Abstract
The use of antimicrobials in the food animal industry has caused an increased prevalence of antimicrobial-resistant bacteria and antimicrobial resistance genes, which can be transferred to the microbiota of humans through the food chain or the environment. To reduce the development and spread of antimicrobial resistance, restrictions on antimicrobial use in food animals have been implemented in different countries. We investigated the impact of an antimicrobial restriction intervention during two generations of pigs. Fecal samples were collected in five growth phases. The frequency of antimicrobial-resistant coliforms and antimicrobial-resistant bacteria or antimicrobial resistance genes was analyzed. No differences in the richness or abundance of antimicrobial-resistant coliforms or antimicrobial resistance genes were found when animals fed with or without prophylactic antimicrobials were compared. Withholding antimicrobial supplementation did not negatively affect weight gain in pigs. Withdrawal of prophylactic antimicrobial consumption during two generations of pigs was not enough to reduce the prevalence of antimicrobial resistance genes, as measured by richness and abundance markers. This study indicates that the fitness costs associated with bacterial carriage of some antimicrobial resistance genes are low.
Keywords
Introduction
The use of antimicrobials has been effective worldwide in the treatment and control of bacterial infections and contributed to greater human life expectancy. 1 However, a lack of antimicrobial stewardship has created a selective force for the increased frequency of antimicrobial-resistant bacteria (ARB) and antimicrobial resistance genes (ARGs) that threatens current and future antimicrobial therapeutic efficacy.2,3
Approximately 80% of the total antimicrobial production in the world is used in food animals. 4 It is likely that larger amounts of antimicrobials are used in animals in less industrialized countries that lack regulatory policies for antimicrobial use.4–6 Commensal bacteria are ubiquitous in animals’ and humans’ intestines and are considered good antimicrobial resistance (AMR) indicators.7,8 The use of antimicrobials in food-producing animals can cause the proliferation of commensal bacteria with ARGs. Genes that code for AMR can then be horizontally transferred to many other bacterial species (including pathogens) in the intestines.8–11 Antimicrobial-resistant commensal bacteria from food animals can also end up in food animal products such as meat and dairy products; 12 these bacteria can then colonize human intestines and transfer resistance genes to opportunistic human pathogens through mobile genetic elements (MGEs).13–17
Limiting antimicrobial use in food animals reduces selective pressures for ARGs and can contribute to the decrease of ARB in food animals and humans.18–21 However, ARB may persist even in the absence of such selective pressures due to compensatory evolution that allows bacteria to maintain ARGs without apparent fitness costs.22,23 Experiments in which animals are deprived of antimicrobials for growth promotion have revealed contradictory results; some show a decline in ARB, while others report the persistence of intestinal ARB and ARGs even in the absence of such antimicrobials.21,24 In fact, multidrug-resistant (MDR) intestinal bacteria have even been found in animals from organic farms.25–28 The main objective of this study was thus to analyze the effect of the removal of antimicrobial prophylaxis (often given at higher doses than for growth promotion) on the prevalence of ARB and ARGs in fecal microbiota in swine over the span of two generations.
Materials and methods
Ethical approvals
The experimental procedures for this study were approved by the Animal Ethics Committee at la Universidad San Francisco de Quito document Nr 2016-004. A technical committee of the food animal company used in this study also provided their approval prior to beginning research (Document number CnCeCa1604). The company's consent was explicit, as the protocols were designed jointly between the researchers and the company's technical personnel; this agreement was confirmed in a consent letter sent from the company to the Animal Ethics Committee at la Universidad San Francisco de Quito.
This experimental design was approved by farm managers. All facility housing and sampling methods were performed under the supervision of the food animal company's veterinarian. Approval of all protocols and publications was also obtained from the technical managers of the company. The name of the company was kept confidential at the company's request. There is no national requirement to obtain IRB approval for this type of study.
Animals
Overall, 40 pigs were included in this study. Twenty female pigs (sows) were included in the first generation of pigs and selected for fecal sampling with rectal swabs at 180 days of age. Following parturition of the first generation of sows, 20 piglets (10 piglets in each treatment group) were selected to continue the same treatment group as their mothers and followed for 150 days. Fecal samples with rectal swabs were taken from piglets at days 5, 30, 50, 100, and 140. A random, balanced, fixed-effect, double-blind study was conducted in two generations of pigs housed at a swine farm in Ecuador. The entirety of the study was performed under preexisting facility biosafety requirements.
The experimental unit considered in this study was a pig. The number of animals needed was calculated with a statistical power of 87% considering the probable loss of experimental units. Using a simple randomization method, replacement sows donated by a separate commercial swine farm were randomly selected for inclusion in the study's first generation of pigs. From a batch of 70-day-old female PCI1050 pigs, 20 pigs were randomly assigned to one of the 2 groups. One group contained 10 female pigs (the control group) and received conventional feed formulation with antimicrobial additives, identified with a yellow color in the feed bag (Group A). The other group contained 10 female pigs and received feed containing no antimicrobial additives, identified with a purple color in the feed bag (Group B). First-generation pigs received their group's feeding regimen until they were farrowed. These sows’ offspring were considered the second generation of pigs in this study. Each sow gave birth to an average of 12 piglets. Piglets were placed with their corresponding mothers and weaned at 20 days. Piglets were assigned to the same treatment group as their mothers and received their respective treatment (antimicrobials or no antimicrobials) over the course of their entire productive cycles. Each piglet was identified with a numbered earring. Once piglets were 70 days old, they were moved to a larger facility for fattening (71 days–170 days). The same animals were maintained in each experimental group when animals were moved. The transportation of each group was performed in different vehicles within the same day. Overall, animals from each group were monitored for a total of 150 days (Figure 1).

Workflow for each treatment group of pigs (Groups A and B). Treatment Group A was fed with antimicrobial supplements. Treatment Group B was fed with no antimicrobial supplements. Ten young female pigs (70 days) were randomly selected for each treatment. All piglets born were maintained under the same treatment as their mothers. A homogeneous group of 32 piglets (similar age and weight) within each treatment group were selected until the end of the experiment. Rectal swab samples were collected at days 5; 30; 100; and 140. Samples from sows were taken 180 days after they were born (6).
Productive performance was observed based on daily weight gain during the study period and carcass weight after slaughter. The type and concentrations of antimicrobials used in the feed for Group A changed over time and were consistent with the feeding regimen that had been used routinely on the farm for the previous 2 years. Briefly, under conventional farm conditions, antimicrobials are used to prevent infections in pigs caused by opportunistic pathogens. The normal regimen begins with a mixed administration of tilmicosin (macrolide) and colistin to prevent the major health concerns, most often respiratory and gastrointestinal infections, which piglets may face at this stage. Since the completion of this study, this regimen has changed because of a recent national ban on colistin use in food animals. Starting at 29 days of age, chlortetracycline is added to the pig’s’ food and maintained until pigs are 123–139 days old. Additionally, tiamulin is used from 29 to 70 days of age. To complement antimicrobial prophylaxis, trimethoprim–sulfamethoxazole (SXT) and doxycycline were also added to water sources from 37 to 47 days of age. Antimicrobial additives used are described in Table 1.
Antimicrobial additives used in pig’s farm as prophylactics in Group A.

Husbandry
The maternity and weaning phases (piglets until 70 days of age) were carried out in pens with plastic slatted flooring, while the fattening phase was carried out in a different facility with cement floors that were washed daily. In all growing phases, pens were protected from the sun with a roof and ceiling measuring 2.5 m high. During the weaning and fattening phases of the pigs, each pen included 32 piglets. Pig density was 0.45 m2/pig in the weaning phase and 0.90–1.0 m2/pig at the fattening phase. The number of cages is described in Figure 1.
Tap water and feed were administered ad libitum and facility temperature ranged from 20 to 26°C. All experiments were performed under veterinary supervision and vaccines were administered to all animals. Prior to beginning the study, it was specified that antimicrobial treatment would be administered under veterinarian supervision to any animal diagnosed with an infection treatable with antimicrobials over the course of the study. If any sick animal was detected, it would be immediately separated into a different cage to be treated and eliminated from the study. If a pig belonging to the antimicrobial deprived group (Group B) required antimicrobial treatment, they would be eliminated from the study.
Both animal groups (those given and not given antimicrobials) were maintained at the same location with separate feeding and watering supplies. Animals of different treatment groups were fed with different instruments. Pens were kept separated by a 90 cm corridor in all growing phases. There was no airflow control or physical separation of different treatment pens.
Sows that were not included in the study were incorporated into conventional productive cycles within each farm batch. Animals were released from the experiment on day 150 and humanely slaughtered on day 170. Humane slaughter was performed using electric stunning followed by exsanguination, also known as bleeding out.
Samples and phenotypic analysis
At each sampling time point (at 180 days of age for the first-generation sows and then at 5, 30, 50, 100, and 140 days of age for the second-generation piglets), two rectal swabs were taken from each animal (n = 20; 10 pigs within Group A and 10 pigs within Group B). For sample collection, sterile swabs were introduced into the rectum of the animal with circular movements to obtain fecal matter. One swab was stored at −20°C within 2 h of sample collection and transported on dry ice to the Microbiology Institute at la Universidad San Francisco de Quito. Frozen samples were stored in a −80°C ultra-freezer for further analyses. The other swab was placed in a 15 mL sterile tube maintained on ice and used for microbiological analysis within 2 h of collection.
Replacement sows (from the first generation of pigs) were sampled when they were 180 days old, prior to the insemination process. Unfortunately, due to practical restrictions, it was not possible to sample these sows prior to this age. To avoid any animal injury, sampling was always completed at the beginning of the day prior to feeding animals. Group A was sampled first and Group B second. Different farm operators helped catch animals within each treatment group. The same animals were used as subjects of this study along the entire productive cycle, and it was not necessary to replace any of them, as there were no losses to health problems or mortality in either group.
Intestinal coliforms were used as microbial indicators of phenotypic resistance (Figure 1). Antimicrobial-resistant coliform bacteria in the intestinal microbiota of pigs were quantified according to protocols described previously. MacConkey Lactose medium (MKL) was used to culture, isolate, and count coliform colony-forming units (CFUs) from fecal samples.
Briefly, swabs were eluted in 5 mL of sterile tryptic soy broth (TSB) and diluted in phosphate-buffered saline (PBS) (0.0169 M KH2PO4, 0.0719 M K2HPO4, pH 7.2). A 0.1 mL aliquot of this solution was serially diluted in 0.9 mL of PBS up to 10−3, and a 0.1 mL aliquot from each dilution was plated on MKL plates with and without antimicrobials.18,20,29 Some MKL plates were supplemented with antimicrobials in the following concentrations: 4 mg/L of trimethoprim and 76 mg/L of sulfamethoxazole (MKLSXT), 27 32 mg/L of tetracycline (MKLTE), 28 or 16 mg/L of ampicillin (MKLAMP). 30 After 24 h of incubation, lactose-positive colonies were counted from each antimicrobial supplemented MKL plate (n = 30 plates from each sampled group), and the count was compared with the corresponding colony count on MKL plates without antimicrobial supplementation (n = 10 plates from each sampled group). The ratios of resistant coliforms were estimated by counting the number of colonies on MKL plates with antimicrobials divided by the number of colonies on MKL plates without antimicrobials. 31 Only plates with coliform counts greater than four CFUs were used. One lactose fermenting (coliform) colony from each plate (with antimicrobial and without antimicrobial supplementation) was then isolated and stored at −80 °C in TSB with 30% glycerol until used for antimicrobial susceptibility analyses (n = 960).
Antimicrobial susceptibility test
Frozen isolates were revived on MKL agar for 16–24 h. Antimicrobial susceptibility tests were then performed using ampicillin (AMP; 10 μg), tetracycline (TE; 30 μg), trimethoprim–SXT(1.25/23.75 μg), gentamycin (GEN; 10 μg), amoxicillin–clavulanic acid (AMC; 20/10 μg), ciprofloxacin (CIP; 5 μg), chloramphenicol (CHLOR; 30 μg), and ceftriaxone (COX; 30 μg). These antimicrobials were selected as representatives of the most commonly used families of antibacterial drugs in human healthcare settings. The antimicrobial susceptibility test (Kirby-Bauer Disc Diffusion test) was carried out by following Clinical & Laboratory Standards Institute (CLSI) guidelines. Isolates were described as sensitive if the zone of inhibition of growth was greater than that described by the CLSI guidelines.
Molecular analysis
For molecular analysis, five rectal swabs taken from sows and piglets within each treatment group and in the same sampling time frame were frozen at −80°C until use. DNA was isolated using MO BIO Power Soil DNA Isolation Kit (MO BIO, Carlsbad, CA, USA) by following the manufacturer instructions. Nucleic acid quality and quantity were evaluated using NanoDrop (Thermo Fisher Scientific) and Qubit dsDNA HS (Thermo Fisher Scientific, Oregon, USA). For each sampling time point (six for Group A and six for Group B), DNA extracts from fecal samples of five animals were pooled.
8
The concentration of each DNA sample within the pool was 10μg/μL. Each pool was analyzed in duplicate with high-throughput quantitative polymerase chain reaction (qPCR). WaferGen SmartChip real-time PCR system was used to detect 384 genes, 338 of which are informative for ARGs or MGEs. Primers for ARGs, HT-qPCR protocols, and calculations of amplicon abundance and diversity were carried out as described previously32–36 (Supplementary material S1). Briefly, the difference between the cycle threshold (CT) of ARGs and 16S genes (ΔCT) was used to normalize and calculate the fold change in ARGs. The CT value of each amplicon was used to calculate ΔCT by the following formula: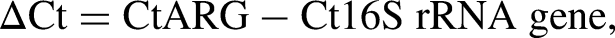
To calculate the difference in ARG abundance and diversity between animal groups (ΔΔCt), the following formula was used:


Statistical analysis
All data collected was recorded in MS EXCEL software. Descriptive and inferential statistical analyses were performed in INFOSTAT (Statistic Software Version 2017). The impact of the antimicrobial restriction on the coliform count, antimicrobial susceptibility, and animal performance was assessed by Student’s t-test and chi-square test, respectively, (p < 0.05). HT-qPCR data were analyzed as previously described.35,37
Results
Within each sampled group, 40 cultured plates were analyzed by counting the number of lactose-positive colonies on each MKL plate. Ratios of AMR (the proportion of antimicrobial-resistant coliforms in MKL agar with and without antimicrobials) showed no significant difference between treatment groups (Group A = 0.722 and Group B = 0.763; p = 0.434) (Table 2). Antimicrobial susceptibility tests of 537 randomly selected strains (Group A = 266 and Group B = 271) revealed resistance to AMP (n = 397; 73.9%), AMC (n = 188; 35%), TE (n = 434; 81.1%), trimethoprim–SXT (n = 301; 56.1%), GEN (n = 125; 23.3%), CIP (n = 71; 13.2%), CHLOR (n = 174; 32.4%), and COX (n = 77; 14.3%), and 65.9% of the strains were MDR (resistance to three or more antimicrobial classes; Table 3).
Ratio of antimicrobial-resistant coliforms carried by pigs.

Notes: Antimicrobial resistance (AMR) ratios for ampicillin (AMP), cotrimoxazole (SXT), and tetracycline (TE) were calculated by dividing the total count of coliform colony-forming units (CFUs) in MacConkey Lactosa (MKL) plates supplemented with antimicrobials by the total count of coliform in MKL without antimicrobials. The average and SD are shown for each treatment.
Analysis of antimicrobial susceptibility test from coliform isolated.

Notes: Strains isolated from pigs fed with antimicrobials are described in “Treatment A” columns and strains from piglets fee without antimicrobials are described in “Treatment B” columns. Within each group, results are organized by piglet’s age (1–5days; 2–30days; 3–50 days; 4–100 days; 5–140 days). p was calculated based on sampling period comparison with chi-square test. Strain resistant to more than two antimicrobial classes was counted as multidrug resistant (MDR). Antimicrobials used in the test are as follows: ampicillin (AM, 10 mg), tetracycline (TE, 30 mg), trimethoprim–sulfamethoxazole (SXT, 1.25/23.75 mg), gentamycin (GEN, 10 mg), amoxicillin–clavulanic acid (AMC, 20/10 mg), ciprofloxacin (CIP, 5 mg), chloramphenicol (CHLOR, 30 mg), and ceftriaxone (COX 30 mg) were used to perform the antimicrobial susceptibility test.
A Spearman correlation test showed no difference in ARG RA% profiles between samples collected during different swine growth phases. Pigs at 30 days of age showed a higher ARG RA%, but it was not statistically significant and declined over time (Figure 3). The ARG richness assessment showed the same tendency in animals within and between groups (Figure 3). The RA% of ARGs decreased as piglets got older, except for TE resistance genes and MGEs, which remained stable in all groups (Figure 3). All data from molecular analyses is available in the supplementary material of this study. Among the most abundant genes detected in this study were aminoglycoside resistance genes (aacA/aphD; aphA3; sat4; aph(2′)-Id; aphA1, aadE, str, aax(6)-im, aadA17, aph (3″)-ia; aac3-Via; aph3-III; ant6-ib; aac(3)-Xa), and MGEs Tp614, IS613, tnpA, int1-a-marko, intl2, intI1F165_ clinical, pBS228-IncP-1, trb-C, IS26, IS256, IS6100, IS91. Genes such as tet and the colistin resistance gene were detected in all samples (RA% 0.00013). PCR amplification revealed the mcr-1 gene (not previously detected in Ecuadorian pigs) in bacteria from fecal samples from 19 of the 20 sows (first-generation) in this study. There were no statistical differences in weight, growth, and health status among Groups A and B (Supplementary material S2).
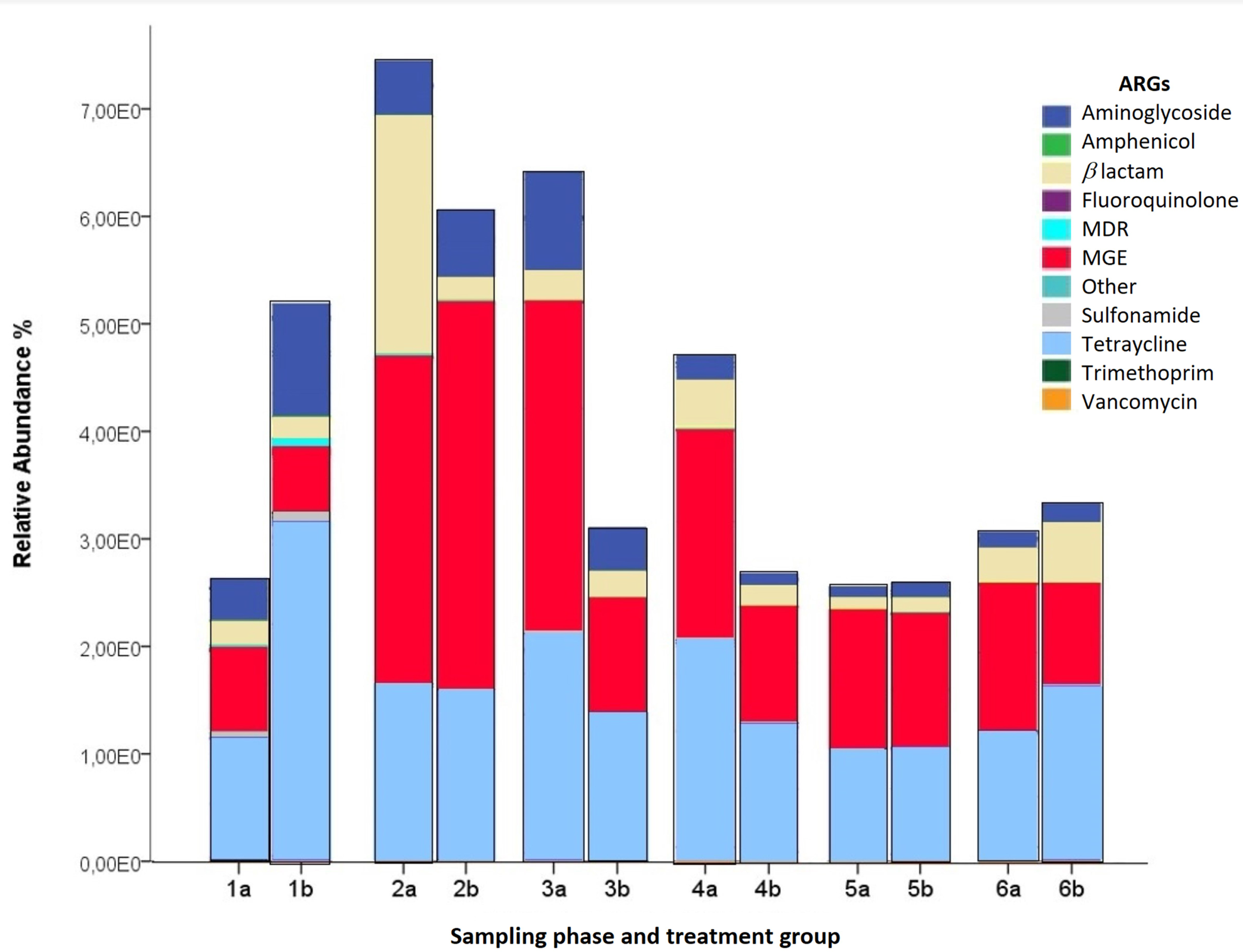
Relative abundance (RA%) of antimicrobial resistance genes (ARGs) are grouped by sampling phase and treatment group (Group A fed with antimicrobials; Group B fed without antimicrobials; 1–5 days; 2–30 days; 3–50 days; 4–100 days, 5–140 days and 6: sows 180 days). Different colors indicate different genes such as ARGs and mobile genetic elements (MGEs). RA% measures the amount of ARGs relative to the housekeeping gene 16Sr RNA.
Discussion
In the present study, we did not detect significant differences in ARB and ARGs in pigs after long deprivation periods of antimicrobial supplementation in feed. We did not find significant differences (p ≥ 0.05) in the total number of antimicrobial-resistant coliforms nor differences in the abundance or diversity of ARGs or MGEs between groups of animals receiving antimicrobials and those not fed with antimicrobials (Figures 2 and 3). In all animals, most AMR was to TE, AMP, trimethoprim–SXT, and CHLOR (Figure 3). High levels of resistance to TE, AMP, and trimethoprim–SXT have been commonly described in pigs elsewhere, and these results are consistent with similar reports showing no effect of antimicrobial restriction on the prevalence of AMR to AMP, TE, and trimethoprim–SXT. 38 The low prevalence of CIP resistance identified here could be explained by the limited use of this antimicrobial on farms. 9 However, while CHLOR had not been used on this pig farm previously, coliforms with CHLOR resistance were detected on this farm. Genes that code for aminoglycoside resistance have also been described in other studies, 9 and the higher prevalence in 30-day-old piglets could be explained by the substantial change in diet during this age. 23
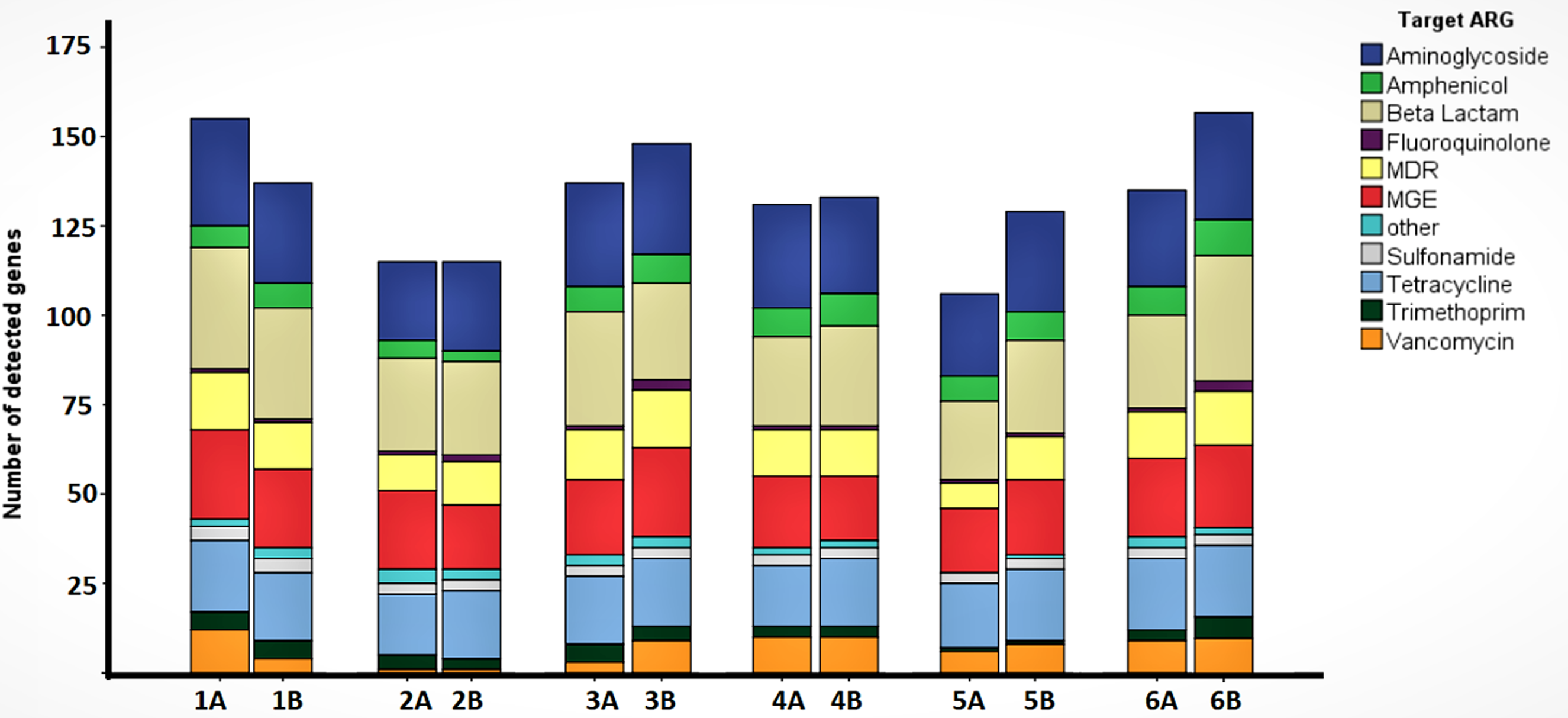
Antimicrobial resistance gene (ARG) abundance and richness. The colors indicate gene type. Different bars represent growing phase of piglets (1–5 days; 2–30 days; 3–50 days; 4–100 days, and 5–140 days) and sows (180 days). Animals fed with antimicrobials were identified as Group A and animals without antimicrobial additives were identified as Group B.
The results of this study are also consistent with reports of similar levels of ARGs found in intestinal bacteria from pigs in organic and conventional farms.24–26,30 In contrast, other studies showed a significant reduction in ARB and ARGs in animals deprived of antimicrobials and on organic farms,32,33,39,40–41 although in many of these studies, the differences were not significant or depended on the specific antimicrobial withdrawn.8,19,22 The apparent reduction in some AMR in such studies could be due to random turnover of ARB in the intestinal microbiome. Consistent with this hypothesis, statistically significant differences in the abundance of antimicrobial-resistant E. coli in humans, not treated with antimicrobials, were found when the members of a community were sampled at different time points over the course of 1 year (Calderon, et al. submitted manuscript).
Our results may indicate that ARGs do not cause a major reduction in fitness in the bacterial population of pig intestines or MGEs carrying ARGs contain addiction mechanisms such as toxin-antitoxin modules. 19 Prolonged compensatory evolution may have lessened such fitness costs.19,21,42 Some AMR genes such as those providing resistance to sulfonamides, TE, and CHLOR appeared in the 1950s, shortly after the introduction of these antimicrobials. These genes have thus been disseminating and coevolving with Enterobacterales for more than 60 years. 43 In contrast, recently emerged resistance genes (e.g. vanA, blaCTX−M, and mcr-1) have shown consistent reduction after the deprivation of corresponding antimicrobials.44–47 Thus, such historic coevolution may have created an association of bacteria and ARGs which cannot be reduced by simply limiting bacterial exposure to antimicrobials. 48
Coliforms isolated from 30-day-old piglets fed without antimicrobials had a higher prevalence of MDR coliforms, including a higher prevalence of resistance to AM, AMC, SXT, CIP, CHLOR, GEN, and COX, though this difference was not statistically significant. MGEs are important actors in the spread of AMR.12,13,48,49 In this study, we also observed a higher RA% of MGEs than specific ARGs in samples from 30-day-old piglets (Figure 3). When under antimicrobial stress, bacteria may turn on their SOS response, which can lead to the mobilization of MGEs to other bacterial hosts and subsequently increased numbers of resistant coliforms.50–52 In this study, the higher RA% of MGEs in samples from 30-day-old pigs corresponds to animals at weaning age. The transition from milk to solid food may cause a dysbiosis 53 and bacterial stress, resulting in SOS responses that could increase the dissemination of MGEs and ARGs. Age is therefore an important factor associated with changes in microbiota composition. 17 Importantly, the withdrawal of antimicrobials did not have any repercussions on the growth or weight of the animals in this study, suggesting changes to industrial standards surrounding antimicrobial use for growth promotion might have limited effects on production (Supplementary material S1).
Our study had several limitations. First, fecal samples (in swabs) were pooled from five animals to investigate the abundance and diversity of ARGs and MGEs in samples from different time frames and groups, limiting the possibility of carrying out some statistical analyses. However, phenotypic data were consistent with molecular findings. Second, we did not have any way to assess the quality or bioavailability of the antimicrobials used in the animals in this study.53,54 In this study, 20 first-generation replacement sows were originally divided into 2 study groups (Groups A and B), and their offspring were then followed during their complete productive cycles (until 150 days of age). This design was used as the study's objective was to detect changes in the prevalence of antimicrobial-resistant coliforms in each pig’s intestine beginning at birth and continuing throughout a normal pig's life in a commercial swine facility. Under industrial workflow, second-generation piglets are raised for the human food supply, and thus it was not possible to follow another generation of piglets within a conventional industry setting. It is possible that ARB present in the environment colonized the intestines of animals deprived of antimicrobials; however, assessment of such intestinal colonization with environmental coliforms was outside of the scope of this study. Because fecal samples were collected directly from each animal’s rectum, environmental contamination of samples themselves during the study procedures was unlikely.
Conclusions
This study found that antimicrobial restriction over two generations of pigs did not reduce the number of ARB in the gastrointestinal tract of pigs raised by a major swine producer in Ecuador, suggesting there are limited fitness costs to ARGs in the bacterial population of swine gastrointestinal tracts. The coevolution of bacterial chromosomes with ARGs and MGEs (such as plasmids) is a phenomenon that requires additional research.
Footnotes
Acknowledgments
The authors would like to thank Germán Romo, Manuel Salgado, Gustavo Nava, Oswaldo Vargas, and Edgar Velarde for their technical advice. Also, we thank Marcela Coba, Juan Castellanos, and Valeria Herrera for their personal support. We also thank Siena Mitman for her editorial improvements to the manuscript.
Author contributions
GT: Funding acquisition, farm access agreement, and project conceptualization.
FL: Experimental design, field and lab techniques, sampling procedures, data register and analysis, write and edition of the publication.
AT: Experimental design, farm permissions, and animal welfare.
LZ: Molecular methodology, technical support, and data analysis.
LZ and GT: Manuscript edition.
Conflict of interest
None of the authors have any conflict of interest.
Funding
This project was funded by la Universidad San Francisco de Quito Grant No. 11182 USFQ and the company (we were not authorized to disclose the name of the company). The funding body did not have any role in the study design, data collection, analysis, data interpretation, or writing of the manuscript.
