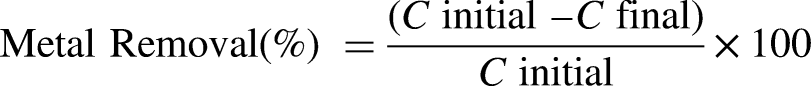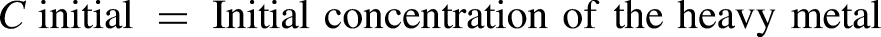Abstract
Objective
Heavy metal pollution is one of the more recent problems of environmental degradation caused by rapid industrialization and human activity. The objective of this study was to isolate, screen, and characterize heavy metal-resistant bacteria from solid waste disposal sites.
Methods
In this study, a total of 18 soil samples were randomly selected from mechanical sites, metal workshops, and agricultural land that received wastewater irrigation. Isolation and screening of the isolates were based on multiple heavy metal (copper (Cu), chromium (Cr), and lead (Pb)) resistance potential. Morphological, biochemical, and 16S rRNA gene sequencing techniques were used to identify the isolates. The minimum inhibitory concentration (MIC) of the potential isolates was further examined at various concentrations (3, 6, 12, 25, 50, and 100 µg/mL). Each isolate's ability to biodegrade and resist antibiotics was also examined.
Results
About 21 bacterial isolates were obtained. Among these, six potential isolates (Agri10−2Is2, Agri10−3Is2, Agri10−3Is1, AL10−1Is2, AL10−3Is1, and GA10−2Is1) were selected. The isolates displayed varied colony morphologies and biochemical characterization features. Phylogenetic tree results revealed that Agri10−2Is2, Agri10−3Is2, Agri10−3Is1, AL10−1Is2, AL10−3Is1, and GA10−2Is1 were identified as Bacillus subtilis, Bacillus subtilis, Bacillus species, Bacillus tequilensis, Klebiesella species, and Cronobacter sakazakii, respectively. Every isolate gave a different MIC value. The biodegradation capacity of Agri10−2Is2, Agri10−3Is2, Agri10−3Is1, AL10−1Is2, AL10−3Is1, and GA10−2Is1 was 92.2%, 92.57%, 92.37%, 92.66%, 92.85%, and 92.52% against Cu; 95.07%, 94.07%, 93.07%, 93.4%, 91.4%, and 94.4% on Cr; and 95.51%, 94.53%, 95.62%, 96.87%, 94.86%, and 97.09% against Pb, respectively. All isolates were found to be resistant to the tested antibiotics.
Conclusion
This study concludes that the selected bacterial isolates could be potential agents for bioremediation of heavy metal-polluted environments.
Introduction
The quote by John Muir emphasizes the profound benefits nature offers to humans, which can be diminished when land pollution is left unchecked. 1 As the world progresses, industrialization leads to an alarming increase in pollution, particularly land pollution, which poses significant risks to both human health and the environment.
Waste, as defined, consists of any unwanted or unuseful materials that have been discarded.2,3 It can take various forms, including construction and demolition waste, mining and hazardous waste, industrial non-hazardous waste, medical waste, radioactive waste, and waste generated from the oil and gas production processes. 4 The surge in waste production is largely due to rapid industrial growth, but it is compounded by the absence of efficient waste disposal systems. 5 Without appropriate management and recycling programs, these materials accumulate on the land, contributing to environmental degradation and threatening biodiversity, soil quality, and public health.
Industrial development is the major cause of environmental pollution with various materials, including heavy metals, antibiotics, pesticides, and wastewater, which is a major source of various heavy metals, pathogens, and organic contaminants. 6 Wastewater has a significant impact on soil productivity and the health of living organisms. Both rapid urbanization and uncontrolled population growth have contributed the lion's share to increasing the level of municipal solid waste (MSW). 4
Waste disposal is a serious global environmental threat resulting in heavy metal pollution. 7 Waste materials that are disposed of in the environment commonly occur in solid and liquid form. 5 Those that are in solid forms (solid wastes) are continually generated from municipal, commercial, and industrial sources. 8 These wastes are then carried from the generation sites to a landfill or dumping site for further processing. 9
The discharges of heavy metals into the environment as a result of human and natural activities and their effects on human health and ecosystems as a whole have been current topics. 10 The pollution by heavy metals is one of the emerging problems of environmental contaminations due to the rapid industrialization of geologic and anthropogenic activities. 11 Heavy metal pollution in MSW is a result of residential, industrial, and agricultural activities. 8 Some of the examples include smelting of metallic ores, industrial processes like electroplating, steel production, leather tanning, wood preservation, ceramics, glass manufacturing, fertilizer applications, military operations, use of metal-containing pesticides, road and transportation runoff, and landfill sites.12,13 The combustion of fossil fuels is just one of the human activities that releases heavy metals into the environment and harms our aquatic ecosystems. 5 And also, through runoff, heavy metals from domestic waste, auto shops, antiseptics and disinfectants, metal insecticides, and agricultural fertilizer enter water bodies. 14
Heavy metals, often known as trace metals, are among the most persistent contaminants found in wastewater. 15 A class of metals with an atomic number larger than 20, a density greater than 5 g/cm³, and a molecular weight greater than 53 g/mol are referred to as heavy metals. 16 In nature, they are widely dispersed and accumulate in food chains. 17 Metallic elements with a comparatively high density in relation to water are known as heavy metals. 18 When combined or in elemental form, heavy metals, including chromium (Cr), cadmium (Cd), mercury (Hg), lead (Pb), nickel (Ni), and thorium (Tl), can be dangerous. 19 Because heavy metals are extremely soluble in watery environments, living things can readily absorb them. 20 In nature, heavy metals are persistent, toxic, poisonous, and non-biodegradable. 21
Heavy metals toxicity is a significant environmental pollution problem of increased significance for ecological, evolutionary, nutritional, and environmental reasons. 15 Metals are divided into two categories, essential and non-essential, based on their function in living things. 22 Microorganism development and growth are attributed to essential metals like Ni, Mn, and Fe. Hg, Pb, and Cd and some other metals, like Co, Cu, Zn, and Ni, are necessary because they serve as cofactors for metalloproteins and enzymes. 23 Heavy metals are non-essential metals that are not required for the growth and development of living things, and these metals serve no biological purpose 24 and have an inhibitory effect on microbes by blocking crucial functional groups, displacing crucial metal ions, or altering the active conformations of biological components. 25 Heavy metals impair the biological growth of an organism and pollute the environment; neurotoxicity, DNA damage, carcinogenesis, cell damage, protein misfolding, and conformational alterations may all be brought on by these contaminants. 26 Heavy metals in industrial effluents are known to pose a serious risk to the health of humans, animals, plants, and natural water resources. 27 High levels of heavy metals discharged into water bodies have a number of negative effects on the environment and human health. 5 It is well recognized that heavy metals disrupt carbon, nitrogen, and organic matter transformation, impair soil enzyme activity, and reduce microorganism variety and biomass, which favors the dominance of heavy metal tolerating microorganisms. 28 Heavy metal poisoning contaminates the microbiota and is linked to producing issues with plant development. 26 Heavy metals like Cu, Ni, and Zn are next in line to offer a major ecological risk to the environment after heavy metals like Hg and Pb. 29 Heavy metals like Hg, Cr, Cu, Zn, Pb, and Cd are known to be toxic, poisonous, mutagenic, and carcinogenic to humans and other living things.16,30
Heavy metal-resistant bacteria might be present in heavy metal-contaminated sites like soils and wastewater. 31 The resistance and efficiency of bacteria to remove heavy metals vary greatly. 32 The heavy metal-polluted soil contains some microorganisms that exhibit tolerance to the metals, which in turn helps in the reduction of heavy metals toxicity accumulated in the soil. 33 Some bacteria not only reduce heavy metal toxicity but also convert them to useful resources. 34
The bacteria in wastewater that are exposed to a lot of heavy metals tend to become resistant to their toxicity. 35 Bacteria that are able to survive well in high concentrations of heavy metals include both Gram-positive genera, Arthrobacter and Corynebacterium, and Gram-negative (Alcaligenes species), which have received growing attention in recent years. 36 The Gram-negative bacteria that are included in the Enterobacteriaceae family, like Cronobacter species (C. sakazakii, C. malonaticus, C. turicensis, C. muytjensii, C. dubliniensis, and C. universalis) as well as Klebsiella species, predominantly K. pneumoniae, have the ability to resist or tolerate high heavy metal concentrations due to the presence of extra plasmids and chromosomal genes. 37
The bacteria species included in the Bacillus genera, such as Bacillus species, B. subtilis, B. tequilensis, B. cereus, B. thuringiensis, etc., also have heavy metal resistance due to a specific resistance mechanism. 38 An extracellular barrier, metal ion reduction, low molecular weight internal sequestration, external sequestration, cysteine-rich proteins, and active metal ion transport (efflux) are a few defense mechanisms linked to bacteria's resistance to heavy metals. 18 This characteristic is potentially useful for bioremediation. 4 Tinatin et al. 39 reported three bacterial species (Alcaligenes faecalis, Delftia tsuruhatensis, and Stenotrophomonas species) with their different taxonomic positions, enzyme systems, and metabolic pathways, which utilized the heavy metals (Pb, As, Co, and Ni) in different ways and complemented each other to affect the biotransformation of the heavy metals. And also, Lysinibacillus fusiformis, Stenotrophomonas species and Enterobacter cloacae are found to be active degradative agents for hydrocarbons degradation and heavy metals like lead, chromium, and nickel transformations.40,41
There is a correlation between antibiotic resistance and metal tolerance. 42 In certain cases, metal tolerance mechanisms contribute to the increase in antibiotic resistance; the occurrence of this phenomenon can be attributed to the clustering of these genes in the same plasmid. 43 Heavy metal ions are known to co-regulate genes responsible for antibiotic resistance and decrease antibiotic susceptibility.39,44 Environmental pollution not only triggers heavy metal co-selection processes but also increases the level of tolerance to some antibiotics due to co-regulation of resistance genes. 45 Therefore, pragmatic measures must be taken to limit the spread of antimicrobial resistance across the environment and to reduce the incidence of healthcare-associated infections.
Heavy metal removal or remediation from hazardous sites has now become mandatory. Bioremediation based on microorganisms is a promising method to remediate heavy metal-contaminated areas due to its ecofriendly, cost-effective, and highly efficient characteristics. 46 Municipal and industrial wastewater that contain heavy metals is dumped straight into streams, rivers, and other bodies of water that are used to irrigate and fertilize crops in developing nations like Ethiopia because there are insufficient treatment facilities and waste disposal systems in place. 47 The bacteria become resistant to harmful heavy metals as a result of being exposed to high concentrations of heavy metals. 4 The bacterial plasmids are recognized to have potential resistance to metals. 48 Therefore, the likelihood of the bacteria carrying heavy metal-resisting characters through horizontal genetic transformation from metal-resisting bacteria can be expected. 27 Additionally, environmental management must address the correlation between heavy metal pollution and antibiotic resistance.
There is no previous study on the isolation and characterization of heavy metal-resistant bacteria in Ethiopia. Hence, taking all such conditions and problems into consideration, the present study aimed to isolate, screen, and characterize heavy metal-resistant bacteria from solid waste dumping sites in the Central Gondar zone, Northwest, Ethiopia.
Materials and methods
Description of study area
The study was carried out in solid waste dumping sites (mechanic sites, metal workshops, and wastewater-irrigated agricultural soils, especially in Maraki and Azezo) in the Central Gondar zone of Northwest Ethiopia. Gondar is located in Northwestern Ethiopia at a latitude of 12°36′N and a longitude of 37°28′E with an elevation of 2133 m above the sea level. The population of Gondar was projected by the Central Statistics Agency to reach 323,875 in 2015 and 360,600 in 2017, according to population projection. 49 Gondar city is 727 km away from Addis Ababa, the capital city of Ethiopia. It has an average annual maximum temperature of 27°C and a minimum temperature of 16°C, and the mean annual rainfall in Gondar city is 1025 mm. Gondar city administration has 23 urban and 11 rural Kebeles (Counties). 50 The map of the Gondar is depicted in Figure 1.

GIS map of the study area (developed by Abebech Dinku).
Study design
An experimental research design was used to conduct this study. The study was conducted from in Microbiology and Molecular Biology Laboratories, Institute of Biotechnology, University of Gondar, Gondar, Ethiopia.
Preparation of stock for selected heavy metals solutions
Stock solutions of Cr (K2Cr2O7, Himedia, extra pure), Cu (CuSO4.5H2O, Himedia, extra pure), and Pb ((CH3COO)2Pb.3H2O, Himedia, extra pure) were prepared by dissolving 10 g of their respective salts in 100 mL distilled water under aseptic conditions. 35
Collection of samples, sampling technique, and processing of samples
About 10 g of heavy metal-contaminated soil samples were collected from Mechanics Sites, Metal and Aluminum Workshops, and Wastewater-irrigated Agricultural areas of the Central Gondar zone (Maraki and Azezo areas), Ethiopia. Purposive sampling was employed to select the sampling sites. 51 Sampling sites were selected by observing the chance of these sites being contaminated by heavy metal coming from near distance or on-site solid waste disposal sites. 36 The samples were aseptically collected in clean, sterilized plastic bags using icebox, and then transported to microbiology laboratories of the Institute of Biotechnology and stored at 4°C in a refrigerator in order to prolong the samples’ stay. 30
Isolation and screening of heavy metal-resistant bacteria
Bacteria were isolated from a heavy metal-contaminated soil sample using the standard dilution plate technique. A six-fold serial dilution of fresh soil (1 g) was made in autoclaved saline water, and 0.1 mL from each of these dilutions was added in metal (0.1 mL) (Luria Bertani, LB, HiMedia) agar plates using the pour plate technique. Plates containing bacterial cultures were incubated at 37°C for 48–72 h. Bacterial colonies having variable morphology were selected to obtain their pure culture. 24 The resulting bacterial colonies were purified by further subculturing in the same media through the streak plate method and were preserved in 20% glycerol stock at −20°C. 21 The bacterial isolates were screened for heavy metal resistance, including Cu, Cr, and Pb. Separate preparations of LB agar media supplemented with 100 µg/mL of Cu, Cr, and Pb were made, autoclaved, and inoculated through the streak plate method. The plates were incubated at 37°C for 48 h, and results were recorded in terms of resistance to Cu, Cr, and Pb after observing growth in culture plates. The control plates, each for Cu, Cr, and Pb, supplemented LB media, were also run simultaneously without inoculating bacterial isolates to check the microbial contamination. 21 Cu-, Cr-, and Pb-resistant bacterial isolates were termed as multiple heavy metal resistant and were purified and preserved at −20°C until further experiments.
Morphological and biochemical characterization of the isolates
Selected bacterial isolates were cultured on LB agar media enriched with 100 μg/mL heavy metal salts ((CH3COO)2Pb, CuSO4, and K2Cr2O7). These isolates were then characterized using biochemical traits like catalase, citrate, and Triple Sugar-Iron Agar, as well as cultural and morphological traits like colony shape, size, color, margin, elevation, texture, motility, and Gram reaction. 52 The biochemical tests were used to identify the bacterial isolates up to the genus level according to Bergey's Manual of Determinative Bacteriology. 52 Based on preliminary screening, isolates were selected for further taxonomic identification using molecular techniques.
Determination of minimum inhibitory concentration (MIC)
The MIC is the lowest metal concentration that prevents bacteria from multiplying observably. 33 The MIC of heavy metals against the bacterial isolates was determined by the LB agar dilution technique by growing bacterial isolates on LB agar plates supplemented with heavy metals Cu, Cr, and Pb. 53 The stock solutions of K2Cr2O7, (CH3COO)2Pb.3H2O, and CuSO4.5H2O were prepared in distilled water and sterilized by autoclaving at 121°C for 15 min. For determining MIC, 20 mL of LB agar was added into each Petri dish, and different concentrations (3, 6, 12, 25, 50, and 100 μg/mL) of Cr, Cu, and Pb were pipetted into the plate using a 1 mL pipette. The molten LB agar and metal solution were combined by swirling the plate, and then the combination was allowed to solidify. Sterile loops were used to streak the isolated cultures of metal-resistant bacteria over the LB agar medium that contained metal salts. The growth of bacteria on the plates was monitored. The culture that grows at a given concentration was transferred to the next (higher) concentration by streaking on the plate. The MIC of the heavy metal-resistant bacterial isolates grown on heavy metal-incorporated media against the corresponding heavy metal was determined by progressively raising the concentration of the heavy metal on an LB agar plate until the strains were unable to form colonies on the plate. After 3 to 5 days of incubation at 37°C, MICs were recorded when the isolates were unable to grow on the plates. 35
Assessment of antibiotic resistance
Heavy metal-resistant bacteria associated with antibiotic resistance were tested employing disc diffusion assays using Mueller Hinton agar media (HiMedia, extra pure). 54 For this test, three antibiotics [ampicillin (10 μg), gentamicin (10 μg), and tetracycline (30 μg)] were selected that have wide-spectrum application. 55 The prepared inoculum for each isolate was put on Mueller Hinton agar plates with standard antibiotic-impregnated discs. The antibiotic discs were provided by HiMedia. Following the standard antibiotic disc chart, the bacterial isolates were categorized as resistant (R), intermediate (I), or susceptible (S) based on measurements of the diameter of the inhibition zones after 24–48 h of incubation at 37°C. 35
Determination of heavy metal removal capacity of isolates
Bacterial isolates were grown in a shake flask containing LB broth medium for 72 h in a rotary shaker at 150 rpm. The temperature and pH were kept at 37°C and 7.0, respectively. Each culture flask was then filled with 100 mg/L of sterilized heavy metals (Cr, Pb, and Cu) in the form of their salts, K2Cr2O7, (CH3COO)2Pb.3H2O, and CuSO4.5H2O. The flasks were then incubated for 72 h under the same conditions. The total culture was centrifuged at 5000 rpm for 15 min. The supernatants were filtered with Whatman 42 filter paper. An atomic absorption spectrophotometer was used to analyze the extract of total heavy metal reduction. The results were compared with the control to determine the heavy metal degradation capacity (%) as follows:
56
Molecular characterization of selected bacterial isolates
DNA extraction
The bacterial isolates were initially cultured for 48 h at 37°C on LB broth medium. Following incubation, 2 mL of the bacterial cell cultures were extracted and centrifuged for 10 min at 15,000 rpm. Genomic DNA was then extracted from the pellets (cells). The genomic DNA of all selected and screened potential bacterial isolates was successfully extracted using a bacterial genomic DNA extraction kit following the instructions of the manufacturer of the DNA extraction kit. For future use, each extracted DNA sample was kept at 4°C.
Measurement of DNA concentration and purity
The concentration of DNA was measured with the help of a NanoDrop spectrophotometer. To the NanoDrop, 1.5 µL of Tris-EDTA elution buffer was used as a blank. After removing the blank using tissue paper, a 1.5 µL sample was added. 24 The concentration of DNA was expressed in ng/µL. The purity of the extracted DNA was also assessed qualitatively using agarose gel electrophoresis. 16
Amplification of 16S rRNA gene region
Amplification using the polymerase chain reaction (PCR) was carried out at the Molecular Biology Laboratory, Institute of Biotechnology, University of Gondar. The 16S rRNA gene was amplified using the universal 16S primers forward [27F (5'-AGAGTTTGATCA TGGCTCAG-3’)] and reverse [1492R (5'-TACGGCTACCTTGTTACGA-3′)] to assess the bacterial identity. 56 A 50 µL reaction mixture comprising 1 µL (10 ng) DNA extract as a template, 5 mM primers, 25 mM MgCl2, and 2 mM dNTPs, as well as 1.5 U of Taq polymerase, nuclease-free water, and buffer, was used to carry out the PCR. 57 The PCR was performed in a thermal cycler. The following cyclic conditions were used: cycle 1 and cycle 2 for denaturation at 95°C for 5 min and 1 min, respectively; cycle 3 for annealing at 53°C for 1 min; cycle 4 and cycle 5 for amplification at 72°C for 1.5 and 10 min, respectively; followed by a 4°C hold temperature. Then, after loading the 1.5% (w/v) agarose gel electrophoresis in 1× TAE buffer and staining it with ethidium bromide (0.5 μg/mL), the PCR products were examined using gel electrophoresis and the Gel Documentation System UC light. 24 The amplicons were then kept for further use at −20°C.
Sequencing and phylogenetic tree analysis
The bacterial isolates of forward and reverse PCR products were sent and sequenced at Macrogen Company from the Netherlands and sequenced using the primers used in PCR. 24 Following receipt of sequenced PCR-amplified products of bacterial isolates, their sequences were edited using BioEdit. The obtained sequence of the 16S rRNA gene sequences of bacterial strains and their consensus region was submitted to the Basic Local Alignment Search Tool (BLAST) (https://blast.ncbi.nlm.nih.gov/) in order to find homologous sequences in GenBank. The 16S rRNA gene sequences of the bacterial strains in the current study were aligned with reference sequences demonstrating sequence similarity from the National Center for Biotechnology Information (NCBI) (https://www.ncbi.nlm.nih.gov) Database using the multiple sequence alignment tool of Molecular Evolution Genetic Analysis (MEGA), version 11.
Gene sequence data were phylogenetically analyzed using the neighbor-joining strategy, and distances were computed using the maximum composite likelihood method. The bootstrap program was used to analyze the branching patterns using sets of 1000 bootstrap repetitions. The BLAST algorithm was used to retrieve homologous sequences in GenBank. 21
Data analysis
The data collected in this study were analyzed using the Statistical Package for Social Sciences software version 21 for Windows. To determine whether there were significant variations between the means, the means and standard deviations of the triplicates were assessed using one-way analysis of variance (ANOVA), which was followed by Duncan's multiple range test. Finally, a variable with a p-value less than or equal to 0.05 (p ≤ 0.05) was considered to be statistically significant. Nucleotide sequence and phylogenetic tree analyses were performed using BioEdit and MEGA11 bioinformatics tools, respectively.
Results
Isolation and screening of heavy metal-resistant bacteria
In this study, from a total of 18 collected heavy metal-contaminated soil samples, 21 bacterial isolates were obtained. The bacterial isolates were coded as Agri10−2Is2, Agri10−3Is2, Agri10−3 Is1, AL10−1Is2, AL10−3Is1, GA10−2Is1, etc. and were subjected to multiple heavy metal resistance tests, and only six potential isolates (Agri10−2Is2, Agri10−3Is2, Agri10−3 Is1, AL10−1Is2, AL10−3Is1, GA10−2Is1) that have multiple metal 100 μg/mL (Cu, Cr, and Pb) resistance potential were selected; however, the remaining 15 isolates were sensitive or did not have multiple metal resistance and were not selected for further activities (Table 1). These six heavy metal-resistant bacterial isolates were subjected to morphological and biochemical characterization tests, MIC analysis, and studies on antibiotic resistance.
Multiple heavy metal resistance responses in potential bacterial isolates.

R indicates resistance to metals.
Characterization of heavy metal-resistant bacterial isolates
Morphological characterization of bacterial isolates
From individual bacterial colonies, which were obtained from serial dilution of soil samples, four had creamy colony color and one had red–yellowish colony color, while the remaining had white colony color. With respect to colony shape, four had an irregular shape, while two exhibited a circular shape, and all isolates had rod cell shape. All isolates exhibited raised elevation and were entire in edge/margin. On the other hand, the remaining isolates were medium in size, with three being small. Rough and smooth textures were exhibited among isolate colonies (Table 2, Figure 2).
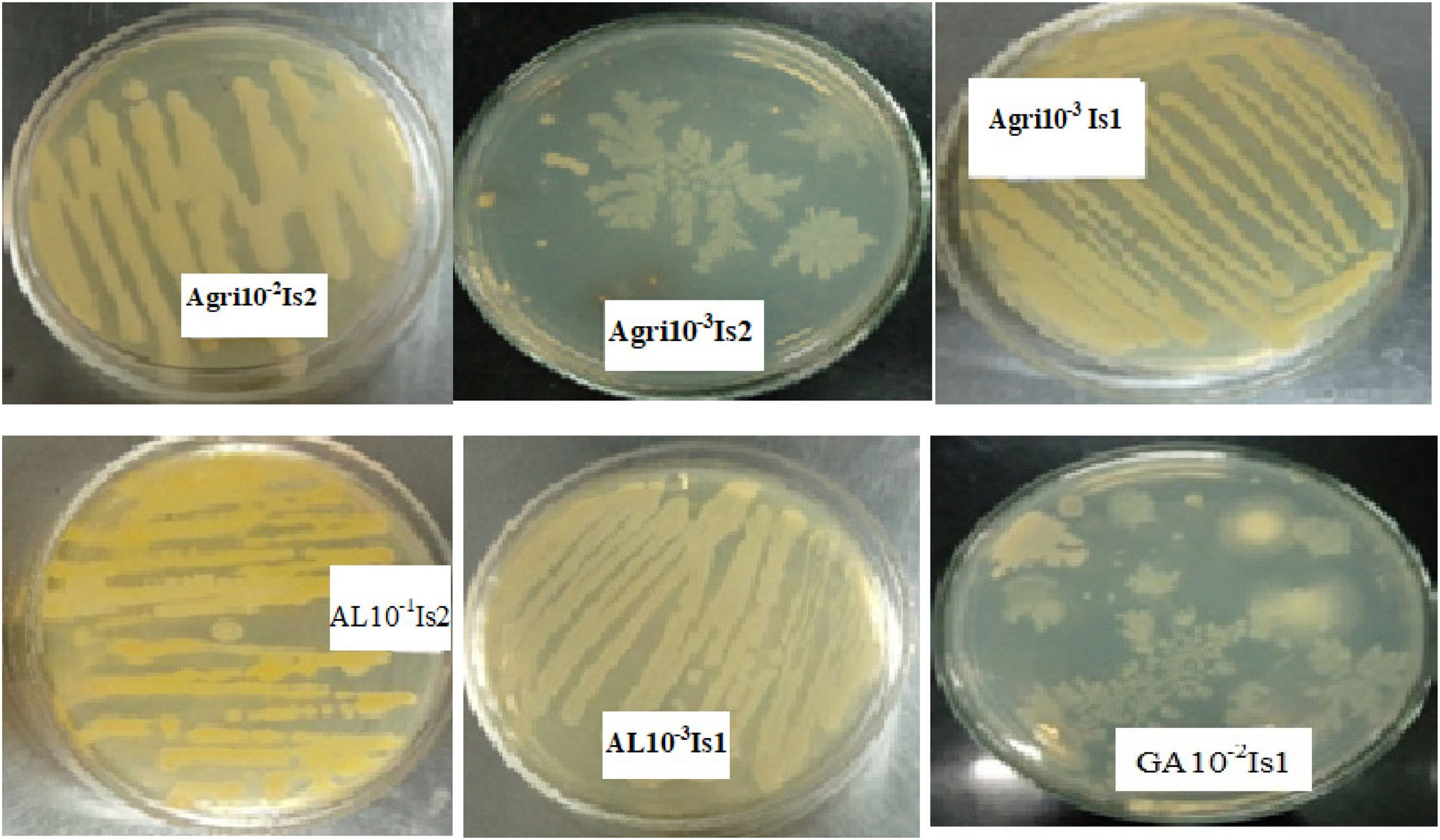
Colony morphological features of selected bacterial isolates.
Morphological characterization of bacterial isolates.

Microscopic characterization of bacterial isolates
Gram staining technique was done to differentiate the Gram-positive and Gram-negative bacterial isolates and viewed under a light microscope. According to microscopic observation of bacterial isolates under oil immersion magnification (100×), all of the six bacterial cells had rod shape, and out of six bacterial isolates, four were Gram-positive rods (Agri10−2Is2, Agri10−3Is2, Agri10−3Is1, and AL10−1Is2), while two were Gram-negative rods (AL10−3Is1 and GA10−2Is1) (Table 3).
The biochemical characteristics test results of the selected bacterial isolates.

+ indicates positive value, while − indicates negative value.
Biochemical characterization of bacterial isolates
In the current study, various biochemical assays were conducted on each isolated heavy metal-resistant bacterium. There were differences among the isolates for the various tests. Table 3 displays the results of each biochemical test. On the other hand, the Gram staining results are depicted in Figure 3.
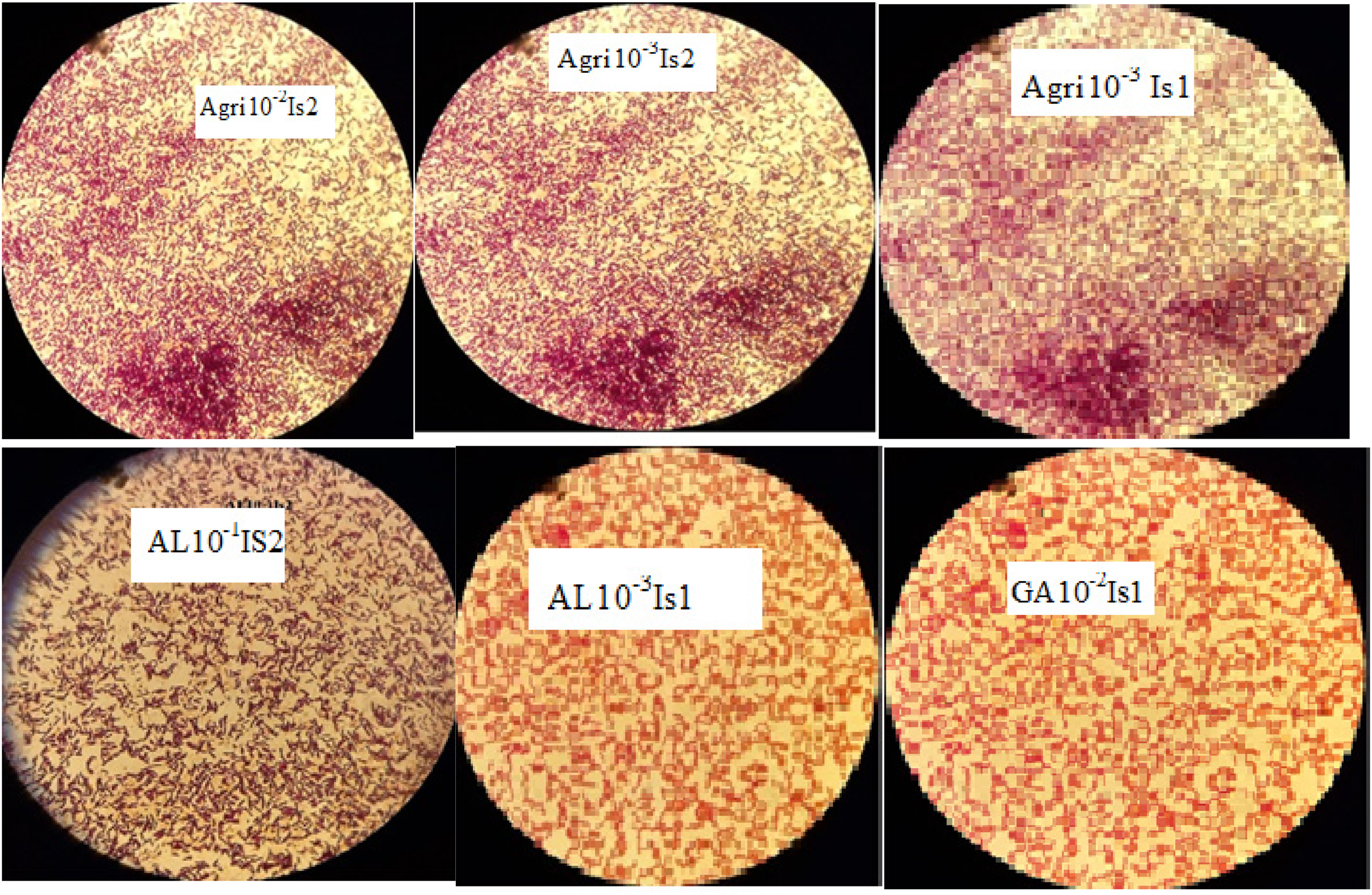
Microscopic view of selected bacterial Gram-stained cells.
Determination of MIC
In this study, the MIC of heavy metals was determined by growing different bacterial isolates on LB medium supplemented with metals (3, 6, 12, 25, 50, and 100 µg/mL) of Pb, Cr, and Cu separately. The six bacteria had different MIC values on different metal concentrations (Cu, Cr, and Pb) separately. The bacterial isolates showed a high degree of resistance to the tested heavy metals as MIC values of the isolates ranged from 12 to 100 µg/ml. There was a progressive decline in the growth of the bacterial isolates as concentrations of heavy metals increased. Finally, no growth was observed. MICs of heavy metals are shown in Table 4.
MIC (μg/mL) of different heavy metals against bacterial isolates.

Assessment of antibiotic resistance
In this study, six potential heavy metal-resistant bacterial isolates (Agri10−2 Is2, Agri10−3 Is2, Agri10−3 Is1, AL10−1 Is2, AL10−3 Is1, and GA10−2 Is1) were further evaluated for their antibiotic resistance using three common broad-spectrum antibiotic discs (tetracycline, ampicillin, and gentamicin). All the six potential bacterial isolates were found to be resistant to all tested antibiotics since no clear zone formed around any of the tested antibiotic discs.
Determination of heavy metal removal capacity
The heavy metal removal efficiency of potential heavy metal-resistant bacterial isolates was determined by measuring the decrease or reduction in concentration of Cr, Cu, and Pb in LB broth medium after 72 h of incubation at a temperature of 37°C and pH 7. The results are shown in Figure 4. In this study, the biodegradation efficiency of the six potential bacterial isolates was measured using an atomic absorption spectrophotometer, which showed considerable degradation of Cr, Cu, and Pb. Isolates had almost similar biodegradability potential to selected heavy metal Cu, while isolates had different removal potential to Pb and Cr. The biodegradation capacity of the different isolates against Cu, Cr, and Pb ranged from 92.2% to 92.85%, 91.4% to 95.07%, and 94.53% to 97.09%, respectively.

Biodegradation % efficiency of heavy metals.
Molecular characterization of selected bacterial isolates
DNA extraction and measurement of DNA concentration and purity
Using a bacterial genomic DNA extraction kit, the genomic DNA of the six bacterial isolates that were selected and screened was successfully extracted. Quantification (DNA concentration) values for isolated bacteria Agri10−2 Is2, Agri10−3Is2, Agri10−3 Is1, AL10−1Is2, AL10−3Is1, and GA10−2Is1 were 25, 36, 45, 65, 55, and 54 ng/µL with a quality of 1.8, 1.9, 1.9, 2, 1.8, and 1.75, respectively, based on their A260/A280 ratios. DNA was qualitatively estimated by running on an agarose gel electrophoresis. This gel was used to determine the concentration and purity of the DNA, which moved along the lanes of wells 1, 2, 3, 4, 5, and 6. This shows that lane L, a 100 bp DNA ladder, is present along with the bacterial genomic DNA of Agri10−2Is2, Agri10−3Is2, Agri10−3Is1, AL10−1Is2, AL10−3Is1, and GA10−2Is1, respectively (Figure 5).

PCR result of selected bacterial isolates. The lanes 1, 2, 3, 4, 5, and 6 indicate genomic DNA of bacterial isolates Agri10−2Is2, Agri10−3Is2, Agri10−3 Is1, AL10−1Is2, AL10−3Is1, and GA10−2Is1, respectively, and together with lane L which is DNA ladder of 100 bp.
PCR amplification of 16S rRNA gene from extracted DNA
Following DNA extraction, the 16S RNA gene was amplified using forward primer 27F and reverse primer 1492R. The amplicon's approximate molecular size, as determined by the PCR amplification result, was 1500 bp. The size and purity of each selected bacterial isolate PCR result employing agarose gel electrophoresis with a 100 bp (L1) DNA ladder are displayed in Figure 5.
Phylogenetic tree analysis
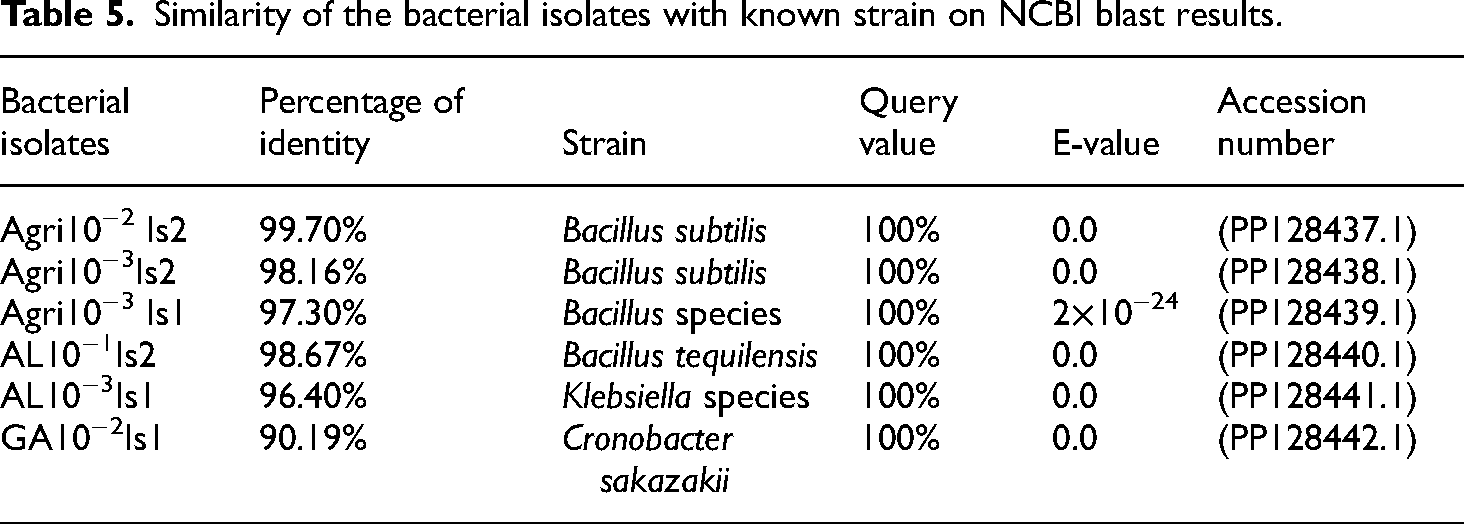
The genomic DNA PCR products of the six potential bacterial isolates were sequenced at the Macrogen Company in the Netherlands. After the PCR-amplified products from each bacterial isolate were sequenced, their forward and reverse sequence products were edited, and their consensus regions were obtained using BioEdit and MEGA11 bioinformatics software. Heavy metal-resistant bacterial isolates Agri10−2Is2 and Agri10−3Is2 were found to be 99.7% and 98.16% identical to B. subtilis, respectively. Agri10−3Is1 was also found to be Bacillus species that have identity 97.30%. On the other hand, isolates AL10−1Is2, AL10−3Is1, and GA10−2Is1 were 98.67%, 96.40%, and 90.19% identical to Bacillus tequilensis, Klebsiella species, and Cronobacter sakazakii, respectively. Their percentage of identity was obtained as shown in Table 5 and Figure 6.

Phylogenetic tree constructed using sequence of bacterial isolates with related sequences from GenBank (NCBI). Numbers indicate evolutionary distance versus branch length.
Similarity of the bacterial isolates with known strain on NCBI blast results.
Discussion
Currently, bacteria have been considered one of the remarkable agents for bioremediation of polluted environments. This has initiated microbiologists to seek potential bioremediating bacteria from different polluted sources, specifically in solid or liquid waste dumping sites, and subsequent evaluations of their specific metal and antibiotic resistance properties have been evaluated. In this regard, Bacillus species are among the predominant bacteria isolated from soil and water and commonly applied as bioremediation agents. Several studies, including the ones described by Yuliani et al., 56 have indicated the ability of Bacillus species to decrease heavy metals such as Cr and Pb. Bacillus species such as B. subtilis, B. megaterium, B. licheniformis, B. cereus, B. circulans, and B. barbaricus have been investigated for their capacity to resist Cr and Pb. 15
Today, various microbes, especially bacteria, have developed resistance mechanisms against heavy metals. Bacteria are resistant to several metals, such as Cu, Cr, Pb, Hg, Zn, and Cd, due to specific detoxifying mechanisms and the presence of resistance genes, which may be mediated by chromosomal DNA or plasmid DNA. 11 Many bacteria have bacterial plasmids that encode resistance mechanisms against toxic metal ions. 57
In this study, from a total of 18 collected heavy metal-contaminated soil samples, 21 bacterial isolates were obtained. From these, six (Agri10-2Is2, Agri10-3Is2, Agri10-3 Is1, AL10-1Is2, AL10-3Is1, GA10-2Is1) potential isolates were screened based on their multiple heavy metal (Cu, Cr, and Pb) resistance properties. These six potential bacterial isolates were declared as multiple heavy metal-resistant isolates. These resistant bacterial isolates were further subjected to additional testing, including molecular analysis, metal removal analysis, antibiotic resistance investigations, MIC analysis, and morphological and biochemical characterization.
Of the bacterial isolates that were obtained from serial dilution of soil samples, four had creamy and one had red–yellowish colony color, while the remaining one had white colony color. With respect to colony shape, four had irregular shape, while two exhibited circular shape, and all isolates had ridden in cell shape. All isolates exhibited raised elevation and entire edges/margins. On the other hand, three of the isolates exhibited small size, whereas the remaining had medium size. Rough and smooth in textures, the colonies were exhibited among isolates. Microscopic observation tests revealed that all of the bacterial cells had a rod shape. These findings are in line with previous research by Hussain and Raza, 21 which showed that Bacillus species account for the majority of these morphological traits.
In this study, each isolated heavy metal-resistant bacterium was subjected to a variety of biochemical tests. Variations were observed for the different tests across the isolates. All six isolates had positive test results for catalase, citrate, hydrogen gas production, and citrate utilization, while negative results for hydrogen sulfide production tests. On the other hand, out of six isolates, five were motile; however, one was non-motile, and finally, among six isolates, four were Gram positive, while two were Gram negative. This result was similar to the research findings by Kalaimurugan et al. 58
In the current study, six of the most potential bacterial isolates were tested for their resistance effect to metals such as Cu, Cr, and Pb to assess their MIC by increasing the concentration of metals until bacteria fail to grow in metal-supplemented LB medium. It was shown that the bacterial isolates were able to grow at different concentrations (3, 6, 12, 25, 50, and 100 μg/mL). All six bacteria had different MIC values on different metal concentrations (Cu, Cr, and Pb) separately. The bacterial isolates showed a high degree of resistance to the tested heavy metals, as the MIC values of the isolates varied from 12 to 100 µg/ml. In general, there was a progressive decline in the growth of the bacterial isolates as concentrations of heavy metals increased. This finding is similar to the research work of Ramesh and Patel. 59 Gangwar et al. 60 isolated two heavy metal-resistant bacteria, Comamonas testosteroni (S4C1) and Bacillus cereus (S5C3). Both isolates were found to be highly resistant to lead (Pb), copper (Cu), chromium (Cr), cadmium (Cd), and nickel (Ni), and they show different MICs against the above heavy metals at different levels.
In this study, antibiotic resistance was also assessed in potential heavy metal-resistant bacterial isolates. All six potential isolates were subjected to the antibiotic susceptibility test, which revealed that the bacteria were resistant to gentamicin, ampicillin, and tetracycline. No clear zone around the antibiotic discs was excited in this study, and this indicates that the bacteria that are resistant to heavy metals also had resistance properties to antibiotics. Reports suggest that metal tolerance and antibiotic resistance are related. 42 Long-term exposure to heavy metals can alter the structure and functions of bacterial isolates, causing them to develop resistance to antibiotics. 5 The grouping of these genes inside the same plasmid is the cause of the phenomena known as metal tolerance mechanisms, which contribute to the increase in antibiotic resistance. 43 Heavy metal ions are known to co-regulate genes responsible for antibiotic resistance and decrease antibiotic susceptibility. 37 Environmental pollution not only triggers heavy metal co-selection processes but also increases the level of tolerance to some antibiotics due to co-regulation of resistance genes. 45 The combined expression of antibiotic and heavy metal resistance by the six isolates in the current study may not be a chance phenomenon but rather a result of selection by heavy metal present in an environment.61–63 Therefore, pragmatic measures must be taken to limit the spread of antimicrobial resistance across the environment and to reduce the incidence of healthcare-associated infections.
In this study, the removal efficiency of the six potential bacterial isolates was measured using an atomic absorption spectrophotometer, which showed considerable degradation of Cr, Cu, and Pb, and the isolates had almost similar biodegradability potential to the selected heavy metal Cu. While isolates had different removal potential to Pb and Cr, they had relatively less removal potential to Cu. Out of six selected potential isolates, Agri10−2Is2 had a 95.07% removal capacity for Cr, and the isolate also had 92.2% and 95.51% removal capacity for Cu and Pb, respectively.
The isolates Agri10−2Is2, Agri10−3Is2, Agri10−3Is1, AL10−1Is2, AL10−3Is1, and GA10−2Is1had removal capacities of 95.07%, 94.07%, 93.07%, 93.4%, 91.4%, and 94.4% of Cr + 3, respectively, and also isolates had 92.2%, 92.57%, 92.37%, 92.66%, 92.85%, and 92.52% Cu + 2 removal, respectively. For Pb, isolates Agri10−2Is2, Agri10−3Is2, Agri10−3 Is1, AL10−1Is2, AL10−3Is1, and GA10−2Is1 had removal potential of 95.51%, 94.53%, 95.62%, 96.87%, 94.86%, and 97.09%, respectively. This result is in concordance with the research reported by Abd Elhady et al. 10 . Tinatin et al. 39 reported that three bacterial species (A. faecalis, D. tsuruhatensis, and Stenotrophomonas species) were found to be highly efficient in removing (remediating) specified heavy metals (Pb, As, Co, and Ni) from soil.
In this study the molecular characterization of bacterial isolates was also conducted. The genomic DNA of the six potential bacterial isolates was successfully extracted using the Sigma-Aldrich (Millipore Sigma) DNA extraction kit. The extracted bacterial genomic DNA was amplified using regions of the 16S rRNA gene of forward primer 27F and reverse primer 1492R. The amplicon's approximate molecular size, as determined by the PCR amplification result, was 1500 bp. The DNA quality was estimated by running on agarose gel that determines the presence of pure DNA in agarose gel electrophoresis. The concentration and purity of DNA were checked using agarose gel electrophoresis, which migrated along the lanes of wells 1, 2, 3, 4, 5, and 6. This indicates that the bacterial genomic DNA of Agri10−2 Is2, Agri10−3Is2, Agri10−3 Is1, AL10−1Is2, AL10−3Is1and GA10−2Is1, respectively, as shown in Figure 5.
At the Macrogen Company in the Netherlands, the 16S rRNA gene PCR-amplified products of the six potential bacterial isolates were sequenced using forward and reverse primers. Each bacterial isolate's forward and reverse sequencing products were edited using MEGA11 and BioEdit software following PCR amplification.
The gene sequences obtained from the current study and those of closely related sequences from GenBank (NCBI) were aligned. Finally, their phylogenetic tree was constructed that displays the phylogenetic relationship of the bacteria strains by a maximum composite likelihood-based neighbor-joining tree method.
The percentage of identity of the bacterial isolates Agri10−2Is2 and Agri10−3Is2 was found to be 99.7% and 98.16% identical to B. subtilis, respectively. Agri10−3Is1 was also found to be Bacillus species. On the other hand, AL10−1Is2, AL10−3Is1, and GA10−2Is1 are 98.67%, 96.40%, and 90.19% identical to B. tequilensis, Klebsiella species, and C. sakazakii, respectively.
Molecular nucleotide sequence analysis was done using software like BioEdit and MEGA11. Based on percentage of identity, the sequences from this study were compared to highly closely matched bacterial species from NCBI. Agri10−2Is2 was matched with B. subtilis (PP128437.1) with its percentage of identity at 99.70%, e-value at 0.0, and query of 100%. Agri10−3Is2 was matched with B. subtilis (PP128438.1) with its percentage of identity at 98.16%, e-value at 0.0, and query of 100%. Agri10−3Is1was was matched with Bacillus species (PP128439.1) with its percentage of identity at 97.30%, e-value at 2e-24, and query of 100%. AL10−1Is2 was matched with B. tequilensis of accession number (PP128440.1) with its percentage of identity at 98.67%, e-value at 0.0, and 100% query. This bacteria strain is confirmed by the information reported by Yuliani et al. 56 states that the bacteria strain Bacillus species is the predominant soil bacterium that has the potential ability to degrade Pb and Cr in the soil. AL10−3Is1 was matched with Klebsiella species (accession number PP128441.1) with its percentage of identity at 96.40%, e-value at 0.0, and 100% query. Among the six bacterial isolates, GA10−2Is1 was more similar to C. sakazakii (PP128442). Among the six bacterial isolates, Agri10−2Is2 and Agri10−3Is2 were similar to B. subtilis, which has the potential ability to reduce Cu and Pb. 56 The phylogenetic tree of these isolates was then analyzed using the MEGA11 software to determine the evolutionary relationships between the acquired bacteria and similar strains deposited in GenBank. All the six potential bacterial isolates shared a common ancestor, and each isolate was evolutionarily related to its corresponding well-known bacterial strains from NCBI GenBank data (Figure 6). Many of the researchers identified heavy metal-tolerant bacterial isolates using 16S rRNA gene sequencing.34,64,65 Martins et al. 66 reported that different bacterial species, namely, B. cereus, S. aureus, E. coli, and Proteus species, isolated from samples collected from a cement company in Nigeria exhibited an impressive tolerance to heavy metals. These findings exhibited that microorganisms can adapt to polluted heavy metal environments and be utilized in bioremediation initiatives. The data obtained in the current study suggested a possible application of the six isolates for detoxification of heavy metals in food processing industries and for wastewater treatment and were considered remarkable agents for bioremediation of heavy metal-contaminated soils or sites.
The limitation of our study was the small number of sample heavy metals and antibiotics tested.
Conclusion
The current study revealed the presence of various potential heavy metal-resistant bacteria in solid waste dumping sites, especially metal-contaminated environments, which are efficient and useful for bioremediation. Six isolates, namely, Agri10−2Is2, Agri10−3Is2, Agri10−3Is1, AL10-1Is2, AL10−3Is1, and GA10−2Is1, were obtained that were resistant to high metal concentrations. All the six bacterial isolates exhibited multiple tolerances to Cu, Pb, and Cr. Therefore, the study concluded the presence of potential heavy metal and antibiotic-resistant bacteria that have potential application for the new technology of bioremediation of contaminated environments and biodegradation of toxic heavy metals. Their safety must be evaluated before their in vivo application in different processing industries. Future research should be done to improve the genetic modification of indigenous bacteria that may help to expand the field of accessible methodologies to identify genes responsible for metal resistance to enhance the bioremediation potential of recombinant as well as indigenous bacteria.
Footnotes
Acknowledgements
Abebech Dinku would like to acknowledge Debre Markos University for sponsoring her MSc study. Both authors would like to extend their gratitude to University of Gondar.
Author contributions
AD performed the experiments as part of her MSc's thesis work. The idea conception was initiated by TMJ. AD and TMJ performed the analysis together. This work was carried out under the supervision of TMJ. TMJ also helped in editing the manuscript.
Data availability
All other data generated or analyzed during this study are included in this published article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical considerations
Permissions were obtained from concerned authorities of the sampling sites to collect samples.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.