Abstract
Objectives
Constitutive promoters have been widely studied and utilized in silkworm resistance breeding and expression of beneficial proteins, but there is limited research comparing the activities of different promoters across various source cell.
Methods
This study screened three promoters, namely, the ie1 gene promoter (ie1p) of Bombyx mori nuclear polyhedrosis virus (BmNPV), the cytoplasmic actin 3 promoter (A3p) and the cytoplasmic actin 4 promoter (A4p). Enhancer fragments hr3 and hr5 were combined with the three promoters to driving the expression of luciferase (luc+) and vacuolar-type ATPase c (Vac). The constructed plasmids were transfected into Sf9, 293 T and BmN cells, and the transient expression of target genes were assessed.
Results
The luc+ activity assays revealed that the enhancer effects of hr3 and hr5 fragments followed a similar trend in both Sf9 and 293 T cell lines. Specifically, hr3 significantly enhanced the ie1p promoter, increasing its activity by 2.44-fold and 2.90-fold, respectively. Similarly, hr5 markedly boosted the A4p promoter, with enhancements of 2.45-fold and 3.00-folds, respectively. There were differences in the relative highest expression levels of Vac gene among the three cell lines, hr5 had a significant effect on all three promoters in Sf9 cells.
Conclusion
This research aims to identify highly active promoters in silkworm cells, refine transgenic methodologies and strategies, and lay the experimental groundwork for developing a transgenic silkworm technology platform.
Introduction
A promoter is defined as the upstream regulatory sequence of the gene that is recognized and bound by RNA polymerase. Based on their function and mode of action, promoter is categorized into three types: constitutive promoters, specific promoters and inducible promoters. Constitutive promoters regulate the expression of structural genes at a constant level, leading to minimal variation in RNA and protein expression across different tissues, and exhibiting no spatiotemporal specificity. These promoters are extensively utilized in genetic improvements in animal and plant.1,2 Bombyx mori, an economically significant and model insect, 3 has been widely employed in genetic transformation, more recently, as a transient expression system, where promoters play a crucial role. Constitutive promoters are primarily utilized in the expression of resistance genes,4,5 the construction of auxiliary plasmids for genetic transformation, 6 and the selection of initiation markers in silkworm genetic control technology.7,8 Different constitutive promoters in silkworm exhibit varying driving activities in vivo or in vitro, 9 making the exploration of efficient promoters crucial for enhancing transgenic efficiency and disease resistance in silkworm.
Bombyx mori cytoplasmic actin gene family comprises four members (actin1 to actin4, A1-A4). 10 Among them, A3 and A4 are constitutive genes responsible for expressing cytoskeletal components in silkworm.11,12 Their upstream promoter sequence can drive the expression of foreign genes, with the A3 promoter (A3p) also exhibiting activity in mammalian cells. 13 A4 is closely related to A3, and the two genes arose from the recent duplication of an intron-containing ancestral gene. Similar to A3, A4 is expressed in various tissues, its mRNA level differs from those of A3.12,14 Using A4IP-Gal4/UAS-EGFP transgenic system, the A4 promoter (A4p) was shown to drive the expression of green fluorescent protein (eGFP) gene in various tissues of the silkworm. 15 The immediate early-1 (ie1) gene, an early expression gene of the BmNPV, plays a critical role in viral infection. Its promoter (ie1p) is active in bacteria, animal, plant cells and is widely used. 16 The ie1p drives the transcription of foreign genes in various tissues and developmental stages of insects, and regulates gene expression without tissue and spatiotemporal specificity. 17 The homologous region hr3 and hr5 are homologous repeat sequences in the BmNPV genome. The hr3 fragment has an enhancing effect on the promoter of the silkworm sericin 1 (ser1) gene, 18 and the two enhancer fragments have varying degrees of enhancing effect on BmNPV ie1p.19,20 The three constitutive promoters, ie1p, A3p and A4p, have been compared and analyzed in the silkworm ovary cell line BmN, but not in the Sf9 and 293 T cell lines. To understand the differences in activities of ie1p, A3p, and A4p in different cell lines and the enhancing effect of the hrs fragment on them, a vector linking the promoter to the red fluorescent protein (DsRed) gene was constructed and transfected into Sf9 cells. Additionally, nine recombinant expression plasmids containing ie1p, A3p, A4p, hr5-ie1p, hr3-A3p, hr3-A4p, hr5-ie1p, hr5-A3p, and hr5-A4p promoters were generated using pGL3-Basic vector. These plasmids were transfected into Sf9, 293 T and BmN cells, and their activity was investigated by detecting luc+ and Vac transient expression levels. The study aims to identify highly expressed promoters in silkworm cells, providing an experimental foundation for establishing a transgenic silkworm technology platform.
Materials and methods
Materials and reagents
The silkworm varieties Dazao (P50) and BmNPV (Baoshan strain) are preserved by the Institute of Sericulture and Apiculture, Yunnan Academy of Agricultural Science. The plasmid pSL1180 [ie1p-DsRed-SV40 af] containing the ie1p promoter fragment, the Vac gene existing in pMD19-Vac plasmid, the pGL3-Basic (Promega, USA) vector containing luc+ and the internal reference plasmid pRL-TK (Promega, USA) were provided by State Key Laboratory of Resource Insect (Southwest University, China). Sf9: Spodoptera frugiperda ovary cell (RRID: CVCL_0549), BmN: Bombyx mori ovary cell (RRID: CVCL_Z088) two cell lines were preserved in our laboratory. The restrictive endonuclease, Ex Taq enzyme and DNA ligase were purchased from Takara. TC-100 cell culture medium, HEK293T: human embryonic kidney cell line (RRID: CVCL_0063), fetal bovine serum (FBS, Gibco. BRL) and Cellfectin™ II Reagent were purchased from ThermoFisher Scientific (USA). DMEM was purchased from Biological Industries (Israel). The dual luciferase detection kit was purchased from YEASEN Biology (Shanghai, China).
Construction of target gene plasmids
The silkworm (P50) tissue was fully ground with liquid nitrogen, and then added to a centrifuge tube containing 1 mL of genomic extract (10 mmol/L Tris-HCl, pH 8.0; 0.1 mol/L EDTA, pH 8.0; 0.5% SDS). Proteinase K (Sangon Biotech, China) was added, and digested in a 55 °C water bath for 2 h. The solution was extracted with Tris-saturated phenol buffer, phenol-chloroform-isoamyl alcohol (25:24:1), and chloroform once each. The genomic DNA was precipitated with anhydrous ethanol, DNA was washed with 70% ethanol, dissolved in Tris-EDTA (TE) buffer and stored at −20 °C for later use. The genome of BmNPV-Baoshan was extracted following the same protocol and similar in TE buffer. The extracted silkworm genome and BmNPV genome were used for cloning promoter and hrs fragment, respectively. For the construction of the promoter activity assay plasmid, the pGL3-Basic vector was employed, the validated A3p and A4p driving activity regions as references.13,15 Relevant primers were designed using Primer 5.0 (Table 1), and hr3 and hr5 fragment amplification primers were designed based on the genome sequence of BmNPV-Baoshan strain (Accession number: MT501299). The A3p and A4p promoter sequences were obtained from the silkworm genome PCR. These primers contained EcoR I, BamH I, Xho I and Hind III restriction sites. The hr3 and hr5 fragments were amplified through PCR using the BmNPV-Baoshan genome as a template, with the primers incorporating Kpn I and Sma I restriction. For validation of the activity of the ie1p promoter, Xho I and Hind III cleavage sites were introduced. After enzyme digestion, A3p and A4p fragments were connected to the vector pSL1180 [ie1p-DsRed-SV40 af] upstream of the DsRed gene to form recombinant plasmids pSL1180 [A3p-DsRed-SV40 af] and pSL1180 [A4p-DsRed-SV40 af]. Using the same method, the promoter fragments ie1p, A3p, and A4p were connected to the pGL3-basic vector to form recombinant plasmids pGL3-ie1p-luc+, pGL3-A3p-luc+, and pGL3-A4p-luc + . Subsequently, the hr3 and hr5 fragments were cloned upstream of the three promoter fragments, respectively. By using Hind III and Xba I enzyme digestion, the luc+ gene was replaced with the Vac gene in the pGL3-promoter-luc+ vector, ultimately constructing the pGL3-promoter-Vac vector. The plasmids were sequenced by Sangon Biotech to confirm the accuracy of the target fragment sequences.
The forward and reverse primer sequences for promoter amplification.

DF/DR: Upstream and downstream primer sequences of the promoter of the red fluorescent protein gene; LF/LR: Upstream and downstream primer sequences of the promoter of the luc+ gene; The enzyme cleavage site is underlined; Lowercase letters represent protective bases.
The Sf9 and BmN cell lines were maintained TC-100 complete medium containing 10% FBS and 1% double antibiotics at 28°Ccondition. For transfection experiments, Sf9 cells were seeded into 12 cm2 flasks at a density of about 1 × 105 cells/mL and cultured for 48 h. Prior to transfection, the culture medium was aspirated, and the cells were washed three times with serum-free TC-100 medium. For preparing 100 μL cell transfection mixture, 5 μL CellfectinTM II Reagent, 2 μg recombinant plasmid and appropriate volume of culture medium was added, and kept at room temperature for 20–30 min. Thereafter, the mixture was added dropwise to the cell culture bottle and incubated at 27 °C for 5 h. Then, the co-transfection solution was discarded, 2 mL of complete culture medium was added, and the cells were cultured for 3–5 days. The expression of red fluorescence under the control of the ie1p, A3p, and A4p promoters was observed with an inverted fluorescence microscope (IX73, Olympus, Japan), Cell transfection efficiency and DsRed fluorescence intensity were detected by flow cytometry (CytoFLEX S, Beckman Coulter, USA). The transfected plasmids ie1p/A3p/A4p-luc+/Vac, hr3-ie1p/A3P/A4p-luc+/Vac, hr5-ie1p/A3p/A4p-luc+/Vac were used for luc+ activity assay and Vac gene expression. 293 T cell culture and transfection: 293 T cells were cultured in DMEM supplemented with 10% FBS under standard conditions (37°C, 5% CO2). For transfection, cells were seeded at a density of 5 × 105 cells/mL in a 6-well plate and cultured for 48 h. The transfection protocol and luciferase activity assay for vectors were the same as described above.
Cell culture and transient transfections
The transfected plasmids ie1p/A3p/A4p-luc+/Vac, hr3-ie1p/A3p/A4p-luc+/Vac, hr5-ie1p/A3p/A4p-luc+/Vac were used for luc+ activity assay and Vac gene expression. 293 T cell culture and transfection: 293 T cells were cultured in DMEM supplemented with 10% FBS under standard conditions (37°C, 5% CO2). For transfection, cells were seeded at a density of 5 × 105 cells/mL in a 6-well plate and cultured for 48 h. The transfection protocol and luciferase activity assay for vectors were the same as described above.
Luciferase assay
Using pGL3-Basic (270 ng/μL) + pRL-TK (283 ng/μL) as the blank control group and different promoter fragments pGL3-luc++pRL-TK as the experimental group, three independent transfection experiments were conducted, and each transfection was repeated three times. After 48 h of transfection, cell lysates were collected, washed once with phosphate-buffered saline (PBS, pH7.4), and the cells were treated with lysis buffer for luc+ determination. Luc+ activity was determined according to the instructions of the detection kit (YEASEN). The data of three independent experiments were expressed as means ± SEM. A two-way analysis of variance (ANOVA) was used to compare the ie1p, A3p and A4p data as well as the promoter enhancement effect of hr3 or hr5 fragments. The pairwise comparisons for various parameters (relative luciferase activities) among three promoters (ie1p, A3p and A4p) was done after post-hoc analysis using Tukey's multiple comparison test of the corresponding factor using main effects in a one-way ANOVA. The significance between hr3 and hr5 enhancement effect was estimated by student's t-test. One-way analysis of variance (ANOVA) and student's t-tests were performed using GraphPad Prism 8 to generate P-values. Statistical significance was assessed using the following criteria (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001).
Quantitative reverse transcription PCR (Rt-qPCR)
The pGL3-promoter-Vac vectors were transiently transfected into three cell lines. Approximately 48 h transient transfection, total RNAs from different cells were prepared using RNA Extraction Kit (Servicebio, Wuhan, China). Total RNAs were prepared using the PrimeScript™ RT reagent Kit with gDNA Eraser (Perfect Real Time) (TaKaRa, Japan) and reverse transcribed into cDNA. To analyze activity difference of promoter, relative expression level of Vac gene was determined by RT-qPCR. The specific primers of Vac (forward 5′-CGGCGTCTGCTATCATCTTCA-3′ and reverse 5′- CACGCACGCCTGCATCTC-3′) were used for RT-qPCR. Primers of the house-keeping gene, Bmactin3 (forward 5′- CCGTATGCGAAAGGAAATCA-3′ and reverse 5′-TTGGAAGGTAGAGAGGGAGG-3′). RT-qPCR reactions were prepared with a SYBR Premix Ex TaqTM II (Tli RNaseH Plus) (TaKaRa, Japan), according to the manufacturer's instructions. In brief, The RT-qPCR reaction was carried out in a 20 μL reaction mixture containing 10 μL of SYBR Premix Ex Taq, 8 μL of ddH2O, 0.4 μL of ROX Reference Dye, 0.8 μL of forward primer, 0.8 μL of reverse primer and 1 μL of cDNA template. The reactions were carried out in the StepOnePlus Real-Time System (Life technologies, USA). The reaction process consisted of an initial denaturation at 95 °C for 5 min and 40 cycles at 95 °C for 15 s and 60 °C for 30 s. Each expression assay was performed in triplicate, and Ct values for both the target gene and internal reference gene were recorded for subsequent data analysis in the experimental and control groups. Relative expression levels were calculated, using the 2−△△Ct method. 21 Statistical analysis was performed using GraphPad Prism 8. The data of three independent experiments were expressed as means ± SEM. A two-way analysis of variance (ANOVA) was used to compare promoter data. The pairwise comparisons for various parameters (relative luciferase activities) among three promoters in a one-way ANOVA. The significance between hr3 and hr5 enhancement effect was estimated by student's t-test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Results
Construction of transient expression plasmids
Using the silkworm (P50) genome as a template, the promoter sequences of the actin3 (A3p) and actin4 (A4p) genes were amplified by PCR. Similarly, the hr3 and hr5 enhancer fragments were obtained using the BmNPV-Baoshan genome as a temple. The A3p and A4p were connected to the DsRed gene vector pSL1180 [ie1p-DsRed-SV40 af]. The promoters and enhancer fragments were cloned to pGL3-Basic luc+/Vac gene vector (Refer to Figure S1 for the schematic diagram of plasmid construction), and sequencing was performed for verification (see Table S1 for details).
Verification of promoter-driven activity
The ie1p-DsRed, A3p-DsRed, and A4p-DsRed plasmids were transfected into Sf9 cells. After 24 h, the transfected cells did not show any red fluorescence. After 48 h, red fluorescence was detected in cells transfected with the three plasmids (Figure 1), indicating that the cloned promoter fragment had driving activity. Flow cytometry showed no significant differences in transfection efficiency or fluorescent expression (DsRed) in Sf9 cells among the three plasmids (Figure S2, Table S2 and Table S3).
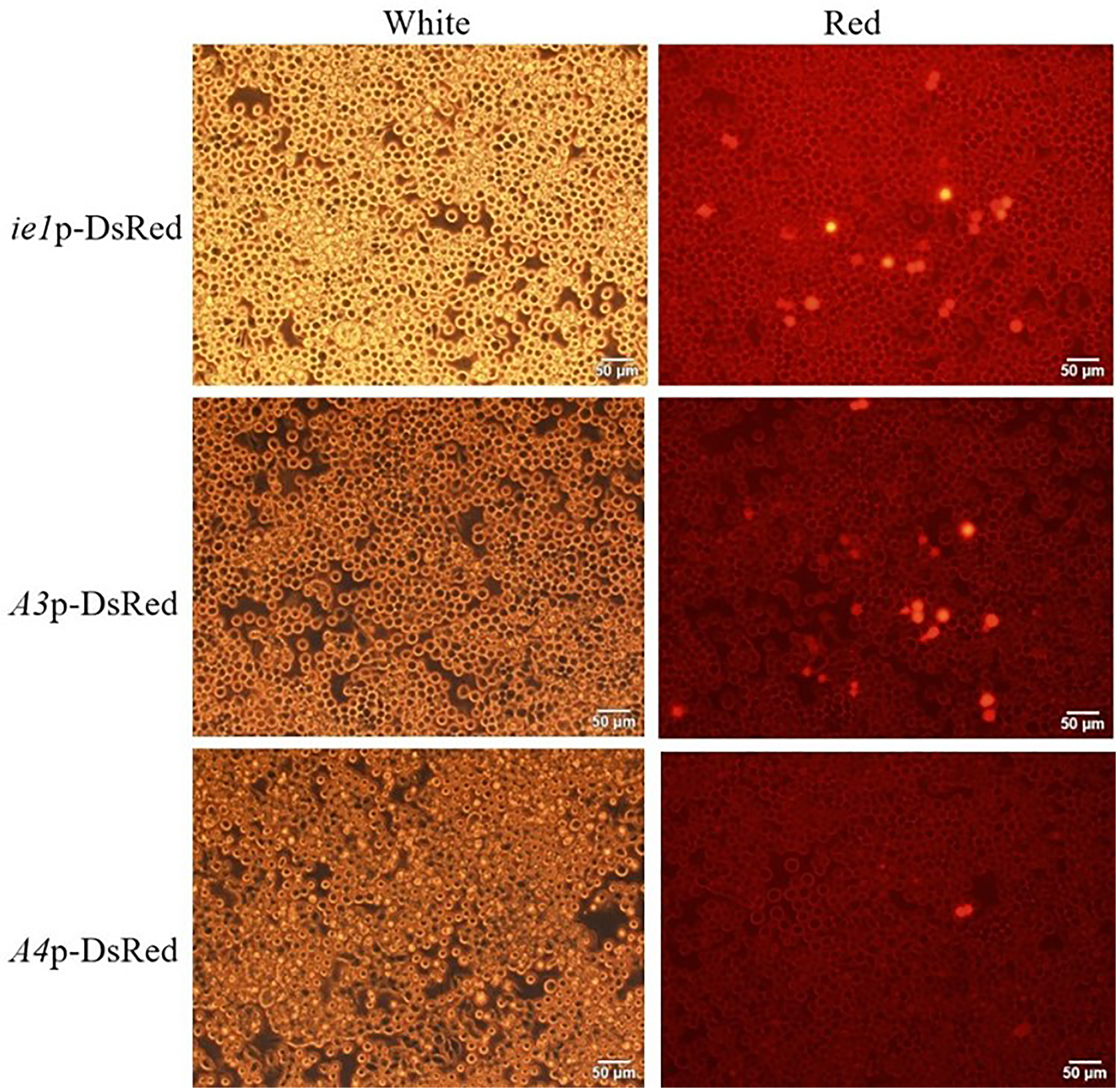
Verification of DsRed expression activity driven by three constitutive promoters. White: Transfected with the three plasmids and detected under white light. Red: Transfected with three plasmids and detected under red fluorescent light. All results were conducted under the same observation conditions. The time of exposure was 800 ms for all picture. Images were acquired at 2000 magnifications (200× objective with 10× eyepiece; scale bar = 50 μm).
Transient expression analysis in Sf9, 293 T and BmN cells
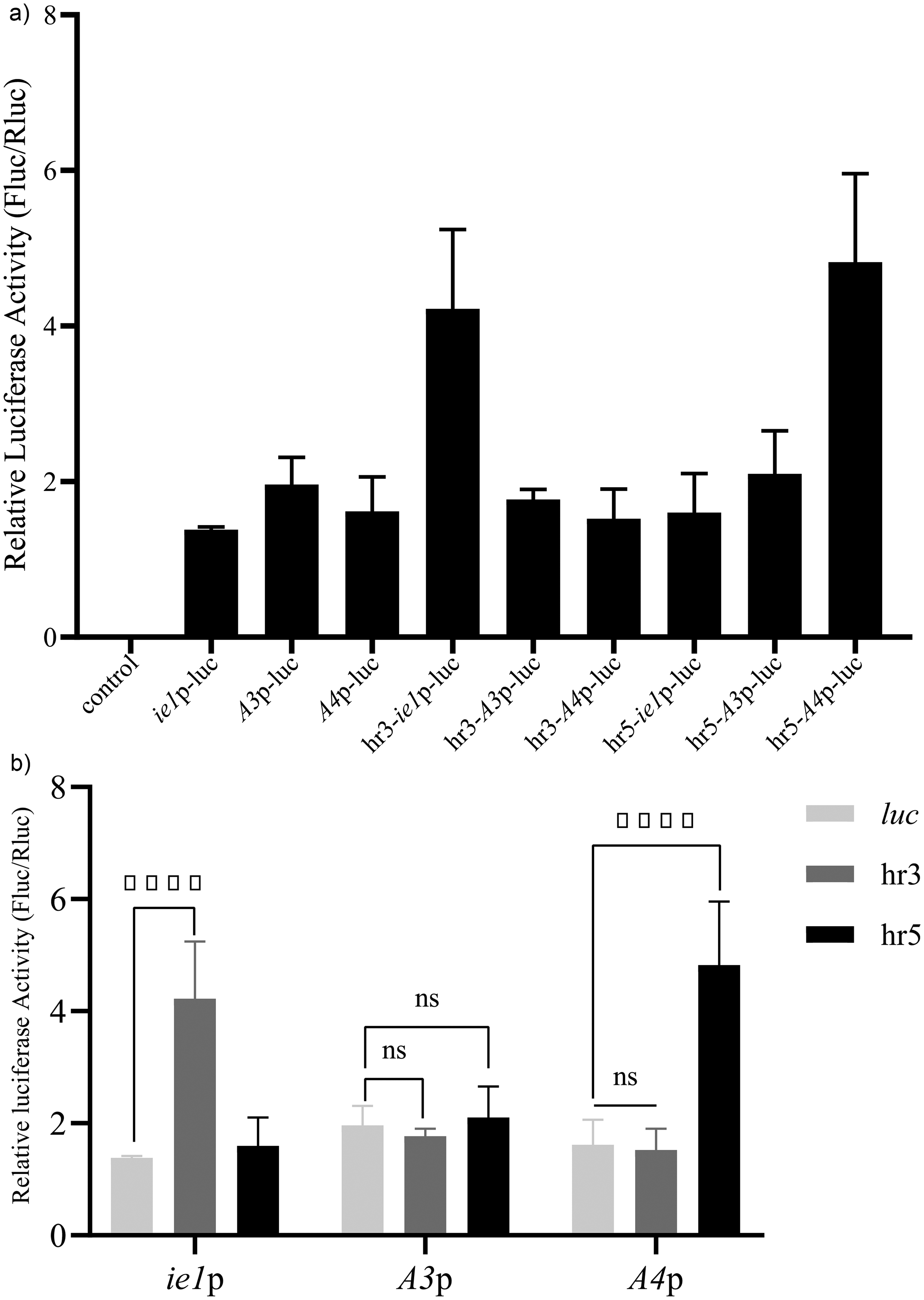
Co-transfection of pGL3-Basic empty vector and nine constructed reporter plasmids into Sf9 and 293 T cells showed transient expression activity of the luc+ reporter gene in both cell types (Figure 2(a) and Figure 3(a)), while hr3 and hr5 enhanced the driving activity of the promoter (Figure 2(a) and Figure 3(b)). The promoter activities of ie1p, A4p, and A3p were sequentially enhanced, and the enhancement effect of hr3 and hr5 fragments on the promoter was similar in both cell lines. The enhancement effect of hr3 fragment on A4p, A3p, and ie1p was sequentially increased, whereas the enhancement effect of hr5 fragment was sequentially weakened. The hr3 fragment had the most pronounced enhancement effect on ie1p, while the hr5 fragment exhibited a more significant enhancement effect on A4p. Nine plasmids containing the Vac gene (pGL3-promoter-Vac) were transfected into Sf9, 293 T, and BmN cells simultaneously with pGL3-Basic empty vector. Transient expression activity of the Vac gene was detected in all three cell species (Figure 4a1, a2, and a3), and hr3 and hr5 enhanced the driving activity of the promoters in three types of cell lines (Figure 4b1, b2, and b3), hr3 and hr5 can enhance the driving activity of A4p promoter in three types of cells; In Sf9 cells, hr5 enhances the expression of ie1p, A3p, and A4p promoters by 11.76 times, 5.96 times, and 35 times, respectively; In 293 T cells, hr3 enhances all three promoters by 1.63 fold, 2.61 fold, and 1.68 fold, respectively; In BmN cells, both hr3 and hr5 have significant enhancing effects on the ie1p promoter, at 13.58 times and 61.61 times levels, respectively.

Comparison of luciferase reporter gene activity driven by ie1p, A3p and A4p promoters in Sf9 cells. a. Comparison of luc+ activity driven by three promoters with or without hrs. b. Analysis of the enhancing effect of hr3 and hr5 on three promoters. Relative luciferase activity (Fluc/Rluc) represents the relative expression level measured after transient transfection of recombinant plasmids for 48 h. Control: Plasmid pGL3-basic was used as negative control. The error bars represent the standard deviation of three replicates, and each experiment was independently repeated three times. The student's t-test was used to evaluate statistical significance (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001), ns: not significant.
Comparison of luciferase reporter gene activity driven by ie1p, A3p and A4p promoters in 293 T cells. a. Comparison of luc+ activity driven by three promoters with or without hrs. b. Analysis of the enhancing effect of hr3 and hr5 on three promoters. Relative luciferase activity (Fluc/Rluc) represents the relative expression level measured after transient transfection of recombinant plasmids for 48 h. Triple experiments were performed to calculate the values for relative activity of Fluc/Rluc. Control: Plasmid pGL3-basic was used as negative control. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.

Transcriptional level comparison of vac gene driven by ie1p, A3p and A4p promoters in Sf9, 293 T and BmN cells. a. Relative expression level of Vac gene by three promoters with or without hrs. a1: Sf9 cells, a2: 293 T cells, a3: BmN cells. b. Comparison of enhancing effect of hr3 and hr5 enhancer fragments on the three promoters. b1: Sf9 cells, b2: 293 T cells, b3: BmN cells. Relative expression level measured after transient transfection of recombinant plasmids for 48 h. Triple experiments were performed to calculate the values for relative levels of Vac transcripts. Control: The group refers to untransfected cells without plasmid, serving as a blank control. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Discussion
In this study, we evaluated the activities of Bombyx mori A3p, A4p and BmNPV ie1p promoters. The three promoters have been previously reported to regulate target genes in various tissues of Bombyx mori.7,15,22 To verify the driving activity of the cloned A3p and A4p promoters in the ovary cell line Sf9 of Spodoptera frugiperda, ie1p-DsRed was used as the positive control. Following transfection of A3p-DsRed and A4p-DsRed constructs into Sf9 cells, red fluorescence was observed 48 h post- transfection. Notably, all three promoters also showed driven activity in 293 T cell line. These findings suggest that the three constitutive promoters can be widely used for heterologous gene expression and functional verification. After 48 h of transfection with the three promoters carrying the DsRed gene, red fluorescence was observed in Sf9 cells. Flow cytometric analysis demonstrated that neither the transfection efficiency of the three DsRed-encoding plasmids nor their fluorescence intensity in Sf9 cells exhibited statistically significant differences. For the three promoters without the influence of hr fragments, red fluorescence apparent relative intensities, although could not be quantified, roughly seemed to agree with the luciferase activity of corresponding luciferase expression vectors observed in the Sf9 cells, with the expressing vector containing hr3-ie1p and hr5-A4p being stronger (Figure 2, Table S4). In the dual luciferase reporter gene assay system (DLRTM), the relative luciferase expression of the three promoters was measured in the two cell lines at 48 h. The promoter activity of A3p, A4p and ie1p was evaluated. Zhang et al. compared the activity of the actin3 gene promoter and the ie1 gene promoter using recombinant AcMNPV, and found that the activity of A3p was 10-fold stronger than the ie1p, 17 this comparable difference was consistent with the author's research findings, but not as significant. The A3 promoter (BmA3p) of the silkworm, Bombyx mori, was used for the first time for stable genetic transformation in silkworms. 7 The BmA3p also plays a constitutive role in the cotton bollworm (Pectinophora gossypiella) 23 and in the Hymenoptera insect sawfly (Athalia rosae). 24 Although the Bombyx mori A3 promoter successfully drives the expression of marker genes in heterologous insects, its activity is species-dependent. It is speculated that the activity of Bombyx mori A4 promoter in Sf9 cells may be lower than that of the silkworm tissue-derived cell line. 15 In this study, the activity of three promoters was assessed at the 48 h time point through red fluorescence observation and luc+ quantitative analysis. Due to the length of cloned promoter fragments or the source of the selected cell line in the experiments, the detection results may differ from previous reports. 25 The Sf9 cell line established from the ovarian tissue of Spodoptera frugiperda does not have cell totipotency, the target gene was expressed instantly in vitro, and the expression level of the gene is influenced by multiple factors.
The hrs sequence has been proven to be the start site of viral DNA replication.26,27 As a cis-regulatory element of the early baculovirus promoter, hrs plays an enhanced role, such as 39 K,28,29 vp39 30 and ie-N,31,32 as well as constitutive promoters.18,33 In this study, the enhancing effects of hr3 and hr5 on the three promoters ie1p, A3p and A4p was evaluated, both hr3 and hr5 enhanced the driving activity of the promoters (Figure 2, Figure 3 and Figure 4). Specifically, the promoter activities of ie1p, A4p and A3p were sequentially increased, as in detailed in Table S4. The trend of promoter enhancement by hr3 and hr5 fragments was the same in Sf9 and 293 T cell lines. Notably, the hr3 fragment had the most significant enhancement effect on ie1p, while the hr5 fragment had a more significant enhancement effect on A4p. Furthermore, the expression level of luc+ had increased 2–4 times between Sf9 and 293 T cells, indicating a robust and consistent enhancement effect mediated by these hr fragments.
The construction of various promoter fragment plasmids in DLRTM detection system provides a robust tool for investigating promoter regulation and assessing the impact of different cis-regulatory elements on transcription. This system serves as an in vitro expression platform to analyze promoter activity, with findings that can substantiate promoter-driven in vivo expression of target genes. In insects, constitutive promoters can drive the expression of marker genes throughout the larva and pupa stages.34,35 Given that BmNPV hrs significantly increase the transcription of constitutive promoters, their integration with promoters can amplify gene expression. Applications of enhancers in entomological research, the combination of hr3 with BmNPV 39Kp and A4p has been shown to markedly boost promoter activity, thereby elevating the expression level of the target gene Hyphantria cunea NPV hycu-ep32 (an antiviral gene), which enhances the disease resistance of silkworms. 36 Additionally, inserting the hr5 fragment into the promoters of ie1, SfHSC70-P1780, and SfPub-P2009 genes significantly increased promoter activity, resulting in higher luciferase expression. When fall armyworm larvae were infected with a recombinant baculovirus containing these promoters to drive EGFP expression, a notable enhancement in fluorescence across the entire larval body was observed. 33 This study highlights the differential regulatory roles of hr3 and hr5 in modulating promoter activity, providing valuable insights into their potential applications in gene expression regulation.
The reliance on plasmid transfection introduces potential variability due to differences in transfection efficiency across cell populations. While we standardized protocols to minimize inconsistencies, uneven plasmid uptake may affect observed phenotypic outcomes. Alternative approaches, such as viral transduction or the use of clonal cell lines, could improve reproducibility in future studies. A potential limitation is the reliance on transient transfection, which may result in heterogeneous expression. Although this approach was suitable for our initial goals, future studies employing stable cell lines would reduce variability.
Conclusion
Concluding, BmNPV hrs demonstrated a notable enhancing effect on constitutive promoters. The luc+ was used for promoter activity validation, the enhancing effects of hrs were consistent in Sf9 and 293 T cells. Hr3 had a significant enhancement effect on the ie1p promoter, while hr5 exhibited a pronounced effect on the A4p promoter. When comparing Sf9 and BmN cells, with Vac served as the target gene, the enhancement effect of hrs fragments on the promoter were also consistent, with hr5 showing a stronger effect on the ie1p promoter. Overall, the A4p promoter display relatively stable activity, and both hr3 and hr5 effectively enhanced its performance. This study systematically compared the activities of three constitutive promoters, utilizing hrs fragments to enhance their driving efficiency. The findings provide valuable insights for identifying optimal promoter combinations, which can serve as a reference for the expression of insect-resistant genes and beneficial proteins in future applications.
Supplemental Material
sj-docx-1-sci-10.1177_00368504251347466 - Supplemental material for Construction of synthetic constitutive promoters and evaluation of their expression strengths in Sf9, HEK293T, and BmN cells
Supplemental material, sj-docx-1-sci-10.1177_00368504251347466 for Construction of synthetic constitutive promoters and evaluation of their expression strengths in Sf9, HEK293T, and BmN cells by Yonghong Zhang, Jiafu Luo, Yulan Shao, Zhengqin Wang and Hongrui Zhang in Science Progress
Supplemental Material
sj-docx-2-sci-10.1177_00368504251347466 - Supplemental material for Construction of synthetic constitutive promoters and evaluation of their expression strengths in Sf9, HEK293T, and BmN cells
Supplemental material, sj-docx-2-sci-10.1177_00368504251347466 for Construction of synthetic constitutive promoters and evaluation of their expression strengths in Sf9, HEK293T, and BmN cells by Yonghong Zhang, Jiafu Luo, Yulan Shao, Zhengqin Wang and Hongrui Zhang in Science Progress
Supplemental Material
sj-docx-3-sci-10.1177_00368504251347466 - Supplemental material for Construction of synthetic constitutive promoters and evaluation of their expression strengths in Sf9, HEK293T, and BmN cells
Supplemental material, sj-docx-3-sci-10.1177_00368504251347466 for Construction of synthetic constitutive promoters and evaluation of their expression strengths in Sf9, HEK293T, and BmN cells by Yonghong Zhang, Jiafu Luo, Yulan Shao, Zhengqin Wang and Hongrui Zhang in Science Progress
Supplemental Material
sj-docx-4-sci-10.1177_00368504251347466 - Supplemental material for Construction of synthetic constitutive promoters and evaluation of their expression strengths in Sf9, HEK293T, and BmN cells
Supplemental material, sj-docx-4-sci-10.1177_00368504251347466 for Construction of synthetic constitutive promoters and evaluation of their expression strengths in Sf9, HEK293T, and BmN cells by Yonghong Zhang, Jiafu Luo, Yulan Shao, Zhengqin Wang and Hongrui Zhang in Science Progress
Footnotes
Acknowledgments
The author sincerely acknowledged the Sericulture and Apiculture Research Institute, Yunnan Academy of Agricultural Science, which provided necessary experimental facilities for this study.
ORCID iD
Yonghong Zhang https://orcid.org/0000-0003-2552-8998
Author contributions
Conceived and designed the study: Yonghong Zhang. Experiment operation: Yonghong Zhang. Analysis of data: Jiafu Luo and Zhengqin Wang. Help to proofread the manuscript: Yulan Shao. Drafted the work: Hongrui Zhang. All the authors have read and approved to the published version of the manuscript.
Funding
This research was supported by Pre Research Project of Yunnan Academy of Agricultural Sciences (Grant number: 2024KYZX-01).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The datasets used and/or analyzed during the current study available from the corresponding author on reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
