Abstract
Cell division cycle-associated (CDCA) genes are dysregulated in carcinomas. Our study aims to identify similarities and differences of the clinical roles of CDCAs in breast cancer (BRCA) and to explore their potential mechanisms. In GEPIA, compared to normal tissues, expressions of CDCAs were higher in BRCA and sub-types. In addition, CDCAs were significantly positively related to stages and predicted worse survival in BRCA. In CancerSEA, expression levels of most CDCAs were strongly positively related to cell cycle, DNA damage, DNA repair, and proliferation. In TIMER, CDCAs were linked with immune infiltration levels of BRCA, including Dendritic cell, B cell and so on, and were positively related to most of the common markers of immune cells, especially CD38 of B cell and IL12RB2 of Th1. In GeneMANIA, there were complex interactions and co-expression relationships between CDCAs and cell division-associated genes. In addition, CDCA1, CDCA3, CDCA5, CDCA6 and CDCA8 had a high proportion of amplification in BRCA, and CDCA1, CDCA2, CDCA5, CDCA7 and CDCA8 had high levels of body DNA methylation. Among 11 transcription factors possibly combining promoters of all CDCAs, FOXP3 and YY1 were significantly higher in BRCA in comparison to normal tissues, and both had a positive relationship with all CDCAs in GEPIA and IHC. In addition, silencing FOXP3 or YY1 decreased levels of CDCAs in MDA-MB-231. In summary, CDCAs have various similarities in clinical functions, functional states, immune infiltration, and mechanisms, and they may become novel potential biomarkers for BRCA.
Introduction
Breast cancer (BRCA) as the most common tumor has been always threatening women's health and safety. 1 Although the comprehensive treatment combined with operation and adjuvant therapy has reduced mortality and improved the prognosis, BRCA is still one of the leading causes of cancer death. 2 Therefore, it is urgent to explore novel biomarkers for the prediction of prognosis and identify potential therapeutic targets.
Cell division cycle-associated (CDCA) gene family, including eight members, CDCA1–8, plays a vital role in various biological processes of cells as well as the occurrence and development of cancers. 3 , For instance, CDCA1, also named NUF2, was essential for the progression of carcinomas and its over-expression predicted worse prognosis of several cancers, including digestive system cancers, genitourinary cancer, and so on.6,7 CDCA4 functioned as a transcription and promoted cancer cell proliferation through the E2F/retinoblastoma pathway. 8 CDCA6, also called chromobox 2 (CBX2), promoted the growth and metastasis of lung cancer and might become a bio-marker for the prognosis and chemoresistance of ovarian cancer.9,10 Moreover, CDCA5 and CDCA8 both increased cancer cell proliferation via the regulation of mitosis.11,12 Therefore, CDCAs were deemed as oncogenes.
In breast carcinoma, previous studies also showed that some of them might become anticancer therapeutic targets and prognostic factors. 13 br However, researches seemed to lack sufficient validation in clinical samples and their mechanisms of actions are still obscure. For example, CDCA2 might serve as a prognostic marker for luminal BRCA, but no further molecular functions were reported in the previous study. 13 Besides, similarities and differences among CDCAs also need to be explored to adequately understand their roles in the tumorigenesis and progression of BRCA. It was reported that only CDCA5 was elevated in BRCA stem cells and increased the stemness capacity through the modulation of cancer stem cell markers. 16 Therefore, in this study, we comprehensively analyzed similarities and differences in clinical functions and potential mechanisms of all CDCAs in BRCA.
Materials and methods
Database analysis of CDCAs expression
Gene Expression Profiling Interactive Analysis 2 (GEPIA2, https://gepia2.cancer-pku.cn/) was performed to analyze CDCAs expression in BRCA and its sub-types. 17 And then, relationships between CDCAs expression and tumor stages were also analyzed by this database.
Prognosis analysis in the database
The relationships between CDCAs expression and prognosis of BRCA patients were estimated via Kaplan–Meier plotter (https://kmplot.com/analysis/) which utilized Affymetrix microarray data from TCGA. 18 The primary outcomes were overall survival (OS) and relapse-free survival (RFS).
Breast cancer samples
Seventy specimens of female invasive BRCA patients were collected from Kunshan First People's Hospital between January 2014 and December 2021. The inclusion criteria: having complete clinical case data, without any radiotherapy or chemotherapy before operation, and the postoperative pathological results were confirmed by pathologists. Among them, 43 cases were Luminal, 19 cases were HER2+ and 8 cases were TNBC. Each case consisted of a breast tumor and its corresponding adjacent normal tissue. We conducted our study in accordance with the Helsinki Declaration of 1975 as revised in 2013. This research received the approval of the Kunshan First People's Hospital Ethics Committee at Room 603, Building Six, in Kunshan First People's Hospital on September 20, 2021 (No. 2021-06-038-K01). Every patient signed the informed consent form. Our study was performed following the relevant guidelines and regulations.
Immunohistochemistry (IHC)
Formalin-fixed paraffin-embedded sections of 70 cases were subjected to immunohistochemical staining by a SP Rabbit & Mouse HRP Kit (CWBIO, China), with antibodies, including anti-CDCA1 (diluted at 1:200, BIOSS), anti-CDCA2 (diluted at 1:200, BIOSS), anti-CDCA3 (diluted at 1:200, BIOSS), anti-CDCA4 (diluted at 1:200, BIOSS), anti-CDCA5 (diluted at 1:200, Proteintech), anti-CDCA6 (diluted at 1:200, Proteintech), anti-CDCA7 (diluted at 1:200, BIOSS), anti-CDCA8 (diluted at 1:200, Proteintech), anti-FOXP3 (diluted at 1:200, BIOSS) and anti-YY1 (diluted at 1:200, BIOSS). Scores of protein expression were evaluated with a semi-quantitative assessment system, immunoreactivity score (IRS).19,20 The IRS was combined by a score of the percentage of tumor cells (0, 0%; 1, 1%–25%; 2, 26%–50%; 3, 51%–75%; 4, 76%–100%), and a score of the staining intensity (0, no staining of the tumor cells; 1, mild staining; 2, moderate staining; and 3, marked staining). The total score ranged from 0 to 12.
CancerSEA analysis
CancerSEA database (https://biocc.hrbmu.edu.cn/) was applied to analyze relationships between CDCAs and different functional states of BRCA. 21 Functional states contained angiogenesis, apoptosis, cell cycle, differentiation, DNA damage, DNA repair, EMT, hypoxia, inflammation, invasion, metastasis, proliferation, quiescence, and stemness.
Tumor immune microenvironment analysis
TIMER database (https://timer.cistrome.org/) was performed to analyze the correlation between CDCAs expression and the abundance of infiltrating immune cell types in BRCA by Spearman correlation analysis.22,23 In addition, we evaluated relationships between CDCAs and immune markers by Spearman correlation analyses.
Exploration of molecules interacting with CDCAs
GeneMANIA database (http://genemania.org/) was utilized to predict molecules interacting with CDCAs and generate hypotheses about their functions on co-expression, co-localization, genetic interactions, pathway, physical interactions, and shared protein domains. 24
Exploration of upstream regulators of CDCAs
EPD database (https://epd.epfl.ch//index.php) was used to identify the promoters of CDCAs. 25 And then, PROMO database (http://alggen.lsi.upc.es/cgi-bin/promo_v3/promo/promoinit.cgi?dirDB = TF_8.3) was performed to predict potential upstream transcription factors (TFs) of CDCAs. 26 Moreover, GEPIA was conducted to analyze relationships between these TFs and CDCAs in BRCA and its sub-types.
Genetic alteration analysis
The cBioPortal web (https://www.cbioportal.org/) was used for queries about the genetic alteration characteristics of CDCAs. 27 We also analyzed prognostic roles of altered CDCAs in the cBioPortal, using Kaplan–Meier plots with log-rank P-value.
DNA methylation analysis
Epigenome-Wide Association Study (EWAS) Data Hub database (https://ngdc.cncb.ac.cn/ewas/datahub/index) was used to analyze the levels of CDCAs body DNA methylation, promoter methylation, and its prognostic roles in BRCA. 28
Cell culture
The human BRCA line MDA-MB-231 (FuHeng Biology, China) was cultured at 37 °C with 5% CO2 in a complete DMEM medium (Gibco, USA) containing 10% fetal bovine serum (Gibco, USA).
siRNA transfection
FOXP3 siRNAs (sc-43569), YY1 siRNAs (sc-36863), and control siRNA (sc-37007) were purchased from Santa Cruz Biotechnology (Santa Cruz, USA). And then, siRNAs were transfected into MDA-MB-231 using Lipofectamine 2000 (Invitrogen, USA) according to the manufacturer's protocol for 48 hours. The cells were then used for qRT-PCR and Western blotting.
Quantitative real-time PCR (qRT-PCR)
Transfected cells were collected to isolate total RNA through Trizol regent (Thermo Fisher Scientific, USA). And then, 2 µg RNA of each sample was reverse transcribed using the HiScript III RT SuperMix for qPCR (Vazyme-innovation in enzyme technology, China). cDNA was subjected to quantitative PCR using primers specific for FOXP3, YY1, and GAPDH. The primers were designed as follow: 5′-CTTGAACCCCATGCCACCAT-3′ (forward) and 5′-TGAAATGTGGCCTGTCCTGG-3′ (reverse) for FOXP3; 5′-CAGGCCAGGTTGGTCATAGG-3′ (forward) and 5′-TGTCAGACAAGCCAAGGTCC-3′ (reverse) for YY1; 5′-CATGAGAAGTATGACAACAGCCT-3′ (forward) and 5′-AGTCCTTCCACGATACCAAAGT-3′ (reverse) for GAPDH. Amplified DNA was measured by ChamQ Universal SYBR qPCR Master Mix (Vazyme-innovation in enzyme technology, China) following the manufacturer's protocol, and qPCR was performed using the real-time fluorescence quantitative PCR detection system (Roche Diagnostics GmbH, Germany). For quantification, we used a 2−ΔΔCt value to calculate the relative expression. We performed three replicates in this experiment.
Western blotting
Proteins of transfected cells were extracted and uploaded in pre-cast protein gels electrophoresis (Thermo Fisher Scientific, USA). And then, proteins were transferred to PVDF membranes (Beyotime Biotechnology, China). Antibodies including anti-CDCA1 (diluted at 1:1000, BIOSS), anti-CDCA2 (diluted at 1:1000, BIOSS), anti-CDCA3 (diluted at 1:1000, BIOSS), anti-CDCA4 (diluted at 1:1000, Proteintech), anti-CDCA5 (diluted at 1:2000, Proteintech), anti-CDCA6 (diluted at 1:2000, Proteintech), anti-CDCA7 (diluted at 1:2000, Proteintech), anti-CDCA8 (diluted at 1:1000, Proteintech), anti-FOXP3 (diluted at 1:1000, BIOSS), anti-YY1 (diluted at 1:1000, BIOSS), and anti-GAPDH (diluted at 1:2000, BIOSS) were used as the primary antibodies. Next, the HRP-conjugated antirabbit/mouse secondary antibody was used to enable detection.
Statistical analysis
Differences in expression levels of CDCAs between breast tumors and normal tissues were analyzed by paired t-test. Differences in relative RNA expression of FOXP3/YY1 between siNC and siRNA groups were analyzed by unpaired t-test. HRs and P value were calculated by log-rank test in Kaplan–Meier plotter. Spearman correlation analyses were utilized for relationships of CDCAs and FOXP3/YY1 detected by IHC. Differences in CDCAs expression among stages were analyzed by ANOVA. The data in this study was analyzed by Statistical Program for Social Sciences 20.0 and GraphPad Prism 9.0, and P < 0.05 was set as a statistically significant threshold.
Results
Overexpression of CDCAs in BRCA
Data from GEPIA showed that except CDCA7, expression levels of other CDCAs in BRCA were obviously higher than those in normal tissues (Figure 1(a)). Notably, compared to normal tissues, expressions of CDCAs, including CDCA7, were all obviously higher in basal-like BRCA (Figure 1(b)). And most of them were also over-expressed in other sub-types (Figure 1(b)). Except CDCA4 (no significance), the others had the highest expressions in basal-like, and the lowest expressions in luminal A (Figure 1(c)). Next, we investigated CDCAs expressions in breast tumors and normal mammary tissues via IHC (Figure 2(a)), and found that their average levels were all significantly higher in BRCA compared to normal tissues (all P < 0.001, Figure 2(b)).

Expressions of CDCAs in BRCA and sub-types in GEPIA. (a) Comparison of CDCAs expressions between BRCA and normal tissues; (b) comparison of CDCAs expressions between BRCA sub-types and normal tissues; (c) comparison of CDCAs expressions among BRCA sub-types. Red represents tumor and gray represents normal tissue. *P < 0.05.

Expressions of CDCAs in BRCA samples through IHC. (a) Microscopic observation (100×); (b) comparison of CDCAs expression scores between breast tumors and normal tissues (n = 70). ***P < 0.001.
Prognostic and clinicopathological roles of CDCAs in BRCA
In the database, GEPIA, except CDCA4 and CDCA7, expression levels of the others were positively related to stages (Figure 3(a)). Expressions of CDCA1-7 of 70 specimens detected by IHC also had a significantly increasing trend (Figure 3(b)). Next, we conducted Kaplan–Meier plotter to assess their prognostic roles in BRCA. All of them played detrimental roles both in OS and RFS (Figure 4). Collectively, higher expression levels of CDCAs predicted worse survival in BRCA.

Clinicopathological roles of CDCAs in BRCA. (a) GEPIA analyses; (b) comparison of CDCAs expressions among different tumor stages in BRCA samples.

Kaplan–Meier plotter analyses: (a) OS; (b) RFS.
Relationships of CDCAs with functional states of BRCA cells at the single-cell level
In CancerSEA, expression levels of most CDCAs were strongly positively related to cell cycle, DNA damage, DNA repair, and proliferation, and were slightly positively related to invasion (Figure 5). However, CDCA6 expression wasn’t correlated with any functional state.

CancerSEA analysis.
Correlations between CDCAs and immune infiltration levels in BRCA
Then, we conducted TIMER database to analyze correlations of CDCAs expressions with immune infiltration levels (Figure 6). Except CDCA4, CDCAs were positively related to infiltration levels of B cell and Neutrophil. Except CDCA3, CDCA4 and CDCA5, CDCAs were positively related to infiltration levels of CD8+ T cell. Except CDCA1 and CDCA3, CDCAs were positively related to infiltration levels of CD4+ T cell. Except CDCA7, CDCAs were negatively related to infiltration levels of Macrophage. All of the CDCAs were positively related to infiltration levels of dendritic cell.

Correlation between CDCAs expressions and the abundance of infiltrating immune cell types in BRCA.
Correlation between CDCAs and immune markers in BRCA
In the TIMER database, we also found that CDCAs were positively related to most of common markers of immune cells, especially CD38 of B cell and IL12RB2 of Th1 (Figure 7(a)). However, CDCAs were negatively related to markers of Th2, and STAT5B and TGFB1 of Treg (Figure 7(a)). Furthermore, CDCAs were positively correlated with most of common checkpoint markers, especially BTLA, IDO1, PDCD1LG2, TIGIT and TNFRSF9 (Figure 7(b)). However, CDCAs were negatively related to TNFRSF14 (Figure 7(b)).

Correlation between CDCAs expressions and immune markers in BRCA. (a) Immune markers of immune cells; (b) checkpoint markers. *P < 0.05, **P < 0.01, ***P < 0.001.
Prediction of molecules interacting with CDCAs
Next, we performed GeneMANIA to explore potential proteins interacting with CDCAs (Figure 8). It showed that there were complex interactions and co-expression relationships between CDCAs and other genes, including BIRC5, CDC20, CENPF, and so on. These genes induced cell division and chromosome segregation, and promoted the growth of cancer cells.29–33 Particularly, CDCA1 (NUF2), CDCA5, CDCA8, AURKB, CDC20, CEP55, PLK1, TACC3, KIF23, and CENPF seemed similar functions on chromosome segregation and mitosis.

GeneMANIA analysis.
Genetic alteration analysis
In the cBioPortal, we found that CDCA1, CDCA3, CDCA5, CDCA6, and CDCA8 all had a high proportion of amplification in cancers, including BRCA, while CDCA2 had a high proportion of deep deletion (Supplement File S1). In addition, mutation frequencies and sites of CDCAs were different (Supplement File S2). Among them, CDCA2 had the most missense mutation sites (Supplement File S2). Compared with the unaltered group, only CDCA3 alteration was associated with worse OS (P = 0.003, Supplement File S3).
DNA methylation analysis
The EWAS database showed that compared with the control group, CDCA1, CDCA2, CDCA5, CDCA7 and CDCA8 all had higher levels of body DNA methylation, while CDCA3, CDCA4 and CDCA6 had lower one (Supplement File S4A). Besides, levels of body CDCAs DNA methylation had no significant relationship with the survival time of BRCA patients (Supplement File S4B). Compared with the control group, CDCA6 and CDCA7 had higher levels of promoter methylation, while others had no obvious differential (Supplement File S4C). There was no analysis of CDCA1 promoter methylation in the EWAS database. Notably, a high level of CDCA7 promoter methylation had a better prognosis in BRCA patients (P = 0.031, Supplement File S4D).
Prediction of upstream regulators of CDCAs in BRCA
Through EPD and PROMO databases, we predicted 11 TFs that could combine promoters of all CDCAs (Figure 9(a)). In the GEPIA database, we found that expression levels of FOXP3 and YY1 in breast tumors were significantly higher than those in breast normal tissues (Figure 9(b)). However, there were no differences in the other 9 TFs expressions between breast tumors and normal tissues. In addition, expression levels of FOXP3 and YY1 were both positively related to all CDCAs in GEPIA (Figure 9(c) and (d)). Therefore, FOXP3 and YY1 might up-regulate the expression of CDCAs through their interaction with promoters of CDCAs.

Prediction of TFs of CDCAs promoters. (a) Results of EPD and PROMO databases; (b) GEPIA showing up-regulation of FOXP3 and YY1 in BRCA in comparison of normal tissues; (c) correlation between CDCAs and FOXP3 expression in GEPIA; (d) correlation between CDCAs and YY1 expression in GEPIA. *P < 0.05.
Silencing of FOXP3/YY1 suppressing expression levels of CDCAs
Through IHC, we verified that expression levels of FOXP3 and YY1 in breast tumors were obviously higher in comparison to breast normal tissues (Figure 10(a) and (b)). In addition, FOXP3 and YY1 expressions were both positively correlated with CDCAs (Figure 10(c) and (d)). After silencing FOXP3 and YY1, respectively, expression levels of CDCAs were all down-regulated in MDA-MB-231 (Figure 11).
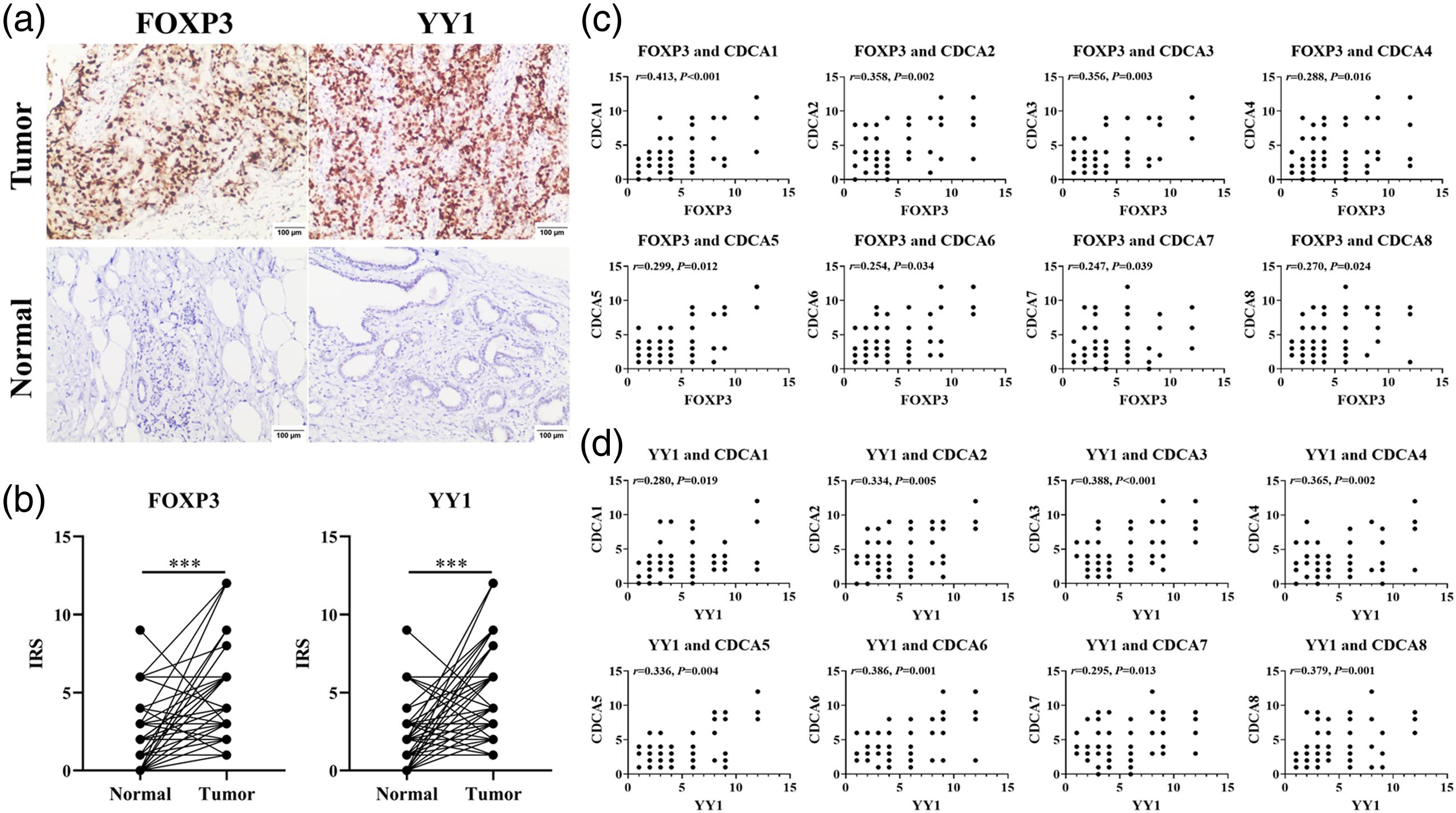
Expressions of FOXP3 and YY1 in BRCA samples through IHC: (a) microscopic observation (100×); (b) comparison of FOXP3 and YY1 expression scores between breast tumors and normal tissues (n = 70); (c) correlation between CDCAs and FOXP3 expression in BRCA samples; (d) correlation between CDCAs and YY1 expression in BRCA samples. ***P < 0.001.

Regulation of CDCAs by silencing FOXP3 and YY1:. (a) qRT-PCR investigating the decrease of FOXP3 and YY1 mRNA via siRNAs; (b) Western blotting analysis detecting expressions of FOXP3, YY1 and CDCAs proteins. ***P < 0.001.
Discussion
CDCAs were putative oncogenes and played critical roles in the growth of carcinomas. 5 Due to this, recent studies explored their clinical value and found that CDCAs might become novel prognostic biomarkers and potential targets for some cancers, such as gastric cancer, prostate cancer, and so on.34,35 Some of these CDCAs, like CDCA3, CDCA5, and CDCA8, also predicted worse prognosis of BRCA in previous studies.36,37 However, more validations in clinical samples and explorations of their molecular mechanisms are still essential for understanding the functions of CDCAs in BRCA. This study analyzed their clinical roles and mechanisms together, and revealed their similarities and differences in BRCA. In our research, these eight genes were abnormally over-expressed in BRCA, and high levels of them indicated poor prognosis. Moreover, compared with the other three sub-types, expression levels of CDCAs in luminal A which had the best prognosis among all sub-types were obviously lower. This sub-type has a low level of Ki-67 which is an objective index to evaluate the cell proliferation state. Therefore, we demonstrated that CDCAs might become a series of biological markers for the evaluation of BRCA prognosis.
Then, we explore their mechanisms from several aspects. In single-cell sequencing analysis of CancerSEA, modulations of cell cycle, DNA damage, DNA repair, and proliferation were mainly functions of CDCAs in BRCA cells. But CDCA6 expression wasn’t related to any functional state. Therefore, CDCAs played vital roles in BRCA cell division and proliferation. Previous studies also showed that several CDCAs, like CDCA4, CDCA7 and CDCA8, promoted cell proliferation and progression of BRCA cell lines.38en, Besides, researchers reported that some of them modulated DNA damage and repair in cells, including several cancers.41,42 Although CDCA5 was related to the stemness capacity of BRCA stem cells, we failed to find that any CDCA protein increased the stemness in the CancerSEA. 16 Thus, further researches are needed to confirm whether all of these CDCAs influenced cell proliferation, DNA damage and repair, cell cycle, and division in BRCA cells.
Notably, CDCAs expressions were closely correlated with immune cell infiltration levels and immune checkpoints. Thus, CDCAs might be not only the crucial molecules of BRCA cell proliferation, but also unexpectedly correlated with the tumor immunity. Several mitotic and transcriptional genes, including CDCA1 and other genes, like CDC20 and CENPF, had been proved to be related to the tumor microenvironment. 43 Their abnormal expression might not only promote cell proliferation, but also interfere the immunotherapy through regulating the tumor microenvironment.44,45 For example, the previous study showed that CDC20 was a prognostic indicator and could evaluate the sensitivity to immunotherapy in patients with prostate cancer. 45 Therefore, identifying the functions of CDCAs on tumor immunity will provide new targets for immunotherapy. However, due to the existing insufficient number of blood samples, we could not accurately evaluate and verify the correlation between CDCAs expression and tumor immunity. Thus, we need to enlarge the number of blood samples.
In the GeneMANIA database, with the complex interactions, CDCAs played critical roles in cell division and chromosome segregation. Especially, three of them, CDCA1, CDCA5 and CDCA8, might serve some similar functions to other mitotic genes. In addition, several genes, like CDC20 and CENPF, not only modulated mitosis through the interactions with CDCAs, but also influenced the tumor microenvironment. 43 In addition, they might be related to the release of a series of angiogenic factors and extracellular matrix proteins. 43 Therefore, further researches are needed to verify the mutual effect of CDCAs and related mitotic genes.
In this study, promoters of all CDCAs had binding sites of 11 TFs, but only FOXP3 and YY1 expressions were abnormally expressed in BRCA and were correlated with CDCAs. In addition, after silencing of FOXP3 or YY1, expression levels of CDCAs in MDA-MB-231 were obviously decreased. Previous studies showed that FOXP3 and YY1 could promote the proliferation of cancer cells through the modulation of promoters of downstream targets.46,47 Thus, we demonstrated that FOXP3 and YY1 were both the upstream regulators of CDCAs. Moreover, it was reported that FOXP3 promoted CCR4(+) Treg infiltration and was related to tumor immunity. 48 And YY1 was reported to be involved in immune evasion, resistance to apoptosis, and tumor progression. 49 Therefore, abnormal expression of FOXP3 and YY1 might become initial factors resulting in CDCAs-related tumor immune escape and cancer progression. However, further researches, like dual-luciferase reporter assay and chromatin immunoprecipitation assay, are necessary to perform to elaborate mechanisms of the regulation of CDCAs. Since we just used MDA-MB-231 cell line in this study, more cell lines can be used to validate our findings in the future. Moreover, we couldn’t identify whether FOXP3/YY1 interacted with CDCAs directly. We need to conduct co-immunoprecipitation assay to explore their possible interactions.
Admittedly, there were several limitations in our research. Firstly, due to the limitation of the sample size in each sub-type, we failed to analyze the clinicopathological roles of CDCAs in sub-types of BRCA. Secondly, due to no multivariate analyses in public databases, we only use univariate analyses when assessing their roles in the prognosis of BRCA. Thus, we need to expand the samples’ quantity and collect follow-up information on patients to evaluate the prognostic roles of CDCAs in BRCA and its sub-types. Since we failed to extract proteins from paraffin-embedded tissues for western blot, we only conducted IHC to detect their protein expression levels. In future research, we need to collect adequate fresh tumor tissues. Although we found the functions of CDCAs on cancer progression and tumor immunity, our results lack external validation in vitro and in vivo researches. Therefore, this theoretical work remains to be verified.
Conclusion
In conclusion, CDCAs possess diverse similar characteristics in expression, clinicopathological feathers, prognosis, cell functional states, tumor immunity, and molecular and genetic mechanisms of BRCA. Therefore, CDCAs may become novel potential biomarkers for BRCA.
Supplemental Material
sj-docx-1-sci-10.1177_00368504241312305 - Supplemental material for Roles of the CDCA gene family in breast carcinoma
Supplemental material, sj-docx-1-sci-10.1177_00368504241312305 for Roles of the CDCA gene family in breast carcinoma by Wei Ding, Wei Han, Chun-Tao Shi, Li-Qian Yao, Zhi-Wei Liang and Ming-Hui Zhou, Hao-Nan Wang in Science Progress
Supplemental Material
sj-docx-2-sci-10.1177_00368504241312305 - Supplemental material for Roles of the CDCA gene family in breast carcinoma
Supplemental material, sj-docx-2-sci-10.1177_00368504241312305 for Roles of the CDCA gene family in breast carcinoma by Wei Ding, Wei Han, Chun-Tao Shi, Li-Qian Yao, Zhi-Wei Liang and Ming-Hui Zhou, Hao-Nan Wang in Science Progress
Supplemental Material
sj-docx-3-sci-10.1177_00368504241312305 - Supplemental material for Roles of the CDCA gene family in breast carcinoma
Supplemental material, sj-docx-3-sci-10.1177_00368504241312305 for Roles of the CDCA gene family in breast carcinoma by Wei Ding, Wei Han, Chun-Tao Shi, Li-Qian Yao, Zhi-Wei Liang and Ming-Hui Zhou, Hao-Nan Wang in Science Progress
Supplemental Material
sj-docx-4-sci-10.1177_00368504241312305 - Supplemental material for Roles of the CDCA gene family in breast carcinoma
Supplemental material, sj-docx-4-sci-10.1177_00368504241312305 for Roles of the CDCA gene family in breast carcinoma by Wei Ding, Wei Han, Chun-Tao Shi, Li-Qian Yao, Zhi-Wei Liang and Ming-Hui Zhou, Hao-Nan Wang in Science Progress
Footnotes
Authors’ contributions
WD and HNW conceived and designed the experiments. WD, WH, CTS, LQY, ZWL, and MHZ performed the experiments. WD, WH, and HNW analyzed and interpreted the data. WD, WH, and CTS contributed reagents, materials, analysis tools or data. WD, WH, and CTS wrote the paper.
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval and consent to participate
This study was approved by the Kunshan First People's Hospital Ethics Committee (2021-06-038-K01) and all patients signed the informed consent form.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Special Science and Technology Project of Kunshan First People's Hospital (No. KRY-YN002); Suzhou Municipal Bureau on Science and Technology (No. KJXW2023074); Wuxi Municipal Bureau on Science and Technology (No. Y20212039); and Kunshan High-level Health Talents Plan the second batch of training talents—Young top talents (No. 28).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
