responses were screened, the expression level of the PpADC gene showed a unique decreasing-increasing-decreasing trend over the period of drought stress. This suggests that the PpADC gene is involved in the responses of peach plants to drought stress. The findings provide a theoretical basis for the cloning and functional analysis of genes conferring drought resistance, and the cultivation of more drought resistant varieties of peach.
Footnotes
Acknowledgements
The authors would like to thank TopEdit (www. topeditsci. com) for its linguistic assistance during the preparation of this manuscript.
Ethical considerations
No ethical approval was required for this study, as it involved physiological and molecular analyses of Prunus davidiana seedlings, which do not involve human subjects, endangered species, or animal experimentation. All plant materials were collected from non-protected areas in compliance with local regulations.
Author contributions
Conceptualization, R.Z., B.W.; methodology, R.Z., B.W. and L.Z.; validation, B.W.; formal analysis, R.Z., B.W., Q.L. and L.Z.; investigation, S.L. and L.Z.; data curation, R.Z. and L.Z.; writing-original draft preparation, L.Z.; writing-review and editing, R.Z. and Q.L.; visualization, B.W.; supervision, R.Z. All authors have read and agreed to the published version of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Key Research and Development Program of China, Henan Provincial Department of Science and Technology Research Project, (grant number 2018YFD1000300, 232102111087).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding authors.
