Abstract
Lung cancer (LC) is a highly lethal cancer worldwide. Research on the distribution and nature of extrachromosomal DNA molecules (EcDNAm) in early LC is scarce. In this study, after removing linear DNA and mitochondrial circular DNA, EcDNAm were extracted from two paired LC tissue samples and amplified using rolling circle amplification. High throughput extrachromosomal DNA (EcDNA) or RNA sequencing and bioinformatics analysis were subsequently utilized to explore the distribution and nature of the EcDNAm. Additionally, to elucidate the role of oncogenes with large EcDNAm sizes, gene ontology and Kyoto Encyclopedia of Genes and Genomes pathway analyses were performed. The RNA sequencing results revealed significant differences in certain genes between tumors and corresponding normal samples. At the same time, slight distinctions were observed between relapsed and non-relapsed tumor samples. The nature of the EcDNAm was compared between LC samples and matched normal samples. There was a tendency for the number of EcDNAm with longer size (EcDNA) and its containing driver oncogenes to be higher in cancer samples. Enrichment analysis of the cancer samples revealed enrichment in biological processes, such as positive regulation of protein localization, axon development, and in-utero embryonic development. This study highlights the universal distribution and characteristics of EcDNAm in early LC. Moreover, our work fills the investigation of the EcDNAm gap and future studies should focus on the application of EcDNA as a potential biomarker in patients with early LC.
Keywords
Introduction
Lung cancer (LC) has the highest morbidity and mortality rate in the world.1,2 Non-small cell LC (NSCLC) accounts for three-quarters of all LC cases. 3 With the popularization of computed tomography (CT) examination and low-dose spiral CT screening for LC, the number of patients with early NSCLC is increasing year by year. 4 Compared to targeted therapy and immunotherapy for patients with advanced LC, surgical therapy is the main treatment for patients with early LC, but the prognosis is different.5–7 According to the eighth edition of the staging classification, the 2-year recurrence and metastasis rate of surgically resected stage I NSCLC patients is 38%, and the 5-year overall survival rate is 60% to 74%.8,9 To improve the survival rate of patients with early NSCLC, it is particularly important to find potential biomolecular targets and to elucidate their roles and mechanisms in the process of recurrence and metastasis.
Apart from 22 linear autosome pairs and a pair of sex chromosomes, extrachromosomal DNA molecules (EcDNAm) were found in the human genome.10,11 With the explosive growth of next-generation sequencing biological data in recent years, EcDNAm have become a research hotspot in the field of biomedical oncology.12–15 EcDNAm were found to be a closed loop structure.16,17 Wu et al. reported that EcDNAm with longer sizes are often enriched in the process of tumor formation and aging, and participate in the occurrence and development of tumor aging in a special way.18–20 At the same time, the chromatin of EcDNAm with longer sizes is in a highly open state, in which the transcription of genes is abnormally active, and the mutation of the oncogenic driver gene mediates drug resistance.19,21 These findings indicated that EcDNAm play an important role in tumorigenesis. However, the mechanism of action of EcDNAm in patients with early stage LC remains unclear.
In this study, we explored the mechanism of action of EcDNAm in the early stage of LC. To the best of our knowledge, this is the first report to investigate the distribution, function, and mechanism of EcDNAm in matched early LC samples.
Materials and methods
Tissue DNA preparation and EcDNAm sequencing
The study protocol was approved by the institutional review board at Zhejiang Cancer Hospital, Hangzhou, China (IRB-2020-63). Written informed consent was obtained from two male patients with early stage LC undergoing surgery at the Department of Thoracic Surgery, Zhejiang Cancer Hospital. The samples were collected in March 2022 at Zhejiang Cancer Hospital, Hangzhou, China. High throughput extrachromosomal DNA (EcDNA) sequencing was performed by Novogene Biotech Inc. (Beijing, China). The method of circular DNA purification is the Circle-Seq protocol, which involves column purification, removal of remaining linear chromosomal DNA, rolling circle amplification, sequencing, and mapping.22,23 The detailed EcDNA purification and sequencing were performed according to a previously published method.16,22
Sequencing analysis of EcDNAm
Sequencing data were obtained using an Illumina NovaSeq 6000 sequencer in 150 bp paired-end mode. The fragmented DNA was subjected to library preparation with NEBNext Ultra II DNA Library Prep Kit for Illumina (New England Biolabs, MA, USA). Subsequently, the raw sequencing data were further quality-controlled by Cutadapt software (v1.9.1). For instance, low-quality reads (Q < 10; reads containing more than 10% of the total length of N base) were discarded, and the adaptor sequences were removed. The clean reads were further aligned to the reference genome (UCSC hg19). Various bioinformatics tools, such as circle-map,24,25 samtools, 26 and bedtools, 26 have been used to detect EcDNAm. Gene function and pathway analyses were performed in a single sample using the clusterProfiler package (version 4.2.0).
Statistical analysis
All statistical analyses and diagram drawings were processed by the R software (version 4.1.2, R Foundation for Statistical Computing, Vienna, Austria). DEseq2 software (version 1.26.0) was used to analyze the expression of 59 early LC patients and their corresponding normal samples using RNA transcriptome data. P < 0.05 was considered for the statistical significance.
Results
Differential expression between tumor and corresponding normal samples using RNA sequencing data
RNA sequencing data and EcDNAm sequencing data were performed to explore the expression levels of an oncogene on linear chromosomes and extra-chromosomes in patients with early LC. In this part, to investigate the gene expression patterns on linear chromosomes in patients with early LC, we used transcriptome data 27 to assess the expression profiles of 59 early LC patients and their corresponding normal samples. As shown in Figure 1, we found a difference between the tumor and corresponding normal samples for certain genes, such as epidermal growth factor receptor (EGFR). At the same time, slight distinctions were observed between relapsed and non-relapsed tumor samples.

Differential expression of genes between tumor and normal samples.
Genome-wide detection of EcDNAm in paired Lc samples
EcDNAm were detected in 23 pairs of chromosomes by mapping the clean reads to the genome. As illustrated in Figure 2, these results indicate that the presence of EcDNAm is prevalent in both tumor and matched normal samples.

The distribution of EcDNAm in the tumor and matched normal samples.
The genomic distribution analysis of EcDNAm revealed its widespread presence across all 23 pairs of chromosomes. Few EcDNAm were detected in the mitochondria.
According to the previous classification criteria,
28
EcDNAm is generally divided into two types. One type is extrachromosomal circular DNA (EccDNA
Comparing the discrepancy in EcDNA between LC and matched normal samples
The majority of the large EcDNAm (>1 MB) were detected in both the LC and matched normal samples. As depicted in Figure 3, the number of genes within EcDNAm varied in chromosomes between LC and matched normal samples. Previous studies have demonstrated that large EcDNAm harbor one or multiple driver oncogenes.18,28,29 Consequently, the distribution of driver oncogenes derived from EcDNAm was examined. Compared to the matched normal samples, there was a trend for the number of EcDNAm that contained driver oncogenes to be higher in cancer samples (Figure 4).

The distribution of the number of gene counts derived from EcDNAm between LC and matched normal samples.
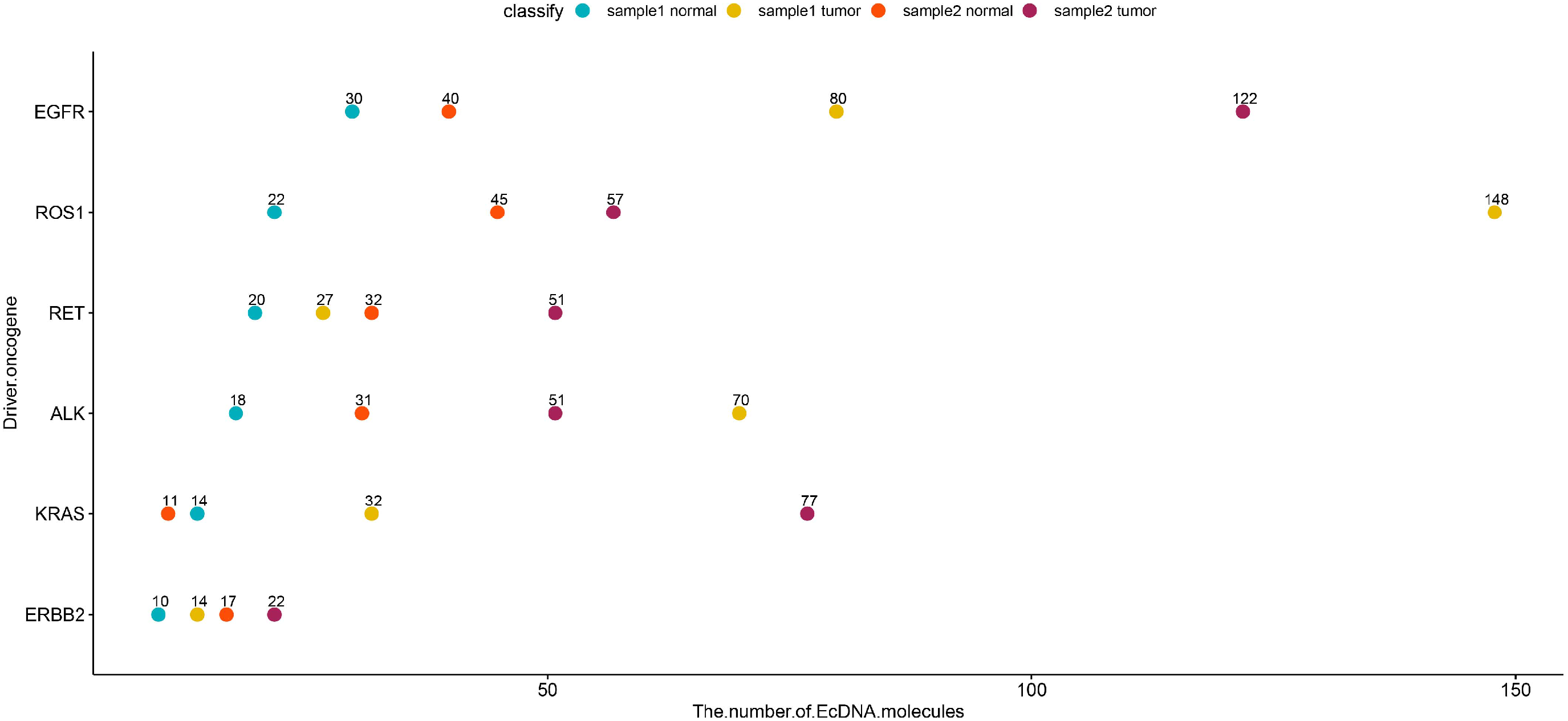
The distribution of the driver oncogenes derived from EcDNAm between LC and matched normal samples.
Gene ontology and Kyoto encyclopedia of genes and genomes pathway analysis of genes in EcDNAm
Gene ontology (GO) analysis was performed to examine the function of genes associated with EcDNAm, focusing on related cellular components (CCs), molecular functions (MFs), and biological processes (BPs), as outlined in Table 1. The predominant biological processes were related to the positive regulation of protein localization, axon development, and in-utero embryonic development in the tumor samples. Detailed results of gene functions are shown in Figure 5.
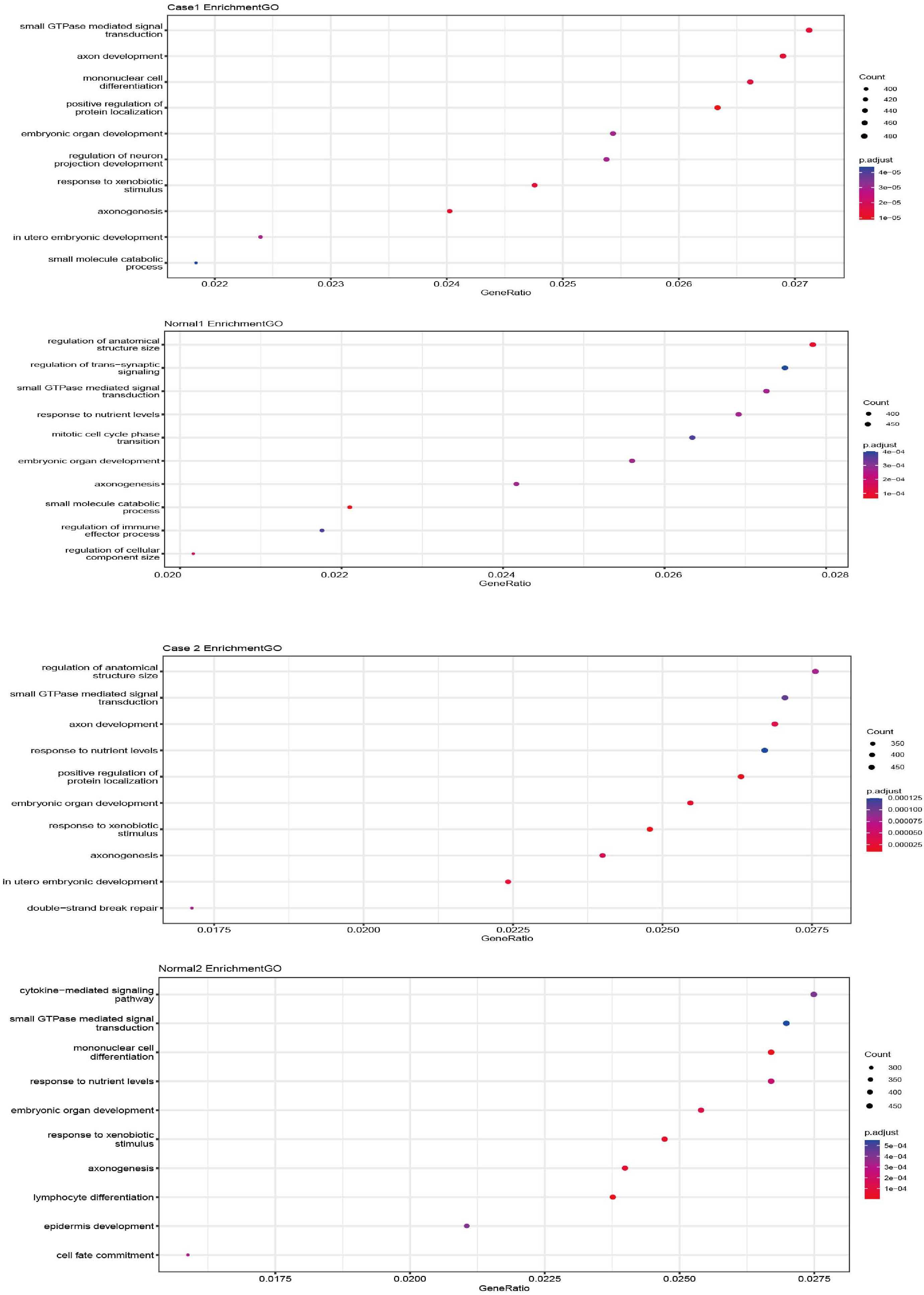
GO analysis of genes in EcDNAm.
GO analysis of the genes associated with EcDNA in paired early LC samples.

GO: gene ontology: EcDNAm: extrachromosomal DNA molecules; BP: biological process; CC: cellular component; MF: molecular function.
In addition, the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis indicated that the primary pathway was involved in the cell cycle indicating a deviation from the patterns observed in normal samples (Figure 6).

KEGG pathway analyses of genes in EcDNAm.
Discussion
An EcDNAm is a circular DNA that plays an important role in the development and heterogeneity of cancer.13,30,31 However, the comprehensive profiling of the structure, composition, and genome-wide frequency of EcDNA has not been extensively profiled. In this study, high throughput sequencing demonstrated the presence of EcDNAm in paired early LC samples, consistent with previous studies in other cancer types.32,33 In addition, our data demonstrated the presence of EcDNAm could be found in the human genome, except mitochondria, in early LC.
To investigate the expression levels of an oncogene on linear chromosomes and extra-chromosomes in patients with early LC, RNA sequencing data and EcDNAm sequencing data were performed in this study. The RNA sequencing results revealed differential genes between tumor and normal samples, such as EGFR.
Based on size, two classes of EcDNAm exist in human cells: small (EccDNA) and large (EcDNA). Previous studies have shown that oncogenes in EcDNA are highly expressed.19,34 In comparison with the corresponding normal samples, there was a trend for the number of EcDNAm with a larger size (namely EcDNA), which contains driver oncogenes (e.g. EGFR) was relatively elevated. Furthermore, there was a tendency for the quantity of EcDNA to be higher in tumor samples, although no statistical analysis has been conducted to confirm the difference.
GO analysis of genes associated with EcDNA showed that the predominant biological processes were related to positive regulation of protein localization, axon development, and in-utero embryonic development in tumor samples. Additionally, KEGG pathway analysis of genes associated with EcDNA indicated a primary involvement in the cell cycle, diverging from normal samples. In this study, we found a difference between tumor and corresponding normal samples for certain genes, such as EGFR. However, due to the limited data and experimental validation, the carcinogenic mechanism of the relationship between chromosomal DNA and EcDNA remains unclear. Furthermore, a direct statistical comparison of oncogenes associated with EcDNA was not performed between the LC samples and matched normal samples. We firmly believe that the oncogenes (such as EGFR) associated with EcDNA might play a crucial role in early LC.
In the future, we will expand the sample size and delve deeper into the functional mechanism of EcDNAm in early LC. We believe that our findings address the current gap in the understanding of EcDNAm in early LC, and we advocate for future studies to emphasize the potential application of EcDNA as a biomarker and therapeutic target in patients with early LC.
Conclusions
In conclusion, our study confirmed the genome-wide presence of EcDNAm in paired LC samples. In addition, our work revealed the potential mechanisms of EcDNAm in early LC. This work provides further insights into our understanding of genome plasticity and the role of EcDNAm in early LC, contributing to the development of potential clinical therapies.
Footnotes
Author contributions
JF and DS conceived, designed, and revised the study. JF analyzed the data and wrote the manuscript. LY, ZM, YY, and RZ analyzed the data. All authors have read and approved the final manuscript.
Availability of data and materials
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval and consent to participate
This study protocol was approved by the institutional review board at Zhejiang Cancer Hospital (IRB-2020-63). Written informed consent was obtained from two patients with early LC at the Department of Thoracic Surgery at Zhejiang Cancer Hospital. This study was performed in line with the principles of the Declaration of Helsinki.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Zhejiang Provincial Natural Science Foundation, (grant number LQ21F010001).
