Abstract
Purpose:
To propose a multi-output fully convolutional network (MOFCN) to segment bilateral lung, heart and spinal cord in the planning thoracic computed tomography (CT) slices automatically and simultaneously.
Methods:
The MOFCN includes two components: one main backbone and three branches. The main backbone extracts the features about lung, heart and spinal cord. The extracted features are transferred to three branches which correspond to three organs respectively. The longest branch to segment spinal cord is nine layers, including input and output layers. The MOFCN was evaluated on 19,277 CT slices from 966 patients with cancer in the thorax. In these slices, the organs at risk (OARs) were delineated and validated by experienced radiation oncologists, and served as ground truth for training and evaluation. The data from 61 randomly chosen patients were used for training and validation. The remaining 905 patients’ slices were used for testing. The metric used to evaluate the similarity between the auto-segmented organs and their ground truth was Dice. Besides, we compared the MOFCN with other published models. To assess the distinct output design and the impact of layer number and dilated convolution, we compared MOFCN with a multi-label learning model and its variants. By analyzing the not good performances, we suggested possible solutions.
Results:
MOFCN achieved Dice of 0.95
Conclusion:
The results demonstrated the MOFCN’s effectiveness. It uses less parameters to delineate three OARs simultaneously and automatically, and thus shows a relatively low requirement for hardware and has potential for broad application.
Keywords
Introduction
An effective radiotherapy is to kill tumor using a high prescribed dose while sparing organs at risk (OARs). 1 Delineating OARs in computed tomography (CT) scans is an essential step of optimizing a radiation treatment plan and evaluating clinical goals quantitatively. Several studies have shown that the delineation accuracy is highly correlated to tumor control and radiotherapy toxicities.2–4 Additionally, the inconsistencies arising from inter- and intra-observer variations have an impact on quantitative5–9 and dosimetric10–13 analysis. Therefore, the task of delineating OARs requires accuracy and less inter- and intra-observer variations.14–17
Although a lot of conventional image segmentation approaches have been used in the task of automatic or semi-automatic OARs delineation,18–20 such as those based on image grayscale information, 21 statistical shape modeling22,23 and body atlas-based methods,24–26 they can’t guarantee the accuracy, consistency and reproducibility. The grayscale information-based algorithms are sensitive to imaging artifacts, low contrast-to-noise ratio (CNR), and so on. A segmentation based on a statistical shape model uses prior knowledge about a generic structure to constrain segmentation of individual organs. Such models may be challenged by any abnormalities. The body atlas-based methods are actually deformable image registrations referencing to an image with all segmented organs (i.e. the atlas). For them, the anatomical variation and vanishing volumes may be problematic. More importantly, those segmentations have no incorporation of the essential human clinical decision-making. The prior-knowledge and/or experience about what voxels are included as part of the segmented OAR can’t just come from image and usually has an inherent correlation to the treatment quality. Above all, a mankind’s work on the definition of OAR contours still serves as the gold criterion. 21
As an artificial intelligence (AI) method of mimicking human beings’ activity, the fully convolutional network (FCN) is proposed to delineate OARs automatically.27–31 The first attempt of applying a convolutional neural network (CNN) on OARs’ segmentation was reported by Ibragimov and Xing. 31 Their work focused on the OARs in head and neck region. Their input image was an image patch that was around voxels belonging to the interested OAR, instead of a whole CT scan. For the thoracic OARs, a 11-layer FCN was adopted to label the voxels of lung and achieved Dice of 0.96. 28 Dice is the common similarity metric between two images. According to Zhu et al., 29 a 13-layer and 5-channel CNN delineated the spinal cord with a Dice of 0.71–0.79, and lung and heart with a Dice of 0.87–0.95 in a series of images with the size of 96 × 96 pixels. A 96 × 96 CT usually involves 106 mm × 106 mm image, it is unable to show a normal human’s whole transverse section. A large amount of methods 32 used a single-input and single-output FCN with many layers to delineate one OAR in down-sampled or cropped CT scans. Therefore, they lack the appropriateness for clinical application and show a high computation requirement to delineate all OARs for a lot of patients. Corresponding to the high computation requirement, the high cost may limit their widespread application in most hospitals in China.
This paper proposes a multi-output FCN (MOFCN) and reports our experience of using it on a large testing set. Specifically, the contributions of this paper are as follows:
We propose a network consisting of a shared main backbone and multiple distinct output branches. It segments bilateral lung, heart and spinal cord automatically and simultaneously in planning CTs that include the whole body’s transverse section.
This design tries to combine the information sharing of various neighboring organs and independent feature extraction for individual one.
The proposed model is evaluated on a large testing dataset. It is more like a clinical application scenario where a model is applied on a large data, but is trained on a limited quantity of images.
The testing dataset includes abnormal cases, such as different body type (high/short, fat/thin), spongy bone and lung collapse. It is different to most public reports.
Based on the above test results, we analyze for the failure causes and give possible solutions.
The remainder of this paper is organized as follows. Section 2 introduces the MOFCN architecture and experiment details. The experiment results are shown and compared with other models in section 3. The study on the design of MOFCN is also shown in section 3 by comparing with its variants. In section 4, we further discuss the results and the exceptive cases in which MOFCN fails and suggest the potential solutions. The study’s conclusion is presented in section 5.
Materials and methods
Data acquisition
The data (i.e. 19,277 CT scans totally) was collected from 966 patients with cancers in the thorax receiving radiotherapy from January 2015 to December 2018 in our department. All patients underwent CT scans on Light Speed (GE Healthcare, Chicago, America) or Brilliance CT Big Bore system (Philips Healthcare, Best, the Netherlands) on helical scan mode with or without contrast. All CT images were reconstructed into matrix size of 512 × 512 with a thickness of 5 mm. The OARs were delineated in the planning CTs by experienced radiation oncologists and were regarded as the ground truth.
Multi-output fully convolutional network (MOFCN) for segmentation
The proposed MOFCN’s workflow and structure are shown in Figure 1.

Illustration of the: (a) workflow and (b) architecture of the proposed multi-output fully convolutional network (MOFCN).
Image preprocessing
To discard unnecessary air intensities, all images were cut into 512 × 256. Then their pixel intensities were linearly scaled from the tissue radiodensity range of [−135HU, 215HU] to grayscale range of [0, 255].
Architecture of MOFCN
In Figure 1(b), MOFCN’s convolution kernel size decreases with the increase of the network depth. It is based on the hypothesis that when the clinicians delineate OARs in the planning CTs, they first find the rough edges of OARs in their mind, and then determine their contour carefully. The big-size convolution kernel in MOFCN is used first to extract rough features from the big-size patches of a CT image. Then a small-size kernel is adopted to derive the subtle features.
Based on the above designing thought and limited by the computing hardware, the first network layer is a dilated convolutional one.33,34 The difference between a dilated and a conventional convolution is that they extract features from image patches with different sizes when using the same-size kernel. The dilated one’s image patch is larger than the conventional one. Each convolution layer contains an activation function of ReLU, except the last one. The function in the last convolution layer is sigmoid to produce a probability for a binary classification (i.e. background and foreground).
The MOFCN includes two components: one main backbone and three branches. The main backbone extracts the image features about 3 OARs. Then the features are transferred to three branches to generate three output images. This design is to ensure that the feature maps can involve the neighboring organs’ spatial information, because an organ’s location not only relates to its gray value, but also relates to its neighboring tissues. This point is discussed detailly in the last subsection of Discussion.
Postprocessing for final prediction
The three distinct outputs in MOFCN give three probability values (
If in some situations that
Experiments
The proposed MOFCN was implemented using Python (an open-source programming language for deep learning). The program ran on two Intel® E5-2630 processors (2.20 GHz) and four Graphics Cards (GeForce® GTX 1080, NVIDIA, Delaware, America).
In the collected 966 cases, the data from 61 randomly chosen patients were used for training and validation. Wherein, there were 856 CT scans (≈4% of all images) in the training set and 384 CT scans (≈2% of all images) in the validation set. Other 905 cases were used for testing. It encompassed 18,037 scans (≈94% of all images) for assessment. All images included 3 OARs. When splitting these cases, we labeled each case with a unique number from 1 to 966. Then we re-ordered the numbers using the function of random in Python. In the re-ordered sequence, the first 61 numbers corresponded to the identities (IDs) of cases in the training and validation sets. The remaining 905 numbers were the case IDs in the testing set.
The learning algorithm was Adaptive Moment Estimation (Adam). 37 The learning rate and epoch were 10−3 and 3000 respectively. The batch size equaled to 64. Therefore, in each iteration of one epoch, we transferred 64 CT images into the network for training. Then the network outputted three organs’ estimated segmentation images. They were compared with the ground truth for backpropagation. In one epoch, there were 14 (⌈856/64⌉=14, ⌈⌉ means the top integral function) iterations, because there were 856 training images. After each epoch, the loss of validation set was checked and only the model with the minimum loss would be saved.
Loss function and performance evaluation
The similarity between the auto-segmented OAR and its ground truth is scored using Dice similarity coefficient:
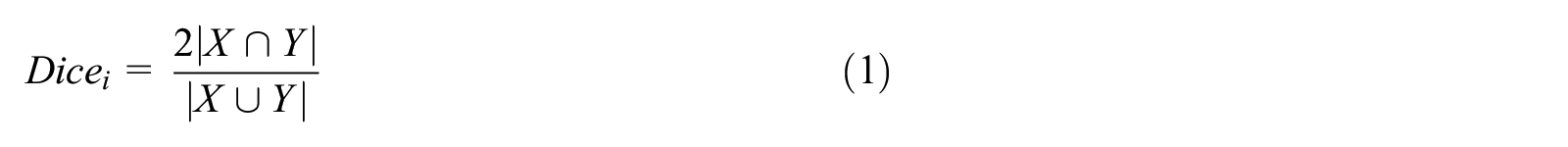
in which X is the model output. Y is the ground truth.
The loss function

where
We applied the proposed MOFCN on each patient’s case and assessed it using Dice in three dimensions (3D). For further analysis on under segmentation, we used the boxplot method
38
to find outliers (i.e. those not good performances). Specifically, the

Results
Testing results
The statistical results on the testing set are listed in Table 1. MOFCN achieves Dice of 0.95 ± 0.02 for lung, 0.91 ± 0.03 for heart and 0.87 ± 0.06 for spinal cord. The number of outlier cases are 1, 15, and 13 for lung, heart and spinal cord respectively.
The segmentation performance of the proposed model on the testing set.

Excluding the low-Dice cases caused by inter-observer variations.
Among the 23 outlier cases listed in Table 1, 4 cases (No. 114, 287, 363, and 710) show abnormality. Specially, No. 114 displays osteoplastic nodule. No. 287, 363, and 710 shows lung collapse. We infer that the abnormality is the reason of under segmentation in the four cases. The remaining 19 cases are further investigated in the Discussion section.
Comparison with other models
In this section, we provide a comparison between our network and other published models in the first subsection for an extensive evaluation. We compare the proposed model with its variants to study our design points: the dilated convolution (in the second subsection) and the distinct multiple outputs (in the third subsection). The effect of layer number on the segmentation performance is also shown in the second subsection.
Comparison with other published methods
In this subsection, we compare the MOFCN with other models, and list them in Table 2. The MOFCN was re-trained on the 36 training cases from 2017 AAPM Thoracic Auto-segmentation Challenges. 32 All methods were assessed on the 12 online testing cases.
Segmentation performance of seven methods. The results are expressed as average ± standard deviation for Dice. Testing time is the average segmentation time for one patient.*

Method 1–5 are the deep learning methods reported in a published work and their lung Dices were obtained by averaging left and right lung Dices. 32 MOFCN is our model.
Table 2 illustrates that MOFCN achieves a comparable segmentation accuracy with other models, but the least time cost.
Comparison with MOFCN variants
In this subsection, we investigate the effect of the MOFCN’s layer number and dilated convolution on the segmentation performance. By removing different layers in MOFCN or replacing the dilated convolution with a conventional one, we constructed multiple variants. These variants were also re-trained on the 36 training cases and tested on the 12 online testing cases from the same challenge in the first comparison. The results are listed in Table 3.
Segmentation performance of MOFCN variants. The results are expressed as average ± standard deviation for Dice (the results with three decimal places are printed in parenthesis for meticulous comparison). Testing time is the average segmentation time for one patient.
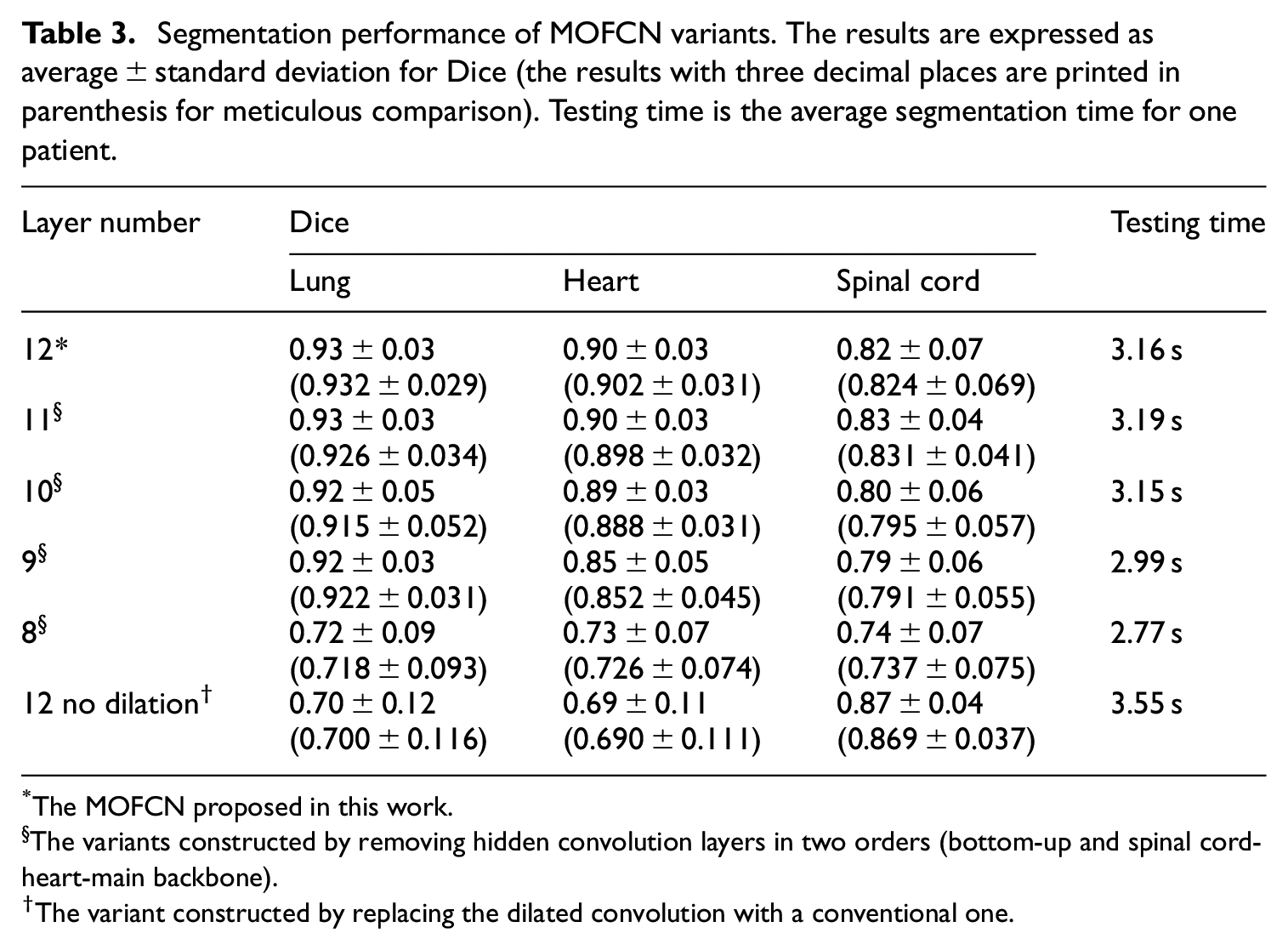
*The MOFCN proposed in this work.
§The variants constructed by removing hidden convolution layers in two orders (bottom-up and spinal cord-heart-main backbone).
The variant constructed by replacing the dilated convolution with a conventional one.
Table 3 shows the trend of decreasing accuracy and time cost when reducing layer number. The layer numbers of 11, 10, and 9 don’t include removing layer in the lung branch. Hence, the lung Dice doesn’t show too much change when comparing with other two organs.
The first and last rows in Table 3 displays that the lung and heart Dice achieved by the no-dilation-convolution MOFCN are significantly lower than the proposed one. It may be caused by the size decrease of receptive filed because of no dilated convolution. As a result, there is no enough knowledge to help the MOFCN variant for classification and it needs more time for correcting discontinuities.
Comparison with multi-label learning-based single-output model
To evaluate the distinct-output design in this subsection, we compare the proposed MOFCN with a single-output model which is based on multi-label learning (abbr. MLFCN). The architecture of MLFCN is shown in Figure 2(a). It is totally same as the spinal cord branch of MOFCN to reduce the impact of network depth on segmentation. The activation function in the last layer of MLFCN is softmax. Softmax produces four probability values of a pixel belonging to four labels (i.e. background, lung, heart and spinal cord) and the largest one gives the final classification. MLFCN was trained using the same data and optimization settings as MOFCN, but its loss function was categorical cross entropy. The MLFCN was also assessed on the same testing data. Its results are compared with MOFCN in Figure 2(b).

(a) Architectures of MOFCN and MLFCN. MOFCN is the proposed model. MLFCN, a multi-label learning-based model, is the comparative one. The structure of MLFCN is totally same as the spinal cord branch (in the blue dotted box) in MOFCN, except the activation function in the last layer. The abbreviations indicate the same as in Figure 1.
Figure 2(b) suggests that (a) MOFCN gets a better spinal cord segmentation than MLFCN and (b) MLFCN and MOFCN achieve equally well results when segmenting lung and heart. For (a), we infer that the enhancement is beneficial from the distinct output, because the design concentrates the last several layers’ learning capability on segmenting spinal cord, rather than distributing to other tasks. This is also why we employ multiple outputs. It is to combine the information sharing of various neighboring organs and independent feature extraction for individual one. For (b), it may be caused by the small contribution of a small-volume organ (e.g. spinal cord) on recognizing large-volume organs (e.g. lung and heart). Besides, even with deeper network in MLFCN, the delineation of lung and heart don’t show too much improvement.
Discussion
We analyze the testing result of spinal cord, further investigate the 19 outlier cases, and take spinal cord for example to explain why we adopt the structure of multiple outputs.
Analysis on the testing result of spinal cord
Table 1 suggests that the spinal cord, corresponding to the longest network branch in MOFCN, shows the lowest Dice among the three organs. We infer that the main reason is the inter-observer variations. Figure 3 gives some representative slices in which the spinal cord segmentation reaches a Dice of 0.48–0.63, but they are clinically accepted according to the recommended atlas of spina cord. 36

Illustration of spinal cord segmentation with a Dice of <0.7, but clinically accepted (a-f) are 6 such cases Denotation: No., patient number; s., slice number.
Detailed investigation of 19 exceptional cases
In this section, we give a detailed investigation on the above 19 exceptions. By reviewing each case, we categorize the under-segmentation into misrecognition and missed recognition (as shown in Table 4). The former one means that the non-object pixels are recognized as part of object. The latter one represents that the object pixels are not recognized as part of object.
Under-segmentation category for 19 outlier testing cases.

Analysis on the failure cause of misrecognition
For a CNN, the misrecognition directly results from the similar features of the non-object pixel, compared to an object one. Figure 4 illustrates the misrecognition images of No. 75, 104, and 439. Compared to the reference scan in Figure 4(d), there are non-zero neighboring pixels around heart (indicated by the yellow thick arrows in Figure 4) in the three failure cases. Accordingly, we infer that the misrecognition reason is the lack or less of such samples in training data, hence the model didn’t learn how to distinguish heart from its uncommon neighbors.

Illustration of the misrecognition cases (No. 75, 104, and 439 in (a–c)). (d) shows a common image with heart for reference. Yellow thick arrows indicate the difference between (a–c) and (d). “s.” represents the slice number.
Analysis on the failure cause of missed recognition
The segmentation task of a CNN is a pixel-wise classification task. 40 The classification is only based on the knowledge of the pixel’s reception field. Thus, any variations in the reception field, such as geometric and grayscale variations, are high likely to cause an under segmentation. To validate our conjecture on the missed recognition causes, we conducted the geometric or grayscale adjustments on the original input images for improving performance and display them in Figure 5.

Bar graph of segmentation accuracy before and after adjustment for the missed recognition cases. (a–c) relate to the heart, spinal cord (abbr. sp) and both respectively. (d) is the adjustment reference (No. 10 slice for No. 1 patient). The body size = 300 × 207.The left-right direction = horizon. The gray value (
Figure 5(a–c) show that most cases receive improvement, when we adjust the input image according to the reference (as shown in Figure 5(d)). The adjustments include: (a) resizing the body to have the same size; (b) rotating the body until its left-right direction approximates horizontal and (c) rescaling the mapping between tissue radiodensity and gray value to increase the number of 255-grayvalue pixels around the spinal cord.
Future work
Above all, there are two factors impacting the proposed MOFCN. They are (a) special radiodensity of tissues and (b) geometric change, such as body size change and rotation. The factor (a) may be caused by the spongy bone, pulmonary atelectasis and other special cases that are not or less included in the training data.
In our future work, to overcome factor (a-b), we would include more special radiodensity and geometry samples in training to improve the model’s anti-interference performance or adopt an adaptive grayscale transformation. 3D models are also potential in leading to better performances. By expanding the convolution kernel to 3D and hence involving inter-slice information, it is likely to learn more about the comprehensively spatial relationship among different voxels and survive from multiple disturbance. To consider both performance and cost for a widespread clinical application, we would investigate the correlation of segmentation improvement to various number of 3D model parameters. Further modification would be a tradeoff between performance and efficiency (e.g. cost on hardware, running time).
Another possible solution for factor (b) is to develop a geometric correction in the image preprocessing to guarantee desirable segmentations.
The three OARs segmented by our model are the dose-limited organs for conventional radiation treatment. For stereotactic body radiotherapy (SBRT), more than three organs are involved in concerns. 36 In such scenario, we can attach other more branches to the proposed MOFCN to build a modified one, and use its parameters as part of the initial values to train the modified network. The pre-trained parameters may accelerate the optimization.
Auto-segmentation of spinal cord using a multi-output structure
There are two advantages of MOFCN on auto-segmenting spinal cord: (a) one is to use a relatively large kernel at the first convolutional layer to involve neighboring organs’ information about spinal cord; (b) the second one is to use the multi-output network structure to avoid over-fitting.
In the thorax, the image features about the spinal cord are supposed to include two points: (i) it appears in gray and is surrounded by a white annular region and (ii) it lies between two large-area black connected regions. The white annular region is the spinal canal and shows white in a CT. The two large-area black connected regions are the left and right lungs. Between them lies the spinal canal.
Without a relatively large-size convolutional kernel, it is hard to distinguish the spinal cord and the humerus marrow. As shown in Figure 6,

Illustration of reception fields for a humerus marrow pixel (A) and a spinal cord pixel (B).
Conclusion
In this article, we propose a MOFCN to segment bilateral lung, heart and spinal cord in the thorax automatically and simultaneously. According to the evaluation results in 18,037 CT images and comparison with other models, the MOFCN is proved effective. Its multi-output structure can perform multiple object recognition and segmentation, which shows a relatively low requirement for computing and thus has potential for broad application. We also suggest potential solutions to further improve its performance on some “unseen” cases, including geometric change, adaptive grayscale transformation and so on.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Key Research and Development Program (2017YFC0113201), Zhejiang Provincial Natural Science Foundation of China (LQ20H180016, LQ17H180003), Zhejiang Key Research and Development Program (2019C03003), Youth Talent Foundation of Zhejiang Medical and Health Project (2019RC023), Appropriate Technology Cultivation and Promotion of Zhejiang Medical and Health Project (2019ZH018), Postdoctoral Program of Zhejiang Province, Chinese Postdoctoral Fund (520000-X91601) and National Natural Science Foundation of China (81230031/H18, 82001928).
Ethics statement
There is no human or animal participants involved in this study. All computed tomography images in our experiment were obtained for the conventional radiation treatment. We just collected and used them to train and validate the proposed model.
