Abstract
Aptamers have attracted exceptional attention in medical field due to their intrinsic properties equivalent to antibodies such as high target affinity, low immunogenicity and toxicity, cost-effectiveness and ease of synthesis and modification, and good stability under extreme conditions, thereby providing new avenues for basic research and clinical application. Protein tyrosine kinase 7 (PTK7) has been proved to be closely linked with the progression of many types of cancer. The aberrant expression of PTK7 has positioned it as a potential theranostic biomarker for multiple cancers. Aptamer sgc8 was initially identified for its high-affinity binding to PTK7 on the T-cell acute lymphoblastic leukemia cell line (CCRF-CEM) through cell-SELEX (systematic evolution of ligands by exponential enrichment) and subsequently has demonstrated the ability to effectively recognize many types of cancer cells that express PTK7 oncogenic target. The easily modifiable nature of sgc8 facilitates its conjugation with functional agents and drugs. This identification mode and modification approach of aptamers against cancer cells provides a potential strategy for cancer diagnosis and treatment. In this review, we discuss the potential of sgc8 aptamers in early cancer diagnosis and targeted therapy, focusing specifically on their interaction with the oncogenic biomarker PTK7.
Introduction
Precision medicine refers to the development of individualized diagnosis and treatment strategies for patients based on personal genomic information in combination with proteomics and other molecular pathological characteristics. 1 As next generation of diagnosis and treatment, the concept of precision medicine is of epoch-making significance and has brought significant changes in contemporary medical practice. And the targeted diagnosis and therapy, serve as a vital component of precision medicine, prompting substantial efforts towards the development of targeted drugs. 2 Research and clinical trials over the past few decades have consistently demonstrated the advantage of targeted therapy, offering great promise for improving the cure rate of diseases especially in malignant cancers and enhancing patient care.
While advancements in diagnostic and therapeutic modalities have significantly improved patient outcomes, notable limitations still remain. The challenges associated with current diagnostic methods include insufficient sensitivity and specificity, limited access to instruments, high costs, and invasiveness of certain procedures. Similarly, the limitations of current therapeutic approaches encompass side effects, treatment resistance, and off-target effects. In the ongoing battle against cancer, targeted therapy has emerged as an alternative to traditional therapies, such as surgery, radiotherapy and chemotherapy. These conventional methods have brought severe side effects due to their inability to discriminate diseased cells from normal ones. However, the implementation of targeted drug delivery systems, which facilitate the direct delivery of specific therapeutic agents to the intended target, can mitigate the issue of unintended damage. Aptamers have emerged as a vital component in the field of targeted diagnosis and therapy, paving a new way for precise diagnosis and treatment. The emergency of cell-SELEX (systematic evolution of ligands by exponential enrichment) seems to largely promote the aptamer development by allowing the selection of ligands from target cells without prior knowledge of target molecules. By using this method, cell-SELEX enables the identification of aptamers that specifically recognize molecular signatures present on the surface of target cells. This technique has been particularly impactful in cancer research, where it has been used to identify aptamers, such as sgc8, can specifically bind to PTK7, which is highly expressed in various cancer cell types, and subsequently undergoes cellular internalization without affecting cell viability. 3 The high specificity of sgc8 allows it to function as a versatile tool in cancer diagnostics and therapeutics, enabling precise targeting and imaging of PTK7-positive cancer cells. Its remarkable selectivity and high binding affinity position sgc8 as a promising candidate for advanced applications in targeted cancer treatment and diagnostic strategies. The data were sourced from PubMed, The Cancer Genome Atlas (TCGA) and The Human Protein Atlas database. This review is guided by the scale for the assessment of narrative review articles (SANRA). 4
Aptamer and SELEX
Nucleic acid aptamers, including single strand DNA (ssDNA) or single strand RNA (ssRNA), possess unique secondary or three-dimensional conformations that enable them to specially bind to targets ranging from small molecules5,6 to various types of cancer cells.7–9 These aptamers have been exploited as versatile molecular probes for cancer diagnostics and treatment. Aptamers typically consist of fewer than 100 nucleotides, allowing for small molecular size compared to antibodies. This smaller size simplifies and reduces the cost of synthesis, while still enabling the aptamers to adopt their specific conformations necessary for target binding.
The selection of aptamers from a huge number of random nucleic acid sequence library involves a complex and iterative in vitro screening process termed SELEX.10,11 Although there may be slight variations in the methods used by different laboratories, they generally follow a similar procedure (Figure 1). Initially, the oligonucleotide library is incubated with a target molecule. Subsequently, sequences with low binding affinity are eliminated through washing, while the sequences exhibiting relatively high binding affinity are collected and amplified using polymerase chain reaction (PCR). This iterative process is repeated for multiple rounds to oligonucleotides are then sequenced and the binding affinity between these oligonucleotides and the target will be determined. This technique allows for the identification of aptamers with high affinity and specificity that bind to targets across different classes, including small molecules, infected cells, and cancer cells.11,12 The small size of aptamers facilitates the rapid tissue penetration and efficient clearance from the body. Additionally, aptamers offer several other advantages, such as effortless selection and chemical modification, high stability, and low immunogenicity. Based on these advantages, aptamers have been applied in diverse areas of target therapeutics, including in vitro bioanalysis, in vitro diagnostics (IVD), bioimaging, biomarker discovery, and aptamer-aided drug discovery.5,13–15 Moreover, it's well worth noting that aptamers can be easily chemically modified to confer the ability to tolerate and recover from extreme conditions. These inherent advantages make aptamer-drug conjugates (ApDCs) a more promising approach compared to traditional therapy. In the latest years, substantial research has been conducted on the application of ApDCs in biomedical research and therapeutics, yielding remarkable achievements.

Scheme of cell-SELEX. An oligonucleotide library is initially incubated with cells express a target molecule. Following incubation, oligonucleotide sequences with low binding affinity are removed through washing, while those with relatively high binding affinity are collected and amplified using polymerase chain reaction (PCR). This iterative process is repeated for multiple times to progressively enrich for sequences with high affinity and specificity. The enriched oligonucleotides are then sequenced and the binding affinity between these oligonucleotides and the target molecule is measured. Negative selection is further used to remove oligonucleotides binding to control cells. The figure was created using curve.
Cell-SELEX
With the advancements in aptamer screening technology, cell-based SELEX offers a promising method to identify cell surface biomarkers even without prior knowledge of gene expression signature of the target cells. In contrast to regular SELEX, cell-SELEX method involves using cells as screening target instead of a specific molecular target. As illustrated in Figure 1, the cell-SELEX process begins with the generation of a randomized single-stranded DNA (ssDNA) or RNA library, typically comprising 1014 to 1016 unique sequences. Each contains a central randomized region flanked by fixed sequences to facilitate PCR amplification. This library is then incubated with target cells, which are typically live cancer cells in their native state, preserving surface markers and conformations. During this step, aptamers with specific binding affinity interact with their target molecules. Unbound and weakly bound sequences are removed through washing steps, leaving high-affinity aptamers attached to the target cells. Bound aptamers are subsequently eluted from target cells. To enhance specificity, a counter-selection step is performed by incubating the eluted aptamer pool with non-target cells, typically normal cells, to eliminate sequences that bind to shared or nonspecific markers. Aptamers that do not bind to the non-target cells are retained and amplified through PCR. The amplified sequences will undergo multiple iterative cycles (usually 12–16 rounds). After the SELEX procedure, the enriched aptamer pool is sequenced to identify individual sequences, high-affinity aptamers are further characterized using binding assays to determine their affinity and specificity.16,17 Furthermore, the biomarker recognized by selected aptamer can effectively be identified via affinity chromatography combined with mass spectrometry (MS).18–20
The development of cell-SELEX has greatly facilitated the generation of many DNA aptamers that exhibit specific binding to various types of cancer cells. Some examples include aptamer III.1, which recognizes rat glioblastoma cells through specific binding to pigpen protein expressed on the cell surface. 21 In another case, aptamer sgc8 was found to recognize human colorectal carcinoma (HCT116), with U87MG used as a negative control. 22 GBI-10, which recognizes human glioma U252 cells via tenascin-C. 23 This selection process has proved to effectively identify biomarkers on the cell surface of tumor cells of clinical patients, even in the absence of detailed prior knowledge about the targets. Moreover, the aptamers selected through cell-SELEX can serve as specific probes not only for cancer diagnosis but also for the detection and characterization of various other diseases. These aptamers hold great potential in advancing the field of diagnostics and can play a crucial role in enhancing the accuracy and efficiency of disease detection and monitoring.
PTK7
PTK7 upregulation in multiple cancers
Aberrant expression of PTK7 has been observed in multiple cancers. In the study of the expression of protein tyrosine kinases (PTKs) in the progress of melanoma, PTK7 mRNA has been detected in normal melanocytes at a considerable level. However, PTK7 was found downregulated during melanoma development, and associated with metastatic progression. 9 In spite of downregulation in melanoma, more evidence have shown that high level of PTK7 expression is associated with various types of cancer such as colon cancer,9,24 gastric cancer,9,25 lung cancer,9,26 acute myeloid leukemia9,27 and triple-negative breast cancer (TNBC).9,28,29 So, we analyzed The Cancer Genome Atlas (TCGA) database and assessed PTK7 mRNA levels in more than 30 cancers, significant upregulation of PTK7 occurred in 23 cancers (Figure 2), suggesting its significant oncogenic role in tumorigenesis. Therefore, PTK7 is a potential biomarker for cancer diagnosis and treatment, as well as promising drug targets.

The expression of PTK7 in various cancers and cell lines. (a) Upregulation of PTK7 occur in various type of tumors. Data derived from The Cancer Genome Atlas (TCGA) database. *p-value < 0.05, **p-value <0.01, ***p-value <0.001. The figure was created with TIMER2.0. ACC, adrenocortical carcinoma; BLCA, bladder urothelial carcinoma; BRCA, breast invasive carcinoma; CESC, cervical cancer; CHOL, cholangiocarcinoma; COAD, colon adenocarcinoma; DLBC, lymphoid Neoplasm Diffuse Large B-cell Lymphoma; ESCA, esophageal carcinoma; GBM, glioblastoma multiforme; HNSC, head and Neck Squamous Cell Carcinoma; KICH; kidney Chromophobe; KIRC; kidney renal clear cell carcinoma; KIRP, kidney renal papillary cell carcinoma; LAML, acute Myeloid Leukemia; LIHC; liver hepatocellular carcinoma; LUAD, lung Adenocarcinoma; LUSC, lung squamous cell carcinoma; MESO, mesothelioma; OV, Ovarian serous cystadenocarcinoma; PAAD, pancreatic adenocarcinoma; PCPG, pheochromocytoma and paraganglioma; PRAD, prostate adenocarcinoma; READ, rectum adenocarcinoma; SARC, sarcoma; SKCM,skin cutaneous melanoma; STAD, stomach adenocarcinoma; TGCT, testicular germ cell tumors; THCA, thyroid carcinoma; THYM, thymoma; UCEC, uterine corpus endometrial carcinoma; UCS, uterine carcinosarcoma; UVM, uveal melanoma. (b) RNA level (nTPM) and Protein level (nRPX) of each cell line in the cell line group of PTK7. Data derived from The Human Protein Atlas database. A-204, Human Rhabdomyosarcoma Cell Line; Rh30, Human Alveolar Rhabdomyosarcoma Cell Line; A-431, Human Epidermoid Carcinoma Cell Line; Hep-G2, Human Hepatocellular Carcinoma Cell Line; BEWO, Human Choriocarcinoma Cell Line; RL95-2, Human Endometrial Carcinoma Cell Line; SK-OV-3, Human Ovarian Adenocarcinoma Cell Line; HuH-28, Human Cholangiocarcinoma Cell Line; AGS, Human Gastric Adenocarcinoma Cell Line; HaCaT, Human Immortalized Keratinocyte Cell Line; T98G, Human Glioblastoma Cell Line; SH-SY5Y, Human Neuroblastoma Cell Line; SuSa, Human Cutaneous T-Cell Lymphoma Cell Line; RD, Human Rhabdomyosarcoma Cell Line; TE-1, Human Esophageal Squamous Cell Carcinoma Cell Line; OE33, Human Esophageal Adenocarcinoma Cell Line; PANC-1, Human Pancreatic Ductal Adenocarcinoma Cell Line; FaDu, Human Hypopharyngeal Squamous Cell Carcinoma Cell Line; SK-N-SH, Human Hypopharyngeal Squamous Cell Carcinoma Cell Line; CAL-27, Human Tongue Squamous Cell Carcinoma Cell Line; CACO-2, Human Colorectal Adenocarcinoma Cell Line; A2780, Human Ovarian Carcinoma Cell Line; SCaBER, Human Bladder Squamous Cell Carcinoma Cell Line; U-251MG, Human Glioblastoma Cell Line; Hep 3B2.1-7, Human Hepatocellular Carcinoma Cell Line; U2OS, Human Osteosarcoma Cell Line; Calu-3, Human Lung Adenocarcinoma Cell Line; HEK293, Human Embryonic Kidney Cell Line; RT-4, Human Bladder Transitional Cell Carcinoma Cell Line; 5637, Human Bladder Carcinoma Cell Line; ACHN, Human Renal Cell Adenocarcinoma Cell Line; HT-29, Human Colorectal Adenocarcinoma Cell Line; HCC1576, Human Colorectal Adenocarcinoma Cell Line; MG-63, Human Osteosarcoma Cell Line; ML-1, Human Acute Myeloid Leukemia Cell Line; BxPC-3, Human Pancreatic Adenocarcinoma Cell Line; HT-1080, Human Fibrosarcoma Cell Line; PC-3, Human Prostate Adenocarcinoma Cell Line; DU145, Human Prostate Carcinoma Cell Line; HeLa, Human Cervical Adenocarcinoma Cell Line; SNU-308, Human Hepatocellular Carcinoma Cell Line; U-87MG ATCC, Human Glioblastoma Cell Line U-87MG; SiHa, Human Cervical Squamous Cell Carcinoma Cell Line; MCF-7, Human Breast Adenocarcinoma Cell Line; THP-1, Human Monocytic Leukemia Cell Line; SK-BR-3, Human Breast Adenocarcinoma Cell Line; A-549, Human Lung Adenocarcinoma Cell Line; 786-O, Human Renal Cell Adenocarcinoma Cell Line; MDA-MB-435, Human Breast Adenocarcinoma Cell Line; HL-60, Human Acute Promyelocytic Leukemia Cell Line; LP-1, Human Plasma Cell Leukemia Cell Line; RPMI-8226, Human Multiple Myeloma Cell Line; Raji, Human Burkitt's Lymphoma Cell Line; Ramos, Human Burkitt's Lymphoma Cell Line.
General characteristics of PTK7 and its signaling pathways
PTK7 is a highly conserved member of receptor tyrosine kinases (RTKs) but without tyrosine kinase activity, known as a pseudo enzyme. RTKs represent the major class of enzyme-linked receptors. 9 They function as both receptors for multiple growth factors and enzymes that catalyze the phosphorylation of downstream target proteins upon binding specific ligands to their extracellular domains. The signaling pathways mediated by growth factor-RTK interactions plays a critical role in comprehensive range of biological functions, consisting of cell growth, migration, differentiation, and metabolism.9,30 For this reason, RTKs are very important tumor drug targets and play a significant role in cancer therapy. Over 80 small molecule kinase inhibitors have been developed and approved by the Food and Drug Administration (FDA) 31 and around half of them are targeting RTKs. 32 Initially, PTK7, as a member of class 1 of the pseudoenzyme family which have no capacity to bind ATP nor Mg2+ (Figure 3), received limited attention until its widespread expression in various tissues and cell types was discovered. Due to lack of catalytic activity, much more studies have revealed that PTK7 functions as a co-receptor and regulator of Wnt pathways and planar cell polarity (PCP).9,33–36

Classification of pseudokinases. Pseudokinases can be categorized into four class based on their binding ability to ATP and Mg2+. Pseudokinases are classified based on ATP and Mg2+ binding: Class 1 (e.g. PTK7, ROR1/2, RYK, IRAK3) binds neither ATP nor Mg2+; Class 2 (e.g. EPHB6, ULK4, TRIB4) binds ATP but not Mg2+; Class 3 (e.g. Sgk223, PEAK1) binds Mg2+ but not ATP; and Class 4 (e.g. HER3, IRAK2, JAK) binds both ATP and Mg2+.
PTK7 can affect tissue morphogenesis and patterning by regulating the PCP pathway, which belongs to the non-canonical Wnt signaling pathways and is crucial for cell orientation.9,33,37,38 The transgenic mice with PTK7 mutation, exhibiting phenotypes such as failure of neural tube closure and inner ear polarity defects, led to the identification of PTK7 as a core regulator of PCP in vertebrates. PTK7 is an evolutionarily conserved transmembrane protein and has orthologs in various species, including mouse PTK7, chicken KLG and Drosophila Off-track (OTK).39–41 The chick KLG protein is required for chick cardiac morphogenesis, while in Drosophila, OTK protein serves as a neuronal cell adhesion molecule for axon guidance regulation,. 42 Collectively, these findings indicate that PTK7 play an evolutionarily conserved and indispensable role in cell movement and polarity.
In addition to its significant role in the PCP pathway, PTK7 can also interact with Wnt ligands and Frizzled (Fz) receptors as a co-receptor to regulate Wnt signaling. 42 The function of PTK7 as a receptor in canonical Wnt signaling pathways remains controversial. It has been observed that PTK7 activates canonical Wnt signaling during the formation of Spemann's organizer and specification of posterior neural tissue.38,43 However, in ptk7 mutant zebrafish, canonical Wnt signaling was enhanced, as reflected by increased expression of ß-catenin, suggesting its functions in suppressing canonical Wnt signaling. 44 This discrepancy may arise from different receptor contexts and warrants further studies to elucidate the functional mechanisms of PTK7 in Wnt signaling pathways.
The full-length PTK7 protein contains 1070 amino acid residues and has a molecular mass of 118 kDa. Similar to other RTKs, the full-length membrane-bound PTK7 consists of three parts: an extracellular region, a transmembrane hydrophobic region, and a catalytically defective cytoplasmic tyrosine kinase domain.45,46 The extracellular domain of PTK7 comprises 7 immunoglobulin-like loops (Ig1-7), and this region, also known as soluble PTK7 (sPTK7), can be enzymatically cleaved by ADAM17, a sheddase enzyme. Subsequently, the remaining fragment is further cleaved into the transmembrane domain and cytoplasmic fragment by the γ-secretase complex.9,47 Apart from ADAM17, membrane type-1 matrix metalloproteinase (MT1-MMP) is another sheddase that cleave the PKP621↓LI sequence in the seventh Ig-like domain of PTK7.9,47,48 This proteolytic processing endows PTK7 with multiple functions, as illustrated in Figure 4 and discussed in this review. PTK7 was relatively abundantly expressed in the placenta, pancreas, kidney, lung, and liver, but expressed at low levels in the brain and heart. 47 This indicates that the function and importance of PTK7 may vary across different tissues.

Schematic illustration of PTK7 proteolysis and downstream signaling pathways that control cell survival and motility. The extracellular domain of PTK7, comprising seven immunoglobulin-like loops, is cleaved by the sheddase ADAM17 and further processed into transmembrane and cytoplasmic fragments by the γ-secretase complex. MT1-MMP also cleaves the PKP621↓LI sequence in PTK7's seventh Ig-like domain.
In addition to its normal physiological function, numerous studies have shown that dysregulation of PTK7 is involved in pathological conditions. Besides its role as a membrane receptor regulating various cellular activities, PTK7 has been shown to be a target of proteolysis involving membrane type-1 matrix metalloproteinase (MT1-MMP), members of the ADAM/ADAMTS family, and γ-secretase. The full-length PTK7 can be cleaved to several fragments, including soluble N-terminal fragments and their respective C-terminal transmembrane and intracellular proteolytic fragments.41,49 Interestingly, these soluble ectodomain fragments have been detected in colon tumor samples but are absent in healthy samples based on analysis of human tissue specimens, confirming that the proteolysis event occurs specifically in colorectal tumor cells rather than in normal tissue. 40 Importantly, the soluble PTK7 proteolytic fragments have been found to play an opposite role to full-length PTK7. The former induces actin cytoskeleton contractility through RhoA signaling, while the latter downregulates myosin light chain phosphorylation, leading to reduced actin cytoskeleton contractility. 39 These mechanisms suggest the significance of PTK7 in cancer cell migration, invasion, and metastasis. Thus, the PTK7 proteolytic fragments hold promise as potential indicators for cancer detection.
As addressed above, numerous studies have demonstrated the close involvement of PTK7 in carcinogenesis. In colorectal cancer (CRC), multiple signaling pathways associated with PTK7 have been uncovered. For example, PTK7 expression can be suppressed by miR-205-5p, and the miR-205-5p/PTK7/CASP10 axis is recognized as a critical pathway mediating PTK7's role in CRC progression.9,50 Besides, it has also been found that PTK7 is a ligand for Macrophage Galactose-type Lectin (MGL) on the surface of CRC cells, suggesting its involvement in CRC immune evasion and tumor growth.9,51 In esophageal squamous cell carcinoma (ESCC), PTK7 has been found involved in PI3 K/ AKT and Ras/MAPK pathways, resulting in the activation of NF-κB and AP-1. Subsequently, NF-κB and AP-1 synergistically transactivate MMP9, thereby promoting the invasive phenotype of ESCC cells.9,52 In lung adenocarcinoma, PTK7 has been shown to enhance cancer cell adhesion via PTK7-NDRG1 axis. Furthermore, PTK7 has also been identified as a direct target gene of miR-503, 53 regulating cytoskeletal dynamics to facilitate cancer cell invasion and migration through FAK and paxillin. 54 In breast cancer (BC), particularly in TNBC, PTK7 acts as a critical role in oncogenic signal transduction by enhancing the activation of FGFR1 and EGFR, leading to BC tumorigenesis and metastasis. 55 Far beyond that, PTK7 has been found to be associated with various other cancers, including melanoma, ovarian cancer (OVCA), and cervical cancer. Jin Zhipeng et al. comprehensively reviewed the molecular mechanisms of PTK7 in various cancers and systematically summarized the diagnostic and therapeutic strategies targeting PTK7, establishing it as a significant therapeutic target. 9
However, the functional mechanism of PTK7 still requires further investigation. While its essential biological functions have been established in Wnt/PCP signaling pathways, these findings do not fully elucidate the underlying mechanisms of PTK7 mediated carcinogenesis. The absence of catalytic activity in PTK7 presents an additional challenge in understanding its specific biological function, compounded by the fact that its ligands have not been fully characterized. Unraveling these mysteries holds the potential to unveil novel biological mechanisms of PTK7.
Given their high affinity and specificity, aptamers have emerged as encouraging agents for cancer diagnosis and therapy. Herein, we will delve into recent advancement in aptamer selection and preclinical application targeting PTK7 receptors in cancer.
The mechanism underlying sgc8's action
Sgc8 targeting PTK7 in CCRF-CEM cells
Aptamer sgc8, along with other aptamers like sgc3, sgd3, sgc4, and sgd2, was identified to exhibit high affinity for CCRF-CEM cell line (T-cell acute lymphoblastic leukemia, T-cell ALL). 8 These aptamers were obtained through cell-SELEX and could detect CCRF-CEM leukemia cells mixed with normal human bone marrow aspirates. Each aptamer possesses a unique structural conformation, enabling it to recognize and bind to specific subpopulations of CCRF-CEM cells. For instance, Shangguan et al. reported that aptamer sgc3 selectively binds to CD5 and CD7 abundantly expressed subpopulations among CCRF-CEM cells. These aptamers were utilized to identify tumor cells in real clinical samples based on their specificity and high affinity. Further results confirmed that all the mentioned aptamers can recognize cancer cells from bone marrow aspirates of T-cell ALL patients, rather than B-cell lymphoma patients, 8 demonstrating the significant potential of these aptamers for molecular diagnosis in clinical practice. More importantly, Sgc8 demonstrates high-affinity binding to ALL T-cells with a dissociation constant (Kd) of 0.8 nM. 3 In real clinical specimens, sgc8 effectively identifies cancer cells closely associated with the ALL cell line. Besides, what distinguish sgc8 from its counterparts is its ability to internalize into its target cells without affecting cell viability in a receptor-dependent manner.56,57 This internalization mechanism allows sgc8 to effectively deliver its cargo to the endosome, leveraging the crucial role of the endosome in intracellular cargo transportation.58,59
Affinity pull-down followed by MS was employed to identify the protein target of aptamer sgc8. 18 Among the 25 protein hits obtained from the MS results, PTK7 ranked second. However, the first protein identified was a ribonucleoprotein that was excluded as a target for aptamer sgc8 due to its nonspecific binding to DNA. Subsequent in vitro study demonstrated that cells with higher expression of PTK7 exhibited stronger binding to sgc8 compared to anti-PTK7 antibodies. Consequently, PTK7 was confirmed as the biomarker for CCRF-CEM cell line. 18 Additionally, sgc8 was tested to specifically bind to leukemia cells in considerable levels in clinical patient samples.8,60 Furthermore, the use of fluorescent probes based on the sgc8 aptamer revealed the upregulation of PTK7 in multiple tumors, which aligns with previous findings using anti-PTK7 monoclonal antibody–fluorescent probe conjugates.61,62 Compared to the PTK7 antibody, the sgc8 aptamer offers the advantage of facile chemical synthesis and low cost, making it a more appealing candidate for use of as delivery agents.
To determine the specific conformational structure critical for target binding, four truncated sequences of Sgc8 were synthesized for the study. Sgc8c, a truncated sequence of the original aptamer Sgc8, possesses a single loop-stem structure that is essential for target recognition and exhibits a reported Kd of 0.78 nM for the PTK7 receptor, like sgc8. Sgc8c comprises only 41 nucleotides but represents the minimal aptamer sequence required for strong target binding (Figure 5(a)). This size reduction facilitates the efficient chemical synthesis of the aptamer. Recently, Chen et al. identified the sgc8c-binding region on PTK7 is within Ig (3-4) through sequential protein truncation (Figure 5(b) to (c)). Compared with other regions, the region between Ig (3) and Ig (4) forms a pocket-like structure with the highest positive charge density which facilitates the binding with sgc8c. 63 Therefore, identification of binding site between PTK7 and sgc8c lay the structure foundation for the design of aptamer-based targeted drugs and diagnostic molecules.

The schematic diagram of binding between PTK7 and sgc8c. (a) The nucleotide sequence of sgc8c. (b) Molecular dynamic simulation results of the Ig (3−4)-sgc8c binding complex. 63 Adapted with permission from Ref. 63 Copyright 2024 American Chemical Society. (c) The cartoon view of sgc8c binding on Ig(3-4) of PTK7 and internalization in endosomes.
Internalization of sgc8 aptamer
Although aptamers are continually being screened out and identified, there are few reports on their intracellular studies and function applications. One of the key reasons is that most aptamers cannot be directly taken up by target cells like small molecules. The discovery of sgc8 appears to be a glimmer of hope. After specifically binding to PTK7, sgc8 was further found to be internalized into CCRF-CEM. Xiao Zeyu et al. used TetraTetramethyl-6-Carboxyrhodamine to label sgc8 (sgc8-TAMAR), and the intracellular fluorescence signals were detected by confocal imaging after having removed specific binding of sgc8 outside the membrane of CCRF-CEM by trypsin treatment. The flow cytometry results further confirmed internalization of sgc8. 3 Subsequently, the intracellular distribution of sgc8 has been examined to be in endosomes by incubating with alexa633-labeled transferrin (transferrin-alexa633) which is commonly used for identification of location in endosomes. 3 Then, the specificity of sgc8 internalization in CCRF-CEM was proved by incubating with different types of cells. Xiao ZY et al. also investigated the cell viability after internalization of sgc8, even at high concentration of sgc8, its cytotoxic effects on cells remain negligible. 3 Therefore, sgc8 aptamer holds great potential as a deliver agent for future applications in cancer diagnosis and targeted therapy.
Application of sgc8 aptamer
The current major cancer treatment modalities involve surgery, radiotherapy, chemotherapy and tumor immunotherapy. 64 Surgery is widely used for resection of solid tumors, but it can cause certain physical damage to patients and carries the potential risk of recurrence. Radiotherapy and chemotherapy use chemical drugs or high-energy ionizing radiation to eliminate tumor cells accompanied with significant side effects, 65 such as the destruction of normal immune cells, leading to immune suppression or tissue damage. Apart from the traditional three approaches mentioned above, the emergency of immunotherapy and targeted therapy have provided new insights into cancer treatment.66,67 Among them, immune checkpoint blockade (ICB) and adoptive cell therapy (ACT) have achieved remarkable achievements.68,69 However, not all patients can benefit from these methods as there are large individual differences in the efficacy and may cause unique immune-related adverse effects, 70 the high cost of immunotherapy limits its widespread application among a large number of patients. The field of cancer treatment has long faced a bottleneck, requiring more precise and effective strategies for early diagnosis and subsequent treatment. Targeted therapy may be a more effective treatment option for patients who respond poorly to traditional treatments or cannot be tolerate their side effects. Since PTK7 has been found highly expressed in many types of cancer cells and can be specifically recognized and bound, as previously discussed, the following discussion will focus on the studies related to the use of sgc8 aptamer in cancer diagnosis and treatment.
Sgc8 aptamer utilized for potential diagnosis
Given these advantages, there has been extensive translational research on applying aptamers for tumor diagnosis and treatment, bridging the gap from bench to clinic. In 1997, an aptamer called NX21909, targeting human neutrophil elastase and labeled with technetium-99 m (99mTc), successfully achieved the desired effect in diagnostic imaging of inflammation in rats. 71 Compared to IgG, NX21909 exhibited advantages such as good tissue penetration, fast clearance due to its low molecular weight, and a high target/non-target ratio. This success demonstrates the feasibility of conjugating aptamers like sgc8 with detection agents and drugs for both diagnosis and treatment purposes (Figure 6). Notably, due to the expression of PTK7 in murine B16F10 melanoma cell line, modifications were made to sgc8c to generate new molecular imaging agents such as sgc8c-NOTA-67Ga and sgc8c-Alexa 647. These aptamer probes exhibited good uptake with low backgrounds in metastatic melanoma, making them promising tools for early detection. 62 Additionally, Calzada et al. developed two probes by conjugating Sgc8c with the fluorophore Alexa 647 or radionuclide 99mTc to recognize tumor cells in mouse model of melanoma and lymphoma. 72 Furthermore, Sgc8 and Alexa 647 conjugate was explored as a molecular imaging probe for colon cancer in a mouse xenograft model and exhibited good tumor-to-background ratios in biodistribution studies. 61 Similarly, in tumor-targeting PET imaging applications, sgc8 was radiolabeled with fluorine-18 (F-18), enabling convenient and highly efficient visualization and tracking in a xenograft mouse model of human colon cancer with high PTK7 expression. This bioconjugation approach led to the development of a unique and groundbreaking in vivo PET imaging tool. 73 Most importantly, the first-in-human pharmacokinetics study of PET tracking of gallium-68 (68Ga) radiolabeled sgc8 aptamer exhibited that radiolabeled aptamers are harmless to normal organs and that the majority accumulated in the kidney and cleared from the body via urine. 74 These encouraging results have proved the potential of 68Ga radiolabeled sgc8 aptamer to be used in PTK-7-positive tumor PET imaging.
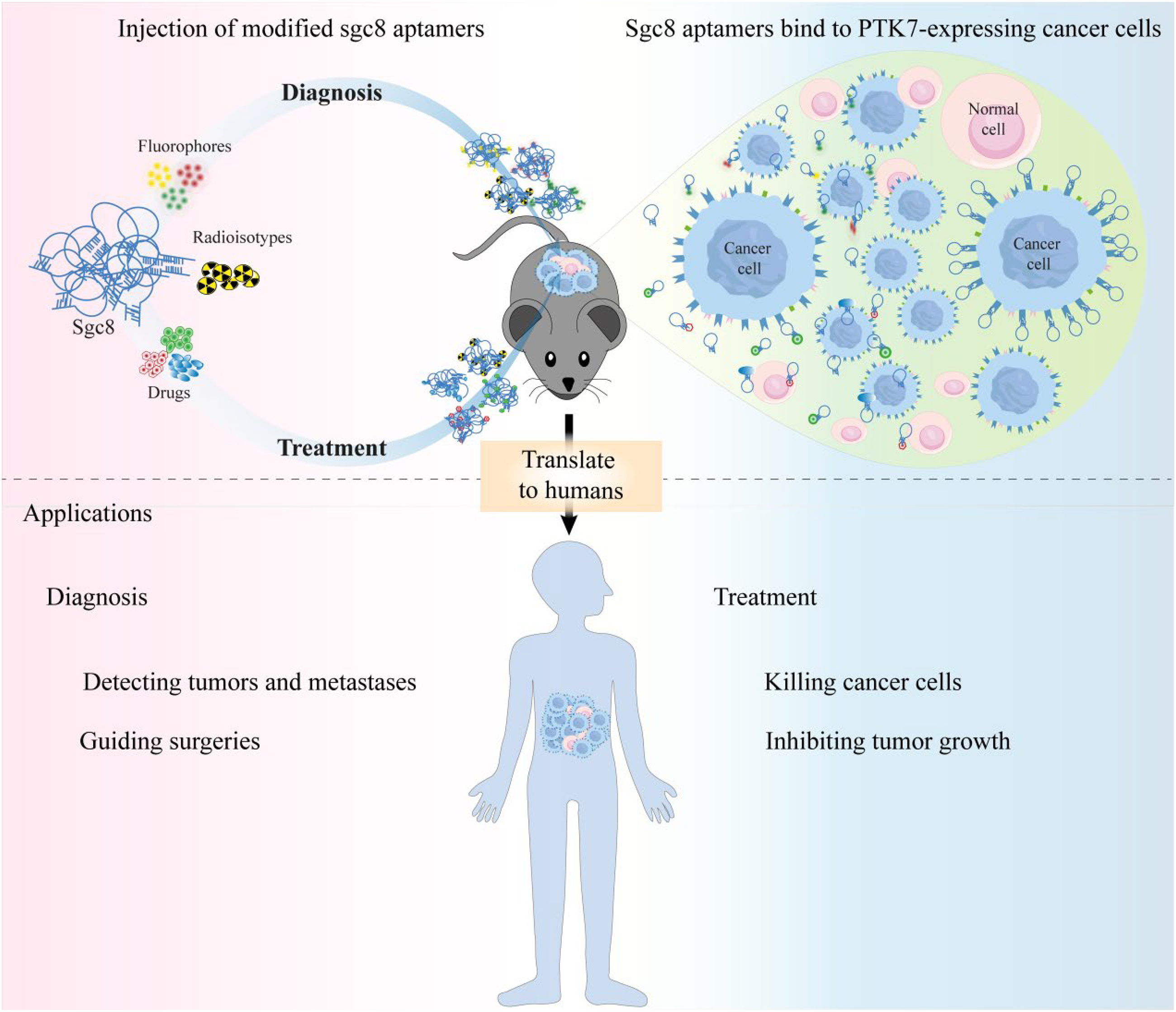
Applications of conjugation sgc8c with agents. Sgc8 conjugates with detection agents and drugs show great potential for cancer diagnosis and treatment. Further translational research is essential to advance aptamers from laboratory studies to clinical applications.
Sgc8 aptamer utilized for treatment
In addition to their application in cancer diagnosis, aptamers have emerged as promising therapeutic drugs since the 1990s. Certain aptamers can function as standalone drugs, such as pegaptanib. 75 AS1411 aptamer, which shows promise as an anticancer therapeutic agent. AS1411 can target and kill cancer cells by binding to nucleolin, which is overexpressed on the surface of cancer cells, thus inhibiting cancer cell proliferation and tumor growth. 76 Moreover, aptamers play a crucial role as guiding molecules in targeted drug delivery. Like antibody-drug conjugates (ADCs), one promising application is ApDCs, where cytotoxic anticancer drugs are conjugated to targeting aptamers.56,59,77,78 This approach allows for specific recognition of tumor cells and targeted drug delivery while minimizing toxic side effects. Compared to ADCs, ApDCs offer several advantages. They are smaller in size, and exhibit improved chemical stability, weaker immunogenicity, enhanced tissue penetration, and easier engineering. 79 An ADC drug known as PF-06647020 is composed of three parts: a humanized anti-PTK7 monoclonal antibody, an auristatin microtubule inhibitor (Aur0101) and a cleavable valine-citrulline-based linker. It has shown promise in the treatment of disease relapse in patients with TNBC, OVCA, and non-small cell lung cancer (NSCLC). 80 These results indicate that PTK7 is a promising target for ADC cancer therapy. Accordingly, sgc8c aptamer was successfully conjugated with the chemotherapeutic drugs such as doxorubicin (Dox). As an anthracycline-based chemotherapeutic drug, Dox is the most widely used anticancer agent against a variety of neoplasms, such as acute lymphoblastic, myeloblastic leukemias, and malignant lymphomas. 81 However, the treatment of Dox is associated with a series of severe side effects like alopecia, mucositis, and cumulative cardiac damage.81,82 Sgc8 aptamer conjugated with Dox reduced its toxic effects on normal cells while maintaining its specificity for killing CCRF-CEM cells. Yang et al. encapsulated Dox in mesoporous silica nanoparticles (MSNs), which possess large surface area, adjustable pore volume and good biocompatibility, and decorated with sgc8. The resulting Sgc8-MSN/Dox nanoparticles exhibited favorable morphology and size for drug delivery applications. These nanoparticles were selectively internalized by leukemia cells in vitro, resulting in efficient tumor cell eradication. 83
The aptamer-functionalized nanoconjugates hold great promise as a novel targeted drug delivery system with potential efficacy and flexibility, offering significant prospects for clinical cancer treatment. For the better targeted delivery of Dox, Hu et al. designed a simple Sgc8 aptamer-modified nucleic acid nanomaterial (Sgc8NM), exhibiting a precise treatment effect on target cells and low toxicity on non-target cells. 84 The aptamer/hairpin DNA-gold nanoparticle (apt/hp-Au NP) conjugate has been devised as a smart drug carrier for targeted delivery of drugs. A single AuNP (13 nm) can load 25 (±3) sgc8c and 305 (±9) Dox molecules on its surface. These constructed nanoconjugates exhibited enhanced drug loading, specific targeting and internalization in PTK7-abundant CCRF-CEM cells. The release of Dox was light-controllable, just triggered by the photothermal effect of Au NPs when exposed to laser illumination. 85 Similarly, Ma et al. modified sgc8 aptamer with F-base(F23), 3,5-bis(trifluoromethyl)benzene, further reacted with paclitaxel (PTX). This modification realized prolonged blood circulation time through hydrophobic properties of F23 and enhanced efficacy. 86 Sgc8 aptamer targeting ability was effectively used to eliminate the limitations of cytarabine (Ara-C) in treatment of leukemia. In this case, sgc8-BSA was designed as a tumor targeting carrier and conjugated to glutathione (GSH)-responsive polymeric micelles (PCL-ss-Ara@Sgc8-BSA). Compared to free Ara-C, this polymeric micell improved plasma half-life and bioavailability, reduced bone marrow and liver toxicity, thereby significantly inhibited tumor growth and increased the survival rate of CCRF-CEM tumor-bearing mice. 87 Huang et al. conjugated Sgc8 aptamer to combretastatin A4 via three types of linkers: a phosphodiester bond linker, a disulfide bond linker and a carbamate linker. All of those conjugates exhibited high precision and cytotoxicity against target cancer cells. 88 Therefore, owing to its good modifiability, sgc8 aptamer can be conjugated to a wide range of imaging agents and drugs to achieve excellent diagnosis and treatment of cancer. The research on the use of Sgc8 as a delivery molecule targeting PTK7 in the past decade is summarized in Table 1. The research using aptamer sgc8 for the targeted therapy and diagnosis represents a microcosm of the broad applications of aptamers identified through SELEX technology. This suggests that there is promising potential for innovative approaches for disease diagnosis and treatment.
Studies on the use of sgc8 or sgc8-c as delivery agents in diagnosis and therapy.

Challenges and future directions
It is also important to note that aptamers including sgc8 face several limitations that hinder its clinical applications. Firstly, in vivo stability, aptamers are prone to enzymatic degradation by nucleases, leading to short biological half-lives. But it seems to be advantageous in terms of diagnosis. Jacobson Orit et al. radiolabeled of sgc8 with 18F-fluorobenzoate as a PET tracer for visualizing PTK7, rapid clearance of 18F-Tr-Sgc8 into urine avoids accumulation in the body. 22 Then, immunogenicity, although aptamers are generally non-immunogenic, the fact that their short half-lives require repeated administration may potentially trigger responses. Delivery efficiency of aptamers to solid tumors or deep tissues remains challenging due to poor penetration or clearance issues. As aptamers could lose their secondary or tertiary under physical conditions, off-target binding may reduce its efficacy in some applications. Additionally, several studies have showed low levels of PTK7 expression was detected to in some organs, such as ovary, mammary gland, lung and digestive tract. 80 Further investigations are necessary to test potential off-target effect to ensure the safe clinical application of sgc8 aptamer.
These challenges currently hinder the translation of aptamers from basic experimental research to clinical applications. To address these challenges, further specific modifications could be designed to enhance the resistance to nucleases and increase their bioavailability. For example, aptamers can also be modified with polyethylene glycol (PEG) or through 2′-O-methylation to achieve increased stability and reduced renal filtration rates. 89 In addition, base replacement with 3,5-bis(trifluoromethyl)benzene (F base) in sgc8 aptamer has showed to extend its circulation time. 90 Ongoing research and advancements continue to address these challenges and improve the potential of aptamers for bio-applications. With further optimization and development, sgc8 aptamers hold great promises as valuable theranostic agents in the field of precision medicine.
In conclusion, this review article summarized the advantages of cell-SELEX based on aptamers and applications of aptamer sgc8 which could specifically bind PTK7 and internalize into target cells. The most significant advantage of cell-SELEX is its ability to screen out aptamers specific for any cells of interest, without relying on prior knowledge of their molecular characteristics, and the selected aptamers exhibit extremely high specificity for target cells. These aptamers offer significant advantages over antibodies, including ease of synthesis, low cost, good stability and low toxicity. Furthermore, PTK7 was found over-expressed in various cancer cells, which makes sgc8 a promising and versatile candidate as a guiding molecule such as conjugating with agents and drugs for targeted cancer therapy, cancer diagnostics, drug delivery system.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by Zhejiang Provincial Natural Science Foundation of China (Grant Number YXD23H0303).
